UCLA Advanced Neuroimaging Summer Program David W. Shattuck, PhD
|
|
|
- Loreen Spencer
- 5 years ago
- Views:
Transcription
1 BrainSuite UCLA Advanced Neuroimaging Summer Program Presented 18 July 2013 David W. Shattuck, PhD Associate Professor Department of Neurology David Geffen School of Medicine at UCLA
2 What is BrainSuite? Collection of image analysis tools designed to process structural and diffusion MRI Automated sequence to extract cortical surface models from T1- MRI Tools to register surface and volume data to an atlas to define anatomical ROIs Tools for processing diffusion imaging data, including coregistration to anatomical T1 image, ODF and tensor fitting, and tractography. Visualization tools for exploring these data. Runs on Windows, Mac, and Linux* * GUI for Linux version is not yet released
3 Overview Presentation Background Cortical Surface Extraction Surface/Volume Registration BrainSuite Diffusion Pipeline Visualization Tools Lab will follow with sample data and exercises
4 Motivation for Mapping It is often the goal to perform comparisons across different brains or brains at different points in time For these comparisons to be meaningful, we must be able to establish spatial anatomical correspondence among the data Once correspondence is established, we can look for significant differences in various neuroanatomical features Size of structures Cortical thickness Cortical complexity White matter architecture Connectivity relationships How these change over time or in the presence of disease or trauma
5 Automate all the things? One approach to comparative neuroimaging is to manually delineate anatomical structures. Drawbacks to manual methods: Raters must be trained to be consistent and to follow a specified protocol Learning effects may bias their processing Raters don t always visualize 3D relationships when viewing slice-based data Human raters still constitute the gold standard for many applications Automated methods can benefit from the expertise of the rater, which may be superior to an automated algorithm. Important to recognize that automated methods may need supervision or correction A manually delineated brain atlas (BrainSuiteAtlas1)
6 Image Registration Goal: identify a transformation that maps from one image to another, such that image features or landmarks are matched. ICBM452 Atlas, aligned to subject image using AIR affine transform and 5 th order warp Subject Image
7 Why use surface models? Cortex is often represented as a high resolution triangulated mesh with ~700,000 triangles Many volumetric-based approaches do not align the cortical anatomy well We are often interested in functional areas in the cortex Surface-based features, e.g., cortical thickness, are of interest in the study of development or disease processes For applications such as EEG/MEG source localization, the location and orientation of the cortical surface can provide additional information Cortical surface mesh representation
8 Why diffusion MRI? Quantify microstructural tissue characteristics Structural connectivity connectomics (Sporns 2005; Wedeen 2008; Hagmann 2007) Clinical investigation of abnormalities in white matter e.g., stroke, Alzheimer's disease (Jones 2011; Johansen- Berg 2009) FA map Fiber tracks
9 BrainSuite Workflow
10 Cortical Surface Extraction
11 Magnetic Resonance Images MRI provide noisy, limited resolution representations of brain anatomy Stained section from a photographic atlas (Roberts et al.) T1 weighted MRI
12 Nonuniformity and Noise The intensity of voxel would ideally be given only by the tissues present in that voxel. Imperfections in the scanner hardware, as well as susceptibility variations in the subject, introduce magnetic field artifacts that produce shading in the image. Other noise in the system will also confound the classification process.
13 Partial Volume Effects Finite resolution of MRI is insufficient for some neuroanatomical details. Each measurement is an average of the tissue signals in the voxel. Tissue mixtures
14 Frequency Image Formation: Ideal Case CSF GM WM Intensity In the ideal case: An extracted brain would contain only GM, WM, and CSF We would measure a single intensity value for each type of tissue present in the image. The resolution would be sufficient that each voxel would be composed of a single tissue type If this were true, then classification would be simple
15 Image Formation: Reality whole-head Some other types of tissue are likely to be present (vessels, sinuses, etc.) skull-stripped skull-stripped, bias corrected Histograms of: (top) a whole-head MRI (middle) a skull-stripped MRI (bottom) non-uniformity corrected skull-stripped MRI Image artifacts produce variations in the measured intensity for the tissues Slow spatial gain variation Spatially independent noise Neuroanatomical detail is much too fine for the mm 3 voxel size typical in MRI
16 Cortical Surface Extraction
17 Brain Surface Extractor (BSE) MRI Filtered MRI Extracts the brain from non-brain tissue (skullstripping) We apply a combination of: anisotropic diffusion filtering edge detection mathematical morphological operators This method can rapidly identify the brain within the MRI Edge Mask Brain Boundary (green)
18 Skull and Scalp Modeling We can apply thresholding, mathematical morphology, and connected component labeling to MRI to identify skull and scalp regions. The method builds upon the BSE skull stripping result. The volumes produced by this algorithm will not intersect. We can produce surface meshes from the label volume. The results are suitable for use in MEG/EEG source localization.
19 Bias Field Corrector Histograms of the two ROIs Two cubic regions of interest (ROIs) Performs non-uniformity correction by analyzing regional histograms Sub-volumes have dramatically different profiles. Regional histograms reflect this. 3D rendering of the ROIs
20 frequency Bias Field Corrector Local histograms Partial volume measurement model CSF CSF CSF GM GM GM WM WM WM CSF/GM GM/WM CSF/Other model density intensity We can fit a tunable model of the tissue profiles to many ROIs spaced throughout the brain. This allows us to estimate the local gain variation.
21 Bias Field Corrector Estimate bias parameter at several points throughout the image. Remove outliers from our collection of estimates. Fit a tri-cubic B-spline to the estimate points. Divide the image by the B-spline to make the correction.
22 Maximum Likelihood Classifier We can compute a maximum likelihood classification from our measurement model. At each voxel, we simply compute the probability of the intensity value belonging to a particular tissue, and then select the label that has the highest likelihood. This maps each intensity value to a specific label, and it thus very fast. Does not take into account any of the surrounding labels. CSF GM WM CSF/GM GM/WM CSF/other
23 Partial Volume Classifier (PVC) We can construct a model that computes a score for local neighborhoods Higher scores for pixel configurations that are similar Lower scores for unlikely combinations (e.g., WM next to CSF) We use this model to produce a maximum a posteriori (MAP) classifier We maximize this function using the Iterated Conditional Modes (ICM) algorithm Initialize ICM with a maximum likelihood labeling. Iteratively update each individual label to maximize CSF GM WM CSF/GM GM/WM CSF/other
24 PVC Tissue Fraction Estimation For each brain voxel, we estimate the tissue fraction as follows: Pure voxels are 100%. Each mixed tissue voxel is assigned a fractional value based on where its signal intensity falls between the class means. WM GM CSF BG
25 Cerebrum Identification For the cortical surface, we are interested in the cerebrum, which we separate from the rest of the brain. We achieve this by registering a multi-subject average brain (ICBM452) to the individual brain using AIR (R. Woods) We have labeled this atlas: cerebrum / cerebellum subcortical regions left / right
26 Cortex Extraction a We combine our registered brain atlas with our tissue map retain subcortical structures, including nuclei identify the inner boundary of the cerebral cortex
27 Topological Errors In normal human brains, the cortical surface can be considered as a single sheet of grey matter. Closing this sheet at the brainstem, we can assume that the topology of the cortical surface is equivalent to a sphere, i.e., it should have no holes or handles. This allows us to represent the cortical surface using a 2D coordinate system. Unfortunately, our segmentation result will produce a surface with many topological defects.
28 Topological Errors We can identify topological loops in the volume segmentation by representing it with two graphs. y z x FG BG If these graphs have cycles, then topological handles exist in the object. y z x
29 Topological Editing By analyzing the graphs, we can identify locations in the object where we can either remove or add voxels in order to break a cycle in the graph. We can make our decisions of where to edit based on making small changes to the object. This method allows us to rapidly remove all topological defects and produce a volumetric segmentation that will yield a genus zero surface mesh Foreground 2 2 Background
30 Topology Correction Cortical surface model produced from binary masks (top right) close-up view of a handle on the surface generated from the volume before topological correction (bottom right) close-up view of the same region on the surface generated from the same volume after topology correction.
