Filogeografía BIOL 4211, Universidad de los Andes 25 de enero a 01 de abril 2006
|
|
|
- Meryl George
- 6 years ago
- Views:
Transcription
1 Laboratory excercise written by Andrew J. Crawford with the support of CIES Fulbright Program and Fulbright Colombia. Enjoy! Filogeografía BIOL 4211, Universidad de los Andes 25 de enero a 01 de abril 2006 Lab 2 Alignment of DNA sequences, visualization and export 04 February 2006 By now you have found your data set to work with for the course. At a minimum, your data include one mitochondrial gene sequenced from multiple individuals from multiple populations within a single species. Your data should also include an outgroup. Your data may also include additional species and additional genes. Today s lab will provide instructions and tips on how to create an alignment using ClustalX and manipulate this alignment in MEGA (Windows OS only) and Mequite. To visualize preliminary trees, we will use TreeEdit. These programs should already be installed on your machines. Otherwise, the programs are available here: Software ClustalX (for all major computer operating systems): ftp://ftp.ebi.ac.uk/pub/software/unix/clustalx/ MEGA version 3.1 (for Windows or with Windows emulator software): Mequite: Additional information Using ClustalX: Use the Help menu in ClustalX, or look at the above URL for detailed information on ClustalX and its various options. TreeEdit: A free program for Macintosh for visualizing and manipulating trees quickly and easily. TreeView: Like TreeEdit, but fewer features. MacClade: Is not free, but it is useful. I believe it is already on the Mac computers in the lab. Feel free to use this program as well. Not available for Windows. Text editors: For preparing data files, I recommend a free, flexible text editor such as TextWrangler. On a Macintosh OSX machine, you can also use a unix-based text editor such as emacs.
2 BIOL 4211 Lab 2, page 2 of 6 On a PC machine, you might use WordPad, but under View > Options > Text (or Rich Text) > Word wrap, select No wrap. GOALS: Organize your sequences into one file. Gain familiarity with ClustalX. Change multiple sequence alignment parameters and note any resulting changes in alignment. Save one or more final alignments for use in the next 8 weeks. For protein-coding sequences, confirm that the data do not contain premature stop codons. I. Gathering you data together. To begin the lab, for each gene fragment or multigene contiguous DNA sequence you should have one file. In passing data from GenBank to ClustalX, the easiest format to use would appear to be FASTA, in which each block of DNA sequences is preceded by one line of text containing the name of the sequence. This line starts with the greater than symbol (>). For example: >gi gb AY Myioborus albifacies isolate ALB_190_1 NADH dehydrogenase subunit 2 (ND2) gene, partial cds; mitochondrial ATAAACCCCCAAGCAAATCTAATTTTCATCATCAGCCTAATCCTAGGAACTACCATTACCATCTCAAGCA ACCATTGAGTCATAGCCTGAACCGGCCTTGAAATCAACACGCTTGCCATCCTCCCACTAATCTCAAAATC CCATCACCCCCGAGC >gi etc etc. All your sequences of any one gene can be in one file, cut and paste from one file into a new, third file if necessary (or you can load the different files one at a time directly into ClustalX). Before proceeding with the next step, you may want to shorten the name of each sequence, leaving only the minimum necessary information, such as GenBank accession number, species name, specimen number, and the population it came from! (The name of the gene can be part of the name of your new file.) However, try to keep the number of characters in the name limited to 26 or fewer, and try not to use any symbols or characters besides English letters and numbers. Use the underscore symbol ( _ ) instead of spaces in the sequence name. If you have multiple genes, make your sequence names such that if you were to alphabetize the sequences in both genes, the individuals would come out in the same order! This will help you a lot later on! You might think to create names for your sequences such that if you were to automatically put them in alphabetical order (such as by using Mequite) the samples would be grouped by population. II. Multiple sequence alignment using ClustalX Launch ClustalX Select File > Load sequence and navigate to your data file and select it. In ClustalX, verify that you have your expected number of sequences. Choose Alignment > Multiple Alignment > Multiple Alignment Parameters Check the following parameters and note their default values: Gap Opening: Too high = few gaps but also fewer matching bases. Too low = lots of gaps but with more matches. Gap Extention [Extension]: Penalty for additional gaps in a row. Delay Divergent Sequences: Align weird sequences last. DNA Transition Weight: Optionally you can downweight transitions 1 no weighting; 0 transitions = mismatch.
3 BIOL 4211 Lab 2, page 3 of 6 For very similar sequences, use no weighting (weight = 1). Select Alignment > Produce Guide Tree Only and follow instructions. If you have TreeEdit or TreeView on your computer, use it to view your new tree. By default his file will end with.dnd Next select Alignment > Do Alignment from Guide Tree. By default, you new alignment file will end with.aln On my Mac laptop, 50 sequences of 1000 bp each took about a half of a minute. The other option, Do Complete Alignment, took 5 minutes. Notice the low horizontal window at the bottom of the screen. This graph indicates by position the relative quality of the alignment. You can change the relative scale via the menu: Quality > Column Score Parameters. If you do not have any gaps inside any DNA sequences (as opposed to on either end of the sequence), then you can skip item II. below and proceed to part III. III. Effect of alignment parameters. If you have gaps in your alignment, you should check whether changing parameters affects your result. First, try to make fewer gaps of larger size using values such as these: Gap Opening: 50 Gap Extension: 1 Remember to save your new alignment under a new name. Look at the distribution of gaps before closing. Next, try to make lots of gaps of small size: Gap Opening: 5 Gap Extension: 5 (Do not bother setting the extension penalty higher than the gap penalty. Did your alignment change among the three trials? If not, your alignment is robust to variation in alignment parameters. If your alignment changed a lot among trials, which alignment do you prefer and why? Note any regions of the alignment that changed a lot among the three trials. Looking at your preferred alignment (you may have to quit & re-lauch ClustalX, and sometimes you have to run the alignment algorithm over again) and the Quality Score window below, note any regions of your alignment that have a low score. Such regions you may want to exclude from all future analyses. IV. Saving and exporting Your preferred alignment is already saved on the computer, along with zero or more other alignments that all end in.aln These files start with the following line of text: CLUSTAL X (1.83) multiple sequence alignment These files are readable directly by programs such as MEGA, Mequite, MacClade, but not by PAUP*. It is always a good idea to save files in various formats, just to be sure that your data will be readable by the next software you may want to use. In ClustalX, you can export your preferred alignment (or alignments) using File > Save Sequences As... Select NEXUS and OK The resulting file starts with lines of text such as:
4 #NEXUS BEGIN DATA; dimensions ntax=53 nchar=1041; format missing=? symbols="abcdefghiklmnpqrstuvwxyz" interleave datatype=dna gap= -; BIOL 4211 Lab 2, page 4 of 6 matrix This file should be readable by PAUP*, MacClade, Mesquite, MEGA, etc. However, for our purposes, interleave format is not preferred for single-gene data. (Later, if you combine multiple gene sequences in one NEXUS file, you will need the word interleave at that time.) For now, working with one gene at a time, you should open your data (either version) in one of the above programs and then export it again, making sure that the interleave option is not selected. If you have protein-coding sequences, you can just do this re-exporting at the end of step V below. V. Checking amino acid translation. Using MEGA (also MacClade does this, but Mequite does not, as far as I can tell.): Launch MEGA, and click on Click me to activate a data file. Navigate to your file, and click Open. Utilities > Convert to MEGA Format, then in the Select File and Format window select the Data Format box and choose type of format your data was in (either.aln (CLUSTAL) or.nexus (PAUP*, MacClade). Note,.nexus and.nxs refer to the same format, as does.nex Finally, select OK Close the new data window and the old data window and re-open your new.meg file. Select Nucleotide Sequences. Reply Yes or No as to whether the sequence is protein coding. If yes, next select appropriate genetic code. To translate the DNA sequence into the inferred amino acid sequence, click on the button UUC Phe. MEGA automatically tries to identify the correct reading frame by minimizing the number of stop codons. Now check for stop codons! This is important!! I believe MEGA indicates a stop codon with an asterisk (*). For easier visualization, select Display > Color Cells. If your protein-coding gene has stop codons anywhere besides the end, your sequence might be in the reverse complement state. MEGA I don t think makes this conversion, but Mesquite and MacClade do. Option B, MacClade: Of course you can use this program, but I prefer to use free software (although I m still using PAUP*). If you need to check the reverse complement of your DNA sequences to see if this way you see no premature stop codons: Launch the application, Mesquite, select File > Open File and choose either your.nxs file (NEXUS or PAUP* format) or your.aln file (CLUSTAL format), and then select the appropriate format to be read in, if necessary. Select Matrix > Alter/Transform > Other Choices... and select Nucleotide Complement Repeat the above command but chose Reverse Sequence instead.
