User Guide PD map 01 September 2017
|
|
|
- Iris Hamilton
- 5 years ago
- Views:
Transcription
1 User Guide PD map 01 September 2017
2 Contents 1. How to start Introduction Pathways and Compartments view Biological overview Submaps How to browse and search Browse Search: GENERIC Search: How to use drug interface Search: How to use chemical interaction interface Search: How to use mirna interface How to display implemented data sets How to display custom data sets on the PD map Upload and highlight a list of elements Upload of a data file: Data preparation Upload of a data file: Data upload How to provide comments and feedback How to extract information from the PD map Export function Select mode Help /09/2017 PD map user guide 1
3 1. How to start 1.1. Introduction The Parkinson s disease (PD) map ( displays disease-related molecular and biochemical pathways within a neuronal cell system, focused on a dopaminergic neuron, adjacent cells (i.e., astrocytes, microglia) and cellular structures (i.e., blood-brain-barrier, synaptic terminals) integrates more than 1300 manually curated publications and over 250 entries from public databases such as Reactome or KEGG (as of Sep, 2017) has an intuitive Google Maps display that enables easy navigation is composed of species (here: elements, entities) such as genes, proteins, molecules, complexes, annotated with relevant database entries reactions (or interactions) describing relations between species, annotated with relevant literature and/or database entries LEGEND button (right upper side, blue panel) provides details on the format of elements and interactions 1.2. Pathways and Compartments view Pathways and compartments is the default view providing an overview on the content. Fig. 1: Pathways (grey) and compartments (coloured) view of the PD map provides hierarchical information on the cellular structures, processes and complexes click in a compartment provides additional information in the left panel 01/09/2017 PD map user guide 2
4 zooming out/in provides higher level compartment view or a more detailed view on molecular species and interaction, respectively (see also chapter 2) select Network for a pure network layout select Empty for a colourless network view, useful within data sets overlay on the PD map (see also chapter 3 and 4) Biological overview To get a view of the biological systems that are modelled in the PD map click on the button SHOW OVERVIEW (Fig. 2; red rectangle). Fig. 2: Biological overview of the PD map Each figure contains three standard buttons in the left upper corner ( home, back, to map ) to navigate through the pictures or to go to the respective area in the PD map Clicking on the marked areas leads you to more detailed pictures of the surrounded area or if no further downstream picture is available it links to the respective areas in the PD map. Important for tablet and smart phone use: Do not tip to the links in the case you have enlarged the biological view. The links to the PD map only work proper when the biological view has his original size. 01/09/2017 PD map user guide 3
5 1.4. Submaps To maintain overall structure of the PD map, some parts of the main network where displayed in separate, more detailed submaps. The submaps are fully operational; i.e., search functions and data display works simultaneously on the whole PD map and on the submaps. Enter submaps by clicking the "SUBMAPS" tab. Click on magnifier to enter the receptive submap. Alternatively, you may click on dedicated elements in the map and open it from the left side information panel. 2. How to browse and search 2.1. Browse An intuitive Google Maps" display enables easy navigation through the PD map. Clicking on any element in the PD map (e.g., proteins, molecules or phenotypes) provides detailed annotations and links to external databases, which are displayed in the left panel (Fig. 3), presumably SERCH GENERIC is active. Otherwise, the information is visible on the bottom of the left panel (see Fig. 11: CREPPB protein). Annotations Fig. 3: Screenshot showing annotations of the protein: MARK2 (MAP/microtubule affinity-regulating kinase 2) 01/09/2017 PD map user guide 4
6 By clicking on the bubble pinned to the element (Fig. 4A red arrow), additional information is displayed in a pop up box. For genes, mirnas and proteins a special pop up box enables the search for interacting drugs, chemicals and mirnas. First click on the element, then on the coloured number above and then the related checkbox ( Show all ) (Fig. 4). For more information on the different tools for drugs, chemicals and mirnas interaction see related chapters 2.3, 2.4, and 2.5. (A) (B) Fig. 4: Additional information and functions in relation to MARK2 displayed as pop up. Clicking on a connection line between elements (Fig. 5) provides detailed information on the underlying reaction (coloured in blue), displayed in the left panel. This includes information on reactants and links to the underlying sources (i.e., PubMed, Reactome or KEGG). Annotations Fig. 5: Screenshot showing annotations of reaction re4026: MUL1 mediates ubiquitination of MFN2. Activated reaction is coloured in blue. 01/09/2017 PD map user guide 5
7 2.2. Search: GENERIC The "GENERIC SEARCH" function is located in the upper part of the left panel. All elements (e.g., genes, proteins, small molecules, complexes) are searchable. Search function also works with synonyms. Search for multiple elements: separate names by semicolon (see Fig. 6). Click on PERFECT MATCH reduces the search to those elements that exactly correspond to the input. The location of the searched species is displayed on the PD map by coloured circles. Results from searches for multiple elements are displayed with different colours. The left panel provides annotations on the elements and the reaction types including links to external databases. Search for reactions: type in: reaction: rexxxx (XXXX = reaction number). Search for publications and related interaction: type pubmed:xxxxxxx (where XXXXXXX = PubMed ID). Search for publications also works with the specific publication list. Under INFO you find the link to the list of the current publications curated into the PD map. This list is fully searchable and contains links back to the PD map. Clicking on CLEAR (right upper corner) will remove all search results from the PD map Fig. 6: Search for Mitofusin2 (MFN2) and Mulan1 (MUL1): All locations of MFN2 (red circle) and MUL1 (blue circle) are displayed in the PD map. Left panel shows annotations. 01/09/2017 PD map user guide 6
8 2.3. Search: How to use drug interface The DRUG interface allows to search for potential drug targets within the PD map. Activate the DRUG tab under SEARCH. Type drug names (synonyms or brand names also work) in the search field. Separate multiple search by semicolon Potential targets are fetched from the drug databases DrugBank and ChEMBL, displayed and marked in the PD map by numbers within a coloured rectangle. Additional information concerning the drugs and the potential targets, including links to drug databases and PubMed is provided in the left panel. (A) (B) 01/09/2017 PD map user guide 7
9 (C) Fig. 7: Search for the drug Mirapex. (A) Information panel displays information on Pramipexole as Mirapex is a brand name for the drug Pramipexole. Drug targets are displayed in the map. (B) Pramipexole targets dopamine receptors (DRD2 and DRD1). (C) The bubble pinned to the target (e.g., DRD1) provides information on drug interaction and offers additional functionalities to search for other drugs, chemical or mirna that interacts with the DRD1 protein (see also Fig. 4B, Fig. 8, Fig. 9). Please note: If within a multiple search several drugs hit the same element, the bubbles will overlap and only the one from the second search is visible on that element (see Fig. 8) The same happens when search queries from different tools targeting the same element in the PD map. Only the bubble from the first search is visible on the element that is targeted by the two or more search queries. All results are always displayed in the left panel. All results are displayed simultaneously in a pop up information field when clicking on the bubble pinned to the target (see Fig. 4; Fig. 7C, Fig. 8 and Fig 11). At the end of the list, the left panel shows also targets that are not in the PD map (see Fig. 9). 01/09/2017 PD map user guide 8