31 Digital Object Filtering
32 Pial Surface Expand inner cortex to outer boundary Produces a surface with 1-1 vertex correspondence from GM/WM to GM/CSF Preserves the surface topology Provides direct thickness computation Data from each surface maps directly to the other
33 Pial Surface Contour view showing the inner (blue) and outer (orange) boundaries of the cortex.
34 SVReg: Surface-constrained Volumetric Registration
35 Surface-constrained Volumetric Registration
36 BrainSuite T1 MRI and label overlay Atlas Single subject atlas labeled at USC by expert neuroanatomist left and right hemispheres 26 sulcal curves per hemisphere 98 volumetric regions of interest (ROIs), 35*2=70 cortical ROIs Included with BrainSuite13 as BrainSuiteAtlas1 flat maps
37 Cortical Surface Parameterization corpus callosum Mapping Flat-map color coded by curvature Each hemisphere is mapped to the unit square using an energy-minimization technique.
38 Curvature-based Registration subject atlas
39 3D Alignment Input mid surface Smoothed surfaces Atlas Subject Atlas Subject AIR transformation Matching based on L2 penalty
40 Multi-resolution Surface Matching Input mid surface Smoothed surfaces Cumulative curvature computation for multi-resolution representation atlas subject warped subject color-coded labels Elastic matching for atlas and subject flat maps
41 Curvature Weighting Shown is the color-coded curvature variance, as computed by aligning 100 normal adult brains. Inverse of curvature variance is used as a weighting on the curvature cost function to reduce the influence of highly variable areas.
42 Curvature Weighting Results No curvature variance weighting With curvature-variance weighting
43 Refinement of labels and sulci Original labels plotted on a smoothened representation of a cortical surface Labels after geodesic curvature Flow plotted on a smoothened representation of a cortical surface Animation of the geodesic curvature flow for sulcal refinement
44 Surface Registration Methods?? + Accurate sulcal alignment - Doesn t define volumetric correspondence
45 Motivation for Surface-constrained Volumetric Registration Alignment of 2 brains by AIR (5 th order) + Good alignment of subcortical structures - Sulcal alignment inaccurate
46 Extension to Volumetric Registration Accurate Sulcal Alignment Accurate Subcortical Feature Alignment Intensity-based Alignment Surface registration Extrapolation to volume Solves the difficult problem of surface/sulcal registration in 3D volume Volumetric intensity registration Surface and Volume Registration (SVReg) method performs accurate alignment of both cortical surfaces as well as subcortical volumes. Joshi et al. (2007) IEEE Trans. Med. Imag.
47 Atlas Subject BrainSuite ROI Labeling (top) Surface and volume views of the BrainSuite13 anatomical atlas, delineated into anatomical regions of interest. (bottom) Similar views of an automatically labeled subject dataset.
48 SVReg Software Workflow SVReg Outputs - Labeled cortical surfaces - Labeled brain volume - Measurements for each ROI (area, volume) - Mappings to atlas space - Mapped sulcal curves
49 Integration with BrainStorm BrainSuite Cortical Surface Model with ROIs Labeling imported into BrainStorm. The BrainSuite parcellation can be directly imported into BrainSuite, where the ROIs are useful for interpreting current sources. see also:
50 BDP: BrainSuite Diffusion Pipeline
51 Diffusion Image Acquisition A set of diffusion-weighted images is acquired with diffusion-sensitizing magnetic field gradients Gradients are oriented in different directions A 3D volume image is acquired for each direction Reconstruction methods are used to estimate the local diffusion properties Image from: Chiang M et al. (2009) Genetics of Brain Fiber Architecture and Intellectual Performance J. Neurosci. 29:
52 Diffusion Tensor Imaging (DTI) With at least six directions and a baseline image, a tensor model can be estimated. Different types of tissue will have different diffusion properties Oriented along nerve fibers Free diffusion in CSF and grey matter Visualization of scalar properties (e.g., fractional anisotropy) Visualization of major eigenvector using direction encoded color (DEC) maps Red: x, left/right Green: y, anterior/posterior Blue: z, inferior/superior A/P L/R I/S
53 DTI Visualization Often visualized using ellipsoids Spherical shapes indicate isotropic diffusion Elongated shapes indicate directionality Flat discs are suggestive of the crossing or junction of nerve fibers A/P L/R I/S
54 High Angular Resolution Diffusion Imaging (HARDI) The tensor model is limited in what it can resolve Fiber tracts may cross in a voxel, presenting ambiguities in determining the meaning of the diffusion pattern By sampling in many more directions, we can get a more complete picture of the diffusion pattern Examples include Q-Ball imaging (Tuch, 2004) Can be processed and visualized using spherical harmonics Image from Shattuck et al. (2008) Visualization Tools for High Angular Resolution Diffusion Imaging, Medical Image Computing and Computer Assisted Intervention (MICCAI) 2008
55 BrainSuite Diffusion Pipeline DICOM to NifTi Distortion Correction and Registration Tensor and ODF Estimation Tractography and Quantitative Analysis A framework to: Read diffusion data from DICOM images Align diffusion and MPRAGE image Correct diffusion data for distortions Fit different diffusion models tensor and ODFs Compute different quantitative diffusion parameters Compute diffusion tracks and connectivity matrix
56 DICOM to NifTi Distortion Correction and Registration Tensor and ODF Estimation Tractography and Quantitative Analysis Scanner saved DICOM images, as input De-mosaic the diffusion images Extracts diffusion parameters bmat, bval, bvec Re-orients diffusion gradients to voxel coordinates Writes standard 4D nifti files
57 DICOM to NifTi Distortion correction and Registration Tensor and ODF Estimation Tractography and Quantitative Analysis Diffusion MRI uses fast acquisition Echo planar Imaging (EPI) EPI is sensitive to magnetic field (B 0 ) inhomogeneity Localized geometric distortion MPRAGE image b=0 image Field inhomogeneity map
58 DICOM to NifTi Distortion correction and Registration Tensor and ODF Estimation Tractography and Quantitative Analysis Distortions results in misalignment with structural scans by several millimeters Limits the accuracy of multi-modal analysis b=0 image MPRAGE image Overlay with edges
59 DICOM to NifTi Distortion correction and Registration Tensor and ODF Estimation Tractography and Quantitative Analysis Corrects the distortion in diffusion (EPI) images using non-rigid registration No fieldmap is required for correction Before After Before After Each figure shows (left) distorted and (right) corrected b=0 image, overlaid with the edge-map (red outline) generated from the T1-weighted image. Arrows indicate areas of significant correction. Bhushan et al Correcting Susceptibility-Induced Distortion in Diffusion- Weighted MRI using Constrained Nonrigid Registration, APSIPA 2012.
60 DICOM to NifTi Distortion Correction and Registration Tensor and ODF Estimation Tractography and Quantitative Analysis Estimates diffusion tensors FA, MD, color-fa Axial, Radial Diffusivity ODFs using FRT and FRACT FRACT (Haldar and Leahy, 2013) improved accuracy higher angular resolution
61 DICOM to NifTi Distortion Correction and Registration Tensor and ODF Estimation Tractography and Quantitative Analysis Fiber tracking in T1-structural space ROI-based connectivity analysis
62 Acknowledgments Richard Leahy, PhD Anand Joshi, PhD Chitresh Bhushan Justin Haldar, PhD Soyoung Choi Andrew Krause Jessica Wisnowski, PhD Hanna Damasio, MD This work was supported in part by NIH grants R01-NS074980, R01-EB009048, P41-EB015922, and U01-MH93765.
