Segmentation of Thalamic Nuclei from DTI Using Spectral Clustering
|
|
|
- Marian Tate
- 5 years ago
- Views:
Transcription
1 Segmentation of Thalamic Nuclei from DTI Using Spectral Clustering Ulas Ziyan 1,DavidTuch 2, and Carl-Fredrik Westin 1,3 1 MIT Computer Science and Artificial Intelligence Lab, Cambridge MA, USA ulas@mit.edu 2 Novartis Pharma AG, Basel, Switzerland david.tuch@novartis.com 3 Laboratory of Mathematics in Imaging, Brigham and Women s Hospital, Harvard Medical School, Boston MA, USA westin@bwh.harvard.edu Abstract. Recent work shows that diffusion tensor imaging (DTI) can help resolving thalamic nuclei based on the characteristic fiber orientation of the corticothalamic/thalamocortical striations within each nucleus. In this paper we describe a novel segmentation method based on spectral clustering. We use Markovian relaxation to handle spatial information in a natural way, and we explicitly minimize the normalized cut criteria of the spectral clustering for a better optimization. Using this modified spectral clustering algorithm, we can resolve the organization of the thalamic nuclei into groups and subgroups solely based on the voxel affinity matrix, avoiding the need for explicitly defined cluster centers. The identification of nuclear subdivisions can facilitate localization of functional activation and pathology to individual nuclear subgroups. 1 Introduction All of the sensory pathways of the human brain (with the exception of the olfactorypathway)project to the cortex via relay neurons in the thalamus. These relay neurons are clustered into discrete clusters or nuclei that can be delineated based on histological or functional criteria. Delineation of the nuclei is important for surgical treatment of the thalamus and for localization of functional brain activation to specific nuclei. Recently, it has been shown that the thalamic nuclei can be resolved using a type of magnetic resonance imaging (MRI) called diffusion tensor imaging (DTI) [1, 2]. DTI measures the molecular diffusion, i.e., Brownian motion, of the endogenous water in tissue. In fibrous biological tissues such as cerebral white matter, water diffusion is anisotropic. In nerve tissue, the diffusion anisotropy is thought to be due to the diffusion barrier presented by the cell membrane, although the physical mechanisms for diffusion anisotropy in nerve tissue are not entirely understood [3]. On DTI, the thalamus shows distinct clusters of diffusion orientation. These clusters correspond in anatomic location and fiber orientation to the thalamic R. Larsen, M. Nielsen, and J. Sporring (Eds.): MICCAI 2006, LNCS 4191, pp , c Springer-Verlag Berlin Heidelberg 2006
2 808 U. Ziyan, D. Tuch, and C.-F. Westin nuclei [2]. Diffusion orientation thus provides a new anatomic criterion for distinguishing thalamic nuclei. Although DTI can resolve thalamic nuclei, a segmentation algorithm is required to explicitly delineate the nuclei from the DTI data. Several methods have been proposed to resolve the thalamic nuclei, including the use of k-means [2], level-sets [4] and tract reconstructions from manually defined regions on the cortical surface [1]. A weakness of the k-means algorithm is its geometric bias towards ellipsoidal clusters. Further, both k-means and level sets show sensitivity to initialization, and susceptibility to local minima. The present report describes a new approach for segmentation of thalamic nuclei using the spectral segmentation algorithm described by Shi and Malik [5] with some modifications. The spectral segmentation algorithm has attracted considerable interest in the pattern recognition community due its computational simplicity, strong empirical performance, and rich underlying theory [6, 7, 8]. The algorithm is based on a classical result from spectral graph partitioning which relates the second smallest eigenvector of the Laplacian matrix of a graph to optimal partitions. The spectral clustering algorithm has a number of desirable features. For example, the algorithm consists entirely of direct matrix operations and is therefore computationally efficient. Furthermore, the algorithm does not require a geometric representation of the clusters and therefore contains no explicit geometric biases. DTI data from 10 healthy participants were collected and segmented using the modified spectral clustering to demonstrate feasibility of the proposed method. 2 Theory Spectral clustering reduces segmentation into a graph partitioning problem. The nodes of the graph are chosen to be the data points, and the links connecting the nodes, commonly referred as edges, are assigned weights based on similarities of the data points being linked together. In the case of DTI data, the nodes can be chosen as the diffusion tensors, and the edge weights can be related to the similarities of the neighboring tensors. In Section 2.1, we will describe the details of the graph construction for DTI data, and in Section 2.2, we will describe a procedure to solve the graph partitioning problem via explicit minimization of an objective function defined on the graph. Once the graph partitioning is completed, the algorithm produces a hierarchical tree, from which different levels of segmentations can be extracted. 2.1 Graph Construction Our approach to nuclei segmentation is to partition the DTI data into compact regions with homogeneous diffusion properties, since similar diffusion properties are likely to be indicative of belonging to the same nuclei. For that purpose we are looking to construct a graph in which the nodes within a homogeneous region are well connected and the nodes in different homogeneous regions are not. This
3 Segmentation of Thalamic Nuclei from DTI Using Spectral Clustering 809 graph is constructed based on the spatial distances between the voxels as well as the tensor dissimilarities. The first step in the construction is to choose an appropriate metric to quantify the tensor dissimilarities. There are a number of possible metrics for this purpose, such as angular difference between the principle eigenvector directions, f 1 (T i, T j ) = arccos ( v i v j ), (1) where v i is the principal eigenvector of tensor T i. The absolute value of the dot product solves the problem with the sign ambiguity of the eigenvectors (if v 1 is an eigenvector, so is v 1 ). Another option is to use the full tensor information, f 2 (T i, T j )= trace((t i T j ) 2 ). (2) f 2 (T i, T j ) is explored in several DTI studies under names such as generalized tensor dot product [9], Frobenius norm [2] and Euclidean distance metric [10]. We also experimented with a recently proposed information theoretic measure, symmetric K-L divergence [11], f 3 (T i, T j )= trace(t 1 i T j + T 1 j T i ) 2n, (3) where n = 3 for diffusion tensors. After choosing a tensor dissimilarity metric, we need to incorporate spatial relations of the voxels into the graph. One way to do so is to linearly combine the tensor dissimilarity metric, f(t i, T j ), with a spatial distance, s(x i, x j ), as in [2]: d((x i, T i ), (x j, T j )) = f(t i, T j )+γs(x i, x j ), (4) where γ is a weighting factor to control the trade-off between the tensor dissimilarity and the spatial distances. However, selecting an appropriate weighting factor is complicated by the fact that the spatial distances and the tensor dissimilarities are inherently different types of features. To avoid this problem, we use Markovian relaxation, which provides a natural way to incorporate the spatial information. This is done by constructing a sparse affinity matrix, W s,with non-zero similarities only between face neighbors: f(t i,t j)2 {exp W s (i, j) = σ, if s(x 2 i, x j ) 1 0, otherwise. (5) The parameter σ is chosen to be the sample variance of f(t i, T j ) for neighboring voxels. We then calculate the full affinity matrix, W,usingMarkovian relaxation [12]. Specifically, we first convert the sparse weight matrix into a one-step transition probability matrix whose rows and columns add up to one: P 1 (i, j) = { 1 max l d(l) W s (i, j), if i j max l d(l) d(i), if i = j, (6)
4 810 U. Ziyan, D. Tuch, and C.-F. Westin where d is a vector containing row sums of W s, d(i) = j W s(i, j). The diagonal adjustment ensures that the underlying random walk has a uniform steady state distribution, and therefore every part of the corresponding Markov field is explored at the same speed during the relaxation. Once the one-step transition probability matrix is computed, it is possible to calculate the n-step transition probabilities through Markovian relaxation, which is equivalent to raising the one-step transition probability matrix to the power n: P n = P n 1 [12]. n is chosen to be smallest integer that will produce a non-zero transition probability between every node. Finally the diagonal elements of P n are set to zero to produce the full affinity matrix W. 2.2 Graph Cuts Once the graph is constructed, one can calculate the k-way graph partition using Normalized Cuts criteria. Formally, the problem is now reduced to the following: Given a graph, G =(V, E) and its edge weight matrix W = {ω ij }, find a set of disjoint sets V 1, V 2,..., V k, such that k i=1 V i = V, which minimizes the normalized cut (NCut): NCut (V 1, V 2,..., V k )= k i=1 asso(v i, V ) asso(v i, V i ) asso(v i, V ), (7) where asso(a, B) = i A,j B ω ij [5]. Finding the normalized cuts, even for the case where k =2,isNP-complete due to the combinatorial nature of the discrete permutations, therefore, we need a polynomial time algorithm that will provide a reasonable solution. Shi and Malik suggest a number of possibilities for such an algorithm. In this paper, we use a combination of the methods suggested. Specifically, we first normalize the weight matrix; M = D 1 W,whereD is a diagonal matrix with D(i, i) = j ω ij. We then calculate the two largest eigenvectors of the normalized weight matrix, M. The largest eigenvector is a vector of constants due to the normalization and therefore discarded. We then consider cutting W into two clusters by thresholding the second eigenvector of M at every possible threshold level, looking for the 2-way cut with the minimum NCut value. Once the cut is made, we repeat this process for each of the two newly created clusters, until a very high threshold of a 2-way NCut value is reached. This threshold is typically close to 1, which is the level at which every point is clustered by itself regardless of the weights. This splitting is followed by a greedy merging algorithm, minimizing NCut at every merging step, to reduce the number of clusters to k. During this merging, the algorithm produces a hierarchical tree, from which different levels of segmentation can be extracted by varying k. At the final stage of our algorithm we allow single data node swaps between clusters to further reduce the NCut value. At every iteration of this stage, the algorithm considers the possibility of reassigning a single node to another cluster. If the resulting cut has a smaller NCut value than before, that node is reassigned to its new cluster. The algorithm stops if there are nodes that can be reassigned.