10 2.4. Search: How to use chemical interaction interface The CHEMICAL interface allows to search for potential interactions of chemicals with the PD map elements based on the Comparative Toxicogenomics Database (ctdbase.org). Only interactions that are annotated to the disease term Parkinson s disease (MeSH ID: D010300) in the CTD database are considered. Activate CHEMICAL tab under SEARCH. Type in the full name of a chemical according to the ctdbase/chem database; abbreviations or synonyms do not work (e.g., MTPT does not work but 1-Methyl-4-phenyl- 1,2,3,6-tetrahydropyridine). If the chemical name includes comma(s), place a semicolon behind the name to avoid a segmentation of the name during processing. Always separate multiple search by semicolon. Potential targets are fetched from the Comparative Toxicogenomics Database, displayed and marked in the PD map by a number within a coloured circle. Additional information concerning the chemical and the potential targets, including links to CTD databases and PubMed are provided in the left panel. Fig. 8: Search for the interaction targets of rotenone and hydrazine. Information panel displays information on hydrazine. Interacting species are labelled in the map (green bubbles for hydrazine; blue bubbles for rotenone). Clicking on framed number above the target (e.g., green framed 3 ) provides information on the target (i.e., HSPA9) and chemical interactions and offers additional functionalities to search for other drugs, chemical or mirna that interacts with the target (see also Fig. 4; Fig. 7C, and Fig 11). Please note: If within a multiple search several drugs hit the same element, the bubbles will overlap and only the one from the second search is visible on that element. 01/09/2017 PD map user guide 9
11 2.5. Search: How to use mirna interface The MiRNA" interface allows to search for potential targets of mirna within the PD map elements based on the mirna database: mirtarbase. Only target interactions that have strong evidence according to mirtarbase criteria (i.e., reporter assays, western blot or qpcr analysis) are considered. Activate MiRNA tab under SEARCH. Type in the mirna ID, use only mature sequence IDs e.g., hsa-mir-125a-3p (see: Separate multiple search by semicolon. Matching targets from the mirtarbase are displayed in the PD map by pinned bubbles (here: coloured triangles). Additional information concerning the mirnas and the potential targets, including links to mirtarbase and PubMed is provided in the left panel. Fig. 9: Search for targets of hsa-mir-125a-3p. Left panel displays information on hsa-mir-125a-3p and its targets. Targets are marked by pinned bubbles (green triangles) in the PD map. Clicking on the pinned bubble provides information on mirna interactions and offers additional functionalities to search for other mirna, drugs, chemical that interacts with the target. Left panel shows also targets that are not in the PD map (here: GPC4). 01/09/2017 PD map user guide 10
12 3. How to display implemented data sets The visualization of specific data sets on differential gene regulation is accessible via the OVERLAYS tab (left panel). Different data sets are indicated under GENERAL OVERLAYS. Ageing brain: Differentially expressed genes in healthy aging brain. Enrico Glaab (LCSB) contributed this data derived from Allen Brain Atlas database. red: stronger expression in aged brain; blue: lower expression in aged brain. See: Glaab and Schneider, PD substantia nigra: Differential gene expression (FDR <= 0.05) from the analysis of eight transcriptome data sets, comparing human post mortem brain samples from PD vs. healthy controls. red: stronger expression in PD; blue: lower expression in PD. See: Fujita et al., PD UK GO Project genes: Genes annotated by the Parkinson's disease GO Annotation Project. dark green: high priority gene list, light green: Parkinson s-relevant protein. See: Foulger e al., The expression data could be downloaded under Data (note: the value column in the downloaded list shows normalized expression values) Click boxes to display the respective data set on the PD map. Pathways and compartment view provides an overview on cellular processes affected by the visualized data (Fig. 10A) Detailed differential expression (red rectangle: up regulated, blue rectangle: down regulated) is best visible using the colourless overlay Empty (Fig. 10B). (A) 01/09/2017 PD map user guide 11
13 (B) Fig. 10: Overlay of the PD substantia nigra data set to the PD map. (A) Pathways and compartment view. (B) Empty view Displaying multiple data set is possible by ticking several checkboxes in the View column. Results from different data sets are simultaneously displayed by subdivided coloured rectangles (Fig. 11) Additional information is provided when clicking on the species (Fig. 11) Fig. 11: Overlay of PD substantia nigra data set and ageing brain data set to the PD map. Clicking on CREBBP provided detailed information in a pop up box and on the bottom of the information panel. In addition, information on Interacting drugs, chemical or microrna can be displayed (see Chapters: 4, 5, and 6) 01/09/2017 PD map user guide 12
14 4. How to display custom data sets on the PD map A personal account is needed to upload data. The account is for free. Click on the INFO tab. Click on Request an account under USER DATA. User name and password will be provided by . The account enables the upload of 100 separate data sets. After login, user personal data are displayed in the INFO tab. To upload own data sets, click on the OVERLAYS tab. This opens the menu for data set upload and management ( USER-PROVIDED OVERLAYS ) Under Add overlays you find two possibilities to upload data sets and highlight elements on the PD map (see Fig. 12) Upload and highlight a list of elements Upload a list of element names and related expression values (as txt.file) Fig. 12: Screenshot: Add overlays form Upload and highlight a list of elements Click on Add overlays and fill in the form. Name (default) Description (free text) Provide list of elements (one per line) 01/09/2017 PD map user guide 13
15 This list must contain the name of elements as it appears in the PD map (e.g., HGNC name for proteins and genes; CHEBI name for simple molecules) The list may contain names for different element types at the same time (see Fig. 13A) Click UPLOAD starts the process The new file will appear in USER-PROVIDED OVERLAYS. A short report will be send by mail ( MapViewer notification ). Data are visualized by green colour on the PD map after clicking the view box (Fig. 13B). (A) (B) Fig. 13: Upload (A) a list of elements to the PD map and visualisation (B) 4.2. Upload of a data file: Data preparation Data file needs to be uploaded as a txt. file that consists of two columns first column (headline = name) contains HGNC gene symbols second column (headline = value) contains values between -1 and 1 As both proteins and genes are defined in the PD map by the uniform HGNC identifier and HGNC gene symbol, data derived from protein or gene expression will be displayed on gene and protein species in the PD map. To upload data from other species than human, the species-specific identifiers need to be translated to HGNC identifiers. The differential expression values (fold changes) needs to be normalized into a range between -1 to 1 (see Fig. 14). 01/09/2017 PD map user guide 14
16 HGNC symbol fold change normalization HMOX SRXN GSTA DDIT ANXA NFE2L GCLM GSR SRXN TXNRD NQO MFN GSS BCL NDUFA BECN GSK3B DNM1L ATP KEAP CYCS AMBRA MFN BCL2L ATP5D UQCRC BAD CAMK2G DNAJC CAMK2A HSPA CAMK2B SLC1A Fig. 14: Differential gene expression data (excel file) were edited and transformed into a txt. file before upload. Normalization applied here: fold change (f.c.) >5 is set 1; f.c. <-5 set -1; f.c. between -1.5 and 1.5 is set 0; f.c. between -5 and -1.5 as well as 1.5 and 5 are normalized by dividing the value by Upload of a data file: Data upload Click on the OVERLAYS tab. This opens the menu for data set upload and management ( USER-PROVIDED OVERLAYS ) Click on Add overlays Fill in form Name (default) Description (free text) Click on Browse an select the file to be uploaded Click UPLOAD to start the process The new file will appear in USER-PROVIDED OVERLAYS. A short report will be send by mail ( MapViewer notification ). Data could be visualized on the PD map by clicking on view. Color code: Blue for down regulation and Red for up regulation Elements are automatically coloured by the following colour codes in accordance to their dedicated values 01/09/2017 PD map user guide 15
