All have parameters: Voxel dimensions: 90 * 108 * 78, Voxel size: 1.5mm * 1.5mm * 1.5mm, Data type: float, Orientation: Axial (S=>I).
|
|
|
- Loren Murphy
- 5 years ago
- Views:
Transcription
1 Week 5 Exercises This week we will discuss some concepts related to the idea of fmri data as sets of numbers that can be manipulated computationally. We will also introduce the concept of "statistical maps" (or "overlays"). We will be working with statistical maps throughout the course, so we are introducing them early in the laboratories. We will talk about automated ways of loading data, processing, and viewing data using the BIAC XML header format ( *.bxh ). You will use the same basic viewing tool, Showsrs2, but with a new way of loading the data. Exercise 1: Understanding overlays In this next exercise, you will work with data collected in a biological motion paradigm. Subjects watched three types of motion (of cartoon figures): mouth, eye, and hand. The data are stored in the directory \\Huxley\data\Class.01\Examples\BIOMOT. The AVI files in this directory show the stimuli that were used. Each of the ####_overlay files indicates the value of a statistical test (t) at each voxel. If you have not already done so, you will need to create a directory for your group under Class.01\Students. You will not need to copy any files to your new directory in this lesson; you will just use it for storing the headers that you create. Then change to that directory in MATLAB. 1.1 Create BXH files for each image type You will need to create four BXH files. Each should point to the data in the examples directory. To create the BXH file, you will need to: Change to your group s directory in MATLAB (cd command) Use command bxhabsorb. Once you type bxabsorb, a graphic user interface will pop up. You then select the file you want; remember that you may need to change file type in the dialog to make it visible. You will need to put in the parameters for the data you want to load. The four BXH files you should create in your directory are: 1) base.bxh : for the base.img file. 2) mouth.bxh : for the mouth_overlay.img file 3) eye.bxh : for the eye_overlay.img file 4) hand.bxh : for the hand_overlay.img file All have parameters: Voxel dimensions: 90 * 108 * 78, Voxel size: 1.5mm * 1.5mm * 1.5mm, Data type: float, Orientation: Axial (S=>I). After you have created these files, load in the data to MATLAB. You should end up with four variables (base, mouth, eye, and hand). Note that you can load the data, once a BXH file has been created, either by using the readmr interface or by just typing the following (if base.bxh was in your current directory). p. 1 of 8
2 base = readmr( base.bxh ); % repeat for the other files 1.2 Display the data using Showsrs2 To display the data, you can just type: showsrs2(base, mouth, eye, hand) You should change to the 3 Orthogonal Planes view under the Orientation menu. Q1. Describe the data. What do you see as you move around the brain? Q2. What do you think that the different values in the bottom right window represent? Are they generally similar or different across the three conditions? 1.3 Understanding overlays These color maps are often called overlays because they are laid on top of the original anatomical data. The single most important thing to understand about such overlays is that they represent statistical tests of whether the data are consistent/inconsistent with an experimental hypothesis. They are not snapshots of brain activity. To illustrate this point, look at the data in showsrs2, with all thresholds set to the default values (as loaded automatically). Q3. How many discrete areas are active in the brain? Give a rough estimate of the number of large clusters that you see. Also, about what percentage of the brain is active? Now, lower all of your thresholds for the different overlays to a lower limit of 1.0. To do so, you can 1) click on the W+L button and select the different overlays using the pull-down menu at upper right, or 2) click on the Config menu and select adjust colormaps and clipping. Q4. After adjusting the threshold downward, how did the brain activity change from the previous threshold? Finally, raise all of the thresholds to a lower limit of 6.5. Q5. After adjusting the threshold upward, how many areas are now active? What is different about the activity? Note that the same brain overlays were used in every case. The only difference was in the statistical value that we called significant. p. 2 of 8
3 1.4 Quantifying the effects of threshold We can very quickly count how many voxels have significance values above different threshold levels in MATLAB. Suppose that you want to count how many voxels have a t value > 0.5, a t value > 1.0, a t value > 1.5, and so forth. You could define a set of such steps (0.5, 1.0, 1.5,, 7.5, 8.0) using the command: steps = [0.5: 0.5 : 8]; Then you could set up a place to store data: a=zeros(length(steps),1); Finally, you can quickly count how many voxels had significance values greater than each step. (You can plot the output using the figure and plot commands). for i = 1:length(steps);a(i)=length(find(mouth.data>steps(i)));end Q6. How does the number of active voxels change with significance value? BONUS Exercise: You could also plot how many voxels are within small ranges (e.g., from 0 to 0.2, from 0.2 to 0.4, etc.) in order to map out the distribution of significance values across voxels. 1.5 Finding the maximum and minimum values You should next find points in the brain with the maximum values for each overlay (note that there may be more than one point with a given value). Hint: adjust the thresholds in the configuration window so that you display one overlay at a time, and adjust the range for that overlay so that you only see the top of the range. The movement keys (A&D, W&S, and Q&E) may be useful. Write those values below: Q7. What are the coordinates of the maximum significance values, estimated visually: eye_overlay: maximum (X,Y,Z): t = hand_overlay: maximum (X,Y,Z): t = mouth_overlay: maximum (X,Y,Z): t = 1.6 Using transparencies Using the transparency function on the "W+L" screen, determine whether each of your maxima are found in gray matter or white matter. Put a G or W next to the points listed above. 1.7 Check your answer You can also quickly find out the minimum and maximum values in each image: p. 3 of 8