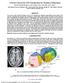
5 Segmentation of Thalamic Nuclei from DTI Using Spectral Clustering Methods 3.1 Image Acquisition and Pre-processing DTI data were acquired using a twice-refocused spin-echo EPI sequence [13] on a 3 Tesla Siemens Trio MRI scanner using an 8-channel head coil. The sequence parameters were TR/TE=8400/82 ms, b=700 s/mm 2, gmax=26 mt/m, 10 T2 images, 60 diffusion gradient directions, 1 average, total acquisition time 9 min 59 seconds. The field-of-view was mm and the matrix size was to give 2 2 mm in-plane resolution. The slice thickness was 2 mm with 0 mm gap. Correction for motion and residual eddy current distortion was applied by registering all of the scans to the first acquired non-diffusion-weighted scan for each participant. The registration used a 12 degree-of-freedom global affine transformation and a mutual information cost function [14]. Trilinear interpolation was used for the resampling. The diffusion tensor, the tensor eigensystem, and the FA metric were calculated for each voxel using the formulas of [15] and [16]. The diffusion tensor and FA volumes were normalized to MNI-space (Montreal Neurological Institute) by registering each participant s T2 volume to a skullstripped version of the MNI 152-subject T2 template [17] and then applying the transformation to the diffusion tensor and FA volumes with 12 degree-of-freedom global affine transformation and a mutual information cost function [14]. The registration transformation was then applied separately to the FA and tensor volumes. The FA volume was resampled using trilinear interpolation and the tensor volume was resampled using nearest neighbor interpolation. The tensors were reoriented using the rotational portion of the atlas transformation. 3.2 Segmentation Thalamus masks were drawn manually for each individual by a trained neuroanatomist. The masks were drawn for each hemisphere on each individual s MNInormalized FA map. Each hemisphere further segmented into its seven nuclei on the corresponding tensor map by the neuro-anatomist. The masked thalamus DTI data was later segmented separately for each hemisphere using the spectral algorithm described in Section 2. 4 Results For all thalamic data sets, spectral reorderingofthe voxelaffinity matrix revealed significant clustering structure (Figure 1D). The network created by the algorithm provides necessary information for the clustering and the spectral ordering followed by the modified spectral clustering algorithm was able to identify the clusters. These clusters are presented in Figure 2 (Right) for one of the individuals along with the expert labels (Left). The automatically segmented clusters are similar in both hemispheres which were processed independently, and they match well with the expert segmentation. All 10 subjects were segmented using the modified spectral clustering algorithm at two different levels (of the hierarchical tree) with k =7andk = 12. The
6 812 U. Ziyan, D. Tuch, and C.-F. Westin A C E B D F Fig. 1. A schematic outline of spectral segmentation algorithm. (A) A single slice tensor data (B) Initial graph corresponding to W s (C) Unordered W (D) Ordered and clustered W (E) Clusters on the slice (F) Clusters in 3D. Ant Ant MD MD VL/VP VL/VP LGN IL/CM IL/CM LGN MGN Pu Pu MGN Fig. 2. Left: 3D rendering of expert segmentation of both hemispheres from one subject. Right: The same subject segmented by spectral clustering with k = 12. experiment is repeated with W s and W to quantify the improvement achieved by the Markovian relaxation. The resulting clusters are then named in accordance with the expert labels. At this stage several clusters are allowed to inherit the same labels for a quantitative comparison. These final segmentations are then compared with the expert labels using the Dice volume overlap measure [18]. Mean and standard deviation of the volume overlaps from the 10 subjects are presented in Table 1.
7 Segmentation of Thalamic Nuclei from DTI Using Spectral Clustering 813 Table 1. Dice volume overlap comparison of spectral clustering results with expert labels for three tensor distance metrics f 1, f 2, f 3. The experiment is repeated with the sparse affinity matrix, W s, and with the product of Markovian relaxation, W,at clustering levels of k =7andk = 12. A score of 100 would indicate a perfect match. W s W W s W k =7 k =7 k =12 k =12 Tensor Dist. f 1 f 2 f 3 f 1 f 2 f 3 f 1 f 2 f 3 f 1 f 2 f 3 Mean St. Dev Conclusions In this paper, we presented a modified spectral clustering method for segmenting the thalamic nuclei using DTI data. Our method can automatically identify nuclear groups and subgroups solely based on the voxel affinity without any prior knowledge of cluster centers. Our experiments indicate that the angular distances between principle eigenvector directions of tensors produce more consistent segmentations compared to full tensor based measures. Acknowledgements The authors would like to thank Jonathan J Wisco for drawing the thalamus masks and assigning anatomical labels to the clusters. This work was supported by NINDS NS46532, NCRR RR14075, NCI CA09502, P41-RR13218 (NAC), GlaxoSmithKline, the Athinoula A. Martinos Foundation, the Mental Illness and Neuroscience Discovery (MIND) Institute, and the National Alliance for Medical Image Computing (NAMIC) (NIBIB U54 EB005149) which is funded through the NIH Roadmap for Medical Research. References [1] Behrens, T.E., Johansen-Berg, H., Woolrich, M.W., Smith, S.M., Wheeler- Kingshott, C.A., Boulby, P.A., Barker, G.J., Sillery, E.L., Sheehan, K., Ciccarelli, O., Thompson, A.J., Brady, J.M., Matthews, P.M.: Non-invasive mapping of connections between human thalamus and cortex using diffusion imaging. Nat Neurosci 6(7) (2003) [2] Wiegell, M.R., Tuch, D.S., Larsson, H.B., Wedeen, V.J.: Automatic segmentation of thalamic nuclei from diffusion tensor magnetic resonance imaging. Neuroimage 19 (2003) [3] Beaulieu, C.: The basis of anisotropic water diffusion in the nervous system - a technical review. NMR Biomed 15(7-8) (2002) [4] Jonasson, L., Hagmann, P., Richero Wilson, C., Bresson, X., Pollo, C., Meuli, R., Thiran, J.P.: Coupled, region based level sets for segmentation of the thalamus and its subnuclei in DT-MRI. In: ISMRM. (2005) 731