17 Please note: Combining display of multiple data sets from GENERAL OVERLAYS and USER-PROVIDED OVERLAYS is possible by clicking different data sets under View. Results are displayed by subdivided coloured rectangles (see Fig. 11) In general, all PD map elements can be individually coloured on the PD map. If you like to colour elements individually contact us directly To upload data from other species (e.g., mouse), gene and protein names has to be translated to human HGNC identifier. All uploaded data are accessible by the system administrators due to technical reasons but they are not visible by any other user. 5. How to provide comments and feedback Right click with the mouse in the PD map at the position where the comment is related to (note: this feature does not work on tablets and smart phones). Click on: Add comment, fill in the form (use the dropdown list for Type ) and press send button (Fig. 15). The curation team will be notified on new comments and respond to them immediately. The comment (without personal information) will be visible on the PD map if Pinned is checked (set by default) Fig. 15: Screenshot shows the comment field 01/09/2017 PD map user guide 16
18 To see all pinned comments, click on the button COMMENTS (red circle, left upper corner, Fig. 16). Clicking on the comment icon in the PD map displays the content (see Fig. 16) Fig. 16: Screenshot after activating the COMMENTS button (red circle), displaying position of all pinned comments. Details of the specific comment on re4026 is visible after clicking on the related icon. 6. How to extract information from the PD map 6.1. Export function Go to INFO tap and click on EXPORT button (red arrow) Fig. 17: Export function 01/09/2017 PD map user guide 17
19 Use the tab ELEMENT EXPORT to select specific elements for download. Use the tab NETWORK EXPORT to select specific networks for download (only SIF files). Use the tab GRAPHICS to download whole PD map and the single sub maps as SVG file Note: The whole PD map could be downloaded under source file. Pathway Cell death HGNC_SYMBOL: ADD1 UNIPROT: P35611 Cell death HGNC_SYMBOL: AKT1 UNIPROT: P31749 Cell death HGNC_SYMBOL: AKT2 UNIPROT: P31751 Cell death HGNC_SYMBOL: APAF1 UNIPROT: O14727 Cell death HGNC_SYMBOL: ATG12 UNIPROT: O94817 Cell death HGNC_SYMBOL: ATG5 UNIPROT: Q9H1Y0 Cell death HGNC_SYMBOL: BAD UNIPROT: Q92934 Cell death HGNC_SYMBOL: BAK1 UNIPROT: Q16611 Cell death HGNC_SYMBOL: BAX UNIPROT: Q07812 Cell death HGNC_SYMBOL: BBC3 UNIPROT: Q96PG8 Cell death HGNC_SYMBOL: BCL2 UNIPROT: P10415 Cell death HGNC_SYMBOL: BCL2L1 UNIPROT: Q07817 Cell death HGNC_SYMBOL: BCL2L11 UNIPROT: O43521 Cell death HGNC_SYMBOL: BID UNIPROT: P55957 Cell death HGNC_SYMBOL: BMF UNIPROT: Q96LC9 Cell death HGNC_SYMBOL: CAPN1 UNIPROT: P07384 Cell death HGNC_SYMBOL: CAPN2 UNIPROT: P17655 Cell death HGNC_SYMBOL: CASP2 UNIPROT: P42575 Cell death HGNC_SYMBOL: CASP3 UNIPROT: P42574 Cell death HGNC_SYMBOL: CASP6 UNIPROT: P55212 Cell death HGNC_SYMBOL: CASP7 UNIPROT: P55210 Cell death HGNC_SYMBOL: CASP8 UNIPROT: Q14790 Cell death HGNC_SYMBOL: CASP9 UNIPROT: P55211 Cell death HGNC_SYMBOL: CHCHD2 UNIPROT: Q9Y6H1 Cell death HGNC_SYMBOL: CRADD UNIPROT: P78560 Cell death HGNC_SYMBOL: CTNNB1 UNIPROT: P35222 Fig. 18: Export function: Screenshot shows example for ELEMENTS EXPORT. Here proteins related to Cell Death pathway were extracted. HGNC symbol and Uniport identifier were used for annotation. Note: only green coloured annotations are feasible Select mode Enables the export of certain areas of PD map area as model (CellDesigner xml. file) or as image (PNG, PDF). Right click with the mouse in the PD map in the area you like to export. Click on Select mode (Fig. 19). Clicking around the area of interest will generate a frame (Fig. 20). As soon as the frame is closed the content can be exported. Clicking right mouse button within the frame (Fig. 20). 01/09/2017 PD map user guide 18
20 Fig. 19: Generation of a frame to define region for export Fig. 20: Export of the selected area (grey coloured) as SBML file Click on Export as model to download the corresponding xml file (CellDesigner SBML) or SBGN-ML file. Click on Export as image to download an image file (PNG or PDF). Please note Export as image always generates rectangles 7. Help In case of any problems or questions regarding the PD map functionalities please contact marek.ostaszewski@uni.lu or stephan.gebel@uni.lu. 01/09/2017 PD map user guide 19
Pathway Analysis using Partek Genomics Suite 6.6 and Partek Pathway
 Pathway Analysis using Partek Genomics Suite 6.6 and Partek Pathway Overview Partek Pathway provides a visualization tool for pathway enrichment spreadsheets, utilizing KEGG and/or Reactome databases for
Pathway Analysis using Partek Genomics Suite 6.6 and Partek Pathway Overview Partek Pathway provides a visualization tool for pathway enrichment spreadsheets, utilizing KEGG and/or Reactome databases for
The genexplain platform. Workshop SW2: Pathway Analysis in Transcriptomics, Proteomics and Metabolomics
 The genexplain platform Workshop SW2: Pathway Analysis in Transcriptomics, Proteomics and Metabolomics Saturday, March 17, 2012 2 genexplain GmbH Am Exer 10b D-38302 Wolfenbüttel Germany E-mail: olga.kel-margoulis@genexplain.com,
The genexplain platform Workshop SW2: Pathway Analysis in Transcriptomics, Proteomics and Metabolomics Saturday, March 17, 2012 2 genexplain GmbH Am Exer 10b D-38302 Wolfenbüttel Germany E-mail: olga.kel-margoulis@genexplain.com,
User Guide. v Released June Advaita Corporation 2016
 User Guide v. 0.9 Released June 2016 Copyright Advaita Corporation 2016 Page 2 Table of Contents Table of Contents... 2 Background and Introduction... 4 Variant Calling Pipeline... 4 Annotation Information
User Guide v. 0.9 Released June 2016 Copyright Advaita Corporation 2016 Page 2 Table of Contents Table of Contents... 2 Background and Introduction... 4 Variant Calling Pipeline... 4 Annotation Information
Customisable Curation Workflows in Argo
 Customisable Curation Workflows in Argo Rafal Rak*, Riza Batista-Navarro, Andrew Rowley, Jacob Carter and Sophia Ananiadou National Centre for Text Mining, University of Manchester, UK *Corresponding author:
Customisable Curation Workflows in Argo Rafal Rak*, Riza Batista-Navarro, Andrew Rowley, Jacob Carter and Sophia Ananiadou National Centre for Text Mining, University of Manchester, UK *Corresponding author:
RLIMS-P Website Help Document
 RLIMS-P Website Help Document Table of Contents Introduction... 1 RLIMS-P architecture... 2 RLIMS-P interface... 2 Login...2 Input page...3 Results Page...4 Text Evidence/Curation Page...9 URL: http://annotation.dbi.udel.edu/text_mining/rlimsp2/
RLIMS-P Website Help Document Table of Contents Introduction... 1 RLIMS-P architecture... 2 RLIMS-P interface... 2 Login...2 Input page...3 Results Page...4 Text Evidence/Curation Page...9 URL: http://annotation.dbi.udel.edu/text_mining/rlimsp2/
HymenopteraMine Documentation
 HymenopteraMine Documentation Release 1.0 Aditi Tayal, Deepak Unni, Colin Diesh, Chris Elsik, Darren Hagen Apr 06, 2017 Contents 1 Welcome to HymenopteraMine 3 1.1 Overview of HymenopteraMine.....................................
HymenopteraMine Documentation Release 1.0 Aditi Tayal, Deepak Unni, Colin Diesh, Chris Elsik, Darren Hagen Apr 06, 2017 Contents 1 Welcome to HymenopteraMine 3 1.1 Overview of HymenopteraMine.....................................
Reactome Error! Bookmark not defined. Reactome Tools
 Reactome This document introduces Reactome, the user interface and the database content. Further information can be found in the online Reactome user guide at http://www.reactome.org/userguide/usersguide.html.
Reactome This document introduces Reactome, the user interface and the database content. Further information can be found in the online Reactome user guide at http://www.reactome.org/userguide/usersguide.html.
mirnet Tutorial Starting with expression data
 mirnet Tutorial Starting with expression data Computer and Browser Requirements A modern web browser with Java Script enabled Chrome, Safari, Firefox, and Internet Explorer 9+ For best performance and
mirnet Tutorial Starting with expression data Computer and Browser Requirements A modern web browser with Java Script enabled Chrome, Safari, Firefox, and Internet Explorer 9+ For best performance and
WAIPA DISTRICT COUNCIL. Maps Online 9. Updated January This document contains an overview of IntraMaps/Maps Online version 9.
 WAIPA DISTRICT COUNCIL Maps Online 9 Updated January 2018 This document contains an overview of IntraMaps/Maps Online version 9.0 Contents Starting Maps Online... 3 Menu Bar... 4 Tools... 5 View Tab...
WAIPA DISTRICT COUNCIL Maps Online 9 Updated January 2018 This document contains an overview of IntraMaps/Maps Online version 9.0 Contents Starting Maps Online... 3 Menu Bar... 4 Tools... 5 View Tab...
Working with PDF s. To open a recent file on the Start screen, double click on the file name.