4 mm = minmax(mouth.data) To determine the X, Y, and Z coordinates of these values, use the following: mm_min = find(mouth.data == mm(1)) mm_max = find(mouth.data == mm(2)) [x,y,z]= ind2sub(size(mouth.data), mm_min) [x,y,z]= ind2sub(size(mouth.data), mm_max) Q8. What are the coordinates of the minimum and maximum significance values, determined in MATLAB? What are the actual t values? eye_overlay: min: t = max: t = hand_overlay: min: t = max: t = mouth_overlay: min: t = max: t = Exercise 2: Working with Time Series of data 2.1. Creating BXH files for time series A "time series" of data is a sequence of similar images showing changes in the brain over time. You are going to work with "averaged epochs", which indicate the changes in the brain time-locked to the occurrence of the different stimuli. These data consist of 12 time points, from 3 time points before (6s) through 8 time points after the stimulus (16 sec). The data are saved in the same directory as files in the form : Eye_0001.img. You will need to create BXH files for each of the three time series. The only thing that you need to do differently is to select use wildcard read ; if you then select one file in a series, it will associate that BXH file with all of them. The examples below will assume that you use the file convention: eye_all.bxh. When you are finished creating the BXH files, 1) load all three time series into MATLAB and 2) display them in showsrs2. NOTE: WHEN SELECTING THE IMAGES FOR THE TIME SERIES, CLICK ON THE FIRST ONE IN THE SEQUENCE, THEN SELECT USE WILDCARD READ. AFTER YOU VE DONE THAT, YOU LL SEE THAT IT TRIES TO LOAD 13 IMAGES. TO FIX, JUST PUT A 0 BEFORE THE *.IMG ; E.G., Eye_0*.img. showsrs2(base, mouth_all, eye_all, hand_all) 2.2. Changing the colormap limits You will need to change the values displayed on the colormap, as we will now be dealing with relative MR signal change instead of t values. Change the lower limit of each overlay to 2.0 and the upper limit to 3.5. p. 4 of 8
5 2.3. Looking at time series of data Find the points that you identified as being active in all three conditions. What does the time course of activation look like at those points? You can view the data in the window at bottom right, and/or play the data using the controls at bottom left. Q9. Write a short description of the time course of activity at the most active voxels in the space below Finding the peak response Q10. At what point does the brain response peak? Is this similar for all of the conditions? Is it similar across all of the points? Write a short description of your findings in the space below Looking at the spatial pattern of activation Q11. How does the spatial pattern of activation change across time points? At what time point is there the most activation? Write down your findings in the space below. Summary: Q12. Write a general summary of the data below. You looked at (real) data showing patterns and time courses of activity when the subjects were watching eye, hand, and mouth movements. What brain areas were activated by these stimuli? How did the activation pattern in those areas change over time? How similar were the activation patterns for the different conditions? p. 5 of 8
6 Questions: Q1. Describe the data. What do you see as you move around the brain? Q2. What do you think that the different values in the bottom right window represent? Are they generally similar or different across the three conditions? Q3. How many discrete areas are active in the brain? Give a rough estimate of the number of large clusters that you see. Also, about what percentage of the brain is active? Q4. After adjusting the threshold downward, how did the brain activity change from the previous threshold? Q5. After adjusting the threshold upward, how many areas are now active? What is different about the activity? Q6. How does the number of active voxels change with significance value? p. 6 of 8
7 Q7. What are the coordinates of the maximum significance values, estimated visually: eye_overlay: maximum (X,Y,Z): t = hand_overlay: maximum (X,Y,Z): t = mouth_overlay: maximum (X,Y,Z): t = Q8. What are the coordinates of the minimum and maximum significance values, determined in MATLAB? What are the actual t values? eye_overlay: min: t = max: t = hand_overlay: min: t = max: t = mouth_overlay: min: t = max: t = Q9. Write a short description of the time course of activity at the most active voxels in the space below. Q10. At what point does the brain response peak? Is this similar for all of the conditions? Is it similar across all of the points? Write a short description of your findings in the space below. Q11. How does the spatial pattern of activation change across time points? At what time point is there the most activation? Write down your findings in the space below. p. 7 of 8
8 Q12. Write a general summary of the data below. You looked at (real) data showing patterns and time courses of activity when the subjects were watching eye, hand, and mouth movements. What brain areas were activated by these stimuli? How did the activation pattern in those areas change over time? How similar were the activation patterns for the different conditions? p. 8 of 8
0.1. Setting up the system path to allow use of BIAC XML headers (BXH). Depending on the computer(s), you may only have to do this once.
 Week 3 Exercises Last week you began working with MR data, both in the form of anatomical images and functional time series. This week we will discuss some concepts related to the idea of fmri data as
Week 3 Exercises Last week you began working with MR data, both in the form of anatomical images and functional time series. This week we will discuss some concepts related to the idea of fmri data as
This exercise uses one anatomical data set (ANAT1) and two functional data sets (FUNC1 and FUNC2).
 Exploring Brain Anatomy This week s exercises will let you explore the anatomical organization of the brain to learn some of its basic properties, as well as the location of different structures. The human
Exploring Brain Anatomy This week s exercises will let you explore the anatomical organization of the brain to learn some of its basic properties, as well as the location of different structures. The human
CS/NEUR125 Brains, Minds, and Machines. Due: Wednesday, April 5
 CS/NEUR125 Brains, Minds, and Machines Lab 8: Using fmri to Discover Language Areas in the Brain Due: Wednesday, April 5 In this lab, you will analyze fmri data from an experiment that was designed to
CS/NEUR125 Brains, Minds, and Machines Lab 8: Using fmri to Discover Language Areas in the Brain Due: Wednesday, April 5 In this lab, you will analyze fmri data from an experiment that was designed to
SPM Course! Single Subject Analysis
 SPM Course! Single Subject Analysis Practical Session Dr. Jakob Heinzle & Dr. Frederike Petzschner & Dr. Lionel Rigoux Hands up: Who has programming experience with Matlab? Who has analyzed an fmri experiment
SPM Course! Single Subject Analysis Practical Session Dr. Jakob Heinzle & Dr. Frederike Petzschner & Dr. Lionel Rigoux Hands up: Who has programming experience with Matlab? Who has analyzed an fmri experiment
Microsoft Excel 2007 Lesson 7: Charts and Comments
 Microsoft Excel 2007 Lesson 7: Charts and Comments Open Example.xlsx if it is not already open. Click on the Example 3 tab to see the worksheet for this lesson. This is essentially the same worksheet that