8 814 U. Ziyan, D. Tuch, and C.-F. Westin [5] Shi, J., Malik, J.: Normalized cuts and image segmentation. PAMI 22(8) (2000) [6] Higham, D., Kibble, M.: A unified view of spectral clustering. Mathematics Research Report (2004) 1 17 [7] Kannan, R., Vempala, S., Vetta, A.: On clusterings: Good, bad and spectral. J ACM 51(3) (2004) [8] Weiss, Y.: Segmentation using eigenvectors: A unifying view. In: ICCV. (1999) 975 [9] Jones, D.K., Horsfield, M.A., Simmons, A.: Optimal strategies for measuring diffusion in anisotropic systems by magnetic resonance imaging. MRM 42(3) (1999) [10] Alexander, D.C., Gee, J.C., R., B.: Similarity measures for matching diffusion tensor images. In: BMVC. (1999) [11] Wang, Z., Vemuri, B.C.: An affine invariant tensor dissimilarity measure and its applications to tensor-valued image segmentation. In: CVPR (1). (2004) [12] Tishby, N., Slonim, N.: Data clustering by markovian relaxation and the information bottleneck method. In: NIPS. (2000) [13] Reese, T.G., Heid, O., Weisskoff, R.M., Wedeen, V.J.: Reduction of eddy-currentinduced distortion in diffusion MRI using a twice-refocused spin echo. MRM 49(1) (2003) [14] Jenkinson, M., Bannister, P., Brady, M., Smith, S.: Improved optimization for the robust and accurate linear registration and motion correction of brain images. Neuroimage 17(2) (2002) [15] Basser, P.J., Mattiello, J., LeBihan, D.: Estimation of the effective self-diffusion tensor from the NMR spin echo. J Magn Reson B 103(3) (1994) [16] Pierpaoli, C., Basser, P.J.: Toward a quantitative assessment of diffusion anisotropy. MRM 36(6) (1996) [17] Mazziotta, J.C., Toga, A.W., Evans, A., Fox, P., Lancaster, J.: A probabilistic atlas of the human brain: theory and rationale for its development. the international consortium for brain mapping (icbm). Neuroimage 2(2) (1995) [18] Dice, L.R.: Measures of the amount of ecologic association between species. Ecology 26 (1945)
FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING. Francesca Pizzorni Ferrarese
 FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING Francesca Pizzorni Ferrarese Pipeline overview WM and GM Segmentation Registration Data reconstruction Tractography
FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING Francesca Pizzorni Ferrarese Pipeline overview WM and GM Segmentation Registration Data reconstruction Tractography
Fiber Selection from Diffusion Tensor Data based on Boolean Operators
 Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhof 1, G. Greiner 2, M. Buchfelder 3, C. Nimsky 4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhof 1, G. Greiner 2, M. Buchfelder 3, C. Nimsky 4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Acceleration of Probabilistic Tractography Using Multi-GPU Parallel Processing. Jungsoo Lee, Sun Mi Park, Dae-Shik Kim
 Acceleration of Probabilistic Tractography Using Multi-GPU Parallel Processing Jungsoo Lee, Sun Mi Park, Dae-Shik Kim Introduction In particular, probabilistic tractography requires relatively long computation
Acceleration of Probabilistic Tractography Using Multi-GPU Parallel Processing Jungsoo Lee, Sun Mi Park, Dae-Shik Kim Introduction In particular, probabilistic tractography requires relatively long computation
Evaluation of Local Filter Approaches for Diffusion Tensor based Fiber Tracking
 Evaluation of Local Filter Approaches for Diffusion Tensor based Fiber Tracking D. Merhof 1, M. Buchfelder 2, C. Nimsky 3 1 Visual Computing, University of Konstanz, Konstanz 2 Department of Neurosurgery,
Evaluation of Local Filter Approaches for Diffusion Tensor based Fiber Tracking D. Merhof 1, M. Buchfelder 2, C. Nimsky 3 1 Visual Computing, University of Konstanz, Konstanz 2 Department of Neurosurgery,
A Method for Registering Diffusion Weighted Magnetic Resonance Images
 A Method for Registering Diffusion Weighted Magnetic Resonance Images Xiaodong Tao and James V. Miller GE Research, Niskayuna, New York, USA Abstract. Diffusion weighted magnetic resonance (DWMR or DW)
A Method for Registering Diffusion Weighted Magnetic Resonance Images Xiaodong Tao and James V. Miller GE Research, Niskayuna, New York, USA Abstract. Diffusion weighted magnetic resonance (DWMR or DW)
A Novel Contrast for DTI Visualization for Thalamus Delineation
 A Novel Contrast for DTI Visualization for Thalamus Delineation Xian Fan a, Meredith Thompson a,b, John A. Bogovic a, Pierre-Louis Bazin c, Jerry L. Prince a,c a Johns Hopkins University, Baltimore, MD,
A Novel Contrast for DTI Visualization for Thalamus Delineation Xian Fan a, Meredith Thompson a,b, John A. Bogovic a, Pierre-Louis Bazin c, Jerry L. Prince a,c a Johns Hopkins University, Baltimore, MD,
Atelier 2 : Calcul Haute Performance et Sciences du Vivant Forum er juillet, Paris, France
 From Diffusion MR Image Analysis to Whole Brain Connectivity Simulation Jean-Philippe Thiran EPFL Lausanne, Switzerland EPFL - Lausanne HPC in life sciences at EPFL The Blue Brain project: create a biologically
From Diffusion MR Image Analysis to Whole Brain Connectivity Simulation Jean-Philippe Thiran EPFL Lausanne, Switzerland EPFL - Lausanne HPC in life sciences at EPFL The Blue Brain project: create a biologically
NIH Public Access Author Manuscript Med Image Comput Comput Assist Interv. Author manuscript; available in PMC 2009 December 4.
 NIH Public Access Author Manuscript Med Image Comput Comput Assist Interv. Author manuscript; available in PMC 2009 December 4. Published in final edited form as: Med Image Comput Comput Assist Interv.
NIH Public Access Author Manuscript Med Image Comput Comput Assist Interv. Author manuscript; available in PMC 2009 December 4. Published in final edited form as: Med Image Comput Comput Assist Interv.
HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Similarity Measures for Matching Diffusion Tensor Images
 Similarity Measures for Matching Diffusion Tensor Images Daniel Alexander, James Gee, Ruzena Bajcsy GRASP Lab. and Dept. Radiology U. of Penn. 40 Walnut St., Philadelphia, PA 904, USA daniel2@grip.cis.upenn.edu
Similarity Measures for Matching Diffusion Tensor Images Daniel Alexander, James Gee, Ruzena Bajcsy GRASP Lab. and Dept. Radiology U. of Penn. 40 Walnut St., Philadelphia, PA 904, USA daniel2@grip.cis.upenn.edu
The organization of the human cerebral cortex estimated by intrinsic functional connectivity
 1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Generation of Hulls Encompassing Neuronal Pathways Based on Tetrahedralization and 3D Alpha Shapes
 Generation of Hulls Encompassing Neuronal Pathways Based on Tetrahedralization and 3D Alpha Shapes Dorit Merhof 1,2, Martin Meister 1, Ezgi Bingöl 1, Peter Hastreiter 1,2, Christopher Nimsky 2,3, Günther
Generation of Hulls Encompassing Neuronal Pathways Based on Tetrahedralization and 3D Alpha Shapes Dorit Merhof 1,2, Martin Meister 1, Ezgi Bingöl 1, Peter Hastreiter 1,2, Christopher Nimsky 2,3, Günther
Clustering Fiber Traces Using Normalized Cuts
 Clustering Fiber Traces Using Normalized Cuts Anders Brun 1,4, Hans Knutsson 1, Hae-Jeong Park 2,3,4, Martha E. Shenton 2, and Carl-Fredrik Westin 4 1 Department. of Biomedical Engineering, Linköping University,
Clustering Fiber Traces Using Normalized Cuts Anders Brun 1,4, Hans Knutsson 1, Hae-Jeong Park 2,3,4, Martha E. Shenton 2, and Carl-Fredrik Westin 4 1 Department. of Biomedical Engineering, Linköping University,
Uncertainty in White Matter Fiber Tractography
 Uncertainty in White Matter Fiber Tractography Ola Friman and Carl-Fredrik Westin Laboratory of Mathematics in Imaging, Department of Radiology Brigham and Women s Hospital, Harvard Medical School Abstract.