63 Questions
64 References The software is available online at: Additional documentation at: User forum: More details for the methods described in this talk can be found in the following papers: Segmentation Shattuck DW, Sandor-Leahy SR, Schaper KA, Rottenberg DA and Leahy RM (2001) Magnetic Resonance Image Tissue Classification Using a Partial Volume Model NeuroImage, 13(5): Shattuck DW and Leahy RM (2001) Graph Based Analysis and Correction of Cortical Volume Topology, IEEE Transactions on Medical Imaging, 20(11) Shattuck DW and Leahy RM (2002) BrainSuite: An Automated Cortical Surface Identification Tool Medical Image Analysis, 8(2): Dogdas B, Shattuck DW, and Leahy RM (2005) Segmentation of Skull and Scalp in 3D Human MRI Using Mathematical Morphology Human Brain Mapping26(4):273-85
65 References Registration and Curve Delineation Joshi AA, Leahy RM, Thompson PM, Shattuck DW (2004) Cortical Surface Parameterization by P-Harmonic Energy Minimization. ISBI 2004: Joshi AA, Shattuck DW, Thompson PM, and Richard M. Leahy (2007) Surface- Constrained Volumetric Brain Registration Using Harmonic Mappings IEEE Trans. on Medical Imaging 26(12): Joshi AA, Shattuck DW, Thompson PM, and Leahy RM (2009) A Parameterizationbased Numerical Method for Isotropic and Anisotropic Diffusion Smoothing on Non- Flat Surfaces, IEEE Trans. on Image Processing18(6): Shattuck DW, Joshi AA, Pantazis D, Kan E, Dutton RA, Sowell ER, Thompson PM, Toga AW, Leahy RM (2009) Semi-automated Method for Delineation of Landmarks on Models of the Cerebral Cortex, Journal of Neuroscience Methods 178(2): Joshi AA, Pantazis D, Li Q, Damasio H, Shattuck D, Toga A, Leahy RM (2010) Sulcal set optimization for cortical surface registration NeuroImage 50(3):950-9 Pantazis D, Joshi AA, Jintao J, Shattuck DW, Bernstein LE, Damasio H, and Leahy RM (2010) Comparison of landmark-based and automatic methods for cortical surface registration, NeuroImage 49(3): Joshi AA, Shattuck DW and Leahy RM, A Fast and Accurate Method for Automated Cortical Surface Registration and Labeling, Proc. WBIR LNCS Springer 2012,
66 References Diffusion MRI Haldar JP and Leahy RM (2013) Linear transforms for Fourier data on the sphere: Application to high angular resolution diffusion MRI of the brain NeuroImage 71: Bhushan C, Haldar JP, Joshi AA, and Leahy RM (2012) Correcting susceptibility-induced distortion in diffusion-weighted MRI using constrained nonrigid registration, Asia-Pacific Signal & Information Processing Association Annual Summit and Conference (APSIPA ASC), 1-9, Los Angeles, California, 3-6 Dec 2012
BrainSuite. presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014
 BrainSuite presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 David Shattuck Ahmanson-Lovelace Brain Mapping Center Department of Neurology David Geffen School of Medicine at
BrainSuite presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 David Shattuck Ahmanson-Lovelace Brain Mapping Center Department of Neurology David Geffen School of Medicine at
BDP: BrainSuite Diffusion Pipeline. Chitresh Bhushan
 BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
BDP: BrainSuite Diffusion Pipeline. Chitresh Bhushan
 BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
A Fast and Accurate Method for Automated Cortical Surface Registration and Labeling
 A Fast and Accurate Method for Automated Cortical Surface Registration and Labeling Anand A Joshi 1, David W Shattuck 2 and Richard M Leahy 1 1 Signal and Image Processing Institute, University of Southern
A Fast and Accurate Method for Automated Cortical Surface Registration and Labeling Anand A Joshi 1, David W Shattuck 2 and Richard M Leahy 1 1 Signal and Image Processing Institute, University of Southern
BrainSuite Lab Exercises. presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014
 BrainSuite Lab Exercises presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 1. Opening and Displaying an MRI Start BrainSuite Drag and drop the T1 image from the native space
BrainSuite Lab Exercises presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 1. Opening and Displaying an MRI Start BrainSuite Drag and drop the T1 image from the native space
Lilla Zöllei A.A. Martinos Center, MGH; Boston, MA
 Lilla Zöllei lzollei@nmr.mgh.harvard.edu A.A. Martinos Center, MGH; Boston, MA Bruce Fischl Gheorghe Postelnicu Jean Augustinack Anastasia Yendiki Allison Stevens Kristen Huber Sita Kakonoori + the FreeSurfer
Lilla Zöllei lzollei@nmr.mgh.harvard.edu A.A. Martinos Center, MGH; Boston, MA Bruce Fischl Gheorghe Postelnicu Jean Augustinack Anastasia Yendiki Allison Stevens Kristen Huber Sita Kakonoori + the FreeSurfer
Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm
 Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm Lin Chen and Gudrun Wagenknecht Central Institute for Electronics, Research Center Jülich, Jülich, Germany Email: l.chen@fz-juelich.de
Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm Lin Chen and Gudrun Wagenknecht Central Institute for Electronics, Research Center Jülich, Jülich, Germany Email: l.chen@fz-juelich.de
HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge
 Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
The organization of the human cerebral cortex estimated by intrinsic functional connectivity
 1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
MR IMAGE SEGMENTATION
 MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases
 Quantities Measured by MR - Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases Static parameters (influenced by molecular environment): T, T* (transverse relaxation)
Quantities Measured by MR - Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases Static parameters (influenced by molecular environment): T, T* (transverse relaxation)
An Introduction To Automatic Tissue Classification Of Brain MRI. Colm Elliott Mar 2014
 An Introduction To Automatic Tissue Classification Of Brain MRI Colm Elliott Mar 2014 Tissue Classification Tissue classification is part of many processing pipelines. We often want to classify each voxel
An Introduction To Automatic Tissue Classification Of Brain MRI Colm Elliott Mar 2014 Tissue Classification Tissue classification is part of many processing pipelines. We often want to classify each voxel
Subvoxel Segmentation and Representation of Brain Cortex Using Fuzzy Clustering and Gradient Vector Diffusion
 Subvoxel Segmentation and Representation of Brain Cortex Using Fuzzy Clustering and Gradient Vector Diffusion Ming-Ching Chang Xiaodong Tao GE Global Research Center {changm, taox} @ research.ge.com SPIE
Subvoxel Segmentation and Representation of Brain Cortex Using Fuzzy Clustering and Gradient Vector Diffusion Ming-Ching Chang Xiaodong Tao GE Global Research Center {changm, taox} @ research.ge.com SPIE
NEURO M203 & BIOMED M263 WINTER 2014
 NEURO M203 & BIOMED M263 WINTER 2014 MRI Lab 2: Neuroimaging Connectivity Lab In today s lab we will work with sample diffusion imaging data and the group averaged fmri data collected during your scanning
NEURO M203 & BIOMED M263 WINTER 2014 MRI Lab 2: Neuroimaging Connectivity Lab In today s lab we will work with sample diffusion imaging data and the group averaged fmri data collected during your scanning
Advanced MRI Techniques (and Applications)
 Advanced MRI Techniques (and Applications) Jeffry R. Alger, PhD Department of Neurology Ahmanson-Lovelace Brain Mapping Center Brain Research Institute Jonsson Comprehensive Cancer Center University of
Advanced MRI Techniques (and Applications) Jeffry R. Alger, PhD Department of Neurology Ahmanson-Lovelace Brain Mapping Center Brain Research Institute Jonsson Comprehensive Cancer Center University of
Fmri Spatial Processing
 Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
Diffusion model fitting and tractography: A primer
 Diffusion model fitting and tractography: A primer Anastasia Yendiki HMS/MGH/MIT Athinoula A. Martinos Center for Biomedical Imaging 03/18/10 Why n how Diffusion model fitting and tractography 0/18 Why
Diffusion model fitting and tractography: A primer Anastasia Yendiki HMS/MGH/MIT Athinoula A. Martinos Center for Biomedical Imaging 03/18/10 Why n how Diffusion model fitting and tractography 0/18 Why
Automated MR Image Analysis Pipelines