Microsoft Excel 2007 Lesson 7: Charts and Comments Open Example.xlsx if it is not already open. Click on the Example 3 tab to see the worksheet for this lesson. This is essentially the same worksheet that
SPM99 fmri Data Analysis Workbook
 SPM99 fmri Data Analysis Workbook This file is a description of the steps needed to use SPM99 analyze a fmri data set from a single subject using a simple on/off activation paradigm. There are two parts
SPM99 fmri Data Analysis Workbook This file is a description of the steps needed to use SPM99 analyze a fmri data set from a single subject using a simple on/off activation paradigm. There are two parts
Data Loading & 3D Visualization
 Neuroimage Analysis Center Data Loading & 3D Visualization Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School Leonardo da Vinci (1452-1519), Virgin and Child Alte Pinakothek, München
Neuroimage Analysis Center Data Loading & 3D Visualization Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School Leonardo da Vinci (1452-1519), Virgin and Child Alte Pinakothek, München
Table of Contents. IntroLab < SPMLabs < Dynevor TWiki
 Table of Contents Lab 1: Introduction to SPM and data checking...1 Goals of this Lab...1 Prerequisites...1 An SPM Installation...1 SPM Defaults...2 L/R Brain Orientation...2 Memory Use for Data Processing...2
Table of Contents Lab 1: Introduction to SPM and data checking...1 Goals of this Lab...1 Prerequisites...1 An SPM Installation...1 SPM Defaults...2 L/R Brain Orientation...2 Memory Use for Data Processing...2
SIVIC GUI Overview. SIVIC GUI Layout Overview
 SIVIC GUI Overview SIVIC GUI Layout Overview At the top of the SIVIC GUI is a row of buttons called the Toolbar. It is a quick interface for loading datasets, controlling how the mouse manipulates the
SIVIC GUI Overview SIVIC GUI Layout Overview At the top of the SIVIC GUI is a row of buttons called the Toolbar. It is a quick interface for loading datasets, controlling how the mouse manipulates the
fmri Basics: Single Subject Analysis
 fmri Basics: Single Subject Analysis This session is intended to give an overview of the basic process of setting up a general linear model for a single subject. This stage of the analysis is also variously
fmri Basics: Single Subject Analysis This session is intended to give an overview of the basic process of setting up a general linear model for a single subject. This stage of the analysis is also variously
Intro to Animation. Introduction: Frames and Keyframes. Blender Lesson: Grade Level: Lesson Description: Goals/Objectives: Materials/Tools: 4th and up
 Blender Lesson: Intro to Animation Grade Level: 4th and up Lesson Description: This lesson serves as an introduction to animation with Blender. The lesson begins by talking about some core concepts of
Blender Lesson: Intro to Animation Grade Level: 4th and up Lesson Description: This lesson serves as an introduction to animation with Blender. The lesson begins by talking about some core concepts of
Structural MRI of Amygdala Tutorial: Observation, Segmentation, Quantification
 Structural MRI of Amygdala Tutorial: Observation, Segmentation, Quantification The FMRIB Software Library (FSL) is a powerful tool that allows users to observe the human brain in various planes and dimensions,
Structural MRI of Amygdala Tutorial: Observation, Segmentation, Quantification The FMRIB Software Library (FSL) is a powerful tool that allows users to observe the human brain in various planes and dimensions,
SnPM is an SPM toolbox developed by Andrew Holmes & Tom Nichols
 1 of 14 3/30/2005 9:24 PM SnPM A Worked fmri Example SnPM is an SPM toolbox developed by Andrew Holmes & Tom Nichols This page... introduction example data background design setup computation viewing results
1 of 14 3/30/2005 9:24 PM SnPM A Worked fmri Example SnPM is an SPM toolbox developed by Andrew Holmes & Tom Nichols This page... introduction example data background design setup computation viewing results
GLM for fmri data analysis Lab Exercise 1
 GLM for fmri data analysis Lab Exercise 1 March 15, 2013 Medical Image Processing Lab Medical Image Processing Lab GLM for fmri data analysis Outline 1 Getting Started 2 AUDIO 1 st level Preprocessing
GLM for fmri data analysis Lab Exercise 1 March 15, 2013 Medical Image Processing Lab Medical Image Processing Lab GLM for fmri data analysis Outline 1 Getting Started 2 AUDIO 1 st level Preprocessing
Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question.
 (1) conn - Functional connectivity toolbox v1.0 Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question. 2) in "Basic" enter "1" subject, "6" seconds
(1) conn - Functional connectivity toolbox v1.0 Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question. 2) in "Basic" enter "1" subject, "6" seconds
Organizing Design Data
 Organizing Design Data Module Overview This module explains how to use the data in different files for reference purposes. Module Prerequisites Knowledge of MicroStation s interface Some knowledge about
Organizing Design Data Module Overview This module explains how to use the data in different files for reference purposes. Module Prerequisites Knowledge of MicroStation s interface Some knowledge about
Version. Getting Started: An fmri-cpca Tutorial
 Version 11 Getting Started: An fmri-cpca Tutorial 2 Table of Contents Table of Contents... 2 Introduction... 3 Definition of fmri-cpca Data... 3 Purpose of this tutorial... 3 Prerequisites... 4 Used Terms
Version 11 Getting Started: An fmri-cpca Tutorial 2 Table of Contents Table of Contents... 2 Introduction... 3 Definition of fmri-cpca Data... 3 Purpose of this tutorial... 3 Prerequisites... 4 Used Terms
Tutorial BOLD Module
 m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial BOLD Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial BOLD Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
SPM Introduction. SPM : Overview. SPM: Preprocessing SPM! SPM: Preprocessing. Scott Peltier. FMRI Laboratory University of Michigan
 SPM Introduction Scott Peltier FMRI Laboratory University of Michigan! Slides adapted from T. Nichols SPM! SPM : Overview Library of MATLAB and C functions Graphical user interface Four main components:
SPM Introduction Scott Peltier FMRI Laboratory University of Michigan! Slides adapted from T. Nichols SPM! SPM : Overview Library of MATLAB and C functions Graphical user interface Four main components:
SPM Introduction SPM! Scott Peltier. FMRI Laboratory University of Michigan. Software to perform computation, manipulation and display of imaging data
 SPM Introduction Scott Peltier FMRI Laboratory University of Michigan Slides adapted from T. Nichols SPM! Software to perform computation, manipulation and display of imaging data 1 1 SPM : Overview Library
SPM Introduction Scott Peltier FMRI Laboratory University of Michigan Slides adapted from T. Nichols SPM! Software to perform computation, manipulation and display of imaging data 1 1 SPM : Overview Library
Transform Introduction page 96 Spatial Transforms page 97
