Faster STORM using compressed sensing. Supplementary Software. Nature Methods. Nature Methods: doi: /nmeth.1978
|
|
|
- Janis James
- 6 years ago
- Views:
Transcription
1 Nature Methods Faster STORM using compressed sensing Lei Zhu, Wei Zhang, Daniel Elnatan & Bo Huang Supplementary File Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure 3 Supplementary Figure 4 Supplementary Figure 5 Supplementary Figure 6 Supplementary Figure 7 Supplementary Figure 8 Supplementary Figure 9 Supplementary Note Supplementary Video 1 Supplementary Software Title Analyzing single molecule images by compressed sensing. Effect of ε on compressed sensing. Analysis of the limitation of molecular distance for perfect recovery using compressed sensing. Comparing compressed sensing to DAOSTORM in the high photon count case corresponding to the Alexa Fluor 647 dye. Localization precision of the compressed sensing algorithm as a function of molecule density. Comparing compressed sensing and single-molecule fitting in analyzing simulated data corresponding to the fluorescent protein meos2 images. Compressed sensing analysis in a low signal-to-noise case. Comparing compressed sensing and single-molecule fitting in analyzing simulated low signal-to-noise images. Minimum number of frames to achieve a given overall image resolution for a continuous 2D sample based on the simulation results. Algorithm implementation, evaluation, and analysis of the fundamental limits in compressed sensing. STORM movie of microtubules in a living Drosophila S2 cell stably expressing meos2-fused tubulin, with a time resolution of 3 sec. The movie is reconstructed from 4349 camera frames (77 sec) and plays 11 times as fast as real time. Three snapshots from the movie are shown in Figure 2b. Scale bar: 1 µm. Matlab code for STORM data analysis using compressed sensing
2 Supplementary Notes 1. Implementation of the compressed sensing algorithm A major consideration of our implementation of the compressed sensing algorithm is the computation time. Therefore, we subdivided the original image into smaller patches, which not only reduces to total time to analyze the entire image, but also opens the possibility for parallel computing to further improve the computation speed. On the other hand, merging results from different patches becomes important. Our implementation took a simple approach that is facilitated by a small overlap between adjacent patches: Given: camera image B, super-resolution image x, patch size u-byv pixels, oversampling factor R, reduced chi-squared target ε, the imaging matrix A and the vector c for a small u-by-v image. Dividing the camera image, B, into a set of small u-by-v image patches, b, with two pixels shared by adjacent patches. For each small image, b, do: 1. Create an oversample grid, x, with a size of R (u + 4) by R (v + 4). The margin of extra 2 pixels on each edge of the patch accounts for the contribution from molecules outside. 2. Minimize c T x subject to x i and Ax b 2 ε sqrt(sum(b)) For each patch of the optimization result, x, set the outmost 3- pixel boundary to zero to avoid overlap (2 pixel extra marging + one pixel overlap between patches). Add all x together to form the result of the full image. The computation time of our compressed sensing algorithm depends on the number of molecules in an image frame. Analyzing a pixel 2 image typically takes about 3 seconds on our Intel Xeon X556 (2.8 GHz) workstation using a single computation thread. The slow computation speed is inherent for such a large optimization problem, and can also be partially attributed to our choice of MATLAB programming language instead of the faster C/C++. On the other hand, the fact that our algorithm performs independent optimizations on small, uniformly
3 sized image patches makes it easily adaptable to massive parallel computing, especially GPU computation 1,2, for much enhanced speed. 2. Evaluation of the compressed sensing method We first analyzed the density of molecules identifiable by different methods. In the high photon number simulation which corresponds to bright organic fluorophores (Figure 1b), compressed sensing is much more efficient in identifying densely located molecules. The number of identified molecules using the single-molecule fitting algorithm peaks at.58 per μm 2 when the original image has approximately 2 molecules per μm 2. On the contrary, compressed sensing can identify as many as 8.8 molecules per μm 2 out of 12.5 molecules per μm 2 in the original image. Next, we examined the localization precisions (Figure 1c). At low molecule density, the precision of compressed sensing is just slightly worse than that of single-molecule fitting, presumably due to the finite size of the grid and the unweighted least square constraints used. In simulations with only one molecule in the view field, both methods approach the theoretical limit (9.2 nm for fitting, 11.1 nm for compressed sensing, whereas the Cremer-Rao lower bound (CRLB) 2,3,4 is 8.8 nm; all numbers are FWHM). As expected, the localization precision starts to degrade with high molecule densities. At a density higher than 2 molecules per μm 2, compressed sensing gives slightly better precision than the fitting method. We also analyzed the same set of simulated data with the previous reported DAOSTORM method using the Python code provided by its authors 5. Supplementary Figure 4 compares the performance in molecule identification and localization precision. The number of molecules identified by DAOSTORM plateaus at 2.4 to 2.7 per µm 2 when the molecule density exceeds 4 per µm 2, At the same time, the localization error of DAOSTORM in this range of molecule density is about the same as the single-molecule fitting method. At relatively low molecule density (< 2 per µm 2 ), all three methods show similar localization precision. We note that our DAOSTORM results are consistent with those previously published 5 considering that our simulation uses a smaller average photon number per molecule (3, versus 8,333 (5, effective)), includes a higher background (7 photons per pixel versus 1), and considers the variation of photon numbers among molecules. Finally, for the overall temporal resolution, Figure 1d and Supplementary Figure 9 summarizes the minimum number of image frames to attain a given overall spatial resolution. At a resolution of 4 nm (the best allowed by our grid size) to 11 nm (close to what can be achieved by the faster Structured Illumination Microscopy 6 ), compressed sensing allows 1/6 to 1/15 of the number of camera frames to be used compared to the single-molecule fitting method. That means a factor of 6 to 15 of improvement in time resolution depending on the desired spatial resolution. For example, only 45 camera frames are needed to support a 1 nm spatial resolution, corresponding to.75 sec integration time if the camera operates at 6 frames / sec. In the two additional sets of simulations with photon statistics corresponding to fluorescent protein meos2 and a very low photon case (see Supplementary Figs. 6 and 8), we have also seen
4 the substantial improvement in molecule identification efficiency by compressed sensing. At 75 photons per molecule, compressed sensing identifies 6.7 molecules per µm 2 with a precision of 113 nm. At 2 photons per molecule, compressed sensing identifies 3.8 molecules per µm 2 with a precision of 126 nm. These values suggest an imaging speed that is approximately 75% and 4%, respectively, compared to the 3, photons per molecule case if one targets ~ 1 nm resolution. We must emphasize that the resolution calculation above is for a 2D continuous sample. In real biological samples where the molecules are clustered into one-dimensional (e.g. microtubule) or zero-dimensional (e.g. clathrin-coated pits) structures, the number of frames needed to achieve a given molecule density can be even smaller Fundamental limit in molecule density Compressed sensing is able to recover sparse or compressed signals using very few measurements from a linear system, without knowing in advance the support of the signal (i.e. where the signals are exactly zeros). An important concept in the compressed sensing theory is the restricted isometry principle (RIP) 8, which determines the accuracy and the stability of the signal recovery. A linear system A is said to obey RIP with a sparsity s (i.e., s non-zero elements in x) and an error tolerance s if one has (1 δ s ) x 2 2 Ax 2 2 (1 + δ s ) x 2 2 (1) for all s-sparse vector x, where s is the smallest value such that the inequation holds and s-sparse means that x has at most s non-zero elements. To recover s-sparse vectors, it is important to consider 2s, since the difference between two s-sparse vectors must be a 2s-sparse vector. If 2s is sufficiently smaller than 1 (practical implementations require 2s.4) 8, then any pair of s-sparse vectors will have a degree of distinguishability in the data space. The compressed sensing theory shows that s-sparse vectors can be exactly recovered via L1 norm minimization on the estimated signals 9. Our problem of localizing sparse molecules fits the application scenario of compressed sensing. The condition on the molecule sparsity for exact signal recovery, however, requires a complete RIP analysis. Although such an analysis is complicated in general, here we try to use a simplified case to provide insights into the limit of molecular density that compressed sensing can detect. Specifically, we investigate what is the smallest distance of two molecules (s = 2) for them to be perfectly detected using compressed sensing. For this purpose, we calculate 4 as shown in Supplementary Figure 3 for different minimum distances between molecules. The system PSF is assumed to be a 2D symmetric Gaussian function, and the molecule distances are plotted in unit of PSF standard deviation. The 4 values are below.4 for molecule distances larger than 2.5 (approximately the FWHM of the PSF), which indicates that the compressed sensing algorithm is able to achieve exact molecule localization for a molecule density up to approximately.16/ 2. Given that our observed PSF has = 143 nm, this limit corresponds to a molecular density of 7.8 µm -2, consistent with fact that in the high photon number case, our molecule identification starts to lose efficiency at a molecule density around 9 µm -2. It is also
5 consistent with the maximum density of recovered molecules (around 8.8 µm -2 ) in our simulations despite our overly simplified derivations. The conventional fitting-based algorithms may allow a similar minimum molecule distance for accurate detection if only a few molecules are present in the imaging field and the total number is known. Compressed sensing is able to automatically detect all the molecule locations without prior knowledge of their total number, as long as their mutual distance (therefore density) is not higher than the value suggested by the 4 plot. The advantage is more obvious when one image frame contains a large number of molecules. Finally, we would like to emphasize that a rigorous RIP analysis should calculate s of all s values for the given linear system A. The above description should be considered as only a rough estimation of the maximum density of detected molecules in the STORM imaging using compressed sensing, which will hopefully be replaced by future, more solid work. It is also an interesting topic to compare the compressed sensing approach and the statistical approach. The goal of STORM imaging in general is to determine the molecule locations with some prior knowledge. The available prior knowledge in this problem includes the sparsity of the true signals, the probability distribution of the detected photons and the detector PSF. The advantage of compressed sensing is efficient recovery of sparse signals even if the measured data are heavily corrupted by noise or errors. On the other hand, statistics-based approaches improve the imaging performance by accurate modeling of the probability distribution of the detected photons. These two categories of methods use different prior knowledge in the same application, with different pros and cons. In the study presented in this paper, we adopt a classic form of compressed sensing reconstruction, using very limited prior knowledge of the statistics. As shown in the general framework (Eq. 1), only an estimated noise standard deviation of the measured data is required in our algorithm. However, compressed sensing is compatible with statistics-based approaches. We can further improve our algorithm performance, especially in the case of high noise, by modeling the noise statistics in our optimization framework. This improved framework will achieve the theoretical limit as shown in Ref. 1. It will be one of our future research directions. References 1. Quan, T.W. et al., Ultra-fast, high-precision image analysis for localization-based super resolution microscopy, Opt. Express, 18, (21). 2. Smith, C. S., Joseph, N., Rieger B., & Lidke, K. A, Fast, single-molecule localization that achieves theoretically minimum uncertainty, Nat. Methods, 7, (21). 3. Mortensen, K. I., Churchman, L. S., Spudich, J. A., & Flyvbjerg, H., Optimized localization analysis for single-molecule tracking and super-resolution microscopy, Nat. Methods, 7, (21). 4. Abraham, A.V., Ram, S., Chao, J., Ward, E.S. & Ober, R.J., Quantitative study of single molecule location estimation techniques, Opt. Express, 17, (29).