 Working with PDF s Acrobat DC Start Screen (Home Tab) When Acrobat opens, the Acrobat Start screen (Home Tab) populates displaying a list of recently opened files. The search feature on the top of the
Working with PDF s Acrobat DC Start Screen (Home Tab) When Acrobat opens, the Acrobat Start screen (Home Tab) populates displaying a list of recently opened files. The search feature on the top of the
User Manual Mobile client User Interface Version 5.0. Powered by
 User Manual Mobile client User Interface Version 5.0 Powered by Cartographic browser Gomap 4 1 Access control 5 1.1 Public access 5 1.2 Secured access 5 1.3 Multiple applications 5 2 Organisation 6 3 Parameters
User Manual Mobile client User Interface Version 5.0 Powered by Cartographic browser Gomap 4 1 Access control 5 1.1 Public access 5 1.2 Secured access 5 1.3 Multiple applications 5 2 Organisation 6 3 Parameters
Web Resources. iphemap: An atlas of phenotype to genotype relationships of human ipsc models of neurological diseases
 Web Resources iphemap: An atlas of phenotype to genotype relationships of human ipsc models of neurological diseases Ethan W. Hollingsworth 1, 2, Jacob E. Vaughn 1, 2, Josh C. Orack 1,2, Chelsea Skinner
Web Resources iphemap: An atlas of phenotype to genotype relationships of human ipsc models of neurological diseases Ethan W. Hollingsworth 1, 2, Jacob E. Vaughn 1, 2, Josh C. Orack 1,2, Chelsea Skinner
The Allen Human Brain Atlas offers three types of searches to allow a user to: (1) obtain gene expression data for specific genes (or probes) of
 Microarray Data MICROARRAY DATA Gene Search Boolean Syntax Differential Search Mouse Differential Search Search Results Gene Classification Correlative Search Download Search Results Data Visualization
Microarray Data MICROARRAY DATA Gene Search Boolean Syntax Differential Search Mouse Differential Search Search Results Gene Classification Correlative Search Download Search Results Data Visualization
BovineMine Documentation
 BovineMine Documentation Release 1.0 Deepak Unni, Aditi Tayal, Colin Diesh, Christine Elsik, Darren Hag Oct 06, 2017 Contents 1 Tutorial 3 1.1 Overview.................................................
BovineMine Documentation Release 1.0 Deepak Unni, Aditi Tayal, Colin Diesh, Christine Elsik, Darren Hag Oct 06, 2017 Contents 1 Tutorial 3 1.1 Overview.................................................
Genome Browsers Guide
 Genome Browsers Guide Take a Class This guide supports the Galter Library class called Genome Browsers. See our Classes schedule for the next available offering. If this class is not on our upcoming schedule,
Genome Browsers Guide Take a Class This guide supports the Galter Library class called Genome Browsers. See our Classes schedule for the next available offering. If this class is not on our upcoming schedule,
Pathway Analysis of Untargeted Metabolomics Data using the MS Peaks to Pathways Module
 Pathway Analysis of Untargeted Metabolomics Data using the MS Peaks to Pathways Module By: Jasmine Chong, Jeff Xia Date: 14/02/2018 The aim of this tutorial is to demonstrate how the MS Peaks to Pathways
Pathway Analysis of Untargeted Metabolomics Data using the MS Peaks to Pathways Module By: Jasmine Chong, Jeff Xia Date: 14/02/2018 The aim of this tutorial is to demonstrate how the MS Peaks to Pathways
Partner Side SMART Guide
 Partner Side SMART Guide Table of Contents 1. Introduction... 3 2. Partner Registration Process... 3 3. Additional Form... 12 4. Scorecard... 13 5. View Buyer Profile... 14 Partner Side User Manual 31
Partner Side SMART Guide Table of Contents 1. Introduction... 3 2. Partner Registration Process... 3 3. Additional Form... 12 4. Scorecard... 13 5. View Buyer Profile... 14 Partner Side User Manual 31
XMReality 6. User Manual for Windows XMReality AB Teknikringen 10, 8 fl SE Linköping Sweden
 XMReality 6 User Manual for Windows - 6.3 1 XMReality AB Teknikringen 10, 8 fl SE-583 30 Linköping Sweden Introduction This is a user manual for XMReality Remote Guidance Generation 6 for Windows. An account
XMReality 6 User Manual for Windows - 6.3 1 XMReality AB Teknikringen 10, 8 fl SE-583 30 Linköping Sweden Introduction This is a user manual for XMReality Remote Guidance Generation 6 for Windows. An account
Tutorial:OverRepresentation - OpenTutorials
 Tutorial:OverRepresentation From OpenTutorials Slideshow OverRepresentation (about 12 minutes) (http://opentutorials.rbvi.ucsf.edu/index.php?title=tutorial:overrepresentation& ce_slide=true&ce_style=cytoscape)
Tutorial:OverRepresentation From OpenTutorials Slideshow OverRepresentation (about 12 minutes) (http://opentutorials.rbvi.ucsf.edu/index.php?title=tutorial:overrepresentation& ce_slide=true&ce_style=cytoscape)
WAITOMO DISTRICT COUNCIL ONLINE MAPS. Updated June This document contains an overview of Waitomo District Council Online Maps
 WAITOMO DISTRICT COUNCIL ONLINE MAPS V8 Updated June2017 - This document contains an overview of Waitomo District Council Online Maps Table of Contents Starting Online Maps... 3 Main Screen... 4 Menu Bar...
WAITOMO DISTRICT COUNCIL ONLINE MAPS V8 Updated June2017 - This document contains an overview of Waitomo District Council Online Maps Table of Contents Starting Online Maps... 3 Main Screen... 4 Menu Bar...
FunRich Tool Documentation
 FunRich Tool Documentation Version 2.1.2 Shivakumar Keerthikumar, Mohashin Pathan, Johnson Agbinya and Suresh Mathivanan Mathivanan Lab http://www.mathivananlab.org La Trobe University LIMS1 Department
FunRich Tool Documentation Version 2.1.2 Shivakumar Keerthikumar, Mohashin Pathan, Johnson Agbinya and Suresh Mathivanan Mathivanan Lab http://www.mathivananlab.org La Trobe University LIMS1 Department
Annotating a Genome in PATRIC
 Annotating a Genome in PATRIC The following step-by-step workflow is intended to help you learn how to navigate the new PATRIC workspace environment in order to annotate and browse your genome on the PATRIC
Annotating a Genome in PATRIC The following step-by-step workflow is intended to help you learn how to navigate the new PATRIC workspace environment in order to annotate and browse your genome on the PATRIC
MINERVA a platform for visualization and curation of molecular interaction networks
 www.nature.com/npjsba ARTICLE OPEN MINERVA a platform for visualization and curation of molecular interaction networks Piotr Gawron 1, Marek Ostaszewski 1, Venkata Satagopam 1, Stephan Gebel 1, Alexander
www.nature.com/npjsba ARTICLE OPEN MINERVA a platform for visualization and curation of molecular interaction networks Piotr Gawron 1, Marek Ostaszewski 1, Venkata Satagopam 1, Stephan Gebel 1, Alexander
Skills Funding Agency
 Provider Data Self-Assessment Toolkit (PDSAT) v17 User Guide Contents Introduction... 2 Compatibility and prerequisites... 2 1. Installing PDSAT... 3 2. Opening PDSAT... 6 2.1 Opening Screen... 6 2.2 Updates...
Provider Data Self-Assessment Toolkit (PDSAT) v17 User Guide Contents Introduction... 2 Compatibility and prerequisites... 2 1. Installing PDSAT... 3 2. Opening PDSAT... 6 2.1 Opening Screen... 6 2.2 Updates...
Quick Tutorial. Running CellDesigner TM Simulation with ControlPanel
 Running CellDesigner TM Simulation with ControlPanel Quick Tutorial 2005/09/01 The Systems Biology Institute http://www.systems-biology.org/ Mitsui Knowledge Industry Co. Ltd. http://bio.mki.co.jp/en/
Running CellDesigner TM Simulation with ControlPanel Quick Tutorial 2005/09/01 The Systems Biology Institute http://www.systems-biology.org/ Mitsui Knowledge Industry Co. Ltd. http://bio.mki.co.jp/en/
Ontology-based annotation of multiscale imaging data: Utilizing and building the Neuroscience Information Framework. Maryann E.
 Ontology-based annotation of multiscale imaging data: Utilizing and building the Neuroscience Information Framework Maryann E. Martone University of California, San Diego What does this mean? 3D Volumes
Ontology-based annotation of multiscale imaging data: Utilizing and building the Neuroscience Information Framework Maryann E. Martone University of California, San Diego What does this mean? 3D Volumes
EGAN Tutorial: A Basic Use-case
 EGAN Tutorial: A Basic Use-case July 2010 Jesse Paquette Biostatistics and Computational Biology Core Helen Diller Family Comprehensive Cancer Center University of California, San Francisco (AKA BCBC HDFCCC
EGAN Tutorial: A Basic Use-case July 2010 Jesse Paquette Biostatistics and Computational Biology Core Helen Diller Family Comprehensive Cancer Center University of California, San Francisco (AKA BCBC HDFCCC
SEEK User Manual. Introduction
 SEEK User Manual Introduction SEEK is a computational gene co-expression search engine. It utilizes a vast human gene expression compendium to deliver fast, integrative, cross-platform co-expression analyses.