 Transform Introduction page 96 Spatial Transforms page 97 Pad page 97 Subregion page 101 Resize page 104 Shift page 109 1. Correcting Wraparound Using the Shift Tool page 109 Flip page 116 2. Flipping
Transform Introduction page 96 Spatial Transforms page 97 Pad page 97 Subregion page 101 Resize page 104 Shift page 109 1. Correcting Wraparound Using the Shift Tool page 109 Flip page 116 2. Flipping
fmri/dti analysis using Dynasuite
 fmri/dti analysis using Dynasuite Contents 1 Logging in 2 Finding patient session 3 Viewing and adjusting images 4 Checking brain segmentation 5 Checking image registration 6 Seeing fmri results 7 Saving
fmri/dti analysis using Dynasuite Contents 1 Logging in 2 Finding patient session 3 Viewing and adjusting images 4 Checking brain segmentation 5 Checking image registration 6 Seeing fmri results 7 Saving
Basic Introduction to Data Analysis. Block Design Demonstration. Robert Savoy
 Basic Introduction to Data Analysis Block Design Demonstration Robert Savoy Sample Block Design Experiment Demonstration Use of Visual and Motor Task Separability of Responses Combined Visual and Motor
Basic Introduction to Data Analysis Block Design Demonstration Robert Savoy Sample Block Design Experiment Demonstration Use of Visual and Motor Task Separability of Responses Combined Visual and Motor
Section 9. Human Anatomy and Physiology
 Section 9. Human Anatomy and Physiology 9.1 MR Neuroimaging 9.2 Electroencephalography Overview As stated throughout, electrophysiology is the key tool in current systems neuroscience. However, single-
Section 9. Human Anatomy and Physiology 9.1 MR Neuroimaging 9.2 Electroencephalography Overview As stated throughout, electrophysiology is the key tool in current systems neuroscience. However, single-
USER's GUIDE for PLOT_FINDIF_1
 USER's GUIDE for PLOT_FINDIF_1 S. T. Bolmer & R. A. Stephen November 2004 Page 1 PLOT_FINDIF_1 Plot_findif_1.m is a MATLAB 6 script, which is used to plot output from the Woods Hole Oceanographic Institution
USER's GUIDE for PLOT_FINDIF_1 S. T. Bolmer & R. A. Stephen November 2004 Page 1 PLOT_FINDIF_1 Plot_findif_1.m is a MATLAB 6 script, which is used to plot output from the Woods Hole Oceanographic Institution
Structural MRI of Amygdala Tutorial: Observation, Segmentation, Quantification
 Structural MRI of Amygdala Tutorial: Observation, Segmentation, Quantification The FMRIB Software Library (FSL) is a powerful tool that allows users to observe the human brain in various planes and dimensions,
Structural MRI of Amygdala Tutorial: Observation, Segmentation, Quantification The FMRIB Software Library (FSL) is a powerful tool that allows users to observe the human brain in various planes and dimensions,
Display. Introduction page 67 2D Images page 68. All Orientations page 69 Single Image page 70 3D Images page 71
 Display Introduction page 67 2D Images page 68 All Orientations page 69 Single Image page 70 3D Images page 71 Intersecting Sections page 71 Cube Sections page 72 Render page 73 1. Tissue Maps page 77
Display Introduction page 67 2D Images page 68 All Orientations page 69 Single Image page 70 3D Images page 71 Intersecting Sections page 71 Cube Sections page 72 Render page 73 1. Tissue Maps page 77
Practice Exam Sample Solutions
 CS 675 Computer Vision Instructor: Marc Pomplun Practice Exam Sample Solutions Note that in the actual exam, no calculators, no books, and no notes allowed. Question 1: out of points Question 2: out of
CS 675 Computer Vision Instructor: Marc Pomplun Practice Exam Sample Solutions Note that in the actual exam, no calculators, no books, and no notes allowed. Question 1: out of points Question 2: out of
NEURO M203 & BIOMED M263 WINTER 2014
 NEURO M203 & BIOMED M263 WINTER 2014 MRI Lab 2: Neuroimaging Connectivity Lab In today s lab we will work with sample diffusion imaging data and the group averaged fmri data collected during your scanning
NEURO M203 & BIOMED M263 WINTER 2014 MRI Lab 2: Neuroimaging Connectivity Lab In today s lab we will work with sample diffusion imaging data and the group averaged fmri data collected during your scanning
BrainSuite Lab Exercises. presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014
 BrainSuite Lab Exercises presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 1. Opening and Displaying an MRI Start BrainSuite Drag and drop the T1 image from the native space
BrainSuite Lab Exercises presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 1. Opening and Displaying an MRI Start BrainSuite Drag and drop the T1 image from the native space
The alignment process uses two alignment methods: (1) the mrvista function, rxalign, for a rough alignment, and then (2) KNK's alignment for
 Alignment (Allegra) Download example alignment script here. Anatomical images are collected at much higher resolution (isotropic 1 mm voxels) than functional images (isotropic 2 mm voxels; sometimes 2.5).
Alignment (Allegra) Download example alignment script here. Anatomical images are collected at much higher resolution (isotropic 1 mm voxels) than functional images (isotropic 2 mm voxels; sometimes 2.5).
The Fundamentals. Document Basics
 3 The Fundamentals Opening a Program... 3 Similarities in All Programs... 3 It's On Now What?...4 Making things easier to see.. 4 Adjusting Text Size.....4 My Computer. 4 Control Panel... 5 Accessibility
3 The Fundamentals Opening a Program... 3 Similarities in All Programs... 3 It's On Now What?...4 Making things easier to see.. 4 Adjusting Text Size.....4 My Computer. 4 Control Panel... 5 Accessibility
Slicer4Minute Tutorial. Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School
 Slicer4Minute Tutorial Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School 1 Slicer4 minute tutorial This tutorial is a 4-minute introduction to the 3D visualization capabilities of
Slicer4Minute Tutorial Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School 1 Slicer4 minute tutorial This tutorial is a 4-minute introduction to the 3D visualization capabilities of
Matlab for FMRI Module 1: the basics Instructor: Luis Hernandez-Garcia
 Matlab for FMRI Module 1: the basics Instructor: Luis Hernandez-Garcia The goal for this tutorial is to make sure that you understand a few key concepts related to programming, and that you know the basics
Matlab for FMRI Module 1: the basics Instructor: Luis Hernandez-Garcia The goal for this tutorial is to make sure that you understand a few key concepts related to programming, and that you know the basics
HOW TO SETUP AND RUN SHIVA...
 TABLE OF CONTENTS HOW TO SETUP AND RUN SHIVA... 1 REQUIREMENTS... 1 Java... 1 Architecture... 2 LAUNCHING SHIVA... 2 LOADING THE ATLAS... 2 USAGE INSTRUCTIONS... 2 FILES... 2 Image Volumes... 2 Label Indices...
TABLE OF CONTENTS HOW TO SETUP AND RUN SHIVA... 1 REQUIREMENTS... 1 Java... 1 Architecture... 2 LAUNCHING SHIVA... 2 LOADING THE ATLAS... 2 USAGE INSTRUCTIONS... 2 FILES... 2 Image Volumes... 2 Label Indices...