Uncertainty in White Matter Fiber Tractography Ola Friman and Carl-Fredrik Westin Laboratory of Mathematics in Imaging, Department of Radiology Brigham and Women s Hospital, Harvard Medical School Abstract.
Fiber Selection from Diffusion Tensor Data based on Boolean Operators
 Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhofl, G. Greiner 2, M. Buchfelder 3, C. Nimsky4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhofl, G. Greiner 2, M. Buchfelder 3, C. Nimsky4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
MITK-DI. A new Diffusion Imaging Component for MITK. Klaus Fritzsche, Hans-Peter Meinzer
 MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
An Introduction To Automatic Tissue Classification Of Brain MRI. Colm Elliott Mar 2014
 An Introduction To Automatic Tissue Classification Of Brain MRI Colm Elliott Mar 2014 Tissue Classification Tissue classification is part of many processing pipelines. We often want to classify each voxel
An Introduction To Automatic Tissue Classification Of Brain MRI Colm Elliott Mar 2014 Tissue Classification Tissue classification is part of many processing pipelines. We often want to classify each voxel
NIH Public Access Author Manuscript Med Image Comput Comput Assist Interv. Author manuscript; available in PMC 2013 May 04.
 NIH Public Access Author Manuscript Published in final edited form as: Med Image Comput Comput Assist Interv. 2005 ; 8(0 1): 180 187. A Hamilton-Jacobi-Bellman approach to high angular resolution diffusion
NIH Public Access Author Manuscript Published in final edited form as: Med Image Comput Comput Assist Interv. 2005 ; 8(0 1): 180 187. A Hamilton-Jacobi-Bellman approach to high angular resolution diffusion
Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE)
 Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE) Mahnaz Maddah, Kelly H. Zou, William M. Wells, Ron Kikinis, and Simon K. Warfield
Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE) Mahnaz Maddah, Kelly H. Zou, William M. Wells, Ron Kikinis, and Simon K. Warfield
Fiber Tract Clustering on Manifolds With Dual Rooted-Graphs
 Fiber Tract Clustering on Manifolds With Dual Rooted-Graphs Andy Tsai 1,2 Carl-Fredrik Westin 1 Alfred O. Hero III 3 Alan S. Willsky 2 1 Dept. of Radiology 2 Dept. of Electrical Eng. 3 Dept. of Electrical
Fiber Tract Clustering on Manifolds With Dual Rooted-Graphs Andy Tsai 1,2 Carl-Fredrik Westin 1 Alfred O. Hero III 3 Alan S. Willsky 2 1 Dept. of Radiology 2 Dept. of Electrical Eng. 3 Dept. of Electrical
Streamline Flows for White Matter Fibre Pathway Segmentation in Diffusion MRI
 Streamline Flows for White Matter Fibre Pathway Segmentation in Diffusion MRI Peter Savadjiev 1, Jennifer S.W. Campbell 1,2,G.BrucePike 2, and Kaleem Siddiqi 1 McGill University, Montréal, QC, Canada 1
Streamline Flows for White Matter Fibre Pathway Segmentation in Diffusion MRI Peter Savadjiev 1, Jennifer S.W. Campbell 1,2,G.BrucePike 2, and Kaleem Siddiqi 1 McGill University, Montréal, QC, Canada 1
NEURO M203 & BIOMED M263 WINTER 2014
 NEURO M203 & BIOMED M263 WINTER 2014 MRI Lab 2: Neuroimaging Connectivity Lab In today s lab we will work with sample diffusion imaging data and the group averaged fmri data collected during your scanning
NEURO M203 & BIOMED M263 WINTER 2014 MRI Lab 2: Neuroimaging Connectivity Lab In today s lab we will work with sample diffusion imaging data and the group averaged fmri data collected during your scanning
Generating Fiber Crossing Phantoms Out of Experimental DWIs
 Generating Fiber Crossing Phantoms Out of Experimental DWIs Matthan Caan 1,2, Anne Willem de Vries 2, Ganesh Khedoe 2,ErikAkkerman 1, Lucas van Vliet 2, Kees Grimbergen 1, and Frans Vos 1,2 1 Department
Generating Fiber Crossing Phantoms Out of Experimental DWIs Matthan Caan 1,2, Anne Willem de Vries 2, Ganesh Khedoe 2,ErikAkkerman 1, Lucas van Vliet 2, Kees Grimbergen 1, and Frans Vos 1,2 1 Department
Constrained Reconstruction of Sparse Cardiac MR DTI Data
 Constrained Reconstruction of Sparse Cardiac MR DTI Data Ganesh Adluru 1,3, Edward Hsu, and Edward V.R. DiBella,3 1 Electrical and Computer Engineering department, 50 S. Central Campus Dr., MEB, University
Constrained Reconstruction of Sparse Cardiac MR DTI Data Ganesh Adluru 1,3, Edward Hsu, and Edward V.R. DiBella,3 1 Electrical and Computer Engineering department, 50 S. Central Campus Dr., MEB, University
Automatic Quantification of DTI Parameters along Fiber Bundles
 Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein 1, Simon Hermann 1, Olaf Konrad 1, Horst K. Hahn 1, and Heinz-Otto Peitgen 1 1 MeVis Research, 28359 Bremen Email: klein@mevis.de
Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein 1, Simon Hermann 1, Olaf Konrad 1, Horst K. Hahn 1, and Heinz-Otto Peitgen 1 1 MeVis Research, 28359 Bremen Email: klein@mevis.de
Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00
 Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00 Marcel Prastawa 1 and Guido Gerig 1 Abstract July 17, 2008 1 Scientific Computing and Imaging
Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00 Marcel Prastawa 1 and Guido Gerig 1 Abstract July 17, 2008 1 Scientific Computing and Imaging
Automatic Quantification of DTI Parameters along Fiber Bundles
 Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein, Simon Hermann, Olaf Konrad, Horst K. Hahn, Heinz-Otto Peitgen MeVis Research, 28359 Bremen Email: klein@mevis.de Abstract. We introduce
Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein, Simon Hermann, Olaf Konrad, Horst K. Hahn, Heinz-Otto Peitgen MeVis Research, 28359 Bremen Email: klein@mevis.de Abstract. We introduce
Exploring functional connectivity in fmri via clustering
 Exploring functional connectivity in fmri via clustering The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher
Exploring functional connectivity in fmri via clustering The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher
Complex Fiber Visualization
 Annales Mathematicae et Informaticae 34 (2007) pp. 103 109 http://www.ektf.hu/tanszek/matematika/ami Complex Fiber Visualization Henrietta Tomán a, Róbert Tornai b, Marianna Zichar c a Department of Computer
Annales Mathematicae et Informaticae 34 (2007) pp. 103 109 http://www.ektf.hu/tanszek/matematika/ami Complex Fiber Visualization Henrietta Tomán a, Róbert Tornai b, Marianna Zichar c a Department of Computer
Diffusion model fitting and tractography: A primer
 Diffusion model fitting and tractography: A primer Anastasia Yendiki HMS/MGH/MIT Athinoula A. Martinos Center for Biomedical Imaging 03/18/10 Why n how Diffusion model fitting and tractography 0/18 Why
Diffusion model fitting and tractography: A primer Anastasia Yendiki HMS/MGH/MIT Athinoula A. Martinos Center for Biomedical Imaging 03/18/10 Why n how Diffusion model fitting and tractography 0/18 Why
Appendix E1. Supplementary Methods. MR Image Acquisition. MR Image Analysis
 RSNA, 2015 10.1148/radiol.2015150532 Appendix E1 Supplementary Methods MR Image Acquisition By using a 1.5-T system (Avanto, Siemens Medical, Erlangen, Germany) under a program of regular maintenance (no
RSNA, 2015 10.1148/radiol.2015150532 Appendix E1 Supplementary Methods MR Image Acquisition By using a 1.5-T system (Avanto, Siemens Medical, Erlangen, Germany) under a program of regular maintenance (no
UCLA Advanced Neuroimaging Summer Program David W. Shattuck, PhD
 BrainSuite UCLA Advanced Neuroimaging Summer Program Presented 18 July 2013 David W. Shattuck, PhD Associate Professor Department of Neurology David Geffen School of Medicine at UCLA http://www.loni.ucla.edu/~shattuck/