SEEK User Manual Introduction SEEK is a computational gene co-expression search engine. It utilizes a vast human gene expression compendium to deliver fast, integrative, cross-platform co-expression analyses.
Preferences Table of Contents
 Preferences Table of Contents My Profile... 2 Quick Profile Maintenance... 2 My Names... 3 My Addresses... 3 My E-Mail Addresses... 4 Personal Photo and Logo Maintenance... 4 My Documents... 6 My Phone
Preferences Table of Contents My Profile... 2 Quick Profile Maintenance... 2 My Names... 3 My Addresses... 3 My E-Mail Addresses... 4 Personal Photo and Logo Maintenance... 4 My Documents... 6 My Phone
CellaVision Proficiency Software
 CellaVision Proficiency USER S MANUAL 2.3 CellaVision Proficiency Preface CellaVision is a trademark of CellaVision AB. All other trademarks used in this document are property of their respective owners.
CellaVision Proficiency USER S MANUAL 2.3 CellaVision Proficiency Preface CellaVision is a trademark of CellaVision AB. All other trademarks used in this document are property of their respective owners.
Full Search Map Tab. This map is the result of selecting the Map tab within Full Search.
 Full Search Map Tab This map is the result of selecting the Map tab within Full Search. This map can be used when defining your parameters starting from a Full Search. Once you have entered your desired
Full Search Map Tab This map is the result of selecting the Map tab within Full Search. This map can be used when defining your parameters starting from a Full Search. Once you have entered your desired
Load KEGG Pathways. Browse and Load KEGG Pathways
 Note: This is just partial of VisANT manual related with the NAR 2007 submission. Please visit http://visant.bu.edu for the full user manual. Load KEGG Pathways...1 Browse and Load KEGG Pathways...1 Load
Note: This is just partial of VisANT manual related with the NAR 2007 submission. Please visit http://visant.bu.edu for the full user manual. Load KEGG Pathways...1 Browse and Load KEGG Pathways...1 Load
Climate-Smart New Orleans
 Climate-Smart New Orleans Table of Contents GETTING THERE... 2 Accessing the site... 2 Logging into the site... 2 Navigating the Map... 2 Zoom & Pan... 2 Change the map background... 3 Interacting in the
Climate-Smart New Orleans Table of Contents GETTING THERE... 2 Accessing the site... 2 Logging into the site... 2 Navigating the Map... 2 Zoom & Pan... 2 Change the map background... 3 Interacting in the
About the Edinburgh Pathway Editor:
 About the Edinburgh Pathway Editor: EPE is a visual editor designed for annotation, visualisation and presentation of wide variety of biological networks, including metabolic, genetic and signal transduction
About the Edinburgh Pathway Editor: EPE is a visual editor designed for annotation, visualisation and presentation of wide variety of biological networks, including metabolic, genetic and signal transduction
Bioinformatics Hubs on the Web
 Bioinformatics Hubs on the Web Take a class The Galter Library teaches a related class called Bioinformatics Hubs on the Web. See our Classes schedule for the next available offering. If this class is
Bioinformatics Hubs on the Web Take a class The Galter Library teaches a related class called Bioinformatics Hubs on the Web. See our Classes schedule for the next available offering. If this class is
E-Notebook User Guide
 1. Introduction E-Notebook Features Facilitates daily record-keeping for the scientists (Managing data, organizing information, streamlining workflow). Conducts searches by substructure, key word, date,
1. Introduction E-Notebook Features Facilitates daily record-keeping for the scientists (Managing data, organizing information, streamlining workflow). Conducts searches by substructure, key word, date,
MetScape User Manual
 MetScape 2.3.2 User Manual A Plugin for Cytoscape National Center for Integrative Biomedical Informatics July 2012 2011 University of Michigan This work is supported by the National Center for Integrative
MetScape 2.3.2 User Manual A Plugin for Cytoscape National Center for Integrative Biomedical Informatics July 2012 2011 University of Michigan This work is supported by the National Center for Integrative
MERCATOR TASK MASTER TASK MANAGEMENT SCREENS:- LOGIN SCREEN:- APP LAYOUTS:-
 MERCATOR TASK MASTER TASK MANAGEMENT SCREENS:- LOGIN SCREEN:- APP LAYOUTS:- This is Navigation bar where you have 5 Menus and App Name. This Section I will discuss in brief in the Navigation Bar Section.
MERCATOR TASK MASTER TASK MANAGEMENT SCREENS:- LOGIN SCREEN:- APP LAYOUTS:- This is Navigation bar where you have 5 Menus and App Name. This Section I will discuss in brief in the Navigation Bar Section.
City of La Crosse Online Mapping Website Help Document
 City of La Crosse Online Mapping Website Help Document This document was created to assist in using the new City of La Crosse online mapping sites. When the website is first opened, a map showing the City
City of La Crosse Online Mapping Website Help Document This document was created to assist in using the new City of La Crosse online mapping sites. When the website is first opened, a map showing the City
Overview of ArcGIS Online Applications. Champaign County
 Overview of ArcGIS Online Applications Champaign County Champaign County GIS Consortium Updated: April 2017 Table of Contents ArcGIS Online Application Overview... 3 Map Interface Symbology and Terminology...
Overview of ArcGIS Online Applications Champaign County Champaign County GIS Consortium Updated: April 2017 Table of Contents ArcGIS Online Application Overview... 3 Map Interface Symbology and Terminology...
QPCR User Guide. Version 1.8. Create Date Sep 25, 2007 Last modified Jan 26, 2010 by Stephan Pabinger
 User Guide Version 1.8 Create Date Sep 25, 2007 Last modified Jan 26, 2010 by Table of content Table of content... 2 1 Introduction... 4 1.1 Purpose... 4 1.2 Hints for using the software... 4 1.3 Example
User Guide Version 1.8 Create Date Sep 25, 2007 Last modified Jan 26, 2010 by Table of content Table of content... 2 1 Introduction... 4 1.1 Purpose... 4 1.2 Hints for using the software... 4 1.3 Example
CME E-quotes Wireless Application for Android Welcome
 CME E-quotes Wireless Application for Android Welcome This guide will familiarize you with the application, a powerful trading tool developed for your Android. Table of Contents What is this application?
CME E-quotes Wireless Application for Android Welcome This guide will familiarize you with the application, a powerful trading tool developed for your Android. Table of Contents What is this application?
miscript mirna PCR Array Data Analysis v1.1 revision date November 2014
 miscript mirna PCR Array Data Analysis v1.1 revision date November 2014 Overview The miscript mirna PCR Array Data Analysis Quick Reference Card contains instructions for analyzing the data returned from
miscript mirna PCR Array Data Analysis v1.1 revision date November 2014 Overview The miscript mirna PCR Array Data Analysis Quick Reference Card contains instructions for analyzing the data returned from
CellDesigner 3.0 alpha Preview Version of New Graphical Notation
 CellDesigner 3.0 alpha Preview Version of New Graphical Notation The Systems Biology Institute http://www.systems-biology.org/ Mizuho Information & Research Institute http://www.mizuho-ir.co.jp/ Aug 4
CellDesigner 3.0 alpha Preview Version of New Graphical Notation The Systems Biology Institute http://www.systems-biology.org/ Mizuho Information & Research Institute http://www.mizuho-ir.co.jp/ Aug 4
Welcome to the Rainfall Atlas of Hawai i interactive map!