6 5. Holden, S.J., Uphoff, S. & Kapanidis, A.N. DAOSTORM: an algorithm for highdensity super-resolution microscopy, Nat. Methods 6, (211). 6. Kner, P., Chhun, B. B., Griffis, E. R., Winoto, L., & Gustafsson, M. G. L., Superresolution video microscopy of live cells by structured illumination, Nat. Methods, 6, (29). 7. Jones, S. A., Shim, S.-H., He, J., & Zhuang, X., Fast, three-dimensional superresolution imaging of live cells, Nat. Methods, 8, (211). 8. Candès, E.J., The restricted isometry property and its implications for compressed sensing, Comptes Rendus Mathematique, 346, (28). 9. Candès, E.J., Romberg, J. & Tao, T., Stable signal recovery from incomplete andinaccurate measurements, Comm. Pure Appl. Math., (25). 1. Willett, R. M. & Raginsky, M., Performance bounds on compressed sensing with Poisson noise, Proceedings of the 29 IEEE international conference on Symposium on Information Theory, 1, (29).
7 Supplementary Figures 1 pixel 8x8 grids 32 x 32 pixels 256 x 256 grids 166 nm / pixel 21 nm / grid 4 subdivision 8 subdivision 16 subdivision Supplementary Figure 1. Analyzing single molecule images by compressed sensing. (Upper row) Scheme of dividing image pixels into oversampled grids. (Middle row) Comparing the analysis of the image in Figure 1a using different oversampling factors. (Lower row) Zoomed-in displays of the compressed sensing results for the boxed regions. Note that the difference in molecular identification efficiency is relatively small. The computation times for 4, 8, and 16 oversampling are 15 sec, 28 sec and 134 sec, respectively. Scale bars: 3 nm.
8 3 Identified molecules Supplementary Figure 2. Effect of ε on compressed sensing. Showing the number of molecules identified in the simulated image of Figure 1a (1 molecules in the simulation). Because ε corresponds to the reduced χ 2, an ε value of 1 corresponds to perfect modeling of the image if the noise estimation is exact. With an ε value smaller than 1, the image is over fitted which results in over-identification, whereas an ε value larger than 2 enforces the sparsity condition too strongly. In practice, we choose an ε value of 1.5 for all of our analyses (with an extra factor of 2 in analyzing experimental images from EMCCD to account for its extra noise).
9 1..8 δ d =2.5σ Molecule distance (in units of σ) Supplementary Figure 3. Analysis of the limitation in inter-molecular distances for perfect recovery using compressed sensing. Plotting the value of δ 4 as a function of the separation between two molecules in units of the standard deviation of the PSF, σ. δ 4 reaches a value of.4 (the threshold for perfect recovery, see Candès, E.J., Comptes Rendus Mathematique, 346, (28)) when the separation is larger than 2.5, or approximately the FWHM of the PSF.
10 Identified molecule density ( m -2 ) Single-molecule fitting Compressed sensing DAOSTORM Molecule density ( m -2 ) 6 Localization precision FWHM (nm) Single-molecule fitting Compressed sensing DAOSTORM Molecule density ( m -2 ) 4 2 Localization precision Stdev (nm) Supplementary Figure 4. Comparing compressed sensing to DAOSTORM in the high photon count case corresponding to the Alexa Fluor 647 dye (mean photon number 3, per molecule, background 7 photons per pixel). The same simulation data sets are analyzed by DAOSTORM using the Python code provided by Holden, Uphoff & Kapanidis, Nat. Methods 6, (211). The number of molecules identified and the localization precisions are compared to those of the fitting method and compressed sensing in the same way as in Figures 1b and 1c.
11 N = 1 Density ~ 1 m x or y deviation (nm) x or y deviation (nm) Density ~ 4 m -2 Density ~ 8 m x or y deviation (nm) x or y deviation (nm) Supplementary Figure 5. Localization precision of the compressed sensing algorithm as a function of molecule density. The analysis is done to simulated data in the high photon count case corresponding to the Alexa Fluor 647 dye (mean photon number 3, per molecule, background 7 photons per pixel). Each molecule identified by compressed sensing is matched to the closest true molecule position. The two histograms of the x offsets and y offsets are summed and displayed in each panel. These histograms are fitted with Gaussian functions whose widths are presented in Figure 1c and Supplementary Figure 4.
12 Identified molecule density ( m -2 ) Single-molecule fitting Compressed sensing Molecule density ( m -2 ) Localization precision FWHM (nm) Single-molecule fitting Compressed sensing Molecule density ( m -2 ) Localization precision Stdev (nm) Supplementary Figure 6. Comparing compressed sensing and single-molecule fitting in analyzing simulated data corresponding to the fluorescent protein meos2 images (mean photon number 75 per molecule, background 5 photons per pixel). The dashed line in the localization precision plot represents the CRLB for single-molecule localization (22 nm FWHM).
13 a 3 nm b c Supplementary Figure 7. Compressed sensing analysis in a low signal-to-noise case. The simulation is performed with 5 molecules with 2 photons per molecule and a background of 2 photons per pixel. Showing (a) the simulated camera image, (b) compressed sensing result, and (c) the comparison of the compressed sensing result (red crosses) with the molecule positions in the simulation. Scale bars: 3 nm. It is evident that the result from compressed sensing generally matches well with the simulation input, although closely located molecules are occasionally merged into one because the high noise can make them statistically indistinguishable.
14 Identified molecule density ( m -2 ) Single-molecule fitting Compressed sensing Molecule density ( m -2 ) 6 Localization precision FWHM (nm) Single-molecule fitting Compressed sensing Molecule density ( m -2 ) 4 2 Localization precision Stdev (nm) Supplementary Figure 8. Comparing compressed sensing and single-molecule fitting in analyzing simulated low signal-to-noise images (mean photon number 2 per molecule, background 1 photons per pixel). The dashed line in the localization precision plot represents the CRLB for single-molecule localization (4 nm FWHM).
15 1 Number of frames 1 1 Single-molecule fitting Compressed sensing DAOSTORM Image resolution, FWHM (nm) Supplementary Figure 9. Minimum number of frames to achieve a given overall image resolution for a continuous 2D sample based on the simulation results, displayed on a log-log plot with the image resolution set between 4 and 11 nm. The analysis is done to simulated data in the high photon count case corresponding to the Alexa Fluor 647 dye (mean photon number 3, per molecule, background 7 photons per pixel). The curve for the fitting method is calculated using a constant.58 μm -2 identified molecule density, whereas the curves for the compressed sensing and DAOSTORM are calculated using identified molecule densities that allow the corresponding localization precisions to match the desired image resolution.