BrainSuite UCLA Advanced Neuroimaging Summer Program Presented 18 July 2013 David W. Shattuck, PhD Associate Professor Department of Neurology David Geffen School of Medicine at UCLA http://www.loni.ucla.edu/~shattuck/
Visual Representations for Machine Learning
 Visual Representations for Machine Learning Spectral Clustering and Channel Representations Lecture 1 Spectral Clustering: introduction and confusion Michael Felsberg Klas Nordberg The Spectral Clustering
Visual Representations for Machine Learning Spectral Clustering and Channel Representations Lecture 1 Spectral Clustering: introduction and confusion Michael Felsberg Klas Nordberg The Spectral Clustering
A Ray-based Approach for Boundary Estimation of Fiber Bundles Derived from Diffusion Tensor Imaging
 A Ray-based Approach for Boundary Estimation of Fiber Bundles Derived from Diffusion Tensor Imaging M. H. A. Bauer 1,3, S. Barbieri 2, J. Klein 2, J. Egger 1,3, D. Kuhnt 1, B. Freisleben 3, H.-K. Hahn
A Ray-based Approach for Boundary Estimation of Fiber Bundles Derived from Diffusion Tensor Imaging M. H. A. Bauer 1,3, S. Barbieri 2, J. Klein 2, J. Egger 1,3, D. Kuhnt 1, B. Freisleben 3, H.-K. Hahn
MITK-DI. A new Diffusion Imaging Component for MITK. Klaus Fritzsche, Hans-Peter Meinzer
 MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
Head motion in diffusion MRI
 Head motion in diffusion MRI Anastasia Yendiki HMS/MGH/MIT Athinoula A. Martinos Center for Biomedical Imaging 11/06/13 Head motion in diffusion MRI 0/33 Diffusion contrast Basic principle of diffusion
Head motion in diffusion MRI Anastasia Yendiki HMS/MGH/MIT Athinoula A. Martinos Center for Biomedical Imaging 11/06/13 Head motion in diffusion MRI 0/33 Diffusion contrast Basic principle of diffusion
Computational Neuroanatomy
 Computational Neuroanatomy John Ashburner john@fil.ion.ucl.ac.uk Smoothing Motion Correction Between Modality Co-registration Spatial Normalisation Segmentation Morphometry Overview fmri time-series kernel
Computational Neuroanatomy John Ashburner john@fil.ion.ucl.ac.uk Smoothing Motion Correction Between Modality Co-registration Spatial Normalisation Segmentation Morphometry Overview fmri time-series kernel
Spectral Clustering Algorithms for Ultrasound Image Segmentation
 Spectral Clustering Algorithms for Ultrasound Image Segmentation Neculai Archip 1, Robert Rohling 2, Peter Cooperberg 3, Hamid Tahmasebpour 3, and Simon K. Warfield 1 1 Computational Radiology Laboratory,
Spectral Clustering Algorithms for Ultrasound Image Segmentation Neculai Archip 1, Robert Rohling 2, Peter Cooperberg 3, Hamid Tahmasebpour 3, and Simon K. Warfield 1 1 Computational Radiology Laboratory,
Group Statistics of DTI Fiber Bundles Using Spatial Functions of Tensor Measures
 Group Statistics of DTI Fiber Bundles Using Spatial Functions of Tensor Measures Casey B. Goodlett 1,2, P. Thomas Fletcher 1,2, John H. Gilmore 3, and Guido Gerig 1,2 1 School of Computing, University
Group Statistics of DTI Fiber Bundles Using Spatial Functions of Tensor Measures Casey B. Goodlett 1,2, P. Thomas Fletcher 1,2, John H. Gilmore 3, and Guido Gerig 1,2 1 School of Computing, University
Functional MRI in Clinical Research and Practice Preprocessing
 Functional MRI in Clinical Research and Practice Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization
Functional MRI in Clinical Research and Practice Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization
Basic fmri Design and Analysis. Preprocessing
 Basic fmri Design and Analysis Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial filtering
Basic fmri Design and Analysis Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial filtering
Advanced Visual Medicine: Techniques for Visual Exploration & Analysis
 Advanced Visual Medicine: Techniques for Visual Exploration & Analysis Interactive Visualization of Multimodal Volume Data for Neurosurgical Planning Felix Ritter, MeVis Research Bremen Multimodal Neurosurgical
Advanced Visual Medicine: Techniques for Visual Exploration & Analysis Interactive Visualization of Multimodal Volume Data for Neurosurgical Planning Felix Ritter, MeVis Research Bremen Multimodal Neurosurgical
Technische Universiteit Eindhoven Department of Mathematics and Computer Science. Master s Thesis HIERARCHICAL VISUALIZATION USING FIBER CLUSTERING
 Technische Universiteit Eindhoven Department of Mathematics and Computer Science Master s Thesis HIERARCHICAL VISUALIZATION USING FIBER CLUSTERING by Ing. B. Moberts Supervisors: Dr. A. Vilanova Prof.dr.ir.
Technische Universiteit Eindhoven Department of Mathematics and Computer Science Master s Thesis HIERARCHICAL VISUALIZATION USING FIBER CLUSTERING by Ing. B. Moberts Supervisors: Dr. A. Vilanova Prof.dr.ir.
Estimation of Extracellular Volume from Regularized Multi-shell Diffusion MRI
 Estimation of Extracellular Volume from Regularized Multi-shell Diffusion MRI The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters
Estimation of Extracellular Volume from Regularized Multi-shell Diffusion MRI The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters
SPM8 for Basic and Clinical Investigators. Preprocessing. fmri Preprocessing
 SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
BrainSuite. presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014
 BrainSuite presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 David Shattuck Ahmanson-Lovelace Brain Mapping Center Department of Neurology David Geffen School of Medicine at
BrainSuite presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 David Shattuck Ahmanson-Lovelace Brain Mapping Center Department of Neurology David Geffen School of Medicine at
Joint Reconstruction of Multi-contrast MR Images for Multiple Sclerosis Lesion Segmentation
 Joint Reconstruction of Multi-contrast MR Images for Multiple Sclerosis Lesion Segmentation Pedro A Gómez 1,2,3, Jonathan I Sperl 3, Tim Sprenger 2,3, Claudia Metzler-Baddeley 4, Derek K Jones 4, Philipp
Joint Reconstruction of Multi-contrast MR Images for Multiple Sclerosis Lesion Segmentation Pedro A Gómez 1,2,3, Jonathan I Sperl 3, Tim Sprenger 2,3, Claudia Metzler-Baddeley 4, Derek K Jones 4, Philipp
Diffusion Tensor Processing and Visualization
 NA-MIC National Alliance for Medical Image Computing http://na-mic.org Diffusion Tensor Processing and Visualization Guido Gerig University of Utah Martin Styner, UNC NAMIC: National Alliance for Medical
NA-MIC National Alliance for Medical Image Computing http://na-mic.org Diffusion Tensor Processing and Visualization Guido Gerig University of Utah Martin Styner, UNC NAMIC: National Alliance for Medical
syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets
 syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets Julien Gervais; Lisa Chuah Siemens Healthcare, Magnetic
syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets Julien Gervais; Lisa Chuah Siemens Healthcare, Magnetic
EPI Data Are Acquired Serially. EPI Data Are Acquired Serially 10/23/2011. Functional Connectivity Preprocessing. fmri Preprocessing
 Functional Connectivity Preprocessing Geometric distortion Head motion Geometric distortion Head motion EPI Data Are Acquired Serially EPI Data Are Acquired Serially descending 1 EPI Data Are Acquired
Functional Connectivity Preprocessing Geometric distortion Head motion Geometric distortion Head motion EPI Data Are Acquired Serially EPI Data Are Acquired Serially descending 1 EPI Data Are Acquired
Building an Average Population HARDI Atlas
 Building an Average Population HARDI Atlas Sylvain Bouix 1, Yogesh Rathi 1, and Mert Sabuncu 2 1 Psychiatry Neuroimaging Laboratory, Brigham and Women s Hospital, Harvard Medical School, Boston, MA, USA.