 Automated MR Image Analysis Pipelines Andy Simmons Centre for Neuroimaging Sciences, Kings College London Institute of Psychiatry. NIHR Biomedical Research Centre for Mental Health at IoP & SLAM. Neuroimaging
Automated MR Image Analysis Pipelines Andy Simmons Centre for Neuroimaging Sciences, Kings College London Institute of Psychiatry. NIHR Biomedical Research Centre for Mental Health at IoP & SLAM. Neuroimaging
Generation of Hulls Encompassing Neuronal Pathways Based on Tetrahedralization and 3D Alpha Shapes
 Generation of Hulls Encompassing Neuronal Pathways Based on Tetrahedralization and 3D Alpha Shapes Dorit Merhof 1,2, Martin Meister 1, Ezgi Bingöl 1, Peter Hastreiter 1,2, Christopher Nimsky 2,3, Günther
Generation of Hulls Encompassing Neuronal Pathways Based on Tetrahedralization and 3D Alpha Shapes Dorit Merhof 1,2, Martin Meister 1, Ezgi Bingöl 1, Peter Hastreiter 1,2, Christopher Nimsky 2,3, Günther
Methods for data preprocessing
 Methods for data preprocessing John Ashburner Wellcome Trust Centre for Neuroimaging, 12 Queen Square, London, UK. Overview Voxel-Based Morphometry Morphometry in general Volumetrics VBM preprocessing
Methods for data preprocessing John Ashburner Wellcome Trust Centre for Neuroimaging, 12 Queen Square, London, UK. Overview Voxel-Based Morphometry Morphometry in general Volumetrics VBM preprocessing
Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry
 Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry Nivedita Agarwal, MD Nivedita.agarwal@apss.tn.it Nivedita.agarwal@unitn.it Volume and surface morphometry Brain volume White matter
Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry Nivedita Agarwal, MD Nivedita.agarwal@apss.tn.it Nivedita.agarwal@unitn.it Volume and surface morphometry Brain volume White matter
Automatic Quantification of DTI Parameters along Fiber Bundles
 Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein 1, Simon Hermann 1, Olaf Konrad 1, Horst K. Hahn 1, and Heinz-Otto Peitgen 1 1 MeVis Research, 28359 Bremen Email: klein@mevis.de
Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein 1, Simon Hermann 1, Olaf Konrad 1, Horst K. Hahn 1, and Heinz-Otto Peitgen 1 1 MeVis Research, 28359 Bremen Email: klein@mevis.de
UNC 4D Infant Cortical Surface Atlases, from Neonate to 6 Years of Age
 UNC 4D Infant Cortical Surface Atlases, from Neonate to 6 Years of Age Version 1.0 UNC 4D infant cortical surface atlases from neonate to 6 years of age contain 11 time points, including 1, 3, 6, 9, 12,
UNC 4D Infant Cortical Surface Atlases, from Neonate to 6 Years of Age Version 1.0 UNC 4D infant cortical surface atlases from neonate to 6 years of age contain 11 time points, including 1, 3, 6, 9, 12,
DIFFUSION TENSOR IMAGING ANALYSIS. Using Analyze
 DIFFUSION TENSOR IMAGING ANALYSIS Using Analyze 2 Table of Contents 1. Introduction page 3 2. Loading DTI Data page 4 3. Computing DTI Maps page 5 4. Defining ROIs for Fiber Tracking page 6 5. Visualizing
DIFFUSION TENSOR IMAGING ANALYSIS Using Analyze 2 Table of Contents 1. Introduction page 3 2. Loading DTI Data page 4 3. Computing DTI Maps page 5 4. Defining ROIs for Fiber Tracking page 6 5. Visualizing
Super-Resolution Reconstruction of Diffusion-Weighted Images from Distortion Compensated Orthogonal Anisotropic Acquisitions.
 Super-Resolution Reconstruction of Diffusion-Weighted Images from Distortion Compensated Orthogonal Anisotropic Acquisitions. Benoit Scherrer Ali Gholipour Simon K. Warfield Children s Hospital Boston,
Super-Resolution Reconstruction of Diffusion-Weighted Images from Distortion Compensated Orthogonal Anisotropic Acquisitions. Benoit Scherrer Ali Gholipour Simon K. Warfield Children s Hospital Boston,
Automatic Quantification of DTI Parameters along Fiber Bundles
 Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein, Simon Hermann, Olaf Konrad, Horst K. Hahn, Heinz-Otto Peitgen MeVis Research, 28359 Bremen Email: klein@mevis.de Abstract. We introduce
Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein, Simon Hermann, Olaf Konrad, Horst K. Hahn, Heinz-Otto Peitgen MeVis Research, 28359 Bremen Email: klein@mevis.de Abstract. We introduce
Impact of acquisition protocols and processing streams on tissue segmentation of T1 weighted MR images
 www.elsevier.com/locate/ynimg NeuroImage 29 (2006) 185 202 Impact of acquisition protocols and processing streams on tissue segmentation of T1 weighted MR images Kristi A. Clark, a, * Roger P. Woods, a
www.elsevier.com/locate/ynimg NeuroImage 29 (2006) 185 202 Impact of acquisition protocols and processing streams on tissue segmentation of T1 weighted MR images Kristi A. Clark, a, * Roger P. Woods, a
Discriminative, Semantic Segmentation of Brain Tissue in MR Images
 Discriminative, Semantic Segmentation of Brain Tissue in MR Images Zhao Yi 1, Antonio Criminisi 2, Jamie Shotton 2, and Andrew Blake 2 1 University of California, Los Angeles, USA. zyi@ucla.edu. 2 Microsoft
Discriminative, Semantic Segmentation of Brain Tissue in MR Images Zhao Yi 1, Antonio Criminisi 2, Jamie Shotton 2, and Andrew Blake 2 1 University of California, Los Angeles, USA. zyi@ucla.edu. 2 Microsoft
Supplementary methods
 Supplementary methods This section provides additional technical details on the sample, the applied imaging and analysis steps and methods. Structural imaging Trained radiographers placed all participants
Supplementary methods This section provides additional technical details on the sample, the applied imaging and analysis steps and methods. Structural imaging Trained radiographers placed all participants
A Design Toolbox to Generate Complex Phantoms for the Evaluation of Medical Image Processing Algorithms
 A Design Toolbox to Generate Complex Phantoms for the Evaluation of Medical Image Processing Algorithms Omar Hamo, Georg Nelles, Gudrun Wagenknecht Central Institute for Electronics, Research Center Juelich,
A Design Toolbox to Generate Complex Phantoms for the Evaluation of Medical Image Processing Algorithms Omar Hamo, Georg Nelles, Gudrun Wagenknecht Central Institute for Electronics, Research Center Juelich,
FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING. Francesca Pizzorni Ferrarese
 FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING Francesca Pizzorni Ferrarese Pipeline overview WM and GM Segmentation Registration Data reconstruction Tractography
FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING Francesca Pizzorni Ferrarese Pipeline overview WM and GM Segmentation Registration Data reconstruction Tractography
Diffusion Imaging Visualization
 Diffusion Imaging Visualization Thomas Schultz URL: http://cg.cs.uni-bonn.de/schultz/ E-Mail: schultz@cs.uni-bonn.de 1 Outline Introduction to Diffusion Imaging Basic techniques Glyph-based Visualization
Diffusion Imaging Visualization Thomas Schultz URL: http://cg.cs.uni-bonn.de/schultz/ E-Mail: schultz@cs.uni-bonn.de 1 Outline Introduction to Diffusion Imaging Basic techniques Glyph-based Visualization
Structural Segmentation
 Structural Segmentation FAST tissue-type segmentation FIRST sub-cortical structure segmentation FSL-VBM voxelwise grey-matter density analysis SIENA atrophy analysis FAST FMRIB s Automated Segmentation
Structural Segmentation FAST tissue-type segmentation FIRST sub-cortical structure segmentation FSL-VBM voxelwise grey-matter density analysis SIENA atrophy analysis FAST FMRIB s Automated Segmentation
Where are we now? Structural MRI processing and analysis
 Where are we now? Structural MRI processing and analysis Pierre-Louis Bazin bazin@cbs.mpg.de Leipzig, Germany Structural MRI processing: why bother? Just use the standards? SPM FreeSurfer FSL However:
Where are we now? Structural MRI processing and analysis Pierre-Louis Bazin bazin@cbs.mpg.de Leipzig, Germany Structural MRI processing: why bother? Just use the standards? SPM FreeSurfer FSL However:
SPM8 for Basic and Clinical Investigators. Preprocessing. fmri Preprocessing
 SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
Automatic Registration-Based Segmentation for Neonatal Brains Using ANTs and Atropos
 Automatic Registration-Based Segmentation for Neonatal Brains Using ANTs and Atropos Jue Wu and Brian Avants Penn Image Computing and Science Lab, University of Pennsylvania, Philadelphia, USA Abstract.