16 Supplementary Video STORM movie of microtubules in a living Drosophila S2 cell stably expressing meos2-fused tubulin, with a time resolution of 3 sec. The movie is reconstructed from 4349 camera frames (77 sec) and plays 11 times as fast as real time. Three snapshots from the movie are shown in Figure 2b. Scale bar: 1 µm.
Swammerdam Institute for Life Sciences (Universiteit van Amsterdam), 1098 XH Amsterdam, The Netherland
 Sparse deconvolution of high-density super-resolution images (SPIDER) Siewert Hugelier 1, Johan J. de Rooi 2,4, Romain Bernex 1, Sam Duwé 3, Olivier Devos 1, Michel Sliwa 1, Peter Dedecker 3, Paul H. C.
Sparse deconvolution of high-density super-resolution images (SPIDER) Siewert Hugelier 1, Johan J. de Rooi 2,4, Romain Bernex 1, Sam Duwé 3, Olivier Devos 1, Michel Sliwa 1, Peter Dedecker 3, Paul H. C.
Supplementary Figure 1
 Supplementary Figure 1 Experimental unmodified 2D and astigmatic 3D PSFs. (a) The averaged experimental unmodified 2D PSF used in this study. PSFs at axial positions from -800 nm to 800 nm are shown. The
Supplementary Figure 1 Experimental unmodified 2D and astigmatic 3D PSFs. (a) The averaged experimental unmodified 2D PSF used in this study. PSFs at axial positions from -800 nm to 800 nm are shown. The
Gpufit: An open-source toolkit for GPU-accelerated curve fitting
 Gpufit: An open-source toolkit for GPU-accelerated curve fitting Adrian Przybylski, Björn Thiel, Jan Keller-Findeisen, Bernd Stock, and Mark Bates Supplementary Information Table of Contents Calculating
Gpufit: An open-source toolkit for GPU-accelerated curve fitting Adrian Przybylski, Björn Thiel, Jan Keller-Findeisen, Bernd Stock, and Mark Bates Supplementary Information Table of Contents Calculating
Effect of OTF attenuation on single-slice SR-SIM reconstruction
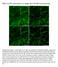 Effect of OTF attenuation on single-slice SR-SIM reconstruction Supplementary Figure 1: Actin filaments in U2OS cells, labelled with Phalloidin-Atto488, measured on a DeltaVision OMX, excited at 488 nm
Effect of OTF attenuation on single-slice SR-SIM reconstruction Supplementary Figure 1: Actin filaments in U2OS cells, labelled with Phalloidin-Atto488, measured on a DeltaVision OMX, excited at 488 nm
Supporting Information for Azimuthal Polarization Filtering for Accurate, Precise, and Robust Single-Molecule Localization Microscopy
 Nano Letters Supporting Information for Azimuthal Polarization Filtering for Accurate, Precise, and Robust Single-Molecule Localization Microscopy Matthew D. Lew, and W. E. Moerner *, Departments of Chemistry
Nano Letters Supporting Information for Azimuthal Polarization Filtering for Accurate, Precise, and Robust Single-Molecule Localization Microscopy Matthew D. Lew, and W. E. Moerner *, Departments of Chemistry
Assembly dynamics of microtubules at molecular resolution
 Supplementary Information with: Assembly dynamics of microtubules at molecular resolution Jacob W.J. Kerssemakers 1,2, E. Laura Munteanu 1, Liedewij Laan 1, Tim L. Noetzel 2, Marcel E. Janson 1,3, and
Supplementary Information with: Assembly dynamics of microtubules at molecular resolution Jacob W.J. Kerssemakers 1,2, E. Laura Munteanu 1, Liedewij Laan 1, Tim L. Noetzel 2, Marcel E. Janson 1,3, and
2, the coefficient of variation R 2, and properties of the photon counts traces
 Supplementary Figure 1 Quality control of FCS traces. (a) Typical trace that passes the quality control (QC) according to the parameters shown in f. The QC is based on thresholds applied to fitting parameters
Supplementary Figure 1 Quality control of FCS traces. (a) Typical trace that passes the quality control (QC) according to the parameters shown in f. The QC is based on thresholds applied to fitting parameters
Supporting Information. Super Resolution Imaging of Nanoparticles Cellular Uptake and Trafficking
 Supporting Information Super Resolution Imaging of Nanoparticles Cellular Uptake and Trafficking Daan van der Zwaag 1,2, Nane Vanparijs 3, Sjors Wijnands 1,4, Riet De Rycke 5, Bruno G. De Geest 2* and
Supporting Information Super Resolution Imaging of Nanoparticles Cellular Uptake and Trafficking Daan van der Zwaag 1,2, Nane Vanparijs 3, Sjors Wijnands 1,4, Riet De Rycke 5, Bruno G. De Geest 2* and
Deconvolution with curvelet-domain sparsity Vishal Kumar, EOS-UBC and Felix J. Herrmann, EOS-UBC
 Deconvolution with curvelet-domain sparsity Vishal Kumar, EOS-UBC and Felix J. Herrmann, EOS-UBC SUMMARY We use the recently introduced multiscale and multidirectional curvelet transform to exploit the
Deconvolution with curvelet-domain sparsity Vishal Kumar, EOS-UBC and Felix J. Herrmann, EOS-UBC SUMMARY We use the recently introduced multiscale and multidirectional curvelet transform to exploit the
Incoherent noise suppression with curvelet-domain sparsity Vishal Kumar, EOS-UBC and Felix J. Herrmann, EOS-UBC
 Incoherent noise suppression with curvelet-domain sparsity Vishal Kumar, EOS-UBC and Felix J. Herrmann, EOS-UBC SUMMARY The separation of signal and noise is a key issue in seismic data processing. By
Incoherent noise suppression with curvelet-domain sparsity Vishal Kumar, EOS-UBC and Felix J. Herrmann, EOS-UBC SUMMARY The separation of signal and noise is a key issue in seismic data processing. By
Chao, J., Ram, S., Ward, E. S., and Ober, R. J. Investigating the usage of point spread functions in point source and microsphere localization.
 Chao, J., Ram, S., Ward, E. S., and Ober, R. J. Investigating the usage of point spread functions in point source and microsphere localization. Proceedings of the SPIE, Three-Dimensional and Multidimensional
Chao, J., Ram, S., Ward, E. S., and Ober, R. J. Investigating the usage of point spread functions in point source and microsphere localization. Proceedings of the SPIE, Three-Dimensional and Multidimensional
Lecture 17 Sparse Convex Optimization
 Lecture 17 Sparse Convex Optimization Compressed sensing A short introduction to Compressed Sensing An imaging perspective 10 Mega Pixels Scene Image compression Picture Why do we compress images? Introduction
Lecture 17 Sparse Convex Optimization Compressed sensing A short introduction to Compressed Sensing An imaging perspective 10 Mega Pixels Scene Image compression Picture Why do we compress images? Introduction
Optical Sectioning. Bo Huang. Pharmaceutical Chemistry
 Optical Sectioning Bo Huang Pharmaceutical Chemistry Approaches to 3D imaging Physical cutting Technical difficulty Highest resolution Highest sensitivity Optical sectioning Simple sample prep. No physical
Optical Sectioning Bo Huang Pharmaceutical Chemistry Approaches to 3D imaging Physical cutting Technical difficulty Highest resolution Highest sensitivity Optical sectioning Simple sample prep. No physical
Live-cell 3D super-resolution imaging in thick biological samples
 Nature Methods Live-cell 3D super-resolution imaging in thick biological samples Francesca Cella Zanacchi, Zeno Lavagnino, Michela Perrone Donnorso, Alessio Del Bue, Lauria Furia, Mario Faretta & Alberto
Nature Methods Live-cell 3D super-resolution imaging in thick biological samples Francesca Cella Zanacchi, Zeno Lavagnino, Michela Perrone Donnorso, Alessio Del Bue, Lauria Furia, Mario Faretta & Alberto
1. ABOUT INSTALLATION COMPATIBILITY SURESIM WORKFLOWS a. Workflow b. Workflow SURESIM TUTORIAL...
 SuReSim manual 1. ABOUT... 2 2. INSTALLATION... 2 3. COMPATIBILITY... 2 4. SURESIM WORKFLOWS... 2 a. Workflow 1... 3 b. Workflow 2... 4 5. SURESIM TUTORIAL... 5 a. Import Data... 5 b. Parameter Selection...
SuReSim manual 1. ABOUT... 2 2. INSTALLATION... 2 3. COMPATIBILITY... 2 4. SURESIM WORKFLOWS... 2 a. Workflow 1... 3 b. Workflow 2... 4 5. SURESIM TUTORIAL... 5 a. Import Data... 5 b. Parameter Selection...