Building an Average Population HARDI Atlas Sylvain Bouix 1, Yogesh Rathi 1, and Mert Sabuncu 2 1 Psychiatry Neuroimaging Laboratory, Brigham and Women s Hospital, Harvard Medical School, Boston, MA, USA.
Efficient population registration of 3D data
 Efficient population registration of 3D data Lilla Zöllei 1, Erik Learned-Miller 2, Eric Grimson 1, William Wells 1,3 1 Computer Science and Artificial Intelligence Lab, MIT; 2 Dept. of Computer Science,
Efficient population registration of 3D data Lilla Zöllei 1, Erik Learned-Miller 2, Eric Grimson 1, William Wells 1,3 1 Computer Science and Artificial Intelligence Lab, MIT; 2 Dept. of Computer Science,
Reconstruction of Fiber Trajectories via Population-Based Estimation of Local Orientations
 IDEA Reconstruction of Fiber Trajectories via Population-Based Estimation of Local Orientations Pew-Thian Yap, John H. Gilmore, Weili Lin, Dinggang Shen Email: ptyap@med.unc.edu 2011-09-21 Poster: P2-46-
IDEA Reconstruction of Fiber Trajectories via Population-Based Estimation of Local Orientations Pew-Thian Yap, John H. Gilmore, Weili Lin, Dinggang Shen Email: ptyap@med.unc.edu 2011-09-21 Poster: P2-46-
Role of Parallel Imaging in High Field Functional MRI
 Role of Parallel Imaging in High Field Functional MRI Douglas C. Noll & Bradley P. Sutton Department of Biomedical Engineering, University of Michigan Supported by NIH Grant DA15410 & The Whitaker Foundation
Role of Parallel Imaging in High Field Functional MRI Douglas C. Noll & Bradley P. Sutton Department of Biomedical Engineering, University of Michigan Supported by NIH Grant DA15410 & The Whitaker Foundation
Hierarchical Fiber Clustering Based on Multi-scale Neuroanatomical Features
 Hierarchical Fiber Clustering Based on Multi-scale Neuroanatomical Features Qian Wang 1, 2, Pew-Thian Yap 2, Hongjun Jia 2, Guorong Wu 2, and Dinggang Shen 2 1 Department of Computer Science, University
Hierarchical Fiber Clustering Based on Multi-scale Neuroanatomical Features Qian Wang 1, 2, Pew-Thian Yap 2, Hongjun Jia 2, Guorong Wu 2, and Dinggang Shen 2 1 Department of Computer Science, University
Preprocessing II: Between Subjects John Ashburner
 Preprocessing II: Between Subjects John Ashburner Pre-processing Overview Statistics or whatever fmri time-series Anatomical MRI Template Smoothed Estimate Spatial Norm Motion Correct Smooth Coregister
Preprocessing II: Between Subjects John Ashburner Pre-processing Overview Statistics or whatever fmri time-series Anatomical MRI Template Smoothed Estimate Spatial Norm Motion Correct Smooth Coregister
Fmri Spatial Processing
 Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
Spatial Regularization of Functional Connectivity Using High-Dimensional Markov Random Fields
 Spatial Regularization of Functional Connectivity Using High-Dimensional Markov Random Fields Wei Liu 1, Peihong Zhu 1, Jeffrey S. Anderson 2, Deborah Yurgelun-Todd 3, and P. Thomas Fletcher 1 1 Scientific
Spatial Regularization of Functional Connectivity Using High-Dimensional Markov Random Fields Wei Liu 1, Peihong Zhu 1, Jeffrey S. Anderson 2, Deborah Yurgelun-Todd 3, and P. Thomas Fletcher 1 1 Scientific
Super-Resolution Reconstruction of Diffusion-Weighted Images from Distortion Compensated Orthogonal Anisotropic Acquisitions.
 Super-Resolution Reconstruction of Diffusion-Weighted Images from Distortion Compensated Orthogonal Anisotropic Acquisitions. Benoit Scherrer Ali Gholipour Simon K. Warfield Children s Hospital Boston,
Super-Resolution Reconstruction of Diffusion-Weighted Images from Distortion Compensated Orthogonal Anisotropic Acquisitions. Benoit Scherrer Ali Gholipour Simon K. Warfield Children s Hospital Boston,
SURFACE RECONSTRUCTION OF EX-VIVO HUMAN V1 THROUGH IDENTIFICATION OF THE STRIA OF GENNARI USING MRI AT 7T
 SURFACE RECONSTRUCTION OF EX-VIVO HUMAN V1 THROUGH IDENTIFICATION OF THE STRIA OF GENNARI USING MRI AT 7T Oliver P. Hinds 1, Jonathan R. Polimeni 2, Megan L. Blackwell 3, Christopher J. Wiggins 3, Graham
SURFACE RECONSTRUCTION OF EX-VIVO HUMAN V1 THROUGH IDENTIFICATION OF THE STRIA OF GENNARI USING MRI AT 7T Oliver P. Hinds 1, Jonathan R. Polimeni 2, Megan L. Blackwell 3, Christopher J. Wiggins 3, Graham
Image Analysis - Lecture 5
 Texture Segmentation Clustering Review Image Analysis - Lecture 5 Texture and Segmentation Magnus Oskarsson Lecture 5 Texture Segmentation Clustering Review Contents Texture Textons Filter Banks Gabor
Texture Segmentation Clustering Review Image Analysis - Lecture 5 Texture and Segmentation Magnus Oskarsson Lecture 5 Texture Segmentation Clustering Review Contents Texture Textons Filter Banks Gabor
CS 534: Computer Vision Segmentation and Perceptual Grouping
 CS 534: Computer Vision Segmentation and Perceptual Grouping Ahmed Elgammal Dept of Computer Science CS 534 Segmentation - 1 Outlines Mid-level vision What is segmentation Perceptual Grouping Segmentation
CS 534: Computer Vision Segmentation and Perceptual Grouping Ahmed Elgammal Dept of Computer Science CS 534 Segmentation - 1 Outlines Mid-level vision What is segmentation Perceptual Grouping Segmentation
SYDE Winter 2011 Introduction to Pattern Recognition. Clustering
 SYDE 372 - Winter 2011 Introduction to Pattern Recognition Clustering Alexander Wong Department of Systems Design Engineering University of Waterloo Outline 1 2 3 4 5 All the approaches we have learned
SYDE 372 - Winter 2011 Introduction to Pattern Recognition Clustering Alexander Wong Department of Systems Design Engineering University of Waterloo Outline 1 2 3 4 5 All the approaches we have learned
White Pixel Artifact. Caused by a noise spike during acquisition Spike in K-space <--> sinusoid in image space
 White Pixel Artifact Caused by a noise spike during acquisition Spike in K-space sinusoid in image space Susceptibility Artifacts Off-resonance artifacts caused by adjacent regions with different
White Pixel Artifact Caused by a noise spike during acquisition Spike in K-space sinusoid in image space Susceptibility Artifacts Off-resonance artifacts caused by adjacent regions with different
SPM8 for Basic and Clinical Investigators. Preprocessing
 SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
Image Segmentation. Srikumar Ramalingam School of Computing University of Utah. Slides borrowed from Ross Whitaker
 Image Segmentation Srikumar Ramalingam School of Computing University of Utah Slides borrowed from Ross Whitaker Segmentation Semantic Segmentation Indoor layout estimation What is Segmentation? Partitioning
Image Segmentation Srikumar Ramalingam School of Computing University of Utah Slides borrowed from Ross Whitaker Segmentation Semantic Segmentation Indoor layout estimation What is Segmentation? Partitioning
Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases
 Quantities Measured by MR - Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases Static parameters (influenced by molecular environment): T, T* (transverse relaxation)
Quantities Measured by MR - Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases Static parameters (influenced by molecular environment): T, T* (transverse relaxation)