Neural Bases of Language, V
 Neural Bases of Language, V61.0043-001 LAB 2 GROUP MEMBERS: SUBJECT ANALYZED: Name of ppt file for final presentation: 1 Lab 2 1. Open BESA (Brain Electric Source Analysis). 2. Open the averaged.fsg data
Neural Bases of Language, V61.0043-001 LAB 2 GROUP MEMBERS: SUBJECT ANALYZED: Name of ppt file for final presentation: 1 Lab 2 1. Open BESA (Brain Electric Source Analysis). 2. Open the averaged.fsg data
evaluation techniques goals of evaluation evaluation by experts cisc3650 human-computer interaction spring 2012 lecture # II.1
 topics: evaluation techniques usability testing references: cisc3650 human-computer interaction spring 2012 lecture # II.1 evaluation techniques Human-Computer Interaction, by Alan Dix, Janet Finlay, Gregory
topics: evaluation techniques usability testing references: cisc3650 human-computer interaction spring 2012 lecture # II.1 evaluation techniques Human-Computer Interaction, by Alan Dix, Janet Finlay, Gregory
Quality Checking an fmri Group Result (art_groupcheck)
 Quality Checking an fmri Group Result (art_groupcheck) Paul Mazaika, Feb. 24, 2009 A statistical parameter map of fmri group analyses relies on the assumptions of the General Linear Model (GLM). The assumptions
Quality Checking an fmri Group Result (art_groupcheck) Paul Mazaika, Feb. 24, 2009 A statistical parameter map of fmri group analyses relies on the assumptions of the General Linear Model (GLM). The assumptions
BrainMask. Quick Start
 BrainMask Quick Start Segmentation of the brain from three-dimensional MR images is a crucial preprocessing step in morphological and volumetric brain studies. BrainMask software implements a fully automatic
BrainMask Quick Start Segmentation of the brain from three-dimensional MR images is a crucial preprocessing step in morphological and volumetric brain studies. BrainMask software implements a fully automatic
Basic principles of MR image analysis. Basic principles of MR image analysis. Basic principles of MR image analysis
 Basic principles of MR image analysis Basic principles of MR image analysis Julien Milles Leiden University Medical Center Terminology of fmri Brain extraction Registration Linear registration Non-linear
Basic principles of MR image analysis Basic principles of MR image analysis Julien Milles Leiden University Medical Center Terminology of fmri Brain extraction Registration Linear registration Non-linear
Statistical Analysis of Neuroimaging Data. Phebe Kemmer BIOS 516 Sept 24, 2015
 Statistical Analysis of Neuroimaging Data Phebe Kemmer BIOS 516 Sept 24, 2015 Review from last time Structural Imaging modalities MRI, CAT, DTI (diffusion tensor imaging) Functional Imaging modalities
Statistical Analysis of Neuroimaging Data Phebe Kemmer BIOS 516 Sept 24, 2015 Review from last time Structural Imaging modalities MRI, CAT, DTI (diffusion tensor imaging) Functional Imaging modalities
NA-MIC National Alliance for Medical Image Computing fmri Data Analysis
 NA-MIC fmri Data Analysis Sonia Pujol, Ph.D. Wendy Plesniak, Ph.D. Randy Gollub, M.D., Ph.D. Acknowledgments NIH U54EB005149 Neuroimage Analysis Center NIH P41RR013218 FIRST Biomedical Informatics Research
NA-MIC fmri Data Analysis Sonia Pujol, Ph.D. Wendy Plesniak, Ph.D. Randy Gollub, M.D., Ph.D. Acknowledgments NIH U54EB005149 Neuroimage Analysis Center NIH P41RR013218 FIRST Biomedical Informatics Research
Documentation for imcalc (SPM 5/8/12) Robert J Ellis
 (_) _ '_ ` _ \ / / _` / Image calculations and transformations (using SPM) (_ (_ ( This software version: 09-Nov-2017 _ _ _ _ \ \,_ _ \ (C) Robert J Ellis (http://tools.robjellis.net) Documentation for
(_) _ '_ ` _ \ / / _` / Image calculations and transformations (using SPM) (_ (_ ( This software version: 09-Nov-2017 _ _ _ _ \ \,_ _ \ (C) Robert J Ellis (http://tools.robjellis.net) Documentation for
Precalculus and Calculus
 4 Precalculus and Calculus You have permission to make copies of this document for your classroom use only. You may not distribute, copy or otherwise reproduce any part of this document or the lessons
4 Precalculus and Calculus You have permission to make copies of this document for your classroom use only. You may not distribute, copy or otherwise reproduce any part of this document or the lessons
syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets
 syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets Julien Gervais; Lisa Chuah Siemens Healthcare, Magnetic
syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets Julien Gervais; Lisa Chuah Siemens Healthcare, Magnetic
n o r d i c B r a i n E x Tutorial DTI Module
 m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DTI Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DTI Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
LESSON 8 COPYING SELECTIONS
 LESSON 8 COPYING SELECTIONS Digital Media I Susan M. Raymond West High School IN THIS TUTORIAL, YOU WILL: COPY AND MOVE SELECTIONS MAKE A COPY OF A SELECTION SO THAT IT OCCUPIES ITS OWN SEPARATE LAYER
LESSON 8 COPYING SELECTIONS Digital Media I Susan M. Raymond West High School IN THIS TUTORIAL, YOU WILL: COPY AND MOVE SELECTIONS MAKE A COPY OF A SELECTION SO THAT IT OCCUPIES ITS OWN SEPARATE LAYER
animation, and what interface elements the Flash editor contains to help you create and control your animation.
 e r ch02.fm Page 43 Wednesday, November 15, 2000 8:52 AM c h a p t 2 Animating the Page IN THIS CHAPTER Timelines and Frames Movement Tweening Shape Tweening Fading Recap Advanced Projects You have totally
e r ch02.fm Page 43 Wednesday, November 15, 2000 8:52 AM c h a p t 2 Animating the Page IN THIS CHAPTER Timelines and Frames Movement Tweening Shape Tweening Fading Recap Advanced Projects You have totally
Working with the Dope Sheet Editor to speed up animation and reverse time.