MetaMorph Super-Resolution Software. Localization-based super-resolution microscopy
 MetaMorph Super-Resolution Software Localization-based super-resolution microscopy KEY BENEFITS Lateral Resolution up to 20 nm & Axial Resolution up to 40nm Real-time and offline superresolution image
MetaMorph Super-Resolution Software Localization-based super-resolution microscopy KEY BENEFITS Lateral Resolution up to 20 nm & Axial Resolution up to 40nm Real-time and offline superresolution image
An Iteratively Reweighted Least Square Implementation for Face Recognition
 Vol. 6: 26-32 THE UNIVERSITY OF CENTRAL FLORIDA Published May 15, 2012 An Iteratively Reweighted Least Square Implementation for Face Recognition By: Jie Liang Faculty Mentor: Dr. Xin Li ABSTRACT: We propose,
Vol. 6: 26-32 THE UNIVERSITY OF CENTRAL FLORIDA Published May 15, 2012 An Iteratively Reweighted Least Square Implementation for Face Recognition By: Jie Liang Faculty Mentor: Dr. Xin Li ABSTRACT: We propose,
CHAPTER 9 INPAINTING USING SPARSE REPRESENTATION AND INVERSE DCT
 CHAPTER 9 INPAINTING USING SPARSE REPRESENTATION AND INVERSE DCT 9.1 Introduction In the previous chapters the inpainting was considered as an iterative algorithm. PDE based method uses iterations to converge
CHAPTER 9 INPAINTING USING SPARSE REPRESENTATION AND INVERSE DCT 9.1 Introduction In the previous chapters the inpainting was considered as an iterative algorithm. PDE based method uses iterations to converge
IDENTIFYING OPTICAL TRAP
 IDENTIFYING OPTICAL TRAP Yulwon Cho, Yuxin Zheng 12/16/2011 1. BACKGROUND AND MOTIVATION Optical trapping (also called optical tweezer) is widely used in studying a variety of biological systems in recent
IDENTIFYING OPTICAL TRAP Yulwon Cho, Yuxin Zheng 12/16/2011 1. BACKGROUND AND MOTIVATION Optical trapping (also called optical tweezer) is widely used in studying a variety of biological systems in recent
DeltaVision OMX SR super-resolution microscope. Super-resolution doesn t need to be complicated
 DeltaVision OMX SR super-resolution microscope Super-resolution doesn t need to be complicated Live-cell super-resolution microscopy DeltaVision OMX SR is a compact super-resolution microscope system optimized
DeltaVision OMX SR super-resolution microscope Super-resolution doesn t need to be complicated Live-cell super-resolution microscopy DeltaVision OMX SR is a compact super-resolution microscope system optimized
SPATIAL DENSITY ESTIMATION BASED SEGMENTATION OF SUPER-RESOLUTION LOCALIZATION MICROSCOPY IMAGES
 SPATIAL DENSITY ESTIMATION ASED SEGMENTATION OF SUPER-RESOLUTION LOCALIZATION MICROSCOPY IMAGES Kuan-Chieh Jackie Chen 1,2,3, Ge Yang 1,2, and Jelena Kovačević 3,1,2 1 Dept. of iomedical Eng., 2 Center
SPATIAL DENSITY ESTIMATION ASED SEGMENTATION OF SUPER-RESOLUTION LOCALIZATION MICROSCOPY IMAGES Kuan-Chieh Jackie Chen 1,2,3, Ge Yang 1,2, and Jelena Kovačević 3,1,2 1 Dept. of iomedical Eng., 2 Center
Full-Colour, Computational Ghost Video. Miles Padgett Kelvin Chair of Natural Philosophy
 Full-Colour, Computational Ghost Video Miles Padgett Kelvin Chair of Natural Philosophy A Quantum Ghost Imager! Generate random photon pairs, illuminate both object and camera SPDC CCD Identical copies
Full-Colour, Computational Ghost Video Miles Padgett Kelvin Chair of Natural Philosophy A Quantum Ghost Imager! Generate random photon pairs, illuminate both object and camera SPDC CCD Identical copies
Detecting Burnscar from Hyperspectral Imagery via Sparse Representation with Low-Rank Interference
 Detecting Burnscar from Hyperspectral Imagery via Sparse Representation with Low-Rank Interference Minh Dao 1, Xiang Xiang 1, Bulent Ayhan 2, Chiman Kwan 2, Trac D. Tran 1 Johns Hopkins Univeristy, 3400
Detecting Burnscar from Hyperspectral Imagery via Sparse Representation with Low-Rank Interference Minh Dao 1, Xiang Xiang 1, Bulent Ayhan 2, Chiman Kwan 2, Trac D. Tran 1 Johns Hopkins Univeristy, 3400
Using. Adaptive. Fourth. Department of Graduate Tohoku University Sendai, Japan jp. the. is adopting. was proposed in. and
 Guan Gui, Abolfazll Mehbodniya and Fumiyuki Adachi Department of Communication Engineering Graduate School of Engineering, Tohoku University Sendai, Japan {gui, mehbod}@mobile.ecei.tohoku.ac..jp, adachi@ecei.tohoku.ac.
Guan Gui, Abolfazll Mehbodniya and Fumiyuki Adachi Department of Communication Engineering Graduate School of Engineering, Tohoku University Sendai, Japan {gui, mehbod}@mobile.ecei.tohoku.ac..jp, adachi@ecei.tohoku.ac.
University of Florida CISE department Gator Engineering. Clustering Part 4
 Clustering Part 4 Dr. Sanjay Ranka Professor Computer and Information Science and Engineering University of Florida, Gainesville DBSCAN DBSCAN is a density based clustering algorithm Density = number of
Clustering Part 4 Dr. Sanjay Ranka Professor Computer and Information Science and Engineering University of Florida, Gainesville DBSCAN DBSCAN is a density based clustering algorithm Density = number of
Cluster Analysis of Super Resolution Fluorescence Images Ki Woong Sung
 NSERC USRA Report Summer 2015 Cluster Analysis of Super Resolution Fluorescence Images Ki Woong Sung 1 Biological Backgound Direct Stochastic Optical Reconstruction Microscopy (dstorm) is a novel, high
NSERC USRA Report Summer 2015 Cluster Analysis of Super Resolution Fluorescence Images Ki Woong Sung 1 Biological Backgound Direct Stochastic Optical Reconstruction Microscopy (dstorm) is a novel, high
Project Updates Short lecture Volumetric Modeling +2 papers
 Volumetric Modeling Schedule (tentative) Feb 20 Feb 27 Mar 5 Introduction Lecture: Geometry, Camera Model, Calibration Lecture: Features, Tracking/Matching Mar 12 Mar 19 Mar 26 Apr 2 Apr 9 Apr 16 Apr 23
Volumetric Modeling Schedule (tentative) Feb 20 Feb 27 Mar 5 Introduction Lecture: Geometry, Camera Model, Calibration Lecture: Features, Tracking/Matching Mar 12 Mar 19 Mar 26 Apr 2 Apr 9 Apr 16 Apr 23
Clustering Part 4 DBSCAN
 Clustering Part 4 Dr. Sanjay Ranka Professor Computer and Information Science and Engineering University of Florida, Gainesville DBSCAN DBSCAN is a density based clustering algorithm Density = number of
Clustering Part 4 Dr. Sanjay Ranka Professor Computer and Information Science and Engineering University of Florida, Gainesville DBSCAN DBSCAN is a density based clustering algorithm Density = number of
Maximum precision closed-form solution for localizing diffraction-limited spots in noisy images
 Maximum precision closed-form solution for localizing diffraction-limited spots in noisy images Joshua D. Larkin and Peter R. Cook * Sir William Dunn School of Pathology, University of Oxford, South Parks
Maximum precision closed-form solution for localizing diffraction-limited spots in noisy images Joshua D. Larkin and Peter R. Cook * Sir William Dunn School of Pathology, University of Oxford, South Parks
Digital Image Processing. Prof. P.K. Biswas. Department of Electronics & Electrical Communication Engineering
 Digital Image Processing Prof. P.K. Biswas Department of Electronics & Electrical Communication Engineering Indian Institute of Technology, Kharagpur Image Segmentation - III Lecture - 31 Hello, welcome
Digital Image Processing Prof. P.K. Biswas Department of Electronics & Electrical Communication Engineering Indian Institute of Technology, Kharagpur Image Segmentation - III Lecture - 31 Hello, welcome
LED holographic imaging by spatial-domain diffraction computation of. textured models
 LED holographic imaging by spatial-domain diffraction computation of textured models Ding-Chen Chen, Xiao-Ning Pang, Yi-Cong Ding, Yi-Gui Chen, and Jian-Wen Dong* School of Physics and Engineering, and
LED holographic imaging by spatial-domain diffraction computation of textured models Ding-Chen Chen, Xiao-Ning Pang, Yi-Cong Ding, Yi-Gui Chen, and Jian-Wen Dong* School of Physics and Engineering, and
COMPRESSED SENSING: A NOVEL POLYNOMIAL COMPLEXITY SOLUTION TO NASH EQUILIBRIA IN DYNAMICAL GAMES. Jing Huang, Liming Wang and Dan Schonfeld
 COMPRESSED SENSING: A NOVEL POLYNOMIAL COMPLEXITY SOLUTION TO NASH EQUILIBRIA IN DYNAMICAL GAMES Jing Huang, Liming Wang and Dan Schonfeld Department of Electrical and Computer Engineering, University
COMPRESSED SENSING: A NOVEL POLYNOMIAL COMPLEXITY SOLUTION TO NASH EQUILIBRIA IN DYNAMICAL GAMES Jing Huang, Liming Wang and Dan Schonfeld Department of Electrical and Computer Engineering, University
MAT 17C - DISCUSSION #4, Counting Proteins
 MAT 17C - DISCUSSION #4, Counting Proteins Visualization of molecules inside a living cell is one of the most important tools of molecular biology in the 21 st century. In fact, it was the subject of the
MAT 17C - DISCUSSION #4, Counting Proteins Visualization of molecules inside a living cell is one of the most important tools of molecular biology in the 21 st century. In fact, it was the subject of the
Contents. 2.5 DBSCAN Pair Auto-Correlation Saving your results References... 18
 Contents 1 Getting Started... 2 1.1 Launching the program:... 2 1.2 Loading and viewing your data... 2 1.3 Setting parameters... 5 1.4 Cropping points... 6 1.5 Selecting regions of interest... 7 2 Analysis...