 Welcome to the Rainfall Atlas of Hawai i interactive map! This guide will walk you through all of the capabilities of the interactive map so that you can make the most of all it has to offer. Conditions
Welcome to the Rainfall Atlas of Hawai i interactive map! This guide will walk you through all of the capabilities of the interactive map so that you can make the most of all it has to offer. Conditions
Product Information Manager PIM. How to Create Smartsheet Single Items
 Product Information Manager PIM How to Create Smartsheet Single Items Smartsheet item information Smartsheet item is for data that is for singe items not for online IF Online use the ECOMM Item smartsheet
Product Information Manager PIM How to Create Smartsheet Single Items Smartsheet item information Smartsheet item is for data that is for singe items not for online IF Online use the ECOMM Item smartsheet
Pathway Studio Quick Start Guide
 Pathway Studio Quick Start Guide This Quick Start Guide is for users of the Pathway Studio 4.0 pathway analysis software. The Quick Start Guide demonstrates the key features of the software and provides
Pathway Studio Quick Start Guide This Quick Start Guide is for users of the Pathway Studio 4.0 pathway analysis software. The Quick Start Guide demonstrates the key features of the software and provides
Ctrack Online User Guide
 Fleetstar Online A Guide to Winter Maintenance Reporting v1.1 Ctrack Online User Guide Title: Ctrack Online Quickstart Guide Date: 18/07/2013 Version: 1.0 Table of Contents 1. Ctrack Online Introduction...
Fleetstar Online A Guide to Winter Maintenance Reporting v1.1 Ctrack Online User Guide Title: Ctrack Online Quickstart Guide Date: 18/07/2013 Version: 1.0 Table of Contents 1. Ctrack Online Introduction...
efip online Help Document
 efip online Help Document University of Delaware Computer and Information Sciences & Center for Bioinformatics and Computational Biology Newark, DE, USA December 2013 K K S I K K Table of Contents INTRODUCTION...
efip online Help Document University of Delaware Computer and Information Sciences & Center for Bioinformatics and Computational Biology Newark, DE, USA December 2013 K K S I K K Table of Contents INTRODUCTION...
SBMLmerge and MIRIAM support
 SBMLmerge and MIRIAM support Marvin Schulz Max Planck Institute for Molecular Genetics Biomodels.net training camp 10. 04. 2006 What exactly does SBMLmerge do? Small parts of the glycolysis SBMLmerge...
SBMLmerge and MIRIAM support Marvin Schulz Max Planck Institute for Molecular Genetics Biomodels.net training camp 10. 04. 2006 What exactly does SBMLmerge do? Small parts of the glycolysis SBMLmerge...
Visualizing Venice Historic Environment Record (Geospatial Database)
 Visualizing Venice Historic Environment Record (Geospatial Database) Table of Contents Introduction... 2 Getting Started opening the sources interface... 3 Searching for a Record... 4 Adding a New Source
Visualizing Venice Historic Environment Record (Geospatial Database) Table of Contents Introduction... 2 Getting Started opening the sources interface... 3 Searching for a Record... 4 Adding a New Source
EUROPEAN ORGANISATION FOR THE SAFETY OF AIR NAVIGATION
 EUROPEAN ORGANISATION FOR THE SAFETY OF AIR NAVIGATION E U R O C O N T R O L TOKAI USER MANUAL Edition: v2.6 DIRECTORATE NETWORK MANAGEMENT 1 Page TOKAI User Manual (Edition v2.6) EUROCONTROL TOKAI Application
EUROPEAN ORGANISATION FOR THE SAFETY OF AIR NAVIGATION E U R O C O N T R O L TOKAI USER MANUAL Edition: v2.6 DIRECTORATE NETWORK MANAGEMENT 1 Page TOKAI User Manual (Edition v2.6) EUROCONTROL TOKAI Application
Geocortex HTML 5 Viewer Manual
 Geocortex HTML 5 Viewer Manual Searching for a feature Use the Search Feature box in the top right hand corner of the viewer window. You can use this to search numerous data types such as property number,
Geocortex HTML 5 Viewer Manual Searching for a feature Use the Search Feature box in the top right hand corner of the viewer window. You can use this to search numerous data types such as property number,
KIN 147 Lab Practical Mid-term: Tibial Acceleration Data Analysis Excel analyses work much better on PCs than on Macs (especially older Macs)
 KIN 147 Lab Practical Mid-term: Tibial Acceleration Data Analysis Excel analyses work much better on PCs than on Macs (especially older Macs) Your goal is to correctly analyze accelerometer data Analyzing
KIN 147 Lab Practical Mid-term: Tibial Acceleration Data Analysis Excel analyses work much better on PCs than on Macs (especially older Macs) Your goal is to correctly analyze accelerometer data Analyzing
Arabidopsis Protein Protein Interaction Analysis Pipeline
 ANAP User Guide The ANAP tool has many useful network biology functions that are demonstrated in this user guide. 1. The ANAP tool http://gmdd.shgmo.org/computational-biology/anap/anap_v1.0/ The ANAP tool
ANAP User Guide The ANAP tool has many useful network biology functions that are demonstrated in this user guide. 1. The ANAP tool http://gmdd.shgmo.org/computational-biology/anap/anap_v1.0/ The ANAP tool
Page 1 of 11
 1800 990 432 Page 1 of 11 Table of Contents Registering Your Business... 3 Eligibility Criteria... 3 Navigating to the Regional Buy Portal... 3 Navigating the Registration Process... 3 The Registration
1800 990 432 Page 1 of 11 Table of Contents Registering Your Business... 3 Eligibility Criteria... 3 Navigating to the Regional Buy Portal... 3 Navigating the Registration Process... 3 The Registration
Overlap Checker & ENC Coverage User Manual
 Overlap Checker & ENC Coverage User Manual Document date: 01.01.2015 Contents Introduction... 3 Access to the VPN Check Overlap Candidates... 3 Coverage... 7 Copyright 2015 ECC AS Page 2 Introduction Overlap
Overlap Checker & ENC Coverage User Manual Document date: 01.01.2015 Contents Introduction... 3 Access to the VPN Check Overlap Candidates... 3 Coverage... 7 Copyright 2015 ECC AS Page 2 Introduction Overlap
Getting Started with the new GIS Map Service Overview:
 Getting Started with the new GIS Map Service Overview: 1. Layer List Widget Shows all available layers. This widget will be open by default. 2. Legend Widget Gives symbology information for all visible
Getting Started with the new GIS Map Service Overview: 1. Layer List Widget Shows all available layers. This widget will be open by default. 2. Legend Widget Gives symbology information for all visible
KaPPA-View 4. Manual for Beginners. ver Kazusa DNA Research Institute. The Kazusa Plant Pathway Viewer, Version 4.0
 KaPPA-View 4 The Kazusa Plant Pathway Viewer, Version 4.0 Manual for Beginners ver. 1.2 Kazusa DNA Research Institute Table of Contents Table of Contents 1. Introduction... 1 1-1. Overview of KaPPA-View4...
KaPPA-View 4 The Kazusa Plant Pathway Viewer, Version 4.0 Manual for Beginners ver. 1.2 Kazusa DNA Research Institute Table of Contents Table of Contents 1. Introduction... 1 1-1. Overview of KaPPA-View4...
Newaygo County Web Map
 Newaygo County Web Map Address/Parcel/Parcel Owner Search Map Overview Zoom Back to default extent Use your current location if allowable Widget Panel At the top of the map is a search function used for
Newaygo County Web Map Address/Parcel/Parcel Owner Search Map Overview Zoom Back to default extent Use your current location if allowable Widget Panel At the top of the map is a search function used for
Page 1 GM-FAQ Team FAQs. Page
 Page 1 Team FAQs Page How do I see my club's teams?... 2 How do I update my team's profile?... 3 How do I add/change my team's picture?... 5 Can I administer more than one team within my club?... 6 How
Page 1 Team FAQs Page How do I see my club's teams?... 2 How do I update my team's profile?... 3 How do I add/change my team's picture?... 5 Can I administer more than one team within my club?... 6 How
BILKENT UNIVERSITY. Bilkent Center for Bioinformatics
 versiion 2.1 BILKENT UNIVERSITY Bilkent Center for Bioinformatics PATİKAweb User s Guide BILKENT UNIVERSITY - CENTER FOR BIOINFORMATICS PATIKAweb User s Guide PATIKAweb 2007 Bilkent University Center for
versiion 2.1 BILKENT UNIVERSITY Bilkent Center for Bioinformatics PATİKAweb User s Guide BILKENT UNIVERSITY - CENTER FOR BIOINFORMATICS PATIKAweb User s Guide PATIKAweb 2007 Bilkent University Center for
WWF CMS Map Tool User Guide
 WWF CMS Map Tool User Guide July 2007 In the latest (1.7) release of the WWF Content Management System (CMS) a dynamic map creation tool is available in all CMS instances. Examples of maps created with
WWF CMS Map Tool User Guide July 2007 In the latest (1.7) release of the WWF Content Management System (CMS) a dynamic map creation tool is available in all CMS instances. Examples of maps created with
Processes Tab Downloads Tab Exercise Disease in Reactome Exercise
 Reactome This tutorial introduces Reactome, the user interface and the database content. Exercises help you practice what you have learned; you will need to refer to the details and screenshots in this
Reactome This tutorial introduces Reactome, the user interface and the database content. Exercises help you practice what you have learned; you will need to refer to the details and screenshots in this
Black Diamond. Black Diamond Advisor Experience: Portfolio View. Activation
 Advisor Experience: Portfolio View Portfolio View is the advisor s home for dynamic client reporting. It features a dashboard of reporting content that can be configured to your preferences. This User
Advisor Experience: Portfolio View Portfolio View is the advisor s home for dynamic client reporting. It features a dashboard of reporting content that can be configured to your preferences. This User
emerge Help Document Table of Contents
 Table of Contents Logging Into emerge... 2 Navigation Bar... 3 Main Menu... 4 Creating a New Order... 6 Order Checklist... 6 Information... 7 Overview... 8 Geography... 9 List Select... 12 Demographics...