Automatic Registration-Based Segmentation for Neonatal Brains Using ANTs and Atropos Jue Wu and Brian Avants Penn Image Computing and Science Lab, University of Pennsylvania, Philadelphia, USA Abstract.
Structural Segmentation
 Structural Segmentation FAST tissue-type segmentation FIRST sub-cortical structure segmentation FSL-VBM voxelwise grey-matter density analysis SIENA atrophy analysis FAST FMRIB s Automated Segmentation
Structural Segmentation FAST tissue-type segmentation FIRST sub-cortical structure segmentation FSL-VBM voxelwise grey-matter density analysis SIENA atrophy analysis FAST FMRIB s Automated Segmentation
Fiber Selection from Diffusion Tensor Data based on Boolean Operators
 Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhof 1, G. Greiner 2, M. Buchfelder 3, C. Nimsky 4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhof 1, G. Greiner 2, M. Buchfelder 3, C. Nimsky 4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
A Genus Bound for Digital Image Boundaries
 A Genus Bound for Digital Image Boundaries Lowell Abrams and Donniell E. Fishkind March 9, 2005 Abstract Shattuck and Leahy [4] conjectured and Abrams, Fishkind, and Priebe [1],[2] proved that the boundary
A Genus Bound for Digital Image Boundaries Lowell Abrams and Donniell E. Fishkind March 9, 2005 Abstract Shattuck and Leahy [4] conjectured and Abrams, Fishkind, and Priebe [1],[2] proved that the boundary
Structural MRI analysis
 Structural MRI analysis volumetry and voxel-based morphometry cortical thickness measurements structural covariance network mapping Boris Bernhardt, PhD Department of Social Neuroscience, MPI-CBS bernhardt@cbs.mpg.de
Structural MRI analysis volumetry and voxel-based morphometry cortical thickness measurements structural covariance network mapping Boris Bernhardt, PhD Department of Social Neuroscience, MPI-CBS bernhardt@cbs.mpg.de
Correction of Partial Volume Effects in Arterial Spin Labeling MRI
 Correction of Partial Volume Effects in Arterial Spin Labeling MRI By: Tracy Ssali Supervisors: Dr. Keith St. Lawrence and Udunna Anazodo Medical Biophysics 3970Z Six Week Project April 13 th 2012 Introduction
Correction of Partial Volume Effects in Arterial Spin Labeling MRI By: Tracy Ssali Supervisors: Dr. Keith St. Lawrence and Udunna Anazodo Medical Biophysics 3970Z Six Week Project April 13 th 2012 Introduction
Introduction to fmri. Pre-processing
 Introduction to fmri Pre-processing Tibor Auer Department of Psychology Research Fellow in MRI Data Types Anatomical data: T 1 -weighted, 3D, 1/subject or session - (ME)MPRAGE/FLASH sequence, undistorted
Introduction to fmri Pre-processing Tibor Auer Department of Psychology Research Fellow in MRI Data Types Anatomical data: T 1 -weighted, 3D, 1/subject or session - (ME)MPRAGE/FLASH sequence, undistorted
Brain Surface Conformal Parameterization with Algebraic Functions
 Brain Surface Conformal Parameterization with Algebraic Functions Yalin Wang 1,2, Xianfeng Gu 3, Tony F. Chan 1, Paul M. Thompson 2, and Shing-Tung Yau 4 1 Mathematics Department, UCLA, Los Angeles, CA
Brain Surface Conformal Parameterization with Algebraic Functions Yalin Wang 1,2, Xianfeng Gu 3, Tony F. Chan 1, Paul M. Thompson 2, and Shing-Tung Yau 4 1 Mathematics Department, UCLA, Los Angeles, CA
NeuroImage 49 (2010) Contents lists available at ScienceDirect. NeuroImage. journal homepage:
 NeuroImage 49 (2010) 2479 2493 Contents lists available at ScienceDirect NeuroImage journal homepage: www.elsevier.com/locate/ynimg Comparison of landmark-based and automatic methods for cortical surface
NeuroImage 49 (2010) 2479 2493 Contents lists available at ScienceDirect NeuroImage journal homepage: www.elsevier.com/locate/ynimg Comparison of landmark-based and automatic methods for cortical surface
A Method for Filling Holes in Objects of Medical Images Using Region Labeling and Run Length Encoding Schemes
 110 Image Processing (NCIMP 2010) Image Processing (NCIMP 2010) Editor: K. Somasundaram Allied Publishers A Method for Filling Holes in Objects of Medical Images Using Region Labeling and Run Length Encoding
110 Image Processing (NCIMP 2010) Image Processing (NCIMP 2010) Editor: K. Somasundaram Allied Publishers A Method for Filling Holes in Objects of Medical Images Using Region Labeling and Run Length Encoding
NeuroQLab A Software Assistant for Neurosurgical Planning and Quantitative Image Analysis
 NeuroQLab A Software Assistant for Neurosurgical Planning and Quantitative Image Analysis Florian Weiler 1, Jan Rexilius 2, Jan Klein 1, Horst K. Hahn 1 1 Fraunhofer MEVIS, Universitätsallee 29, 28359
NeuroQLab A Software Assistant for Neurosurgical Planning and Quantitative Image Analysis Florian Weiler 1, Jan Rexilius 2, Jan Klein 1, Horst K. Hahn 1 1 Fraunhofer MEVIS, Universitätsallee 29, 28359
NIH Public Access Author Manuscript Proc Soc Photo Opt Instrum Eng. Author manuscript; available in PMC 2014 October 07.
 NIH Public Access Author Manuscript Published in final edited form as: Proc Soc Photo Opt Instrum Eng. 2014 March 21; 9034: 903442. doi:10.1117/12.2042915. MRI Brain Tumor Segmentation and Necrosis Detection
NIH Public Access Author Manuscript Published in final edited form as: Proc Soc Photo Opt Instrum Eng. 2014 March 21; 9034: 903442. doi:10.1117/12.2042915. MRI Brain Tumor Segmentation and Necrosis Detection
NIH Public Access Author Manuscript Proc Soc Photo Opt Instrum Eng. Author manuscript; available in PMC 2011 September 7.