Contents 1 Getting Started... 2 1.1 Launching the program:... 2 1.2 Loading and viewing your data... 2 1.3 Setting parameters... 5 1.4 Cropping points... 6 1.5 Selecting regions of interest... 7 2 Analysis...
Back Illuminated Scientific CMOS
 Prime 95B Scientific CMOS Camera Datasheet CMOS, EMCCD AND CCD CAMERAS FOR LIFE SCIENCES Back Illuminated Scientific CMOS Discovery depends on every photon Primary applications: Super-Resolution Microscopy
Prime 95B Scientific CMOS Camera Datasheet CMOS, EMCCD AND CCD CAMERAS FOR LIFE SCIENCES Back Illuminated Scientific CMOS Discovery depends on every photon Primary applications: Super-Resolution Microscopy
Last class. This class. Single molecule imaging Deconvolution. FLIM Confocal
 FLIM, Confocal Last class Single molecule imaging Deconvolution This class FLIM Confocal FLIM Fluorescence Lifetime IMaging FLIM Fluorescence lifetime imaging Easier and more accurate quantitation Much
FLIM, Confocal Last class Single molecule imaging Deconvolution This class FLIM Confocal FLIM Fluorescence Lifetime IMaging FLIM Fluorescence lifetime imaging Easier and more accurate quantitation Much
Randomized sampling strategies
 Randomized sampling strategies Felix J. Herrmann SLIM Seismic Laboratory for Imaging and Modeling the University of British Columbia SLIM Drivers & Acquisition costs impediments Full-waveform inversion
Randomized sampling strategies Felix J. Herrmann SLIM Seismic Laboratory for Imaging and Modeling the University of British Columbia SLIM Drivers & Acquisition costs impediments Full-waveform inversion
Computer Project 3. AA Computational Fluid Dyanmics University of Washington. Mishaal Aleem March 17, 2015
 Computer Project 3 AA 543 - Computational Fluid Dyanmics University of Washington Mishaal Aleem March 17, 2015 Contents Introduction........................................... 1 3.1 Grid Generator.......................................
Computer Project 3 AA 543 - Computational Fluid Dyanmics University of Washington Mishaal Aleem March 17, 2015 Contents Introduction........................................... 1 3.1 Grid Generator.......................................
Virtual Frap User Guide
 Virtual Frap User Guide http://wiki.vcell.uchc.edu/twiki/bin/view/vcell/vfrap Center for Cell Analysis and Modeling University of Connecticut Health Center 2010-1 - 1 Introduction Flourescence Photobleaching
Virtual Frap User Guide http://wiki.vcell.uchc.edu/twiki/bin/view/vcell/vfrap Center for Cell Analysis and Modeling University of Connecticut Health Center 2010-1 - 1 Introduction Flourescence Photobleaching
Compressive Sensing for Multimedia. Communications in Wireless Sensor Networks
 Compressive Sensing for Multimedia 1 Communications in Wireless Sensor Networks Wael Barakat & Rabih Saliba MDDSP Project Final Report Prof. Brian L. Evans May 9, 2008 Abstract Compressive Sensing is an
Compressive Sensing for Multimedia 1 Communications in Wireless Sensor Networks Wael Barakat & Rabih Saliba MDDSP Project Final Report Prof. Brian L. Evans May 9, 2008 Abstract Compressive Sensing is an
Robust Face Recognition via Sparse Representation
 Robust Face Recognition via Sparse Representation Panqu Wang Department of Electrical and Computer Engineering University of California, San Diego La Jolla, CA 92092 pawang@ucsd.edu Can Xu Department of
Robust Face Recognition via Sparse Representation Panqu Wang Department of Electrical and Computer Engineering University of California, San Diego La Jolla, CA 92092 pawang@ucsd.edu Can Xu Department of
COLOCALISATION. Alexia Ferrand. Imaging Core Facility Biozentrum Basel
 COLOCALISATION Alexia Ferrand Imaging Core Facility Biozentrum Basel OUTLINE Introduction How to best prepare your samples for colocalisation How to acquire the images for colocalisation How to analyse
COLOCALISATION Alexia Ferrand Imaging Core Facility Biozentrum Basel OUTLINE Introduction How to best prepare your samples for colocalisation How to acquire the images for colocalisation How to analyse
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature10934 Supplementary Methods Mathematical implementation of the EST method. The EST method begins with padding each projection with zeros (that is, embedding
SUPPLEMENTARY INFORMATION doi:10.1038/nature10934 Supplementary Methods Mathematical implementation of the EST method. The EST method begins with padding each projection with zeros (that is, embedding
SUPPLEMENTARY FILE S1: 3D AIRWAY TUBE RECONSTRUCTION AND CELL-BASED MECHANICAL MODEL. RELATED TO FIGURE 1, FIGURE 7, AND STAR METHODS.
 SUPPLEMENTARY FILE S1: 3D AIRWAY TUBE RECONSTRUCTION AND CELL-BASED MECHANICAL MODEL. RELATED TO FIGURE 1, FIGURE 7, AND STAR METHODS. 1. 3D AIRWAY TUBE RECONSTRUCTION. RELATED TO FIGURE 1 AND STAR METHODS
SUPPLEMENTARY FILE S1: 3D AIRWAY TUBE RECONSTRUCTION AND CELL-BASED MECHANICAL MODEL. RELATED TO FIGURE 1, FIGURE 7, AND STAR METHODS. 1. 3D AIRWAY TUBE RECONSTRUCTION. RELATED TO FIGURE 1 AND STAR METHODS
Coupling of surface roughness to the performance of computer-generated holograms
 Coupling of surface roughness to the performance of computer-generated holograms Ping Zhou* and Jim Burge College of Optical Sciences, University of Arizona, Tucson, Arizona 85721, USA *Corresponding author:
Coupling of surface roughness to the performance of computer-generated holograms Ping Zhou* and Jim Burge College of Optical Sciences, University of Arizona, Tucson, Arizona 85721, USA *Corresponding author:
Automatic Partiicle Tracking Software USE ER MANUAL Update: May 2015
 Automatic Particle Tracking Software USER MANUAL Update: May 2015 File Menu The micrograph below shows the panel displayed when a movie is opened, including a playback menu where most of the parameters
Automatic Particle Tracking Software USER MANUAL Update: May 2015 File Menu The micrograph below shows the panel displayed when a movie is opened, including a playback menu where most of the parameters
Face Recognition via Sparse Representation
 Face Recognition via Sparse Representation John Wright, Allen Y. Yang, Arvind, S. Shankar Sastry and Yi Ma IEEE Trans. PAMI, March 2008 Research About Face Face Detection Face Alignment Face Recognition
Face Recognition via Sparse Representation John Wright, Allen Y. Yang, Arvind, S. Shankar Sastry and Yi Ma IEEE Trans. PAMI, March 2008 Research About Face Face Detection Face Alignment Face Recognition
Sparse Reconstruction / Compressive Sensing
 Sparse Reconstruction / Compressive Sensing Namrata Vaswani Department of Electrical and Computer Engineering Iowa State University Namrata Vaswani Sparse Reconstruction / Compressive Sensing 1/ 20 The
Sparse Reconstruction / Compressive Sensing Namrata Vaswani Department of Electrical and Computer Engineering Iowa State University Namrata Vaswani Sparse Reconstruction / Compressive Sensing 1/ 20 The
MICROTUBULE FILAMENT TRACING AND ESTIMATION. Rohan Chabukswar. Department of Electrical and Computer Engineering Carnegie Mellon University
 MICROTUBULE FILAMENT TRACING AND ESTIMATION Rohan Chabukswar Department of Electrical and Computer Engineering Carnegie Mellon University ABSTRACT The project explores preprocessing of images to facilitate
MICROTUBULE FILAMENT TRACING AND ESTIMATION Rohan Chabukswar Department of Electrical and Computer Engineering Carnegie Mellon University ABSTRACT The project explores preprocessing of images to facilitate
CS4495 Fall 2014 Computer Vision Problem Set 6: Particle Tracking
 CS4495 Fall 2014 Computer Vision Problem Set 6: Particle Tracking DUE Tues, November 25-11:55pm Here you are going to experiment with a particle filter tracker such as was described in class. Recall that
CS4495 Fall 2014 Computer Vision Problem Set 6: Particle Tracking DUE Tues, November 25-11:55pm Here you are going to experiment with a particle filter tracker such as was described in class. Recall that
Towards direct motion and shape parameter recovery from image sequences. Stephen Benoit. Ph.D. Thesis Presentation September 25, 2003