Table of Contents Logging Into emerge... 2 Navigation Bar... 3 Main Menu... 4 Creating a New Order... 6 Order Checklist... 6 Information... 7 Overview... 8 Geography... 9 List Select... 12 Demographics...
emerge Help Document Table of Contents
 Table of Contents Logging Into emerge... 2 Navigation Bar... 3 Main Menu... 4 My Account... 6 My Information... 6 Manage Lists... 7 Manage Seeds... 8 Search/Add Suppress... 9 Update My Suppress... 10 Creating
Table of Contents Logging Into emerge... 2 Navigation Bar... 3 Main Menu... 4 My Account... 6 My Information... 6 Manage Lists... 7 Manage Seeds... 8 Search/Add Suppress... 9 Update My Suppress... 10 Creating
User s Guide: BIOS Online Mapping Tool to access the California Freshwater Species Database
 User s Guide: BIOS Online Mapping Tool to access the California Freshwater Species Database Last updated: December 2017 Tutorial created by: The Nature Conservancy Topics covered Part 1: Exporting a tabular
User s Guide: BIOS Online Mapping Tool to access the California Freshwater Species Database Last updated: December 2017 Tutorial created by: The Nature Conservancy Topics covered Part 1: Exporting a tabular
CPD Essentials User Guide
 CPD Essentials User Guide A practical introduction cii.co.uk/cpdessentials 2 Contents 3 Glossary and terminology 4 The home page 5 My Training Plan 6 Editing time spent on activities 7 Recording and managing
CPD Essentials User Guide A practical introduction cii.co.uk/cpdessentials 2 Contents 3 Glossary and terminology 4 The home page 5 My Training Plan 6 Editing time spent on activities 7 Recording and managing
1 User Guide. 1 Main screen
 1 User Guide 1 Main screen The opening screen appears in figure 1. Please wait until the loading bar (as shown in the bottom left) has filled up and the text changed from loading to completed. From the
1 User Guide 1 Main screen The opening screen appears in figure 1. Please wait until the loading bar (as shown in the bottom left) has filled up and the text changed from loading to completed. From the
40. Sim Module - Common Tools
 HSC Sim Common Tools 15021-ORC-J 1 (33) 40. Sim Module - Common Tools Table of Contents 40.1. Drawing flowsheets and adding tables to flowsheets... 2 40.1.1. Drawing units... 2 40.1.2. Drawing streams...
HSC Sim Common Tools 15021-ORC-J 1 (33) 40. Sim Module - Common Tools Table of Contents 40.1. Drawing flowsheets and adding tables to flowsheets... 2 40.1.1. Drawing units... 2 40.1.2. Drawing streams...
Quick Guide FAST HR. For more resources, including a guide on FAST HR codes, visit # Instructions Screenshot
 Tips & tricks This quick guide describes basic navigation within the FAST HR reporting tool, including how to use filter options, format columns and export reports. For more resources, including a guide
Tips & tricks This quick guide describes basic navigation within the FAST HR reporting tool, including how to use filter options, format columns and export reports. For more resources, including a guide
Getting Started with VicMap
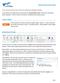 Getting Started with VicMap This is a brief overview of some of the tools and features available on VicMap. At any time you can right click on the map and click Identify What s Here to find more information
Getting Started with VicMap This is a brief overview of some of the tools and features available on VicMap. At any time you can right click on the map and click Identify What s Here to find more information
Script for Administering the Civics EOC Practice Test (epat)
 Script for Administering the Civics EOC Practice Test (epat) This script should be used to administer the Civics EOC Practice Test (epat) to students who will take the Civics EOC Assessment using TestNav
Script for Administering the Civics EOC Practice Test (epat) This script should be used to administer the Civics EOC Practice Test (epat) to students who will take the Civics EOC Assessment using TestNav
How to design and print cards using a database connection with. emedia CS Software
 How to design and print cards using a database connection with emedia CS Software For this exercise, we will use a Database that has been created in EXCEL. The example below shows the database fields populated
How to design and print cards using a database connection with emedia CS Software For this exercise, we will use a Database that has been created in EXCEL. The example below shows the database fields populated
Analysis of Networks with Interactive MOdelling. User s Manual
 Analysis of Networks with Interactive MOdelling User s Manual Contents 1 Requirements and installation 1 1.1 Java............................. 1 1.2 Cytoscape........................... 2 1.3 UPPAAL...........................
Analysis of Networks with Interactive MOdelling User s Manual Contents 1 Requirements and installation 1 1.1 Java............................. 1 1.2 Cytoscape........................... 2 1.3 UPPAAL...........................
Ambra User Guide. If you need help. Ambra Support (any time)
 If you need help Ambra Support 888 315 0790 (any time) support@ambrahealth.com Ambra User Guide Envision Radiology, a Health Images Organization, has provided a list of your site s personnel that need
If you need help Ambra Support 888 315 0790 (any time) support@ambrahealth.com Ambra User Guide Envision Radiology, a Health Images Organization, has provided a list of your site s personnel that need
FlexMLS Maps Quick Reference Guide
 FlexMLS Maps Quick Reference Guide Full Search Map Tab Features Define Search Areas box Map tab in Full Search Radius Search tool from an address Show/Hide Property List, Locate Address, and Define Search
FlexMLS Maps Quick Reference Guide Full Search Map Tab Features Define Search Areas box Map tab in Full Search Radius Search tool from an address Show/Hide Property List, Locate Address, and Define Search
Town of Amherst, NY. GIS Map Machine User Guide. Map Window Search Functions. Help. Toolbar. Layer Control. Scale Bar
 Town of Amherst, NY GIS Map Machine User Guide Toolbar Map Window Search Functions Help Layer Control Scale Bar Map Window Main Elements The MAP WINDOW is the main focus of the screen and where the map
Town of Amherst, NY GIS Map Machine User Guide Toolbar Map Window Search Functions Help Layer Control Scale Bar Map Window Main Elements The MAP WINDOW is the main focus of the screen and where the map
HOW-TO GUIDE. Join or Login. About this Guide!
 HOW-TO GUIDE About this Guide In this guide, you will learn about each section of the online community to help you make the best use of all it has to offer. Here you will find information on: Join or Login
HOW-TO GUIDE About this Guide In this guide, you will learn about each section of the online community to help you make the best use of all it has to offer. Here you will find information on: Join or Login
InSite Prepress. Quick Start Guide. Quick Links. Getting Started Uploading Files View Job Status Smart Review Preview Approving Pages Rejecting Pages
 InSite Prepress Quick Start Guide Quick Links Getting Started Uploading Files View Job Status Smart Review Preview Approving Pages Rejecting Pages BEAUTIFUL PRINT insite.panaprint.com 2015-3-16 Welcome
InSite Prepress Quick Start Guide Quick Links Getting Started Uploading Files View Job Status Smart Review Preview Approving Pages Rejecting Pages BEAUTIFUL PRINT insite.panaprint.com 2015-3-16 Welcome
L Y R A U S E R M A N U A L R A I N O T E S M O D U L E
 L Y R A U S E R M A N U A L R A I N O T E S M O D U L E CONTENTS 1. RAI Summary View... 2 1.1. RAI status... 2 1.2. Rules in RAI Summary View... 3 1.3. Customize RAI Summary View... 3 1.3.1. Show/hide
L Y R A U S E R M A N U A L R A I N O T E S M O D U L E CONTENTS 1. RAI Summary View... 2 1.1. RAI status... 2 1.2. Rules in RAI Summary View... 3 1.3. Customize RAI Summary View... 3 1.3.1. Show/hide
ALES Wordpress Editor documentation ALES Research websites
 ALES Wordpress Editor documentation ALES Research websites Contents Login... 2 Website Dashboard... 3 Editing menu order or structure... 4 Add a new page... 6 Move a page... 6 Select a page to edit...