5 BIOL 4211 Lab 2, page 5 of 6 Now select File > Export... twice, choosing different formats each time, such as FASTA and Old- Fashioned NEXUS, just to be safe, and perhaps adding revcomp to the end of the name. Save data file as well, under a new name. Open one of these new files in MEGA, and follow the instructions above. VI. Final data set. Your final data set should be in NEXUS format and not interleaved. If you have multiple genes, you should both save the genes in separate files, but also you will want to make a master NEXUS file. To make a master file, you will want to put all your individuals in the same order in each data set. Do this by hand or by using the sort (alphabetizing) tool in Mesquite, the tool that looks like a little Mayan pyramid, then re-save or re-export your data. If some individuals are missing from some data sets, you will have to add them in and put a? for every nucleotide site (position). Finally, you will past all your non-interleaved data sets into one giant text file in NEXUS format, one set of aligned sequences BELOW the other. Do not attempt to put all gene sequences of each individual on one line. You will have to change the nchar= parameter at the top of the NEXU file. The new nchar value will be the sum off all the nucleotide sites in all genes in this file. Also, you will have to add the word interleave. Here s an example in which we are missing data from the first gene (COI) for the third individual, and the first individual has no data from the second gene (cyt b). Remember that we can put any comments we want to inside square brackets [ like this ] and they will be ignored by all programs that understand the NEXUS format (e.g., PAUP, dnasp, MrBayes, Mesquite, BioEdit, etc.). Good luck!! #NEXUS BEGIN DATA ; DIMENSIONS NTAX=5 NCHAR=59 [36 plus 23] ; FORMAT interleave DATATYPE=DNA MISSING=? GAP=- ; MATRIX [ [... [ *** THE COI DATA FIRST, NCHAR=36 *** ] azueroen_krl0680_nocytb??????????agaagtctacatcctaatcctgccgg crass_ajc0085_las_cruces_cr TTGGCCACCCGGAAGTATATATTCTTATTCTCCCAG crass_ajc0189_fortuna_nocoi???????????????????????????????????? crass_ajc0209_nusagandi_pa TTGGTCACCCGGAAGTCTACATTCTTATCCTCCCAG crass_ajc0215_campana_pa???????????gaagtctacattcttatcctcccag [ *** THE cyt b DATA BELOW, NCHAR=23 *** ] azueroen_krl0680_nocytb??????????????????????? crass_ajc0085_las_cruces_cr CCTCAGGGCTATTTCTGGCCATA crass_ajc0189_fortuna_nocoi TCTCAGGCCTGTTTCTCGCTATA crass_ajc0209_nusagandi_pa??????actatttctgggccata crass_ajc0215_campana_pa CCTCGGGAGTATTTCTCGCCATA (Lab continues on next page )
6 REPORT: BIOL 4211 Lab 2, page 6 of 6 Remember, the purpose of each lab report is so that by the end of the course each team will be able to prepare its final manuscript very easily. When I write a phylogeography manuscript, I might write the introduction while I m still collecting the data. I definitely start writing the Methods and Results while I am doing the analyses. If you wait until you are done with all the analyses to write the Methods and Results, it is often difficult to find all the details of what you did. So, you can think of each lab report as making some progress on your final manuscript. Some basic information about the data will be repeated on various lab reports, as necessary. Working with your lab partner: Write a very brief summary of your alignment protocol, noting the parameter values you used in your multiple sequence alignment. Note briefly how or why you justify your choice of alignment. Did you check the amino acid translation for premature stop codons? How did you do this? For your Methods section, be sure to always note the version of the software you are using. Also, note the citation. All software packages provide information on the proper way to cite them. Your Results section should include the number of sequences from which genes of which species you have. Mention the total number of positions in your alignment, and mention how many gapped sites you have. Also mention now if you plan to EXCLUDE any sites from your analyses due to skepticism about the alignment in certain regions of the gene. You should start making a Literature Cited section, including the original publication and any other papers associated with data on sequences, specimens, or localities. Also, include the citation for any software you used today. For example, the citation for ClustalX is below. You can cut n paste this reference from the URL mentioned above: Thompson,J.D., Gibson,T.J., Plewniak,F., Jeanmougin,F. and Higgins,D.G. (1997) The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Research, 24: This brief report will be due on the 10 th of February.