 Towards direct motion and shape parameter recovery from image sequences Stephen Benoit Ph.D. Thesis Presentation September 25, 2003 September 25, 2003 Towards direct motion and shape parameter recovery
Towards direct motion and shape parameter recovery from image sequences Stephen Benoit Ph.D. Thesis Presentation September 25, 2003 September 25, 2003 Towards direct motion and shape parameter recovery
High Resolution BSI Scientific CMOS
 CMOS, EMCCD AND CCD CAMERAS FOR LIFE SCIENCES High Resolution BSI Scientific CMOS Prime BSI delivers the perfect balance between high resolution imaging and sensitivity with an optimized pixel design and
CMOS, EMCCD AND CCD CAMERAS FOR LIFE SCIENCES High Resolution BSI Scientific CMOS Prime BSI delivers the perfect balance between high resolution imaging and sensitivity with an optimized pixel design and
Factorization with Missing and Noisy Data
 Factorization with Missing and Noisy Data Carme Julià, Angel Sappa, Felipe Lumbreras, Joan Serrat, and Antonio López Computer Vision Center and Computer Science Department, Universitat Autònoma de Barcelona,
Factorization with Missing and Noisy Data Carme Julià, Angel Sappa, Felipe Lumbreras, Joan Serrat, and Antonio López Computer Vision Center and Computer Science Department, Universitat Autònoma de Barcelona,
Beam Profilier - Beamage 3.0
 Profilier - age 3.0 KEY FEATURES High resolution (160x120 points) 2.2 MPixels resolution gives accurate profile measurements on very small beams Large Area The 11.3 x 6.0 mm sensor allows to measure very
Profilier - age 3.0 KEY FEATURES High resolution (160x120 points) 2.2 MPixels resolution gives accurate profile measurements on very small beams Large Area The 11.3 x 6.0 mm sensor allows to measure very
Median filter. Non-linear filtering example. Degraded image. Radius 1 median filter. Today
 Today Non-linear filtering example Median filter Replace each pixel by the median over N pixels (5 pixels, for these examples). Generalizes to rank order filters. In: In: 5-pixel neighborhood Out: Out:
Today Non-linear filtering example Median filter Replace each pixel by the median over N pixels (5 pixels, for these examples). Generalizes to rank order filters. In: In: 5-pixel neighborhood Out: Out:
Non-linear filtering example
 Today Non-linear filtering example Median filter Replace each pixel by the median over N pixels (5 pixels, for these examples). Generalizes to rank order filters. In: In: 5-pixel neighborhood Out: Out:
Today Non-linear filtering example Median filter Replace each pixel by the median over N pixels (5 pixels, for these examples). Generalizes to rank order filters. In: In: 5-pixel neighborhood Out: Out:
Geometric-Edge Random Graph Model for Image Representation
 Geometric-Edge Random Graph Model for Image Representation Bo JIANG, Jin TANG, Bin LUO CVPR Group, Anhui University /1/11, Beijing Acknowledgements This research is supported by the National Natural Science
Geometric-Edge Random Graph Model for Image Representation Bo JIANG, Jin TANG, Bin LUO CVPR Group, Anhui University /1/11, Beijing Acknowledgements This research is supported by the National Natural Science
BSI scmos. High Quantum Efficiency. Cooled Scientific CMOS Camera
 95 BSI scmos High Quantum Efficiency Cooled Scientific CMOS Camera Backside-illuminated scmos technology Opening a new era of high sensitivity imaging applications! The Dhyana 95 is a highly sensitive
95 BSI scmos High Quantum Efficiency Cooled Scientific CMOS Camera Backside-illuminated scmos technology Opening a new era of high sensitivity imaging applications! The Dhyana 95 is a highly sensitive
Leave-One-Out Support Vector Machines
 Leave-One-Out Support Vector Machines Jason Weston Department of Computer Science Royal Holloway, University of London, Egham Hill, Egham, Surrey, TW20 OEX, UK. Abstract We present a new learning algorithm
Leave-One-Out Support Vector Machines Jason Weston Department of Computer Science Royal Holloway, University of London, Egham Hill, Egham, Surrey, TW20 OEX, UK. Abstract We present a new learning algorithm
CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION
 CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION 6.1 INTRODUCTION Fuzzy logic based computational techniques are becoming increasingly important in the medical image analysis arena. The significant
CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION 6.1 INTRODUCTION Fuzzy logic based computational techniques are becoming increasingly important in the medical image analysis arena. The significant
Objective. A Finite State Machine Approach to Cluster Identification Using the Hoshen-Kopelman Algorithm. Hoshen-Kopelman Algorithm
 Objective A Finite State Machine Approach to Cluster Identification Using the Cluster Identification Want to find and identify homogeneous patches in a D matrix, where: Cluster membership defined by adjacency
Objective A Finite State Machine Approach to Cluster Identification Using the Cluster Identification Want to find and identify homogeneous patches in a D matrix, where: Cluster membership defined by adjacency
Advanced phase retrieval: maximum likelihood technique with sparse regularization of phase and amplitude
 Advanced phase retrieval: maximum likelihood technique with sparse regularization of phase and amplitude A. Migukin *, V. atkovnik and J. Astola Department of Signal Processing, Tampere University of Technology,
Advanced phase retrieval: maximum likelihood technique with sparse regularization of phase and amplitude A. Migukin *, V. atkovnik and J. Astola Department of Signal Processing, Tampere University of Technology,
SPARSE CODE SHRINKAGE BASED ON THE NORMAL INVERSE GAUSSIAN DENSITY MODEL. Department of Physics University of Tromsø N Tromsø, Norway
 SPARSE CODE SRINKAGE ASED ON TE NORMA INVERSE GAUSSIAN DENSITY MODE Robert Jenssen, Tor Arne Øigård, Torbjørn Eltoft and Alfred anssen Department of Physics University of Tromsø N - 9037 Tromsø, Norway
SPARSE CODE SRINKAGE ASED ON TE NORMA INVERSE GAUSSIAN DENSITY MODE Robert Jenssen, Tor Arne Øigård, Torbjørn Eltoft and Alfred anssen Department of Physics University of Tromsø N - 9037 Tromsø, Norway
A SUPER-RESOLUTION MICROSCOPY WITH STANDING EVANESCENT LIGHT AND IMAGE RECONSTRUCTION METHOD
 A SUPER-RESOLUTION MICROSCOPY WITH STANDING EVANESCENT LIGHT AND IMAGE RECONSTRUCTION METHOD Hiroaki Nishioka, Satoru Takahashi Kiyoshi Takamasu Department of Precision Engineering, The University of Tokyo,
A SUPER-RESOLUTION MICROSCOPY WITH STANDING EVANESCENT LIGHT AND IMAGE RECONSTRUCTION METHOD Hiroaki Nishioka, Satoru Takahashi Kiyoshi Takamasu Department of Precision Engineering, The University of Tokyo,
COMPRESSIVE VIDEO SAMPLING
 COMPRESSIVE VIDEO SAMPLING Vladimir Stanković and Lina Stanković Dept of Electronic and Electrical Engineering University of Strathclyde, Glasgow, UK phone: +44-141-548-2679 email: {vladimir,lina}.stankovic@eee.strath.ac.uk
COMPRESSIVE VIDEO SAMPLING Vladimir Stanković and Lina Stanković Dept of Electronic and Electrical Engineering University of Strathclyde, Glasgow, UK phone: +44-141-548-2679 email: {vladimir,lina}.stankovic@eee.strath.ac.uk
Recognition and Measurement of Small Defects in ICT Testing
 19 th World Conference on Non-Destructive Testing 2016 Recognition and Measurement of Small Defects in ICT Testing Guo ZHIMIN, Ni PEIJUN, Zhang WEIGUO, Qi ZICHENG Inner Mongolia Metallic Materials Research
19 th World Conference on Non-Destructive Testing 2016 Recognition and Measurement of Small Defects in ICT Testing Guo ZHIMIN, Ni PEIJUN, Zhang WEIGUO, Qi ZICHENG Inner Mongolia Metallic Materials Research
Modified Iterative Method for Recovery of Sparse Multiple Measurement Problems
 Journal of Electrical Engineering 6 (2018) 124-128 doi: 10.17265/2328-2223/2018.02.009 D DAVID PUBLISHING Modified Iterative Method for Recovery of Sparse Multiple Measurement Problems Sina Mortazavi and