Diffusion Imaging Visualization
 Diffusion Imaging Visualization Thomas Schultz URL: http://cg.cs.uni-bonn.de/schultz/ E-Mail: schultz@cs.uni-bonn.de 1 Outline Introduction to Diffusion Imaging Basic techniques Glyph-based Visualization
Diffusion Imaging Visualization Thomas Schultz URL: http://cg.cs.uni-bonn.de/schultz/ E-Mail: schultz@cs.uni-bonn.de 1 Outline Introduction to Diffusion Imaging Basic techniques Glyph-based Visualization
A Generic Lie Group Model for Computer Vision
 A Generic Lie Group Model for Computer Vision Within this research track we follow a generic Lie group approach to computer vision based on recent physiological research on how the primary visual cortex
A Generic Lie Group Model for Computer Vision Within this research track we follow a generic Lie group approach to computer vision based on recent physiological research on how the primary visual cortex
BDP: BrainSuite Diffusion Pipeline. Chitresh Bhushan
 BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
Blood Particle Trajectories in Phase-Contrast-MRI as Minimal Paths Computed with Anisotropic Fast Marching
 Blood Particle Trajectories in Phase-Contrast-MRI as Minimal Paths Computed with Anisotropic Fast Marching Michael Schwenke 1, Anja Hennemuth 1, Bernd Fischer 2, Ola Friman 1 1 Fraunhofer MEVIS, Institute
Blood Particle Trajectories in Phase-Contrast-MRI as Minimal Paths Computed with Anisotropic Fast Marching Michael Schwenke 1, Anja Hennemuth 1, Bernd Fischer 2, Ola Friman 1 1 Fraunhofer MEVIS, Institute
Diffusion-MRI processing for group analysis
 Diffusion-MRI processing for group analysis Felix Renard IRMaGe: Inserm US 17 / CNRS UMS 3552 University Hospital of Grenoble - France 25/09/2015 felixrenard@gmail.com 1 Diffusion-MRI processing for group
Diffusion-MRI processing for group analysis Felix Renard IRMaGe: Inserm US 17 / CNRS UMS 3552 University Hospital of Grenoble - France 25/09/2015 felixrenard@gmail.com 1 Diffusion-MRI processing for group
Diffusion MRI Acquisition. Karla Miller FMRIB Centre, University of Oxford
 Diffusion MRI Acquisition Karla Miller FMRIB Centre, University of Oxford karla@fmrib.ox.ac.uk Diffusion Imaging How is diffusion weighting achieved? How is the image acquired? What are the limitations,
Diffusion MRI Acquisition Karla Miller FMRIB Centre, University of Oxford karla@fmrib.ox.ac.uk Diffusion Imaging How is diffusion weighting achieved? How is the image acquired? What are the limitations,
Sparse sampling in MRI: From basic theory to clinical application. R. Marc Lebel, PhD Department of Electrical Engineering Department of Radiology
 Sparse sampling in MRI: From basic theory to clinical application R. Marc Lebel, PhD Department of Electrical Engineering Department of Radiology Objective Provide an intuitive overview of compressed sensing
Sparse sampling in MRI: From basic theory to clinical application R. Marc Lebel, PhD Department of Electrical Engineering Department of Radiology Objective Provide an intuitive overview of compressed sensing
C. Leon Partain JOURNAL OF MAGNETIC RESONANCE IMAGING EDITOR-IN-CHIEF
 JOURNAL OF MAGNETIC RESONANCE IMAGING ri m j / m o c. E ning M C ear l lth a he li ey w w. w w VOLUME 37 NUMBER 6 JUNE 2013 VOLUME 37 NUMBER 6 JUNE 2013 PAGES 1257 1504 CORRECTION OF EDDY CURRENT DISTORTIONS
JOURNAL OF MAGNETIC RESONANCE IMAGING ri m j / m o c. E ning M C ear l lth a he li ey w w. w w VOLUME 37 NUMBER 6 JUNE 2013 VOLUME 37 NUMBER 6 JUNE 2013 PAGES 1257 1504 CORRECTION OF EDDY CURRENT DISTORTIONS
Ischemic Stroke Lesion Segmentation Proceedings 5th October 2015 Munich, Germany
 0111010001110001101000100101010111100111011100100011011101110101101012 Ischemic Stroke Lesion Segmentation www.isles-challenge.org Proceedings 5th October 2015 Munich, Germany Preface Stroke is the second
0111010001110001101000100101010111100111011100100011011101110101101012 Ischemic Stroke Lesion Segmentation www.isles-challenge.org Proceedings 5th October 2015 Munich, Germany Preface Stroke is the second
Modern Medical Image Analysis 8DC00 Exam
 Parts of answers are inside square brackets [... ]. These parts are optional. Answers can be written in Dutch or in English, as you prefer. You can use drawings and diagrams to support your textual answers.
Parts of answers are inside square brackets [... ]. These parts are optional. Answers can be written in Dutch or in English, as you prefer. You can use drawings and diagrams to support your textual answers.
A Graph Clustering Algorithm Based on Minimum and Normalized Cut
 A Graph Clustering Algorithm Based on Minimum and Normalized Cut Jiabing Wang 1, Hong Peng 1, Jingsong Hu 1, and Chuangxin Yang 1, 1 School of Computer Science and Engineering, South China University of
A Graph Clustering Algorithm Based on Minimum and Normalized Cut Jiabing Wang 1, Hong Peng 1, Jingsong Hu 1, and Chuangxin Yang 1, 1 School of Computer Science and Engineering, South China University of
COBRE Scan Information
 COBRE Scan Information Below is more information on the directory structure for the COBRE imaging data. Also below are the imaging parameters for each series. Directory structure: var/www/html/dropbox/1139_anonymized/human:
COBRE Scan Information Below is more information on the directory structure for the COBRE imaging data. Also below are the imaging parameters for each series. Directory structure: var/www/html/dropbox/1139_anonymized/human:
Feasibility and Advantages of Diffusion Weighted Imaging Atlas Construction in Q-Space
 Feasibility and Advantages of Diffusion Weighted Imaging Atlas Construction in Q-Space Thijs Dhollander 1,2, Jelle Veraart 3,WimVanHecke 1,4,5, Frederik Maes 1,2, Stefan Sunaert 1,4,JanSijbers 3, and Paul
Feasibility and Advantages of Diffusion Weighted Imaging Atlas Construction in Q-Space Thijs Dhollander 1,2, Jelle Veraart 3,WimVanHecke 1,4,5, Frederik Maes 1,2, Stefan Sunaert 1,4,JanSijbers 3, and Paul
Behavioral Data Mining. Lecture 18 Clustering
 Behavioral Data Mining Lecture 18 Clustering Outline Why? Cluster quality K-means Spectral clustering Generative Models Rationale Given a set {X i } for i = 1,,n, a clustering is a partition of the X i
Behavioral Data Mining Lecture 18 Clustering Outline Why? Cluster quality K-means Spectral clustering Generative Models Rationale Given a set {X i } for i = 1,,n, a clustering is a partition of the X i
6.801/866. Segmentation and Line Fitting. T. Darrell
 6.801/866 Segmentation and Line Fitting T. Darrell Segmentation and Line Fitting Gestalt grouping Background subtraction K-Means Graph cuts Hough transform Iterative fitting (Next time: Probabilistic segmentation)
6.801/866 Segmentation and Line Fitting T. Darrell Segmentation and Line Fitting Gestalt grouping Background subtraction K-Means Graph cuts Hough transform Iterative fitting (Next time: Probabilistic segmentation)
Graph-Shifts Anatomic 3D Segmentation by Dynamic Hierarchical Minimization
 Graph-Shifts Anatomic 3D Segmentation by Dynamic Hierarchical Minimization Jason Corso Postdoctoral Fellow UCLA LONI/CCB jcorso@ucla.edu Motivation The work deals with the problem of automatically labeling