CAOS Documentation and Worked Examples. Neil Sarkar, Paul Planet and Rob DeSalle
 CAOS Documentation and Worked Examples Neil Sarkar, Paul Planet and Rob DeSalle Table of Contents 1. Downloading and Installing p-gnome and p-elf 2. Preparing your matrix for p-gnome 3. Running p-gnome
CAOS Documentation and Worked Examples Neil Sarkar, Paul Planet and Rob DeSalle Table of Contents 1. Downloading and Installing p-gnome and p-elf 2. Preparing your matrix for p-gnome 3. Running p-gnome
INTRODUCTION TO BIOINFORMATICS
 Molecular Biology-2019 1 INTRODUCTION TO BIOINFORMATICS In this section, we want to provide a simple introduction to using the web site of the National Center for Biotechnology Information NCBI) to obtain
Molecular Biology-2019 1 INTRODUCTION TO BIOINFORMATICS In this section, we want to provide a simple introduction to using the web site of the National Center for Biotechnology Information NCBI) to obtain
PRINCIPLES OF PHYLOGENETICS Spring 2008 Updated by Nick Matzke. Lab 11: MrBayes Lab
 Integrative Biology 200A University of California, Berkeley PRINCIPLES OF PHYLOGENETICS Spring 2008 Updated by Nick Matzke Lab 11: MrBayes Lab Note: try downloading and installing MrBayes on your own laptop,
Integrative Biology 200A University of California, Berkeley PRINCIPLES OF PHYLOGENETICS Spring 2008 Updated by Nick Matzke Lab 11: MrBayes Lab Note: try downloading and installing MrBayes on your own laptop,
Tour Guide for Windows and Macintosh
 Tour Guide for Windows and Macintosh 2011 Gene Codes Corporation Gene Codes Corporation 775 Technology Drive, Suite 100A, Ann Arbor, MI 48108 USA phone 1.800.497.4939 or 1.734.769.7249 (fax) 1.734.769.7074
Tour Guide for Windows and Macintosh 2011 Gene Codes Corporation Gene Codes Corporation 775 Technology Drive, Suite 100A, Ann Arbor, MI 48108 USA phone 1.800.497.4939 or 1.734.769.7249 (fax) 1.734.769.7074
Page 1.1 Guidelines 2 Requirements JCoDA package Input file formats License. 1.2 Java Installation 3-4 Not required in all cases
 JCoDA and PGI Tutorial Version 1.0 Date 03/16/2010 Page 1.1 Guidelines 2 Requirements JCoDA package Input file formats License 1.2 Java Installation 3-4 Not required in all cases 2.1 dn/ds calculation
JCoDA and PGI Tutorial Version 1.0 Date 03/16/2010 Page 1.1 Guidelines 2 Requirements JCoDA package Input file formats License 1.2 Java Installation 3-4 Not required in all cases 2.1 dn/ds calculation
INTRODUCTION TO BIOINFORMATICS
 Molecular Biology-2017 1 INTRODUCTION TO BIOINFORMATICS In this section, we want to provide a simple introduction to using the web site of the National Center for Biotechnology Information NCBI) to obtain
Molecular Biology-2017 1 INTRODUCTION TO BIOINFORMATICS In this section, we want to provide a simple introduction to using the web site of the National Center for Biotechnology Information NCBI) to obtain
Introduction to Phylogenetics Week 2. Databases and Sequence Formats
 Introduction to Phylogenetics Week 2 Databases and Sequence Formats I. Databases Crucial to bioinformatics The bigger the database, the more comparative research data Requires scientists to upload data
Introduction to Phylogenetics Week 2 Databases and Sequence Formats I. Databases Crucial to bioinformatics The bigger the database, the more comparative research data Requires scientists to upload data
Lab 12: The What the Hell Do I Do with All These Trees Lab
 Integrative Biology 200A University of California, Berkeley Principals of Phylogenetics Spring 2010 Updated by Nick Matzke Lab 12: The What the Hell Do I Do with All These Trees Lab We ve generated a lot
Integrative Biology 200A University of California, Berkeley Principals of Phylogenetics Spring 2010 Updated by Nick Matzke Lab 12: The What the Hell Do I Do with All These Trees Lab We ve generated a lot
Finding Selection in All the Right Places TA Notes and Key Lab 9
 Objectives: Finding Selection in All the Right Places TA Notes and Key Lab 9 1. Use published genome data to look for evidence of selection in individual genes. 2. Understand the need for DNA sequence
Objectives: Finding Selection in All the Right Places TA Notes and Key Lab 9 1. Use published genome data to look for evidence of selection in individual genes. 2. Understand the need for DNA sequence
2. Take a few minutes to look around the site. The goal is to familiarize yourself with a few key components of the NCBI.
 2 Navigating the NCBI Instructions Aim: To become familiar with the resources available at the National Center for Bioinformatics (NCBI) and the search engine Entrez. Instructions: Write the answers to
2 Navigating the NCBI Instructions Aim: To become familiar with the resources available at the National Center for Bioinformatics (NCBI) and the search engine Entrez. Instructions: Write the answers to
Lab 8: Using POY from your desktop and through CIPRES
 Integrative Biology 200A University of California, Berkeley PRINCIPLES OF PHYLOGENETICS Spring 2012 Updated by Michael Landis Lab 8: Using POY from your desktop and through CIPRES In this lab we re going
Integrative Biology 200A University of California, Berkeley PRINCIPLES OF PHYLOGENETICS Spring 2012 Updated by Michael Landis Lab 8: Using POY from your desktop and through CIPRES In this lab we re going
Wilson Leung 01/03/2018 An Introduction to NCBI BLAST. Prerequisites: Detecting and Interpreting Genetic Homology: Lecture Notes on Alignment
 An Introduction to NCBI BLAST Prerequisites: Detecting and Interpreting Genetic Homology: Lecture Notes on Alignment Resources: The BLAST web server is available at https://blast.ncbi.nlm.nih.gov/blast.cgi
An Introduction to NCBI BLAST Prerequisites: Detecting and Interpreting Genetic Homology: Lecture Notes on Alignment Resources: The BLAST web server is available at https://blast.ncbi.nlm.nih.gov/blast.cgi
Tutorial: Resequencing Analysis using Tracks
 : Resequencing Analysis using Tracks September 20, 2013 CLC bio Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 Fax: +45 86 20 12 22 www.clcbio.com support@clcbio.com : Resequencing
: Resequencing Analysis using Tracks September 20, 2013 CLC bio Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 Fax: +45 86 20 12 22 www.clcbio.com support@clcbio.com : Resequencing
Characters and Character matrices
 Introduction Why? How to Cite Publication Support Credits Help FAQ Web Site Simplicity Files Menus Windows Charts Scripts/Macros Modules How Characters Taxa Trees Glossary New Features Characters and Character
Introduction Why? How to Cite Publication Support Credits Help FAQ Web Site Simplicity Files Menus Windows Charts Scripts/Macros Modules How Characters Taxa Trees Glossary New Features Characters and Character
BIOL591: Introduction to Bioinformatics Alignment of pairs of sequences
 BIOL591: Introduction to Bioinformatics Alignment of pairs of sequences Reading in text (Mount Bioinformatics): I must confess that the treatment in Mount of sequence alignment does not seem to me a model
BIOL591: Introduction to Bioinformatics Alignment of pairs of sequences Reading in text (Mount Bioinformatics): I must confess that the treatment in Mount of sequence alignment does not seem to me a model
Lab 06: Support Measures Bootstrap, Jackknife, and Bremer
 Integrative Biology 200, Spring 2014 Principles of Phylogenetics: Systematics University of California, Berkeley Updated by Traci L. Grzymala Lab 06: Support Measures Bootstrap, Jackknife, and Bremer So
Integrative Biology 200, Spring 2014 Principles of Phylogenetics: Systematics University of California, Berkeley Updated by Traci L. Grzymala Lab 06: Support Measures Bootstrap, Jackknife, and Bremer So
"PRINCIPLES OF PHYLOGENETICS" Spring 2008
 Integrative Biology 200A University of California, Berkeley "PRINCIPLES OF PHYLOGENETICS" Spring 2008 Lab 7: Introduction to PAUP* Today we will be learning about some of the basic features of PAUP* (Phylogenetic
Integrative Biology 200A University of California, Berkeley "PRINCIPLES OF PHYLOGENETICS" Spring 2008 Lab 7: Introduction to PAUP* Today we will be learning about some of the basic features of PAUP* (Phylogenetic
Mitochondrial DNA Typing
 Tutorial for Windows and Macintosh Mitochondrial DNA Typing 2007 Gene Codes Corporation Gene Codes Corporation 775 Technology Drive, Ann Arbor, MI 48108 USA 1.800.497.4939 (USA) +1.734.769.7249 (elsewhere)
Tutorial for Windows and Macintosh Mitochondrial DNA Typing 2007 Gene Codes Corporation Gene Codes Corporation 775 Technology Drive, Ann Arbor, MI 48108 USA 1.800.497.4939 (USA) +1.734.769.7249 (elsewhere)
Uploading sequences to GenBank
 A primer for practical phylogenetic data gathering. Uconn EEB3899-007. Spring 2015 Session 5 Uploading sequences to GenBank Rafael Medina (rafael.medina.bry@gmail.com) Yang Liu (yang.liu@uconn.edu) confirmation
A primer for practical phylogenetic data gathering. Uconn EEB3899-007. Spring 2015 Session 5 Uploading sequences to GenBank Rafael Medina (rafael.medina.bry@gmail.com) Yang Liu (yang.liu@uconn.edu) confirmation
Gegenees genome format...7. Gegenees comparisons...8 Creating a fragmented all-all comparison...9 The alignment The analysis...