Journal of Electrical Engineering 6 (2018) 124-128 doi: 10.17265/2328-2223/2018.02.009 D DAVID PUBLISHING Modified Iterative Method for Recovery of Sparse Multiple Measurement Problems Sina Mortazavi and
Iterative SPECT reconstruction with 3D detector response
 Iterative SPECT reconstruction with 3D detector response Jeffrey A. Fessler and Anastasia Yendiki COMMUNICATIONS & SIGNAL PROCESSING LABORATORY Department of Electrical Engineering and Computer Science
Iterative SPECT reconstruction with 3D detector response Jeffrey A. Fessler and Anastasia Yendiki COMMUNICATIONS & SIGNAL PROCESSING LABORATORY Department of Electrical Engineering and Computer Science
Electromagnetic migration of marine CSEM data in areas with rough bathymetry Michael S. Zhdanov and Martin Čuma*, University of Utah
 Electromagnetic migration of marine CSEM data in areas with rough bathymetry Michael S. Zhdanov and Martin Čuma*, University of Utah Summary In this paper we present a new approach to the interpretation
Electromagnetic migration of marine CSEM data in areas with rough bathymetry Michael S. Zhdanov and Martin Čuma*, University of Utah Summary In this paper we present a new approach to the interpretation
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Performance analysis under four different clustering scenarios. Performance analysis under four different clustering scenarios. i) Standard Conditions, ii) a sparse data set with
Supplementary Figure 1 Performance analysis under four different clustering scenarios. Performance analysis under four different clustering scenarios. i) Standard Conditions, ii) a sparse data set with
A Keypoint Descriptor Inspired by Retinal Computation
 A Keypoint Descriptor Inspired by Retinal Computation Bongsoo Suh, Sungjoon Choi, Han Lee Stanford University {bssuh,sungjoonchoi,hanlee}@stanford.edu Abstract. The main goal of our project is to implement
A Keypoint Descriptor Inspired by Retinal Computation Bongsoo Suh, Sungjoon Choi, Han Lee Stanford University {bssuh,sungjoonchoi,hanlee}@stanford.edu Abstract. The main goal of our project is to implement
Optimised corrections for finite-difference modelling in two dimensions
 Optimized corrections for 2D FD modelling Optimised corrections for finite-difference modelling in two dimensions Peter M. Manning and Gary F. Margrave ABSTRACT Finite-difference two-dimensional correction
Optimized corrections for 2D FD modelling Optimised corrections for finite-difference modelling in two dimensions Peter M. Manning and Gary F. Margrave ABSTRACT Finite-difference two-dimensional correction
ClusterViSu, a method for clustering of protein complexes by Voronoi. tessellation in super-resolution microscopy
 ClusterViSu, a method for clustering of protein complexes by Voronoi tessellation in super-resolution microscopy Leonid Andronov 1,2,3,4, Igor Orlov 1,2,3,4, Yves Lutz 1,2,3,4, Jean-Luc Vonesch 1,2,3,4,
ClusterViSu, a method for clustering of protein complexes by Voronoi tessellation in super-resolution microscopy Leonid Andronov 1,2,3,4, Igor Orlov 1,2,3,4, Yves Lutz 1,2,3,4, Jean-Luc Vonesch 1,2,3,4,
The Benefit of Tree Sparsity in Accelerated MRI
 The Benefit of Tree Sparsity in Accelerated MRI Chen Chen and Junzhou Huang Department of Computer Science and Engineering, The University of Texas at Arlington, TX, USA 76019 Abstract. The wavelet coefficients
The Benefit of Tree Sparsity in Accelerated MRI Chen Chen and Junzhou Huang Department of Computer Science and Engineering, The University of Texas at Arlington, TX, USA 76019 Abstract. The wavelet coefficients
Raster Image Correlation Spectroscopy and Number and Brightness Analysis
 Raster Image Correlation Spectroscopy and Number and Brightness Analysis 15 Principles of Fluorescence Course Paolo Annibale, PhD On behalf of Dr. Enrico Gratton lfd Raster Image Correlation Spectroscopy
Raster Image Correlation Spectroscopy and Number and Brightness Analysis 15 Principles of Fluorescence Course Paolo Annibale, PhD On behalf of Dr. Enrico Gratton lfd Raster Image Correlation Spectroscopy
COLOCALISATION. Alexia Loynton-Ferrand. Imaging Core Facility Biozentrum Basel
 COLOCALISATION Alexia Loynton-Ferrand Imaging Core Facility Biozentrum Basel OUTLINE Introduction How to best prepare your samples for colocalisation How to acquire the images for colocalisation How to
COLOCALISATION Alexia Loynton-Ferrand Imaging Core Facility Biozentrum Basel OUTLINE Introduction How to best prepare your samples for colocalisation How to acquire the images for colocalisation How to
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/1/eaao7005/dc1 Supplementary Materials for Computational discovery of extremal microstructure families The PDF file includes: Desai Chen, Mélina Skouras, Bo Zhu,
advances.sciencemag.org/cgi/content/full/4/1/eaao7005/dc1 Supplementary Materials for Computational discovery of extremal microstructure families The PDF file includes: Desai Chen, Mélina Skouras, Bo Zhu,
Visual object classification by sparse convolutional neural networks
 Visual object classification by sparse convolutional neural networks Alexander Gepperth 1 1- Ruhr-Universität Bochum - Institute for Neural Dynamics Universitätsstraße 150, 44801 Bochum - Germany Abstract.
Visual object classification by sparse convolutional neural networks Alexander Gepperth 1 1- Ruhr-Universität Bochum - Institute for Neural Dynamics Universitätsstraße 150, 44801 Bochum - Germany Abstract.
Characterization of Single Pixel Evaluation PIV for Precise Flow Measurement
 Characterization of Single Pixel Evaluation PIV for Precise Flow Measurement Han-Sheng Chuang 1, Steve T. Wereley 1, Lichuan Gui 2 1: School of Mechanical Engineering, Purdue University, West Lafayette,
Characterization of Single Pixel Evaluation PIV for Precise Flow Measurement Han-Sheng Chuang 1, Steve T. Wereley 1, Lichuan Gui 2 1: School of Mechanical Engineering, Purdue University, West Lafayette,
Sparse coding for image classification
 Sparse coding for image classification Columbia University Electrical Engineering: Kun Rong(kr2496@columbia.edu) Yongzhou Xiang(yx2211@columbia.edu) Yin Cui(yc2776@columbia.edu) Outline Background Introduction
Sparse coding for image classification Columbia University Electrical Engineering: Kun Rong(kr2496@columbia.edu) Yongzhou Xiang(yx2211@columbia.edu) Yin Cui(yc2776@columbia.edu) Outline Background Introduction
APPROACHES IN QUANTITATIVE, MULTI-DIMENSIONAL MICROSCOPY
 APPROACHES IN QUANTITATIVE, MULTI-DIMENSIONAL MICROSCOPY Profs. Zvi Kam and Benjamin Geiger Department of Molecular Cell Biology The Weizmann Institute of Science Rehovot, Israel In this presentation we
APPROACHES IN QUANTITATIVE, MULTI-DIMENSIONAL MICROSCOPY Profs. Zvi Kam and Benjamin Geiger Department of Molecular Cell Biology The Weizmann Institute of Science Rehovot, Israel In this presentation we
Compressed Sensing for Rapid MR Imaging
 Compressed Sensing for Rapid Imaging Michael Lustig1, Juan Santos1, David Donoho2 and John Pauly1 1 Electrical Engineering Department, Stanford University 2 Statistics Department, Stanford University rapid
Compressed Sensing for Rapid Imaging Michael Lustig1, Juan Santos1, David Donoho2 and John Pauly1 1 Electrical Engineering Department, Stanford University 2 Statistics Department, Stanford University rapid
Single-particle electron microscopy (cryo-electron microscopy) CS/CME/BioE/Biophys/BMI 279 Nov. 16 and 28, 2017 Ron Dror
 Single-particle electron microscopy (cryo-electron microscopy) CS/CME/BioE/Biophys/BMI 279 Nov. 16 and 28, 2017 Ron Dror 1 Last month s Nobel Prize in Chemistry Awarded to Jacques Dubochet, Joachim Frank
Single-particle electron microscopy (cryo-electron microscopy) CS/CME/BioE/Biophys/BMI 279 Nov. 16 and 28, 2017 Ron Dror 1 Last month s Nobel Prize in Chemistry Awarded to Jacques Dubochet, Joachim Frank
Based on Regression Diagnostics
 Automatic Detection of Region-Mura Defects in TFT-LCD Based on Regression Diagnostics Yu-Chiang Chuang 1 and Shu-Kai S. Fan 2 Department of Industrial Engineering and Management, Yuan Ze University, Tao