 NIH Public Access Author Manuscript Published in final edited form as: Proc Soc Photo Opt Instrum Eng. 2011 March ; 7962: 796225-1 796225-7. doi:10.1117/12.878405. Automatic Skull-stripping of Rat MRI/DTI
NIH Public Access Author Manuscript Published in final edited form as: Proc Soc Photo Opt Instrum Eng. 2011 March ; 7962: 796225-1 796225-7. doi:10.1117/12.878405. Automatic Skull-stripping of Rat MRI/DTI
Functional MRI in Clinical Research and Practice Preprocessing
 Functional MRI in Clinical Research and Practice Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization
Functional MRI in Clinical Research and Practice Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization
FSL Pre-Processing Pipeline
 The Art and Pitfalls of fmri Preprocessing FSL Pre-Processing Pipeline Mark Jenkinson FMRIB Centre, University of Oxford FSL Pre-Processing Pipeline Standard pre-processing: Task fmri Resting-state fmri
The Art and Pitfalls of fmri Preprocessing FSL Pre-Processing Pipeline Mark Jenkinson FMRIB Centre, University of Oxford FSL Pre-Processing Pipeline Standard pre-processing: Task fmri Resting-state fmri
Automated Graph-Based Analysis and Correction of Cortical Volume Topology
 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 20, NO. 11, NOVEMBER 2001 1167 Automated Graph-Based Analysis and Correction of Cortical Volume Topology David W. Shattuck* and Richard M. Leahy Abstract The
IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 20, NO. 11, NOVEMBER 2001 1167 Automated Graph-Based Analysis and Correction of Cortical Volume Topology David W. Shattuck* and Richard M. Leahy Abstract The
Measuring longitudinal brain changes in humans and small animal models. Christos Davatzikos
 Measuring longitudinal brain changes in humans and small animal models Christos Davatzikos Section of Biomedical Image Analysis University of Pennsylvania (Radiology) http://www.rad.upenn.edu/sbia Computational
Measuring longitudinal brain changes in humans and small animal models Christos Davatzikos Section of Biomedical Image Analysis University of Pennsylvania (Radiology) http://www.rad.upenn.edu/sbia Computational
MITK-DI. A new Diffusion Imaging Component for MITK. Klaus Fritzsche, Hans-Peter Meinzer
 MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
City, University of London Institutional Repository
 City Research Online City, University of London Institutional Repository Citation: Doan, H., Slabaugh, G.G., Unal, G.B. & Fang, T. (2006). Semi-Automatic 3-D Segmentation of Anatomical Structures of Brain
City Research Online City, University of London Institutional Repository Citation: Doan, H., Slabaugh, G.G., Unal, G.B. & Fang, T. (2006). Semi-Automatic 3-D Segmentation of Anatomical Structures of Brain
Basic fmri Design and Analysis. Preprocessing
 Basic fmri Design and Analysis Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial filtering
Basic fmri Design and Analysis Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial filtering
Diffusion Imaging Models 1: from DTI to HARDI models
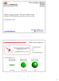 Diffusion Imaging Models 1: from DTI to HARDI models Flavio Dell Acqua, PhD. www.natbrainlab.com flavio.dellacqua@kcl.ac.uk @flaviodellacqua Diffusion Tensor Imaging (DTI) z λ 1 λ 2 The profile of the
Diffusion Imaging Models 1: from DTI to HARDI models Flavio Dell Acqua, PhD. www.natbrainlab.com flavio.dellacqua@kcl.ac.uk @flaviodellacqua Diffusion Tensor Imaging (DTI) z λ 1 λ 2 The profile of the
Brain Warping Via Landmark Points and Curves with a Level Set Representation
 Brain Warping Via Landmark Points and Curves with a Level Set Representation Andrew Y. Wang, Alex D. Leow, 2 Hillary D. Protas, Arthur W. Toga, Paul M. Thompson UCLA Laboratory of Neuro Imaging, Los Angeles,
Brain Warping Via Landmark Points and Curves with a Level Set Representation Andrew Y. Wang, Alex D. Leow, 2 Hillary D. Protas, Arthur W. Toga, Paul M. Thompson UCLA Laboratory of Neuro Imaging, Los Angeles,
Math in image processing
 Math in image processing Math in image processing Nyquist theorem Math in image processing Discrete Fourier Transformation Math in image processing Image enhancement: scaling Math in image processing Image
Math in image processing Math in image processing Nyquist theorem Math in image processing Discrete Fourier Transformation Math in image processing Image enhancement: scaling Math in image processing Image
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit
 Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
SPM8 for Basic and Clinical Investigators. Preprocessing
 SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
IMA Preprint Series # 2078
 A GEOMETRIC METHOD FOR AUTOMATIC EXTRACTION OF SULCAL FUNDI By C.-Y. Kao M. Hofer G. Sapiro J. Stern and D. Rottenberg IMA Preprint Series # 2078 ( November 2005 ) INSTITUTE FOR MATHEMATICS AND ITS APPLICATIONS
A GEOMETRIC METHOD FOR AUTOMATIC EXTRACTION OF SULCAL FUNDI By C.-Y. Kao M. Hofer G. Sapiro J. Stern and D. Rottenberg IMA Preprint Series # 2078 ( November 2005 ) INSTITUTE FOR MATHEMATICS AND ITS APPLICATIONS
Saturn User Manual. Rubén Cárdenes. 29th January 2010 Image Processing Laboratory, University of Valladolid. Abstract
 Saturn User Manual Rubén Cárdenes 29th January 2010 Image Processing Laboratory, University of Valladolid Abstract Saturn is a software package for DTI processing and visualization, provided with a graphic
Saturn User Manual Rubén Cárdenes 29th January 2010 Image Processing Laboratory, University of Valladolid Abstract Saturn is a software package for DTI processing and visualization, provided with a graphic
Image Registration + Other Stuff
 Image Registration + Other Stuff John Ashburner Pre-processing Overview fmri time-series Motion Correct Anatomical MRI Coregister m11 m 21 m 31 m12 m13 m14 m 22 m 23 m 24 m 32 m 33 m 34 1 Template Estimate
Image Registration + Other Stuff John Ashburner Pre-processing Overview fmri time-series Motion Correct Anatomical MRI Coregister m11 m 21 m 31 m12 m13 m14 m 22 m 23 m 24 m 32 m 33 m 34 1 Template Estimate
A Method for Registering Diffusion Weighted Magnetic Resonance Images
 A Method for Registering Diffusion Weighted Magnetic Resonance Images Xiaodong Tao and James V. Miller GE Research, Niskayuna, New York, USA Abstract. Diffusion weighted magnetic resonance (DWMR or DW)
A Method for Registering Diffusion Weighted Magnetic Resonance Images Xiaodong Tao and James V. Miller GE Research, Niskayuna, New York, USA Abstract. Diffusion weighted magnetic resonance (DWMR or DW)
Complex Fiber Visualization
 Annales Mathematicae et Informaticae 34 (2007) pp. 103 109 http://www.ektf.hu/tanszek/matematika/ami Complex Fiber Visualization Henrietta Tomán a, Róbert Tornai b, Marianna Zichar c a Department of Computer
Annales Mathematicae et Informaticae 34 (2007) pp. 103 109 http://www.ektf.hu/tanszek/matematika/ami Complex Fiber Visualization Henrietta Tomán a, Róbert Tornai b, Marianna Zichar c a Department of Computer
EPI Data Are Acquired Serially. EPI Data Are Acquired Serially 10/23/2011. Functional Connectivity Preprocessing. fmri Preprocessing
 Functional Connectivity Preprocessing Geometric distortion Head motion Geometric distortion Head motion EPI Data Are Acquired Serially EPI Data Are Acquired Serially descending 1 EPI Data Are Acquired
Functional Connectivity Preprocessing Geometric distortion Head motion Geometric distortion Head motion EPI Data Are Acquired Serially EPI Data Are Acquired Serially descending 1 EPI Data Are Acquired
1 Introduction Motivation and Aims Functional Imaging Computational Neuroanatomy... 12
 Contents 1 Introduction 10 1.1 Motivation and Aims....... 10 1.1.1 Functional Imaging.... 10 1.1.2 Computational Neuroanatomy... 12 1.2 Overview of Chapters... 14 2 Rigid Body Registration 18 2.1 Introduction.....