 User Manual: Gegenees V 1.1.0 What is Gegenees?...1 Version system:...2 What's new...2 Installation:...2 Perspectives...4 The workspace...4 The local database...6 Populate the local database...7 Gegenees
User Manual: Gegenees V 1.1.0 What is Gegenees?...1 Version system:...2 What's new...2 Installation:...2 Perspectives...4 The workspace...4 The local database...6 Populate the local database...7 Gegenees
Lab 13: The What the Hell Do I Do with All These Trees Lab
 Integrative Biology 200A University of California, Berkeley Principals of Phylogenetics Spring 2012 Updated by Michael Landis Lab 13: The What the Hell Do I Do with All These Trees Lab We ve generated
Integrative Biology 200A University of California, Berkeley Principals of Phylogenetics Spring 2012 Updated by Michael Landis Lab 13: The What the Hell Do I Do with All These Trees Lab We ve generated
Release Notes. Version Gene Codes Corporation
 Version 4.10.1 Release Notes 2010 Gene Codes Corporation Gene Codes Corporation 775 Technology Drive, Ann Arbor, MI 48108 USA 1.800.497.4939 (USA) +1.734.769.7249 (elsewhere) +1.734.769.7074 (fax) www.genecodes.com
Version 4.10.1 Release Notes 2010 Gene Codes Corporation Gene Codes Corporation 775 Technology Drive, Ann Arbor, MI 48108 USA 1.800.497.4939 (USA) +1.734.769.7249 (elsewhere) +1.734.769.7074 (fax) www.genecodes.com
Shortest Path Algorithm
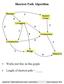 Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
Data Walkthrough: Background
 Data Walkthrough: Background File Types FASTA Files FASTA files are text-based representations of genetic information. They can contain nucleotide or amino acid sequences. For this activity, students will
Data Walkthrough: Background File Types FASTA Files FASTA files are text-based representations of genetic information. They can contain nucleotide or amino acid sequences. For this activity, students will
Mitochondrial DNA Typing
 Tutorial for Windows and Macintosh Mitochondrial DNA Typing 2017 Gene Codes Corporation Gene Codes Corporation 525 Avis Drive, Ann Arbor, MI 48108 USA 1.800.497.4939 (USA) +1.734.769.7249 (elsewhere) +1.734.769.7074
Tutorial for Windows and Macintosh Mitochondrial DNA Typing 2017 Gene Codes Corporation Gene Codes Corporation 525 Avis Drive, Ann Arbor, MI 48108 USA 1.800.497.4939 (USA) +1.734.769.7249 (elsewhere) +1.734.769.7074
Module 1 Artemis. Introduction. Aims IF YOU DON T UNDERSTAND, PLEASE ASK! -1-
 Module 1 Artemis Introduction Artemis is a DNA viewer and annotation tool, free to download and use, written by Kim Rutherford from the Sanger Institute (Rutherford et al., 2000). The program allows the
Module 1 Artemis Introduction Artemis is a DNA viewer and annotation tool, free to download and use, written by Kim Rutherford from the Sanger Institute (Rutherford et al., 2000). The program allows the
Tutorial: De Novo Assembly of Paired Data
 : De Novo Assembly of Paired Data September 20, 2013 CLC bio Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 Fax: +45 86 20 12 22 www.clcbio.com support@clcbio.com : De Novo Assembly
: De Novo Assembly of Paired Data September 20, 2013 CLC bio Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 Fax: +45 86 20 12 22 www.clcbio.com support@clcbio.com : De Novo Assembly
Computational Genomics and Molecular Biology, Fall
 Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
PROTEIN MULTIPLE ALIGNMENT MOTIVATION: BACKGROUND: Marina Sirota
 Marina Sirota MOTIVATION: PROTEIN MULTIPLE ALIGNMENT To study evolution on the genetic level across a wide range of organisms, biologists need accurate tools for multiple sequence alignment of protein
Marina Sirota MOTIVATION: PROTEIN MULTIPLE ALIGNMENT To study evolution on the genetic level across a wide range of organisms, biologists need accurate tools for multiple sequence alignment of protein
Lab 4: Multiple Sequence Alignment (MSA)
 Lab 4: Multiple Sequence Alignment (MSA) The objective of this lab is to become familiar with the features of several multiple alignment and visualization tools, including the data input and output, basic
Lab 4: Multiple Sequence Alignment (MSA) The objective of this lab is to become familiar with the features of several multiple alignment and visualization tools, including the data input and output, basic
CS313 Exercise 4 Cover Page Fall 2017
 CS313 Exercise 4 Cover Page Fall 2017 Due by the start of class on Thursday, October 12, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
CS313 Exercise 4 Cover Page Fall 2017 Due by the start of class on Thursday, October 12, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
Geneious 5.6 Quickstart Manual. Biomatters Ltd
 Geneious 5.6 Quickstart Manual Biomatters Ltd October 15, 2012 2 Introduction This quickstart manual will guide you through the features of Geneious 5.6 s interface and help you orient yourself. You should
Geneious 5.6 Quickstart Manual Biomatters Ltd October 15, 2012 2 Introduction This quickstart manual will guide you through the features of Geneious 5.6 s interface and help you orient yourself. You should
Data Curation Profile Botany / Plant Taxonomy
 Profile Author Sara Rutter Author s Institution University of Hawaii at Manoa Contact srutter@hawaii.edu Researcher(s) Interviewed [withheld], co-pi [withheld], PI Researcher s Institution University of
Profile Author Sara Rutter Author s Institution University of Hawaii at Manoa Contact srutter@hawaii.edu Researcher(s) Interviewed [withheld], co-pi [withheld], PI Researcher s Institution University of
Tutorial: chloroplast genomes
 Tutorial: chloroplast genomes Stacia Wyman Department of Computer Sciences Williams College Williamstown, MA 01267 March 10, 2005 ASSUMPTIONS: You are using Internet Explorer under OS X on the Mac. You
Tutorial: chloroplast genomes Stacia Wyman Department of Computer Sciences Williams College Williamstown, MA 01267 March 10, 2005 ASSUMPTIONS: You are using Internet Explorer under OS X on the Mac. You
DNA sequences obtained in section were assembled and edited using DNA
 Sequetyper DNA sequences obtained in section 4.4.1.3 were assembled and edited using DNA Baser Sequence Assembler v4 (www.dnabaser.com). The consensus sequences were used to interrogate the GenBank database
Sequetyper DNA sequences obtained in section 4.4.1.3 were assembled and edited using DNA Baser Sequence Assembler v4 (www.dnabaser.com). The consensus sequences were used to interrogate the GenBank database
705 Getting Content to Learners on the Cheap Using SharePoint. Jamie Lewis, Archstone
 705 Getting Content to Learners on the Cheap Using SharePoint Jamie Lewis, Archstone Tips and Tricks from Getting Content to Learners on the Cheap Using SharePoint People and Groups You can customize permissions
705 Getting Content to Learners on the Cheap Using SharePoint Jamie Lewis, Archstone Tips and Tricks from Getting Content to Learners on the Cheap Using SharePoint People and Groups You can customize permissions
3.4 Multiple sequence alignment
 3.4 Multiple sequence alignment Why produce a multiple sequence alignment? Using more than two sequences results in a more convincing alignment by revealing conserved regions in ALL of the sequences Aligned
3.4 Multiple sequence alignment Why produce a multiple sequence alignment? Using more than two sequences results in a more convincing alignment by revealing conserved regions in ALL of the sequences Aligned
Tips and Guidance for Analyzing Data. Executive Summary
 Tips and Guidance for Analyzing Data Executive Summary This document has information and suggestions about three things: 1) how to quickly do a preliminary analysis of time-series data; 2) key things to
Tips and Guidance for Analyzing Data Executive Summary This document has information and suggestions about three things: 1) how to quickly do a preliminary analysis of time-series data; 2) key things to
FASTA. Besides that, FASTA package provides SSEARCH, an implementation of the optimal Smith- Waterman algorithm.