Automatic Detection of Region-Mura Defects in TFT-LCD Based on Regression Diagnostics Yu-Chiang Chuang 1 and Shu-Kai S. Fan 2 Department of Industrial Engineering and Management, Yuan Ze University, Tao
Vutara 350. Innovation with Integrity. The Fastest, Super-Resolution Microscope Deep 3D Imaging on Live Cells, Quickly and Easily
 Vutara 350 The Fastest, Super-Resolution Microscope Deep 3D Imaging on Live Cells, Quickly and Easily Innovation with Integrity Fluorescence Microscopy Vutara 350 Don t Get Left Behind Bruker s Vutara
Vutara 350 The Fastest, Super-Resolution Microscope Deep 3D Imaging on Live Cells, Quickly and Easily Innovation with Integrity Fluorescence Microscopy Vutara 350 Don t Get Left Behind Bruker s Vutara
IMAGE DE-NOISING IN WAVELET DOMAIN
 IMAGE DE-NOISING IN WAVELET DOMAIN Aaditya Verma a, Shrey Agarwal a a Department of Civil Engineering, Indian Institute of Technology, Kanpur, India - (aaditya, ashrey)@iitk.ac.in KEY WORDS: Wavelets,
IMAGE DE-NOISING IN WAVELET DOMAIN Aaditya Verma a, Shrey Agarwal a a Department of Civil Engineering, Indian Institute of Technology, Kanpur, India - (aaditya, ashrey)@iitk.ac.in KEY WORDS: Wavelets,
Simultaneous surface texture classification and illumination tilt angle prediction
 Simultaneous surface texture classification and illumination tilt angle prediction X. Lladó, A. Oliver, M. Petrou, J. Freixenet, and J. Martí Computer Vision and Robotics Group - IIiA. University of Girona
Simultaneous surface texture classification and illumination tilt angle prediction X. Lladó, A. Oliver, M. Petrou, J. Freixenet, and J. Martí Computer Vision and Robotics Group - IIiA. University of Girona
y z x SNR(dB) RMSE(mm)
 a b c d y z x SNR(dB) RMSE(mm) 30.16 2.94 20.28 6.56 10.33 13.74 0.98 25.37 Supplementary Figure 1: System performance with different noise levels. The SNRs are the mean SNR of all measured signals corresponding
a b c d y z x SNR(dB) RMSE(mm) 30.16 2.94 20.28 6.56 10.33 13.74 0.98 25.37 Supplementary Figure 1: System performance with different noise levels. The SNRs are the mean SNR of all measured signals corresponding
Supporting Information. Characterization of Porous Materials by. Fluorescence Correlation Spectroscopy Superresolution. Optical Fluctuation Imaging
 Supporting Information Characterization of Porous Materials by Fluorescence Correlation Spectroscopy Superresolution Optical Fluctuation Imaging Lydia Kisley, 1 Rachel Brunetti, 2 Lawrence J. Tauzin, 1
Supporting Information Characterization of Porous Materials by Fluorescence Correlation Spectroscopy Superresolution Optical Fluctuation Imaging Lydia Kisley, 1 Rachel Brunetti, 2 Lawrence J. Tauzin, 1
Optimal Sampling Geometries for TV-Norm Reconstruction of fmri Data
 Optimal Sampling Geometries for TV-Norm Reconstruction of fmri Data Oliver M. Jeromin, Student Member, IEEE, Vince D. Calhoun, Senior Member, IEEE, and Marios S. Pattichis, Senior Member, IEEE Abstract
Optimal Sampling Geometries for TV-Norm Reconstruction of fmri Data Oliver M. Jeromin, Student Member, IEEE, Vince D. Calhoun, Senior Member, IEEE, and Marios S. Pattichis, Senior Member, IEEE Abstract
ADAPTIVE LOW RANK AND SPARSE DECOMPOSITION OF VIDEO USING COMPRESSIVE SENSING
 ADAPTIVE LOW RANK AND SPARSE DECOMPOSITION OF VIDEO USING COMPRESSIVE SENSING Fei Yang 1 Hong Jiang 2 Zuowei Shen 3 Wei Deng 4 Dimitris Metaxas 1 1 Rutgers University 2 Bell Labs 3 National University
ADAPTIVE LOW RANK AND SPARSE DECOMPOSITION OF VIDEO USING COMPRESSIVE SENSING Fei Yang 1 Hong Jiang 2 Zuowei Shen 3 Wei Deng 4 Dimitris Metaxas 1 1 Rutgers University 2 Bell Labs 3 National University
A Novel Image Super-resolution Reconstruction Algorithm based on Modified Sparse Representation
 , pp.162-167 http://dx.doi.org/10.14257/astl.2016.138.33 A Novel Image Super-resolution Reconstruction Algorithm based on Modified Sparse Representation Liqiang Hu, Chaofeng He Shijiazhuang Tiedao University,
, pp.162-167 http://dx.doi.org/10.14257/astl.2016.138.33 A Novel Image Super-resolution Reconstruction Algorithm based on Modified Sparse Representation Liqiang Hu, Chaofeng He Shijiazhuang Tiedao University,
Structured Light II. Thanks to Ronen Gvili, Szymon Rusinkiewicz and Maks Ovsjanikov
 Structured Light II Johannes Köhler Johannes.koehler@dfki.de Thanks to Ronen Gvili, Szymon Rusinkiewicz and Maks Ovsjanikov Introduction Previous lecture: Structured Light I Active Scanning Camera/emitter
Structured Light II Johannes Köhler Johannes.koehler@dfki.de Thanks to Ronen Gvili, Szymon Rusinkiewicz and Maks Ovsjanikov Introduction Previous lecture: Structured Light I Active Scanning Camera/emitter
DISCRETE DOMAIN REPRESENTATION FOR SHAPE CONCEPTUALIZATION
 DISCRETE DOMAIN REPRESENTATION FOR SHAPE CONCEPTUALIZATION Zoltán Rusák, Imre Horváth, György Kuczogi, Joris S.M. Vergeest, Johan Jansson Department of Design Engineering Delft University of Technology
DISCRETE DOMAIN REPRESENTATION FOR SHAPE CONCEPTUALIZATION Zoltán Rusák, Imre Horváth, György Kuczogi, Joris S.M. Vergeest, Johan Jansson Department of Design Engineering Delft University of Technology
Hyperspectral and Multispectral Image Fusion Using Local Spatial-Spectral Dictionary Pair
 Hyperspectral and Multispectral Image Fusion Using Local Spatial-Spectral Dictionary Pair Yifan Zhang, Tuo Zhao, and Mingyi He School of Electronics and Information International Center for Information
Hyperspectral and Multispectral Image Fusion Using Local Spatial-Spectral Dictionary Pair Yifan Zhang, Tuo Zhao, and Mingyi He School of Electronics and Information International Center for Information
Compressed Sensing for Electron Tomography
 University of Maryland, College Park Department of Mathematics February 10, 2015 1/33 Outline I Introduction 1 Introduction 2 3 4 2/33 1 Introduction 2 3 4 3/33 Tomography Introduction Tomography - Producing
University of Maryland, College Park Department of Mathematics February 10, 2015 1/33 Outline I Introduction 1 Introduction 2 3 4 2/33 1 Introduction 2 3 4 3/33 Tomography Introduction Tomography - Producing
Image Processing
 Image Processing 159.731 Canny Edge Detection Report Syed Irfanullah, Azeezullah 00297844 Danh Anh Huynh 02136047 1 Canny Edge Detection INTRODUCTION Edges Edges characterize boundaries and are therefore
Image Processing 159.731 Canny Edge Detection Report Syed Irfanullah, Azeezullah 00297844 Danh Anh Huynh 02136047 1 Canny Edge Detection INTRODUCTION Edges Edges characterize boundaries and are therefore
2D and 3D Far-Field Radiation Patterns Reconstruction Based on Compressive Sensing
 Progress In Electromagnetics Research M, Vol. 46, 47 56, 206 2D and 3D Far-Field Radiation Patterns Reconstruction Based on Compressive Sensing Berenice Verdin * and Patrick Debroux Abstract The measurement
Progress In Electromagnetics Research M, Vol. 46, 47 56, 206 2D and 3D Far-Field Radiation Patterns Reconstruction Based on Compressive Sensing Berenice Verdin * and Patrick Debroux Abstract The measurement
Using Image's Processing Methods in Bio-Technology
 Int. J. Open Problems Compt. Math., Vol. 2, No. 2, June 2009 Using Image's Processing Methods in Bio-Technology I. A. Ismail 1, S. I. Zaki 2, E. A. Rakha 3 and M. A. Ashabrawy 4 1 Dean of Computer Science
Int. J. Open Problems Compt. Math., Vol. 2, No. 2, June 2009 Using Image's Processing Methods in Bio-Technology I. A. Ismail 1, S. I. Zaki 2, E. A. Rakha 3 and M. A. Ashabrawy 4 1 Dean of Computer Science
Spectral Compression: Weighted Principal Component Analysis versus Weighted Least Squares
 Spectral Compression: Weighted Principal Component Analysis versus Weighted Least Squares Farnaz Agahian a*, Brian Funt a, Seyed Hossein Amirshahi b a Simon Fraser University, 8888 University Dr. V5A 1S6,
Spectral Compression: Weighted Principal Component Analysis versus Weighted Least Squares Farnaz Agahian a*, Brian Funt a, Seyed Hossein Amirshahi b a Simon Fraser University, 8888 University Dr. V5A 1S6,