Contents 1 Introduction 10 1.1 Motivation and Aims....... 10 1.1.1 Functional Imaging.... 10 1.1.2 Computational Neuroanatomy... 12 1.2 Overview of Chapters... 14 2 Rigid Body Registration 18 2.1 Introduction.....
Performance Evaluation of the TINA Medical Image Segmentation Algorithm on Brainweb Simulated Images
 Tina Memo No. 2008-003 Internal Memo Performance Evaluation of the TINA Medical Image Segmentation Algorithm on Brainweb Simulated Images P. A. Bromiley Last updated 20 / 12 / 2007 Imaging Science and
Tina Memo No. 2008-003 Internal Memo Performance Evaluation of the TINA Medical Image Segmentation Algorithm on Brainweb Simulated Images P. A. Bromiley Last updated 20 / 12 / 2007 Imaging Science and
Norbert Schuff VA Medical Center and UCSF
 Norbert Schuff Medical Center and UCSF Norbert.schuff@ucsf.edu Medical Imaging Informatics N.Schuff Course # 170.03 Slide 1/67 Objective Learn the principle segmentation techniques Understand the role
Norbert Schuff Medical Center and UCSF Norbert.schuff@ucsf.edu Medical Imaging Informatics N.Schuff Course # 170.03 Slide 1/67 Objective Learn the principle segmentation techniques Understand the role
Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow
 Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow Abstract. Finding meaningful 1-1 correspondences between hippocampal (HP) surfaces is an important but difficult
Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow Abstract. Finding meaningful 1-1 correspondences between hippocampal (HP) surfaces is an important but difficult
MITK-DI. A new Diffusion Imaging Component for MITK. Klaus Fritzsche, Hans-Peter Meinzer
 MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
EMSegmenter Tutorial (Advanced Mode)
 EMSegmenter Tutorial (Advanced Mode) Dominique Belhachemi Section of Biomedical Image Analysis Department of Radiology University of Pennsylvania 1/65 Overview The goal of this tutorial is to apply the
EMSegmenter Tutorial (Advanced Mode) Dominique Belhachemi Section of Biomedical Image Analysis Department of Radiology University of Pennsylvania 1/65 Overview The goal of this tutorial is to apply the
Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity
 Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity Pahal Dalal 1, Feng Shi 2, Dinggang Shen 2, and Song Wang 1 1 Department of Computer Science and Engineering, University of South
Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity Pahal Dalal 1, Feng Shi 2, Dinggang Shen 2, and Song Wang 1 1 Department of Computer Science and Engineering, University of South
Open Topology: A Toolkit for Brain Isosurface Correction
 Open Topology: A Toolkit for Brain Isosurface Correction Sylvain Jaume 1, Patrice Rondao 2, and Benoît Macq 2 1 National Institute of Research in Computer Science and Control, INRIA, France, sylvain@mit.edu,
Open Topology: A Toolkit for Brain Isosurface Correction Sylvain Jaume 1, Patrice Rondao 2, and Benoît Macq 2 1 National Institute of Research in Computer Science and Control, INRIA, France, sylvain@mit.edu,
Reconstruction of Fiber Trajectories via Population-Based Estimation of Local Orientations
 IDEA Reconstruction of Fiber Trajectories via Population-Based Estimation of Local Orientations Pew-Thian Yap, John H. Gilmore, Weili Lin, Dinggang Shen Email: ptyap@med.unc.edu 2011-09-21 Poster: P2-46-
IDEA Reconstruction of Fiber Trajectories via Population-Based Estimation of Local Orientations Pew-Thian Yap, John H. Gilmore, Weili Lin, Dinggang Shen Email: ptyap@med.unc.edu 2011-09-21 Poster: P2-46-
syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets
 syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets Julien Gervais; Lisa Chuah Siemens Healthcare, Magnetic
syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets Julien Gervais; Lisa Chuah Siemens Healthcare, Magnetic
NIH Public Access Author Manuscript Med Image Comput Comput Assist Interv. Author manuscript; available in PMC 2009 December 4.
 NIH Public Access Author Manuscript Med Image Comput Comput Assist Interv. Author manuscript; available in PMC 2009 December 4. Published in final edited form as: Med Image Comput Comput Assist Interv.
NIH Public Access Author Manuscript Med Image Comput Comput Assist Interv. Author manuscript; available in PMC 2009 December 4. Published in final edited form as: Med Image Comput Comput Assist Interv.
Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00
 Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00 Marcel Prastawa 1 and Guido Gerig 1 Abstract July 17, 2008 1 Scientific Computing and Imaging
Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00 Marcel Prastawa 1 and Guido Gerig 1 Abstract July 17, 2008 1 Scientific Computing and Imaging
ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION
 ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION Abstract: MIP Project Report Spring 2013 Gaurav Mittal 201232644 This is a detailed report about the course project, which was to implement
ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION Abstract: MIP Project Report Spring 2013 Gaurav Mittal 201232644 This is a detailed report about the course project, which was to implement
AN EFFICIENT SKULL STRIPPING ALGORITHM USING CONNECTED REGIONS AND MORPHOLOGICAL OPERATION
 AN EFFICIENT SKULL STRIPPING ALGORITHM USING CONNECTED REGIONS AND MORPHOLOGICAL OPERATION Shijin Kumar P. S. 1 and Dharun V. S. 2 1 Department of Electronics and Communication Engineering, Noorul Islam
AN EFFICIENT SKULL STRIPPING ALGORITHM USING CONNECTED REGIONS AND MORPHOLOGICAL OPERATION Shijin Kumar P. S. 1 and Dharun V. S. 2 1 Department of Electronics and Communication Engineering, Noorul Islam
Multimodal Imaging Brain Connectivity Analysis (MIBCA)
 Multimodal Imaging Brain Connectivity Analysis (MIBCA) Andre Santos Ribeiro, Luis Miguel Lacerda, Hugo Ferreira April 23, 2015 Abstract In recent years, connectivity studies using neuroimaging data have
Multimodal Imaging Brain Connectivity Analysis (MIBCA) Andre Santos Ribeiro, Luis Miguel Lacerda, Hugo Ferreira April 23, 2015 Abstract In recent years, connectivity studies using neuroimaging data have
ASAP_2.0 (Automatic Software for ASL Processing) USER S MANUAL
 ASAP_2.0 (Automatic Software for ASL Processing) USER S MANUAL ASAP was developed as part of the COST Action "Arterial Spin Labelling Initiative in Dementia (AID)" by: Department of Neuroimaging, Institute
ASAP_2.0 (Automatic Software for ASL Processing) USER S MANUAL ASAP was developed as part of the COST Action "Arterial Spin Labelling Initiative in Dementia (AID)" by: Department of Neuroimaging, Institute
Simultaneous Cortical Surface Labeling and Sulcal Curve Extraction
 Simultaneous Cortical Surface Labeling and Sulcal Curve Extraction Zhen Yang a, Aaron Carass a, Chen Chen a, Jerry L Prince a,b a Electrical and Computer Engineering, b Biomedical Engineering, Johns Hopkins
Simultaneous Cortical Surface Labeling and Sulcal Curve Extraction Zhen Yang a, Aaron Carass a, Chen Chen a, Jerry L Prince a,b a Electrical and Computer Engineering, b Biomedical Engineering, Johns Hopkins