 FASTA INTRODUCTION Definition (by David J. Lipman and William R. Pearson in 1985) - Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence
FASTA INTRODUCTION Definition (by David J. Lipman and William R. Pearson in 1985) - Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence
BCRANK: predicting binding site consensus from ranked DNA sequences
 BCRANK: predicting binding site consensus from ranked DNA sequences Adam Ameur October 30, 2017 1 Introduction This document describes the BCRANK R package. BCRANK[1] is a method that takes a ranked list
BCRANK: predicting binding site consensus from ranked DNA sequences Adam Ameur October 30, 2017 1 Introduction This document describes the BCRANK R package. BCRANK[1] is a method that takes a ranked list
Biology 345: Biometry Fall 2005 SONOMA STATE UNIVERSITY Lab Exercise 2 Working with data in Excel and exporting to JMP Introduction
 Biology 345: Biometry Fall 2005 SONOMA STATE UNIVERSITY Lab Exercise 2 Working with data in Excel and exporting to JMP Introduction In this exercise, we will learn how to reorganize and reformat a data
Biology 345: Biometry Fall 2005 SONOMA STATE UNIVERSITY Lab Exercise 2 Working with data in Excel and exporting to JMP Introduction In this exercise, we will learn how to reorganize and reformat a data
TUTORIAL FOR IMPORTING OTTAWA FIRE HYDRANT PARKING VIOLATION DATA INTO MYSQL
 TUTORIAL FOR IMPORTING OTTAWA FIRE HYDRANT PARKING VIOLATION DATA INTO MYSQL We have spent the first part of the course learning Excel: importing files, cleaning, sorting, filtering, pivot tables and exporting
TUTORIAL FOR IMPORTING OTTAWA FIRE HYDRANT PARKING VIOLATION DATA INTO MYSQL We have spent the first part of the course learning Excel: importing files, cleaning, sorting, filtering, pivot tables and exporting
Tutorial 1: Exploring the UCSC Genome Browser
 Last updated: May 12, 2011 Tutorial 1: Exploring the UCSC Genome Browser Open the homepage of the UCSC Genome Browser at: http://genome.ucsc.edu/ In the blue bar at the top, click on the Genomes link.
Last updated: May 12, 2011 Tutorial 1: Exploring the UCSC Genome Browser Open the homepage of the UCSC Genome Browser at: http://genome.ucsc.edu/ In the blue bar at the top, click on the Genomes link.
(Refer Slide Time 6:48)
 Digital Circuits and Systems Prof. S. Srinivasan Department of Electrical Engineering Indian Institute of Technology Madras Lecture - 8 Karnaugh Map Minimization using Maxterms We have been taking about
Digital Circuits and Systems Prof. S. Srinivasan Department of Electrical Engineering Indian Institute of Technology Madras Lecture - 8 Karnaugh Map Minimization using Maxterms We have been taking about
Chapter 2 The SAS Environment
 Chapter 2 The SAS Environment Abstract In this chapter, we begin to become familiar with the basic SAS working environment. We introduce the basic 3-screen layout, how to navigate the SAS Explorer window,
Chapter 2 The SAS Environment Abstract In this chapter, we begin to become familiar with the basic SAS working environment. We introduce the basic 3-screen layout, how to navigate the SAS Explorer window,
1 Introduction to Using Excel Spreadsheets
 Survey of Math: Excel Spreadsheet Guide (for Excel 2007) Page 1 of 6 1 Introduction to Using Excel Spreadsheets This section of the guide is based on the file (a faux grade sheet created for messing with)
Survey of Math: Excel Spreadsheet Guide (for Excel 2007) Page 1 of 6 1 Introduction to Using Excel Spreadsheets This section of the guide is based on the file (a faux grade sheet created for messing with)
Lab 07: Maximum Likelihood Model Selection and RAxML Using CIPRES
 Integrative Biology 200, Spring 2014 Principles of Phylogenetics: Systematics University of California, Berkeley Updated by Traci L. Grzymala Lab 07: Maximum Likelihood Model Selection and RAxML Using
Integrative Biology 200, Spring 2014 Principles of Phylogenetics: Systematics University of California, Berkeley Updated by Traci L. Grzymala Lab 07: Maximum Likelihood Model Selection and RAxML Using
Principles of Bioinformatics. BIO540/STA569/CSI660 Fall 2010
 Principles of Bioinformatics BIO540/STA569/CSI660 Fall 2010 Lecture 11 Multiple Sequence Alignment I Administrivia Administrivia The midterm examination will be Monday, October 18 th, in class. Closed
Principles of Bioinformatics BIO540/STA569/CSI660 Fall 2010 Lecture 11 Multiple Sequence Alignment I Administrivia Administrivia The midterm examination will be Monday, October 18 th, in class. Closed
Overview. Dataset: testpos DNA: CCCATGGTCGGGGGGGGGGAGTCCATAACCC Num exons: 2 strand: + RNA (from file): AUGGUCAGUCCAUAA peptide (from file): MVSP*
 Overview In this homework, we will write a program that will print the peptide (a string of amino acids) from four pieces of information: A DNA sequence (a string). The strand the gene appears on (a string).
Overview In this homework, we will write a program that will print the peptide (a string of amino acids) from four pieces of information: A DNA sequence (a string). The strand the gene appears on (a string).