 Bouncing a Ball Page 1 of 2 Tutorial Bouncing a Ball A bouncing ball is a common first project for new animators. This classic example is an excellent tool for explaining basic animation processes in 3ds
Bouncing a Ball Page 1 of 2 Tutorial Bouncing a Ball A bouncing ball is a common first project for new animators. This classic example is an excellent tool for explaining basic animation processes in 3ds
Lesson 1 Parametric Modeling Fundamentals
 1-1 Lesson 1 Parametric Modeling Fundamentals Create Simple Parametric Models. Understand the Basic Parametric Modeling Process. Create and Profile Rough Sketches. Understand the "Shape before size" approach.
1-1 Lesson 1 Parametric Modeling Fundamentals Create Simple Parametric Models. Understand the Basic Parametric Modeling Process. Create and Profile Rough Sketches. Understand the "Shape before size" approach.
The playhead, shown as a vertical red beam, passes each frame when a movie plays back, much like movie fi lm passing in front of a projector bulb.
 The project: AIRPLANE I will show you a completed version of this project.. Introducing keyframes and the Timeline One of the most important panels in the Flash workspace is the Timeline, which is where
The project: AIRPLANE I will show you a completed version of this project.. Introducing keyframes and the Timeline One of the most important panels in the Flash workspace is the Timeline, which is where
Module: Rasters. 8.1 Lesson: Working with Raster Data Follow along: Loading Raster Data CHAPTER 8
 CHAPTER 8 Module: Rasters We ve used rasters for digitizing before, but raster data can also be used directly. In this module, you ll see how it s done in QGIS. 8.1 Lesson: Working with Raster Data Raster
CHAPTER 8 Module: Rasters We ve used rasters for digitizing before, but raster data can also be used directly. In this module, you ll see how it s done in QGIS. 8.1 Lesson: Working with Raster Data Raster
BME I5000: Biomedical Imaging
 BME I5000: Biomedical Imaging Lecture 1 Introduction Lucas C. Parra, parra@ccny.cuny.edu 1 Content Topics: Physics of medial imaging modalities (blue) Digital Image Processing (black) Schedule: 1. Introduction,
BME I5000: Biomedical Imaging Lecture 1 Introduction Lucas C. Parra, parra@ccny.cuny.edu 1 Content Topics: Physics of medial imaging modalities (blue) Digital Image Processing (black) Schedule: 1. Introduction,
User InterfaceChapter1:
 Chapter 1 User InterfaceChapter1: In this chapter you will learn about several aspects of the User Interface. You will learn about the overall layout of the UI, and then about the details of each element.
Chapter 1 User InterfaceChapter1: In this chapter you will learn about several aspects of the User Interface. You will learn about the overall layout of the UI, and then about the details of each element.
8.2.1 and 2 Lesson Date: Definition of Translation and Three Basic Properties
 8.2.1 and 2 Lesson Date: Definition of Translation and Three Basic Properties Student Objectives I can perform translations of figures along a specific vector. I can label the image of the figure using
8.2.1 and 2 Lesson Date: Definition of Translation and Three Basic Properties Student Objectives I can perform translations of figures along a specific vector. I can label the image of the figure using
CIGAL. CIGAL Workshop. Jim Voyvodic May 13, 2015
 CIGAL CIGAL Workshop Jim Voyvodic May 13, 2015 Advanced topics These CIGAL topics will not be covered in this session: Showplay s modular design - How to replace standard components with your own Structured
CIGAL CIGAL Workshop Jim Voyvodic May 13, 2015 Advanced topics These CIGAL topics will not be covered in this session: Showplay s modular design - How to replace standard components with your own Structured
Naïve Bayes, Gaussian Distributions, Practical Applications
 Naïve Bayes, Gaussian Distributions, Practical Applications Required reading: Mitchell draft chapter, sections 1 and 2. (available on class website) Machine Learning 10-601 Tom M. Mitchell Machine Learning
Naïve Bayes, Gaussian Distributions, Practical Applications Required reading: Mitchell draft chapter, sections 1 and 2. (available on class website) Machine Learning 10-601 Tom M. Mitchell Machine Learning
Laboratory. Disk Scheduling. Objectives. References. Watch how the operating system schedules the reading or writing of disk tracks.
 Laboratory Disk Scheduling 11 Objectives Watch how the operating system schedules the reading or writing of disk tracks. References Software needed: 1) A web browser (Internet Explorer or Netscape) 2)
Laboratory Disk Scheduling 11 Objectives Watch how the operating system schedules the reading or writing of disk tracks. References Software needed: 1) A web browser (Internet Explorer or Netscape) 2)
GUIDE TO View3D. Introduction to View3D
 View3D Guide Introduction to View3D... 1 Starting Hampson-Russell Software... 2 Starting View3D... 4 A Brief Summary of the View3D Process... 8 Loading the Seismic and Horizon Data... 8 Viewing the Data...
View3D Guide Introduction to View3D... 1 Starting Hampson-Russell Software... 2 Starting View3D... 4 A Brief Summary of the View3D Process... 8 Loading the Seismic and Horizon Data... 8 Viewing the Data...