EMSegment Tutorial. How to Define and Fine-Tune Automatic Brain Compartment Segmentation and the Detection of White Matter Hyperintensities
 EMSegment Tutorial How to Define and Fine-Tune Automatic Brain Compartment Segmentation and the Detection of White Matter Hyperintensities This documentation serves as a tutorial to learn to customize
EMSegment Tutorial How to Define and Fine-Tune Automatic Brain Compartment Segmentation and the Detection of White Matter Hyperintensities This documentation serves as a tutorial to learn to customize
Learning-based Neuroimage Registration
 Learning-based Neuroimage Registration Leonid Teverovskiy and Yanxi Liu 1 October 2004 CMU-CALD-04-108, CMU-RI-TR-04-59 School of Computer Science Carnegie Mellon University Pittsburgh, PA 15213 Abstract
Learning-based Neuroimage Registration Leonid Teverovskiy and Yanxi Liu 1 October 2004 CMU-CALD-04-108, CMU-RI-TR-04-59 School of Computer Science Carnegie Mellon University Pittsburgh, PA 15213 Abstract
A Novel Contrast for DTI Visualization for Thalamus Delineation
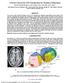 A Novel Contrast for DTI Visualization for Thalamus Delineation Xian Fan a, Meredith Thompson a,b, John A. Bogovic a, Pierre-Louis Bazin c, Jerry L. Prince a,c a Johns Hopkins University, Baltimore, MD,
A Novel Contrast for DTI Visualization for Thalamus Delineation Xian Fan a, Meredith Thompson a,b, John A. Bogovic a, Pierre-Louis Bazin c, Jerry L. Prince a,c a Johns Hopkins University, Baltimore, MD,
Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE)
 Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE) Mahnaz Maddah, Kelly H. Zou, William M. Wells, Ron Kikinis, and Simon K. Warfield
Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE) Mahnaz Maddah, Kelly H. Zou, William M. Wells, Ron Kikinis, and Simon K. Warfield
Network connectivity via inference over curvature-regularizing line graphs
 Network connectivity via inference over curvature-regularizing line graphs Asian Conference on Computer Vision Maxwell D. Collins 1,2, Vikas Singh 2,1, Andrew L. Alexander 3 1 Department of Computer Sciences
Network connectivity via inference over curvature-regularizing line graphs Asian Conference on Computer Vision Maxwell D. Collins 1,2, Vikas Singh 2,1, Andrew L. Alexander 3 1 Department of Computer Sciences
Modified Normal Vector Voting Estimation in neuroimage
 Modified Normal Vector Voting Estimation in neuroimage Neuroimage Processing (339.632) Instructor: Moo. K. Chung Seung-goo KIM Dec 15, 2009 Abstract The normal vector is one of the metrics that are sensitive
Modified Normal Vector Voting Estimation in neuroimage Neuroimage Processing (339.632) Instructor: Moo. K. Chung Seung-goo KIM Dec 15, 2009 Abstract The normal vector is one of the metrics that are sensitive
Optimization of Brain Conformal Mapping with Landmarks
 Optimization of Brain Conformal Mapping with Landmarks Yalin Wang 1,LokMingLui 1,TonyF.Chan 1, and Paul M. Thompson 2 Mathematics Department, UCLA, Los Angeles, CA 90095, USA Lab. of Neuro Imaging, UCLA
Optimization of Brain Conformal Mapping with Landmarks Yalin Wang 1,LokMingLui 1,TonyF.Chan 1, and Paul M. Thompson 2 Mathematics Department, UCLA, Los Angeles, CA 90095, USA Lab. of Neuro Imaging, UCLA
Assessing Accuracy Factors in Deformable 2D/3D Medical Image Registration Using a Statistical Pelvis Model
 Assessing Accuracy Factors in Deformable 2D/3D Medical Image Registration Using a Statistical Pelvis Model Jianhua Yao National Institute of Health Bethesda, MD USA jyao@cc.nih.gov Russell Taylor The Johns
Assessing Accuracy Factors in Deformable 2D/3D Medical Image Registration Using a Statistical Pelvis Model Jianhua Yao National Institute of Health Bethesda, MD USA jyao@cc.nih.gov Russell Taylor The Johns
Advanced Visual Medicine: Techniques for Visual Exploration & Analysis
 Advanced Visual Medicine: Techniques for Visual Exploration & Analysis Interactive Visualization of Multimodal Volume Data for Neurosurgical Planning Felix Ritter, MeVis Research Bremen Multimodal Neurosurgical
Advanced Visual Medicine: Techniques for Visual Exploration & Analysis Interactive Visualization of Multimodal Volume Data for Neurosurgical Planning Felix Ritter, MeVis Research Bremen Multimodal Neurosurgical
Brain Portion Peeling from T2 Axial MRI Head Scans using Clustering and Morphological Operation
 159 Brain Portion Peeling from T2 Axial MRI Head Scans using Clustering and Morphological Operation K. Somasundaram Image Processing Lab Dept. of Computer Science and Applications Gandhigram Rural Institute
159 Brain Portion Peeling from T2 Axial MRI Head Scans using Clustering and Morphological Operation K. Somasundaram Image Processing Lab Dept. of Computer Science and Applications Gandhigram Rural Institute
Surface-based Analysis: Inter-subject Registration and Smoothing
 Surface-based Analysis: Inter-subject Registration and Smoothing Outline Exploratory Spatial Analysis Coordinate Systems 3D (Volumetric) 2D (Surface-based) Inter-subject registration Volume-based Surface-based
Surface-based Analysis: Inter-subject Registration and Smoothing Outline Exploratory Spatial Analysis Coordinate Systems 3D (Volumetric) 2D (Surface-based) Inter-subject registration Volume-based Surface-based
SURFACE RECONSTRUCTION OF EX-VIVO HUMAN V1 THROUGH IDENTIFICATION OF THE STRIA OF GENNARI USING MRI AT 7T
 SURFACE RECONSTRUCTION OF EX-VIVO HUMAN V1 THROUGH IDENTIFICATION OF THE STRIA OF GENNARI USING MRI AT 7T Oliver P. Hinds 1, Jonathan R. Polimeni 2, Megan L. Blackwell 3, Christopher J. Wiggins 3, Graham
SURFACE RECONSTRUCTION OF EX-VIVO HUMAN V1 THROUGH IDENTIFICATION OF THE STRIA OF GENNARI USING MRI AT 7T Oliver P. Hinds 1, Jonathan R. Polimeni 2, Megan L. Blackwell 3, Christopher J. Wiggins 3, Graham
Histograms. h(r k ) = n k. p(r k )= n k /NM. Histogram: number of times intensity level rk appears in the image
 Histograms h(r k ) = n k Histogram: number of times intensity level rk appears in the image p(r k )= n k /NM normalized histogram also a probability of occurence 1 Histogram of Image Intensities Create
Histograms h(r k ) = n k Histogram: number of times intensity level rk appears in the image p(r k )= n k /NM normalized histogram also a probability of occurence 1 Histogram of Image Intensities Create
Case Study: Interacting with Cortical Flat Maps of the Human Brain
 Case Study: Interacting with Cortical Flat Maps of the Human Brain Monica K. Hurdal Department of Mathematics Florida State University and International Neuroimaging Consortium VA Medical Center University
Case Study: Interacting with Cortical Flat Maps of the Human Brain Monica K. Hurdal Department of Mathematics Florida State University and International Neuroimaging Consortium VA Medical Center University
Automatic Generation of Training Data for Brain Tissue Classification from MRI
 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. COCOSCO, Alex P. ZIJDENBOS, and Alan C. EVANS http://www.bic.mni.mcgill.ca/users/crisco/ McConnell Brain Imaging
Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. COCOSCO, Alex P. ZIJDENBOS, and Alan C. EVANS http://www.bic.mni.mcgill.ca/users/crisco/ McConnell Brain Imaging