INTRODUCTION TO CONSED
 INTRODUCTION TO CONSED OVERVIEW: Consed is a program that can be used to visually assemble and analyze sequence data. This introduction will take you through the basics of opening and operating within
INTRODUCTION TO CONSED OVERVIEW: Consed is a program that can be used to visually assemble and analyze sequence data. This introduction will take you through the basics of opening and operating within
Matlab notes Matlab is a matrix-based, high-performance language for technical computing It integrates computation, visualisation and programming usin
 Matlab notes Matlab is a matrix-based, high-performance language for technical computing It integrates computation, visualisation and programming using familiar mathematical notation The name Matlab stands
Matlab notes Matlab is a matrix-based, high-performance language for technical computing It integrates computation, visualisation and programming using familiar mathematical notation The name Matlab stands
Week - 01 Lecture - 04 Downloading and installing Python
 Programming, Data Structures and Algorithms in Python Prof. Madhavan Mukund Department of Computer Science and Engineering Indian Institute of Technology, Madras Week - 01 Lecture - 04 Downloading and
Programming, Data Structures and Algorithms in Python Prof. Madhavan Mukund Department of Computer Science and Engineering Indian Institute of Technology, Madras Week - 01 Lecture - 04 Downloading and
MacVector for Mac OS X
 MacVector 10.6 for Mac OS X System Requirements MacVector 10.6 runs on any PowerPC or Intel Macintosh running Mac OS X 10.4 or higher. It is a Universal Binary, meaning that it runs natively on both PowerPC
MacVector 10.6 for Mac OS X System Requirements MacVector 10.6 runs on any PowerPC or Intel Macintosh running Mac OS X 10.4 or higher. It is a Universal Binary, meaning that it runs natively on both PowerPC
Creating Reports in Access 2007 Table of Contents GUIDE TO DESIGNING REPORTS... 3 DECIDE HOW TO LAY OUT YOUR REPORT... 3 MAKE A SKETCH OF YOUR
 Creating Reports in Access 2007 Table of Contents GUIDE TO DESIGNING REPORTS... 3 DECIDE HOW TO LAY OUT YOUR REPORT... 3 MAKE A SKETCH OF YOUR REPORT... 3 DECIDE WHICH DATA TO PUT IN EACH REPORT SECTION...
Creating Reports in Access 2007 Table of Contents GUIDE TO DESIGNING REPORTS... 3 DECIDE HOW TO LAY OUT YOUR REPORT... 3 MAKE A SKETCH OF YOUR REPORT... 3 DECIDE WHICH DATA TO PUT IN EACH REPORT SECTION...
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014
 Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
Salvador Capella-Gutiérrez, Jose M. Silla-Martínez and Toni Gabaldón
 trimal: a tool for automated alignment trimming in large-scale phylogenetics analyses Salvador Capella-Gutiérrez, Jose M. Silla-Martínez and Toni Gabaldón Version 1.2b Index of contents 1. General features
trimal: a tool for automated alignment trimming in large-scale phylogenetics analyses Salvador Capella-Gutiérrez, Jose M. Silla-Martínez and Toni Gabaldón Version 1.2b Index of contents 1. General features
Tutorial for Windows and Macintosh SNP Hunting
 Tutorial for Windows and Macintosh SNP Hunting 2017 Gene Codes Corporation Gene Codes Corporation 525 Avis Drive, Ann Arbor, MI 48108 USA 1.800.497.4939 (USA) +1.734.769.7249 (elsewhere) +1.734.769.7074
Tutorial for Windows and Macintosh SNP Hunting 2017 Gene Codes Corporation Gene Codes Corporation 525 Avis Drive, Ann Arbor, MI 48108 USA 1.800.497.4939 (USA) +1.734.769.7249 (elsewhere) +1.734.769.7074
Tutorial 4 BLAST Searching the CHO Genome
 Tutorial 4 BLAST Searching the CHO Genome Accessing the CHO Genome BLAST Tool The CHO BLAST server can be accessed by clicking on the BLAST button on the home page or by selecting BLAST from the menu bar
Tutorial 4 BLAST Searching the CHO Genome Accessing the CHO Genome BLAST Tool The CHO BLAST server can be accessed by clicking on the BLAST button on the home page or by selecting BLAST from the menu bar
EXCEL BASICS: MICROSOFT OFFICE 2007
 EXCEL BASICS: MICROSOFT OFFICE 2007 GETTING STARTED PAGE 02 Prerequisites What You Will Learn USING MICROSOFT EXCEL PAGE 03 Opening Microsoft Excel Microsoft Excel Features Keyboard Review Pointer Shapes
EXCEL BASICS: MICROSOFT OFFICE 2007 GETTING STARTED PAGE 02 Prerequisites What You Will Learn USING MICROSOFT EXCEL PAGE 03 Opening Microsoft Excel Microsoft Excel Features Keyboard Review Pointer Shapes
Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA.
 Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA. Fasta is used to compare a protein or DNA sequence to all of the
Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA. Fasta is used to compare a protein or DNA sequence to all of the
Simple sets of data can be expressed in a simple table, much like a
 Chapter 1: Building Master and Detail Pages In This Chapter Developing master and detail pages at the same time Building your master and detail pages separately Putting together master and detail pages
Chapter 1: Building Master and Detail Pages In This Chapter Developing master and detail pages at the same time Building your master and detail pages separately Putting together master and detail pages
CLC Sequence Viewer 6.5 Windows, Mac OS X and Linux
 CLC Sequence Viewer Manual for CLC Sequence Viewer 6.5 Windows, Mac OS X and Linux January 26, 2011 This software is for research purposes only. CLC bio Finlandsgade 10-12 DK-8200 Aarhus N Denmark Contents
CLC Sequence Viewer Manual for CLC Sequence Viewer 6.5 Windows, Mac OS X and Linux January 26, 2011 This software is for research purposes only. CLC bio Finlandsgade 10-12 DK-8200 Aarhus N Denmark Contents
2) NCBI BLAST tutorial This is a users guide written by the education department at NCBI.
 Web resources -- Tour. page 1 of 8 This is a guided tour. Any homework is separate. In fact, this exercise is used for multiple classes and is publicly available to everyone. The entire tour will take
Web resources -- Tour. page 1 of 8 This is a guided tour. Any homework is separate. In fact, this exercise is used for multiple classes and is publicly available to everyone. The entire tour will take
Introduction to Bioinformatics Problem Set 3: Genome Sequencing
 Introduction to Bioinformatics Problem Set 3: Genome Sequencing 1. Assemble a sequence with your bare hands! You are trying to determine the DNA sequence of a very (very) small plasmids, which you estimate
Introduction to Bioinformatics Problem Set 3: Genome Sequencing 1. Assemble a sequence with your bare hands! You are trying to determine the DNA sequence of a very (very) small plasmids, which you estimate
Relationship Estimator
 This is a small program that is intended to make the DNA Prediction Chart Spreadsheet a bit easier to use. It is based entirely on the data in this spreadsheet plus some interpolation of missing values.
This is a small program that is intended to make the DNA Prediction Chart Spreadsheet a bit easier to use. It is based entirely on the data in this spreadsheet plus some interpolation of missing values.