ALES Wordpress Editor documentation ALES Research websites Contents Login... 2 Website Dashboard... 3 Editing menu order or structure... 4 Add a new page... 6 Move a page... 6 Select a page to edit...
SAS Report Viewer 8.2 Documentation
 SAS Report Viewer 8.2 Documentation About SAS Report Viewer SAS Report Viewer (the report viewer) enables users who are not report designers to view a report using a web browser. To open a report in the
SAS Report Viewer 8.2 Documentation About SAS Report Viewer SAS Report Viewer (the report viewer) enables users who are not report designers to view a report using a web browser. To open a report in the
This tutorial will guide a curator/user to create the files to upload phenotype experiment annotation and data onto T3.
 This tutorial will guide a curator/user to create the files to upload phenotype experiment annotation and data onto T3. Updated December 2012. Any feedback is welcome! Sections 1. Download the Template
This tutorial will guide a curator/user to create the files to upload phenotype experiment annotation and data onto T3. Updated December 2012. Any feedback is welcome! Sections 1. Download the Template
CUSTOMIZING CHECKPOINT TO WORK FOR YOU
 HOME CUSTOMIZING CHECKPOINT TO WORK FOR YOU QUICK REFERENCE Click Manage my views to modify your Current View or to create a new view. Edit and customize your Quick Links to access links to your frequently
HOME CUSTOMIZING CHECKPOINT TO WORK FOR YOU QUICK REFERENCE Click Manage my views to modify your Current View or to create a new view. Edit and customize your Quick Links to access links to your frequently
User manual for the CRTM LPWA
 User manual for the CRTM 3000 - LPWA September 2018 CONTENTS 1. Introduction 3 2. Specifications 3 a. 3 b. MTinfo 3000 minimum requirements 3 3. Conditions and safety instructions 4 4. MTinfo 3000 features
User manual for the CRTM 3000 - LPWA September 2018 CONTENTS 1. Introduction 3 2. Specifications 3 a. 3 b. MTinfo 3000 minimum requirements 3 3. Conditions and safety instructions 4 4. MTinfo 3000 features
GIS DATA SUBMISSION USER GUIDE. Innovation and Networks Executive Agency
 Innovation and Networks Executive Agency GIS DATA SUBMISSION USER GUIDE Innovation and Networks Executive Agency (INEA) W910 Chaussée de Wavre 910 B-1049 Brussels, Belgium Tel: +32 (0)2 29 95252 Fax: +32
Innovation and Networks Executive Agency GIS DATA SUBMISSION USER GUIDE Innovation and Networks Executive Agency (INEA) W910 Chaussée de Wavre 910 B-1049 Brussels, Belgium Tel: +32 (0)2 29 95252 Fax: +32
Login: Quick Guide for Qualtrics May 2018 Training:
 Qualtrics Basics Creating a New Qualtrics Account Note: Anyone with a Purdue career account can create a Qualtrics account. 1. In a Web browser, navigate to purdue.qualtrics.com. 2. Enter your Purdue Career
Qualtrics Basics Creating a New Qualtrics Account Note: Anyone with a Purdue career account can create a Qualtrics account. 1. In a Web browser, navigate to purdue.qualtrics.com. 2. Enter your Purdue Career
Learning how to use Lexis Red
 Learning how to use Lexis Red This guide takes you through how to use Lexis Red, the innovative new way of accessing looseleaf content from LexisNexis. If you still need assistance after reading this guide
Learning how to use Lexis Red This guide takes you through how to use Lexis Red, the innovative new way of accessing looseleaf content from LexisNexis. If you still need assistance after reading this guide
KIN 147 Lab 02: Acceleration Data Analysis
 KIN 147 Lab 02: Acceleration Data Analysis Excel analyses work much better on PCs than on Macs (especially older Macs) Your goal is to correctly analyze accelerometer data Analyzing the Acceleration Data
KIN 147 Lab 02: Acceleration Data Analysis Excel analyses work much better on PCs than on Macs (especially older Macs) Your goal is to correctly analyze accelerometer data Analyzing the Acceleration Data
Design Review: Fundamentals
 Design Review: Fundamentals Understanding Autodesk Design Review Autodesk Design Review improves team collaboration and communication by using design information the way it is intended to be used by the
Design Review: Fundamentals Understanding Autodesk Design Review Autodesk Design Review improves team collaboration and communication by using design information the way it is intended to be used by the
SAS Report Viewer 8.3 Documentation
 SAS Report Viewer 8.3 Documentation About SAS Report Viewer Introduction to SAS Report Viewer SAS Report Viewer (the report viewer) enables users who are not report designers to view a report using a web
SAS Report Viewer 8.3 Documentation About SAS Report Viewer Introduction to SAS Report Viewer SAS Report Viewer (the report viewer) enables users who are not report designers to view a report using a web
Structural Bioinformatics
 Structural Bioinformatics Elucidation of the 3D structures of biomolecules. Analysis and comparison of biomolecular structures. Prediction of biomolecular recognition. Handles three-dimensional (3-D) structures.
Structural Bioinformatics Elucidation of the 3D structures of biomolecules. Analysis and comparison of biomolecular structures. Prediction of biomolecular recognition. Handles three-dimensional (3-D) structures.
Exercises. Biological Data Analysis Using InterMine workshop exercises with answers
 Exercises Biological Data Analysis Using InterMine workshop exercises with answers Exercise1: Faceted Search Use HumanMine for this exercise 1. Search for one or more of the following using the keyword
Exercises Biological Data Analysis Using InterMine workshop exercises with answers Exercise1: Faceted Search Use HumanMine for this exercise 1. Search for one or more of the following using the keyword
I N S T A L L A T I O N
 USING REFWORKS FOR MACS I N S T A L L A T I O N You can access RefWorks through the University of Notre Dame Library website. Click on RefWorks underneath the Researching Help menu on the first page. Access
USING REFWORKS FOR MACS I N S T A L L A T I O N You can access RefWorks through the University of Notre Dame Library website. Click on RefWorks underneath the Researching Help menu on the first page. Access
Analysis of Networks with Interactive MOdelling. User s Manual
 Analysis of Networks with Interactive MOdelling User s Manual Contents 1 Requirements and installation 1 1.1 Java............................. 1 1.2 Cytoscape........................... 2 1.3 UPPAAL...........................
Analysis of Networks with Interactive MOdelling User s Manual Contents 1 Requirements and installation 1 1.1 Java............................. 1 1.2 Cytoscape........................... 2 1.3 UPPAAL...........................
Vetstreet Web Builder Editor Tool User Guide v2.1. Web Builder. User Guide v2.1
 Web Builder User Guide v2.1 Contact your Account Manager at (888) 799-8387 or email support@vetstreet.com with questions. Page 1 Index... 1 The Editor Tool... 7 Forgot Your Username or Password?... 7 How
Web Builder User Guide v2.1 Contact your Account Manager at (888) 799-8387 or email support@vetstreet.com with questions. Page 1 Index... 1 The Editor Tool... 7 Forgot Your Username or Password?... 7 How
Forms. Section 3: Deleting a Category
 9. If a category was NOT previously published, Authors may modify it by following the same procedures as an Administrator or Publisher. When the category is ready for publishing an Author must Save and
9. If a category was NOT previously published, Authors may modify it by following the same procedures as an Administrator or Publisher. When the category is ready for publishing an Author must Save and
IT Services Financial Services. IT Services Financial Services.
 eledgers IT Services Financial Services IT Services Financial Services http://finserv.uchicago.edu Table of Contents Logging into eledgers... 3 17BThe eledgers Workspace... 4 Basic Search using Custom
eledgers IT Services Financial Services IT Services Financial Services http://finserv.uchicago.edu Table of Contents Logging into eledgers... 3 17BThe eledgers Workspace... 4 Basic Search using Custom