Group (Level 2) fmri Data Analysis - Lab 4
 Group (Level 2) fmri Data Analysis - Lab 4 Index Goals of this Lab Before Getting Started The Chosen Ten Checking Data Quality Create a Mean Anatomical of the Group Group Analysis: One-Sample T-Test Examine
Group (Level 2) fmri Data Analysis - Lab 4 Index Goals of this Lab Before Getting Started The Chosen Ten Checking Data Quality Create a Mean Anatomical of the Group Group Analysis: One-Sample T-Test Examine
Resting state network estimation in individual subjects
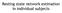 Resting state network estimation in individual subjects Data 3T NIL(21,17,10), Havard-MGH(692) Young adult fmri BOLD Method Machine learning algorithm MLP DR LDA Network image Correlation Spatial Temporal
Resting state network estimation in individual subjects Data 3T NIL(21,17,10), Havard-MGH(692) Young adult fmri BOLD Method Machine learning algorithm MLP DR LDA Network image Correlation Spatial Temporal
Manual image registration in BrainVoyager QX Table of Contents
 Manual image registration in BrainVoyager QX Table of Contents Manual image registration in BrainVoyager QX......1 Performing manual alignment for functional to anatomical images......2 Step 1: preparation......2
Manual image registration in BrainVoyager QX Table of Contents Manual image registration in BrainVoyager QX......1 Performing manual alignment for functional to anatomical images......2 Step 1: preparation......2
BIOSTATISTICS LABORATORY PART 1: INTRODUCTION TO DATA ANALYIS WITH STATA: EXPLORING AND SUMMARIZING DATA
 BIOSTATISTICS LABORATORY PART 1: INTRODUCTION TO DATA ANALYIS WITH STATA: EXPLORING AND SUMMARIZING DATA Learning objectives: Getting data ready for analysis: 1) Learn several methods of exploring the
BIOSTATISTICS LABORATORY PART 1: INTRODUCTION TO DATA ANALYIS WITH STATA: EXPLORING AND SUMMARIZING DATA Learning objectives: Getting data ready for analysis: 1) Learn several methods of exploring the
EMSegment Tutorial. How to Define and Fine-Tune Automatic Brain Compartment Segmentation and the Detection of White Matter Hyperintensities
 EMSegment Tutorial How to Define and Fine-Tune Automatic Brain Compartment Segmentation and the Detection of White Matter Hyperintensities This documentation serves as a tutorial to learn to customize
EMSegment Tutorial How to Define and Fine-Tune Automatic Brain Compartment Segmentation and the Detection of White Matter Hyperintensities This documentation serves as a tutorial to learn to customize
BESA Research. CE certified software package for comprehensive, fast, and user-friendly analysis of EEG and MEG
 BESA Research CE certified software package for comprehensive, fast, and user-friendly analysis of EEG and MEG BESA Research choose the best analysis tool for your EEG and MEG data BESA Research is the
BESA Research CE certified software package for comprehensive, fast, and user-friendly analysis of EEG and MEG BESA Research choose the best analysis tool for your EEG and MEG data BESA Research is the
This is called the vertex form of the quadratic equation. To graph the equation
 Name Period Date: Topic: 7-5 Graphing ( ) Essential Question: What is the vertex of a parabola, and what is its axis of symmetry? Standard: F-IF.7a Objective: Graph linear and quadratic functions and show
Name Period Date: Topic: 7-5 Graphing ( ) Essential Question: What is the vertex of a parabola, and what is its axis of symmetry? Standard: F-IF.7a Objective: Graph linear and quadratic functions and show
MRI Segmentation MIDAS, 2007, 2010
 MRI Segmentation MIDAS, 2007, 2010 Lawrence O. Hall, Dmitry Goldgof, Yuhua Gu, Prodip Hore Dept. of Computer Science & Engineering University of South Florida CONTENTS: 1. Introduction... 1 2. Installing
MRI Segmentation MIDAS, 2007, 2010 Lawrence O. Hall, Dmitry Goldgof, Yuhua Gu, Prodip Hore Dept. of Computer Science & Engineering University of South Florida CONTENTS: 1. Introduction... 1 2. Installing
Lesson 19: The Graph of a Linear Equation in Two Variables Is a Line
 The Graph of a Linear Equation in Two Variables Is a Line Classwork Exercises THEOREM: The graph of a linear equation yy = mmmm + bb is a non-vertical line with slope mm and passing through (0, bb), where
The Graph of a Linear Equation in Two Variables Is a Line Classwork Exercises THEOREM: The graph of a linear equation yy = mmmm + bb is a non-vertical line with slope mm and passing through (0, bb), where
Lesson 19: The Graph of a Linear Equation in Two Variables is a Line
 Lesson 19: The Graph of a Linear Equation in Two Variables is a Line Classwork Exercises Theorem: The graph of a linear equation y = mx + b is a non-vertical line with slope m and passing through (0, b),
Lesson 19: The Graph of a Linear Equation in Two Variables is a Line Classwork Exercises Theorem: The graph of a linear equation y = mx + b is a non-vertical line with slope m and passing through (0, b),
Achieve Planner Quick Start
 Effexis Software Achieve Planner Quick Start Overview of Achieve Planner Copyright 2007 by Effexis Software, LLC. This document is protected by U.S. and international copyright laws. All rights reserved.
Effexis Software Achieve Planner Quick Start Overview of Achieve Planner Copyright 2007 by Effexis Software, LLC. This document is protected by U.S. and international copyright laws. All rights reserved.
DOING PHYSICS WITH MATLAB COMPUTATIONAL OPTICS
 DOING PHYSICS WITH MATLAB COMPUTATIONAL OPTICS RAYLEIGH-SOMMERFELD DIFFRACTION RECTANGULAR APERTURES Ian Cooper School of Physics, University of Sydney ian.cooper@sydney.edu.au DOWNLOAD DIRECTORY FOR MATLAB
DOING PHYSICS WITH MATLAB COMPUTATIONAL OPTICS RAYLEIGH-SOMMERFELD DIFFRACTION RECTANGULAR APERTURES Ian Cooper School of Physics, University of Sydney ian.cooper@sydney.edu.au DOWNLOAD DIRECTORY FOR MATLAB
Assignment 02 (Due: Monday, February 1, 2016)
 Assignment 02 (Due: Monday, February 1, 2016) CSCE 155N 1 Lab Objectives Improve your understanding of arrays and array operations Differentiate array operators and matrix operators Create, access, modify,
Assignment 02 (Due: Monday, February 1, 2016) CSCE 155N 1 Lab Objectives Improve your understanding of arrays and array operations Differentiate array operators and matrix operators Create, access, modify,
BrainMask. Quick Start
 BrainMask Quick Start Segmentation of the brain from three-dimensional MR images is a crucial pre-processing step in morphological and volumetric brain studies. BrainMask software implements a fully automatic
BrainMask Quick Start Segmentation of the brain from three-dimensional MR images is a crucial pre-processing step in morphological and volumetric brain studies. BrainMask software implements a fully automatic
Supplementary Data. in residuals voxel time-series exhibiting high variance, for example, large sinuses.