Graph-Shifts Anatomic 3D Segmentation by Dynamic Hierarchical Minimization Jason Corso Postdoctoral Fellow UCLA LONI/CCB jcorso@ucla.edu Motivation The work deals with the problem of automatically labeling
Whole Body MRI Intensity Standardization
 Whole Body MRI Intensity Standardization Florian Jäger 1, László Nyúl 1, Bernd Frericks 2, Frank Wacker 2 and Joachim Hornegger 1 1 Institute of Pattern Recognition, University of Erlangen, {jaeger,nyul,hornegger}@informatik.uni-erlangen.de
Whole Body MRI Intensity Standardization Florian Jäger 1, László Nyúl 1, Bernd Frericks 2, Frank Wacker 2 and Joachim Hornegger 1 1 Institute of Pattern Recognition, University of Erlangen, {jaeger,nyul,hornegger}@informatik.uni-erlangen.de
Correspondence Detection Using Wavelet-Based Attribute Vectors
 Correspondence Detection Using Wavelet-Based Attribute Vectors Zhong Xue, Dinggang Shen, and Christos Davatzikos Section of Biomedical Image Analysis, Department of Radiology University of Pennsylvania,
Correspondence Detection Using Wavelet-Based Attribute Vectors Zhong Xue, Dinggang Shen, and Christos Davatzikos Section of Biomedical Image Analysis, Department of Radiology University of Pennsylvania,
ECS 234: Data Analysis: Clustering ECS 234
 : Data Analysis: Clustering What is Clustering? Given n objects, assign them to groups (clusters) based on their similarity Unsupervised Machine Learning Class Discovery Difficult, and maybe ill-posed
: Data Analysis: Clustering What is Clustering? Given n objects, assign them to groups (clusters) based on their similarity Unsupervised Machine Learning Class Discovery Difficult, and maybe ill-posed
Direct Registration of White Matter Tractographies with Application to Atlas Construction
 Direct Registration of White Matter Tractographies with Application to Atlas Construction Arnaldo Mayer, Hayit Greenspan Medical image processing lab, Tel-Aviv University, Ramat-Aviv, Israel arnakdom@post.tau.ac.il,
Direct Registration of White Matter Tractographies with Application to Atlas Construction Arnaldo Mayer, Hayit Greenspan Medical image processing lab, Tel-Aviv University, Ramat-Aviv, Israel arnakdom@post.tau.ac.il,
Normalized cuts and image segmentation
 Normalized cuts and image segmentation Department of EE University of Washington Yeping Su Xiaodan Song Normalized Cuts and Image Segmentation, IEEE Trans. PAMI, August 2000 5/20/2003 1 Outline 1. Image
Normalized cuts and image segmentation Department of EE University of Washington Yeping Su Xiaodan Song Normalized Cuts and Image Segmentation, IEEE Trans. PAMI, August 2000 5/20/2003 1 Outline 1. Image
Segmenting Glioma in Multi-Modal Images using a Generative Model for Brain Lesion Segmentation
 Segmenting Glioma in Multi-Modal Images using a Generative Model for Brain Lesion Segmentation Bjoern H. Menze 1,2, Koen Van Leemput 3, Danial Lashkari 4 Marc-André Weber 5, Nicholas Ayache 2, and Polina
Segmenting Glioma in Multi-Modal Images using a Generative Model for Brain Lesion Segmentation Bjoern H. Menze 1,2, Koen Van Leemput 3, Danial Lashkari 4 Marc-André Weber 5, Nicholas Ayache 2, and Polina
Probabilistic Monte Carlo Based Mapping of Cerebral Connections Utilising Whole-Brain Crossing Fibre Information
 Probabilistic Monte Carlo Based Mapping of Cerebral Connections Utilising Whole-Brain Crossing Fibre Information Geoff J.M. Parker 1 and Daniel C. Alexander 2 1 Imaging Science & Biomedical Engineering,
Probabilistic Monte Carlo Based Mapping of Cerebral Connections Utilising Whole-Brain Crossing Fibre Information Geoff J.M. Parker 1 and Daniel C. Alexander 2 1 Imaging Science & Biomedical Engineering,
Detecting Changes In Non-Isotropic Images
 Detecting Changes In Non-Isotropic Images K.J. Worsley 1, M. Andermann 1, T. Koulis 1, D. MacDonald, 2 and A.C. Evans 2 August 4, 1999 1 Department of Mathematics and Statistics, 2 Montreal Neurological
Detecting Changes In Non-Isotropic Images K.J. Worsley 1, M. Andermann 1, T. Koulis 1, D. MacDonald, 2 and A.C. Evans 2 August 4, 1999 1 Department of Mathematics and Statistics, 2 Montreal Neurological
High-Order Diffusion Tensor Connectivity Mapping on the GPU
 High-Order Diffusion Tensor Connectivity Mapping on the GPU Tim McGraw and Donald Herring Purdue University Abstract. We present an efficient approach to computing white matter fiber connectivity on the
High-Order Diffusion Tensor Connectivity Mapping on the GPU Tim McGraw and Donald Herring Purdue University Abstract. We present an efficient approach to computing white matter fiber connectivity on the
Automatic Generation of Training Data for Brain Tissue Classification from MRI
 MICCAI-2002 1 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. Cocosco, Alex P. Zijdenbos, and Alan C. Evans McConnell Brain Imaging Centre, Montreal Neurological
MICCAI-2002 1 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. Cocosco, Alex P. Zijdenbos, and Alan C. Evans McConnell Brain Imaging Centre, Montreal Neurological
BDP: BrainSuite Diffusion Pipeline. Chitresh Bhushan
 BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity
 Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity Pahal Dalal 1, Feng Shi 2, Dinggang Shen 2, and Song Wang 1 1 Department of Computer Science and Engineering, University of South
Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity Pahal Dalal 1, Feng Shi 2, Dinggang Shen 2, and Song Wang 1 1 Department of Computer Science and Engineering, University of South
CS 534: Computer Vision Segmentation II Graph Cuts and Image Segmentation
 CS 534: Computer Vision Segmentation II Graph Cuts and Image Segmentation Spring 2005 Ahmed Elgammal Dept of Computer Science CS 534 Segmentation II - 1 Outlines What is Graph cuts Graph-based clustering
CS 534: Computer Vision Segmentation II Graph Cuts and Image Segmentation Spring 2005 Ahmed Elgammal Dept of Computer Science CS 534 Segmentation II - 1 Outlines What is Graph cuts Graph-based clustering
TRACULA: Troubleshooting, visualization, and group analysis
 TRACULA: Troubleshooting, visualization, and group analysis Anastasia Yendiki HMS/MGH/MIT Athinoula A. Martinos Center for Biomedical Imaging 18/11/13 TRACULA: troubleshooting, visualization, group analysis
TRACULA: Troubleshooting, visualization, and group analysis Anastasia Yendiki HMS/MGH/MIT Athinoula A. Martinos Center for Biomedical Imaging 18/11/13 TRACULA: troubleshooting, visualization, group analysis
Multimodal Visualization of DTI and fmri Data using Illustrative Methods
 Multimodal Visualization of DTI and fmri Data using Illustrative Methods Silvia Born 1, Werner Jainek 1, Mario Hlawitschka 2, Gerik Scheuermann 3, Christos Trantakis 4, Jürgen Meixensberger 4, Dirk Bartz
Multimodal Visualization of DTI and fmri Data using Illustrative Methods Silvia Born 1, Werner Jainek 1, Mario Hlawitschka 2, Gerik Scheuermann 3, Christos Trantakis 4, Jürgen Meixensberger 4, Dirk Bartz
n o r d i c B r a i n E x Tutorial DTI Module
 m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DTI Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DTI Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