Programming assignment for the course Sequence Analysis (2006)
 Programming assignment for the course Sequence Analysis (2006) Original text by John W. Romein, adapted by Bart van Houte (bart@cs.vu.nl) Introduction Please note: This assignment is only obligatory for
Programming assignment for the course Sequence Analysis (2006) Original text by John W. Romein, adapted by Bart van Houte (bart@cs.vu.nl) Introduction Please note: This assignment is only obligatory for
(Refer Slide Time 3:31)
 Digital Circuits and Systems Prof. S. Srinivasan Department of Electrical Engineering Indian Institute of Technology Madras Lecture - 5 Logic Simplification In the last lecture we talked about logic functions
Digital Circuits and Systems Prof. S. Srinivasan Department of Electrical Engineering Indian Institute of Technology Madras Lecture - 5 Logic Simplification In the last lecture we talked about logic functions
ChIP-seq Analysis Practical
 ChIP-seq Analysis Practical Vladimir Teif (vteif@essex.ac.uk) An updated version of this document will be available at http://generegulation.info/index.php/teaching In this practical we will learn how
ChIP-seq Analysis Practical Vladimir Teif (vteif@essex.ac.uk) An updated version of this document will be available at http://generegulation.info/index.php/teaching In this practical we will learn how
Biocomputing II Coursework guidance
 Biocomputing II Coursework guidance I refer to the database layer as DB, the middle (business logic) layer as BL and the front end graphical interface with CGI scripts as (FE). Standardized file headers
Biocomputing II Coursework guidance I refer to the database layer as DB, the middle (business logic) layer as BL and the front end graphical interface with CGI scripts as (FE). Standardized file headers
Programming Project 1: Sequence Alignment
 Programming Project 1: Sequence Alignment CS 181, Fall 2017 Out: Sept. 19 Due: Oct. 3, 11:59 PM 1 Task You must implement the following three alignment algorithms you ve learned in class: 1. Global Alignment
Programming Project 1: Sequence Alignment CS 181, Fall 2017 Out: Sept. 19 Due: Oct. 3, 11:59 PM 1 Task You must implement the following three alignment algorithms you ve learned in class: 1. Global Alignment
Essential Skills for Bioinformatics: Unix/Linux
 Essential Skills for Bioinformatics: Unix/Linux SHELL SCRIPTING Overview Bash, the shell we have used interactively in this course, is a full-fledged scripting language. Unlike Python, Bash is not a general-purpose
Essential Skills for Bioinformatics: Unix/Linux SHELL SCRIPTING Overview Bash, the shell we have used interactively in this course, is a full-fledged scripting language. Unlike Python, Bash is not a general-purpose
TreeCollapseCL 4 Emma Hodcroft Andrew Leigh Brown Group Institute of Evolutionary Biology University of Edinburgh
 TreeCollapseCL 4 Emma Hodcroft Andrew Leigh Brown Group Institute of Evolutionary Biology University of Edinburgh 2011-2015 This command-line Java program takes in Nexus/Newick-style phylogenetic tree
TreeCollapseCL 4 Emma Hodcroft Andrew Leigh Brown Group Institute of Evolutionary Biology University of Edinburgh 2011-2015 This command-line Java program takes in Nexus/Newick-style phylogenetic tree
Working with Mailbox Manager
 Working with Mailbox Manager A user guide for Mailbox Manager supporting the Message Storage Server component of the Avaya S3400 Message Server Mailbox Manager Version 5.0 February 2003 Copyright 2003
Working with Mailbox Manager A user guide for Mailbox Manager supporting the Message Storage Server component of the Avaya S3400 Message Server Mailbox Manager Version 5.0 February 2003 Copyright 2003
TUTORIAL FOR IMPORTING OTTAWA FIRE HYDRANT PARKING VIOLATION DATA INTO MYSQL
 TUTORIAL FOR IMPORTING OTTAWA FIRE HYDRANT PARKING VIOLATION DATA INTO MYSQL We have spent the first part of the course learning Excel: importing files, cleaning, sorting, filtering, pivot tables and exporting
TUTORIAL FOR IMPORTING OTTAWA FIRE HYDRANT PARKING VIOLATION DATA INTO MYSQL We have spent the first part of the course learning Excel: importing files, cleaning, sorting, filtering, pivot tables and exporting
Karlen Communications Citations and Bibliography in Word. Karen McCall, M.Ed.
 Karlen Communications Citations and Bibliography in Word Karen McCall, M.Ed. Table of Contents Introduction... 3 Choose a Document Style Guide... 3 Citations... 4 Manage Sources Master List Dialog... 5
Karlen Communications Citations and Bibliography in Word Karen McCall, M.Ed. Table of Contents Introduction... 3 Choose a Document Style Guide... 3 Citations... 4 Manage Sources Master List Dialog... 5
Intro to Programming. Unit 7. What is Programming? What is Programming? Intro to Programming
 Intro to Programming Unit 7 Intro to Programming 1 What is Programming? 1. Programming Languages 2. Markup vs. Programming 1. Introduction 2. Print Statement 3. Strings 4. Types and Values 5. Math Externals
Intro to Programming Unit 7 Intro to Programming 1 What is Programming? 1. Programming Languages 2. Markup vs. Programming 1. Introduction 2. Print Statement 3. Strings 4. Types and Values 5. Math Externals
Database Searching Using BLAST
 Mahidol University Objectives SCMI512 Molecular Sequence Analysis Database Searching Using BLAST Lecture 2B After class, students should be able to: explain the FASTA algorithm for database searching explain
Mahidol University Objectives SCMI512 Molecular Sequence Analysis Database Searching Using BLAST Lecture 2B After class, students should be able to: explain the FASTA algorithm for database searching explain
Comparative Sequencing
 Tutorial for Windows and Macintosh Comparative Sequencing 2017 Gene Codes Corporation Gene Codes Corporation 525 Avis Drive, Ann Arbor, MI 48108 USA 1.800.497.4939 (USA) +1.734.769.7249 (elsewhere) +1.734.769.7074
Tutorial for Windows and Macintosh Comparative Sequencing 2017 Gene Codes Corporation Gene Codes Corporation 525 Avis Drive, Ann Arbor, MI 48108 USA 1.800.497.4939 (USA) +1.734.769.7249 (elsewhere) +1.734.769.7074
MAXQDA and Chapter 9 Coding Schemes
 MAXQDA and Chapter 9 Coding Schemes Chapter 9 discusses how the structures of coding schemes, alternate groupings are key to moving forward with analysis. The nature and structures of the coding scheme
MAXQDA and Chapter 9 Coding Schemes Chapter 9 discusses how the structures of coding schemes, alternate groupings are key to moving forward with analysis. The nature and structures of the coding scheme
Nexus Files: Total files errors: 62 Possible parser errors: 9 Server issues* 1. *Not reproducible
 Nexus Files: Error type 1. File type errors: 13 examples...2 Error type 2. Parser problems: 4 examples...3 Error type 3. Misspellings, misplaced semi colons, etc: 12 examples...5 Error Type 4. Taxon naming
Nexus Files: Error type 1. File type errors: 13 examples...2 Error type 2. Parser problems: 4 examples...3 Error type 3. Misspellings, misplaced semi colons, etc: 12 examples...5 Error Type 4. Taxon naming
Lecture Overview. Sequence search & alignment. Searching sequence databases. Sequence Alignment & Search. Goals: Motivations:
 Lecture Overview Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating
Lecture Overview Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating
EXCEL BASICS: MICROSOFT OFFICE 2010
 EXCEL BASICS: MICROSOFT OFFICE 2010 GETTING STARTED PAGE 02 Prerequisites What You Will Learn USING MICROSOFT EXCEL PAGE 03 Opening Microsoft Excel Microsoft Excel Features Keyboard Review Pointer Shapes
EXCEL BASICS: MICROSOFT OFFICE 2010 GETTING STARTED PAGE 02 Prerequisites What You Will Learn USING MICROSOFT EXCEL PAGE 03 Opening Microsoft Excel Microsoft Excel Features Keyboard Review Pointer Shapes
COMPARATIVE MICROBIAL GENOMICS ANALYSIS WORKSHOP. Exercise 2: Predicting Protein-encoding Genes, BlastMatrix, BlastAtlas
 COMPARATIVE MICROBIAL GENOMICS ANALYSIS WORKSHOP Exercise 2: Predicting Protein-encoding Genes, BlastMatrix, BlastAtlas First of all connect once again to the CBS system: Open ssh shell client. Press Quick
COMPARATIVE MICROBIAL GENOMICS ANALYSIS WORKSHOP Exercise 2: Predicting Protein-encoding Genes, BlastMatrix, BlastAtlas First of all connect once again to the CBS system: Open ssh shell client. Press Quick
You can use Dreamweaver to build master and detail Web pages, which
 Chapter 1: Building Master and Detail Pages In This Chapter Developing master and detail pages at the same time Building your master and detail pages separately Putting together master and detail pages
Chapter 1: Building Master and Detail Pages In This Chapter Developing master and detail pages at the same time Building your master and detail pages separately Putting together master and detail pages
addition + =5+C2 adds 5 to the value in cell C2 multiplication * =F6*0.12 multiplies the value in cell F6 by 0.12
 BIOL 001 Excel Quick Reference Guide (Office 2010) For your lab report and some of your assignments, you will need to use Excel to analyze your data and/or generate graphs. This guide highlights specific
BIOL 001 Excel Quick Reference Guide (Office 2010) For your lab report and some of your assignments, you will need to use Excel to analyze your data and/or generate graphs. This guide highlights specific
Copyright. (1) Copyright. (2) Trademark
 User Guide Copyright (1) Copyright Copyright 2006, Accelrys Software Inc. All rights reserved. The Accelrys name and logo are registered trademarks of Accelrys Software Inc. This product (software and/or
User Guide Copyright (1) Copyright Copyright 2006, Accelrys Software Inc. All rights reserved. The Accelrys name and logo are registered trademarks of Accelrys Software Inc. This product (software and/or
Sequence Alignment & Search
 Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating the first version
Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating the first version
Basic Tree Building With PAUP
 Phylogenetic Tree Building Objectives 1. Understand the principles of phylogenetic thinking. 2. Be able to develop and test a phylogenetic hypothesis. 3. Be able to interpret a phylogenetic tree. Overview
Phylogenetic Tree Building Objectives 1. Understand the principles of phylogenetic thinking. 2. Be able to develop and test a phylogenetic hypothesis. 3. Be able to interpret a phylogenetic tree. Overview
Tutorial for Windows and Macintosh SNP Hunting
 Tutorial for Windows and Macintosh SNP Hunting 2010 Gene Codes Corporation Gene Codes Corporation 775 Technology Drive, Ann Arbor, MI 48108 USA 1.800.497.4939 (USA) +1.734.769.7249 (elsewhere) +1.734.769.7074
Tutorial for Windows and Macintosh SNP Hunting 2010 Gene Codes Corporation Gene Codes Corporation 775 Technology Drive, Ann Arbor, MI 48108 USA 1.800.497.4939 (USA) +1.734.769.7249 (elsewhere) +1.734.769.7074
Bioinformatics explained: BLAST. March 8, 2007
 Bioinformatics Explained Bioinformatics explained: BLAST March 8, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com Bioinformatics
Bioinformatics Explained Bioinformatics explained: BLAST March 8, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com Bioinformatics
Brief review from last class
 Sequence Alignment Brief review from last class DNA is has direction, we will use only one (5 -> 3 ) and generate the opposite strand as needed. DNA is a 3D object (see lecture 1) but we will model it
Sequence Alignment Brief review from last class DNA is has direction, we will use only one (5 -> 3 ) and generate the opposite strand as needed. DNA is a 3D object (see lecture 1) but we will model it
Advanced UCSC Browser Functions
 Advanced UCSC Browser Functions Dr. Thomas Randall tarandal@email.unc.edu bioinformatics.unc.edu UCSC Browser: genome.ucsc.edu Overview Custom Tracks adding your own datasets Utilities custom tools for
Advanced UCSC Browser Functions Dr. Thomas Randall tarandal@email.unc.edu bioinformatics.unc.edu UCSC Browser: genome.ucsc.edu Overview Custom Tracks adding your own datasets Utilities custom tools for
Heuristic methods for pairwise alignment:
 Bi03c_1 Unit 03c: Heuristic methods for pairwise alignment: k-tuple-methods k-tuple-methods for alignment of pairs of sequences Bi03c_2 dynamic programming is too slow for large databases Use heuristic
Bi03c_1 Unit 03c: Heuristic methods for pairwise alignment: k-tuple-methods k-tuple-methods for alignment of pairs of sequences Bi03c_2 dynamic programming is too slow for large databases Use heuristic
HORIZONTAL GENE TRANSFER DETECTION
 HORIZONTAL GENE TRANSFER DETECTION Sequenzanalyse und Genomik (Modul 10-202-2207) Alejandro Nabor Lozada-Chávez Before start, the user must create a new folder or directory (WORKING DIRECTORY) for all
HORIZONTAL GENE TRANSFER DETECTION Sequenzanalyse und Genomik (Modul 10-202-2207) Alejandro Nabor Lozada-Chávez Before start, the user must create a new folder or directory (WORKING DIRECTORY) for all
Data Structures and Algorithms Dr. Naveen Garg Department of Computer Science and Engineering Indian Institute of Technology, Delhi.
 Data Structures and Algorithms Dr. Naveen Garg Department of Computer Science and Engineering Indian Institute of Technology, Delhi Lecture 18 Tries Today we are going to be talking about another data
Data Structures and Algorithms Dr. Naveen Garg Department of Computer Science and Engineering Indian Institute of Technology, Delhi Lecture 18 Tries Today we are going to be talking about another data
Lesson 13 Molecular Evolution
 Sequence Analysis Spring 2000 Dr. Richard Friedman (212)305-6901 (76901) friedman@cuccfa.ccc.columbia.edu 130BB Lesson 13 Molecular Evolution In this class we learn how to draw molecular evolutionary trees
Sequence Analysis Spring 2000 Dr. Richard Friedman (212)305-6901 (76901) friedman@cuccfa.ccc.columbia.edu 130BB Lesson 13 Molecular Evolution In this class we learn how to draw molecular evolutionary trees
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 04: Variations of sequence alignments http://www.pitt.edu/~mcs2/teaching/biocomp/tutorials/global.html Slides adapted from Dr. Shaojie Zhang (University
EECS730: Introduction to Bioinformatics Lecture 04: Variations of sequence alignments http://www.pitt.edu/~mcs2/teaching/biocomp/tutorials/global.html Slides adapted from Dr. Shaojie Zhang (University
Distance Methods. "PRINCIPLES OF PHYLOGENETICS" Spring 2006
 Integrative Biology 200A University of California, Berkeley "PRINCIPLES OF PHYLOGENETICS" Spring 2006 Distance Methods Due at the end of class: - Distance matrices and trees for two different distance
Integrative Biology 200A University of California, Berkeley "PRINCIPLES OF PHYLOGENETICS" Spring 2006 Distance Methods Due at the end of class: - Distance matrices and trees for two different distance
Sequence alignment algorithms
 Sequence alignment algorithms Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 23 rd 27 After this lecture, you can decide when to use local and global sequence alignments
Sequence alignment algorithms Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 23 rd 27 After this lecture, you can decide when to use local and global sequence alignments
3. Open Vector NTI 9 (note 2) from desktop. A three pane window appears.
 SOP: SP043.. Recombinant Plasmid Map Design Vector NTI Materials and Reagents: 1. Dell Dimension XPS T450 Room C210 2. Vector NTI 9 application, on desktop 3. Tuberculist database open in Internet Explorer
SOP: SP043.. Recombinant Plasmid Map Design Vector NTI Materials and Reagents: 1. Dell Dimension XPS T450 Room C210 2. Vector NTI 9 application, on desktop 3. Tuberculist database open in Internet Explorer