 Supplementary Data Supplementary Materials and Methods Step-by-step description of principal component-orthogonalization technique Below is a step-by-step description of the principal component (PC)-orthogonalization
Supplementary Data Supplementary Materials and Methods Step-by-step description of principal component-orthogonalization technique Below is a step-by-step description of the principal component (PC)-orthogonalization
Fmri Spatial Processing
 Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
How You Use the Timeline
 How You Use the Timeline The Timeline and the Canvas display two different views of the same sequence. The Timeline shows the chronological arrangement of clips and layered video and audio clip items,
How You Use the Timeline The Timeline and the Canvas display two different views of the same sequence. The Timeline shows the chronological arrangement of clips and layered video and audio clip items,
Introduction to Neuroimaging Janaina Mourao-Miranda
 Introduction to Neuroimaging Janaina Mourao-Miranda Neuroimaging techniques have changed the way neuroscientists address questions about functional anatomy, especially in relation to behavior and clinical
Introduction to Neuroimaging Janaina Mourao-Miranda Neuroimaging techniques have changed the way neuroscientists address questions about functional anatomy, especially in relation to behavior and clinical
Creating tables of contents
 Creating tables of contents A table of contents (TOC) can list the contents of a book, magazine, or other publication; display a list of illustrations, advertisers, or photo credits; or include other information
Creating tables of contents A table of contents (TOC) can list the contents of a book, magazine, or other publication; display a list of illustrations, advertisers, or photo credits; or include other information
Getting Started With Squeeze Server
 Getting Started With Squeeze Server & Squeeze Server takes the proven Squeeze encoding engine and makes it available on- premise, in the cloud or both, with a robust application programming interface (API)
Getting Started With Squeeze Server & Squeeze Server takes the proven Squeeze encoding engine and makes it available on- premise, in the cloud or both, with a robust application programming interface (API)
HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Avigilon Control Center Web Client User Guide
 Avigilon Control Center Web Client User Guide Version: 4.12 Standard PDF-WEBCLIENT-S-E-Rev2 Copyright 2013 Avigilon. All rights reserved. The information presented is subject to change without notice.
Avigilon Control Center Web Client User Guide Version: 4.12 Standard PDF-WEBCLIENT-S-E-Rev2 Copyright 2013 Avigilon. All rights reserved. The information presented is subject to change without notice.
Advanced Visual Medicine: Techniques for Visual Exploration & Analysis
 Advanced Visual Medicine: Techniques for Visual Exploration & Analysis Interactive Visualization of Multimodal Volume Data for Neurosurgical Planning Felix Ritter, MeVis Research Bremen Multimodal Neurosurgical
Advanced Visual Medicine: Techniques for Visual Exploration & Analysis Interactive Visualization of Multimodal Volume Data for Neurosurgical Planning Felix Ritter, MeVis Research Bremen Multimodal Neurosurgical
Programming within BrainVISA project
 Programming within BrainVISA project What is the BrainVISA project? Complete development environement RedMine based forge Integration of various programming languages Multiplatform programming Documentation
Programming within BrainVISA project What is the BrainVISA project? Complete development environement RedMine based forge Integration of various programming languages Multiplatform programming Documentation
Playing with data from lab
 Playing with data from lab Getting data off the scanner From the Patient Browser, select the folder for the study you want (or within that study, the set of images you want), and then from the Transfer
Playing with data from lab Getting data off the scanner From the Patient Browser, select the folder for the study you want (or within that study, the set of images you want), and then from the Transfer
Multi-voxel pattern analysis: Decoding Mental States from fmri Activity Patterns
 Multi-voxel pattern analysis: Decoding Mental States from fmri Activity Patterns Artwork by Leon Zernitsky Jesse Rissman NITP Summer Program 2012 Part 1 of 2 Goals of Multi-voxel Pattern Analysis Decoding
Multi-voxel pattern analysis: Decoding Mental States from fmri Activity Patterns Artwork by Leon Zernitsky Jesse Rissman NITP Summer Program 2012 Part 1 of 2 Goals of Multi-voxel Pattern Analysis Decoding
Two-Dimensional Projectile Motion
 Two-Dimensional Projectile Motion I. Introduction. This experiment involves the study of motion using a CCD video camera in which a sequence of video frames (a movie ) is recorded onto computer disk and
Two-Dimensional Projectile Motion I. Introduction. This experiment involves the study of motion using a CCD video camera in which a sequence of video frames (a movie ) is recorded onto computer disk and
SCENE FILE MANIPULATION SCENE FILE MANIPULATION GETTING STARTED MODELING ANIMATION MATERIALS + MAPPING RENDERING. Saving Files. Save.
 SCENE FILE MANIPULATION SCENE FILE MANIPULATION There are several things you can do with a scene file in 3ds Max. You can save a file, save a file temporarily and retrieve it, and combine scene files.
SCENE FILE MANIPULATION SCENE FILE MANIPULATION There are several things you can do with a scene file in 3ds Max. You can save a file, save a file temporarily and retrieve it, and combine scene files.
Measuring baseline whole-brain perfusion on GE 3.0T using arterial spin labeling (ASL) MRI
 Measuring baseline whole-brain perfusion on GE 3.0T using arterial spin labeling (ASL) MRI Revision date: 09/15/2008 Overview This document describes the procedure for measuring baseline whole-brain perfusion
Measuring baseline whole-brain perfusion on GE 3.0T using arterial spin labeling (ASL) MRI Revision date: 09/15/2008 Overview This document describes the procedure for measuring baseline whole-brain perfusion
Matlab 20D, Week 2 Exercise 2.2 (a) : DField drew lines of solutions of points on the plot that I d click on. b)
 Matlab 20D, Week 2 Exercise 2.2 (a) In this exercise, we are going to use DFIELD to plot the direction field of (4). This can be done by entering the following values into the DFIELD setup window. In the
Matlab 20D, Week 2 Exercise 2.2 (a) In this exercise, we are going to use DFIELD to plot the direction field of (4). This can be done by entering the following values into the DFIELD setup window. In the
Introductory Concepts for Voxel-Based Statistical Analysis
 Introductory Concepts for Voxel-Based Statistical Analysis John Kornak University of California, San Francisco Department of Radiology and Biomedical Imaging Department of Epidemiology and Biostatistics
Introductory Concepts for Voxel-Based Statistical Analysis John Kornak University of California, San Francisco Department of Radiology and Biomedical Imaging Department of Epidemiology and Biostatistics
Homework 1: Getting Started with WebGL and Transformations EE267 Virtual Reality 2018
 Homework 1: Getting Started with WebGL and Transformations EE267 Virtual Reality 2018 Due: 04/12/2018, 11:59pm Instruction Students should use JavaScript for this assignment, building on top of the provided
Homework 1: Getting Started with WebGL and Transformations EE267 Virtual Reality 2018 Due: 04/12/2018, 11:59pm Instruction Students should use JavaScript for this assignment, building on top of the provided
SIMULATION LAB #5: Muscle-Actuated Simulation of Kicking
 SIMULATION LAB #5: Muscle-Actuated Simulation of Kicking Modeling and Simulation of Human Movement BME 599 Laboratory Developers: Jeff Reinbolt, Hoa Hoang, B.J. Fregley, Kate Saul Holzbaur, Darryl Thelen,
SIMULATION LAB #5: Muscle-Actuated Simulation of Kicking Modeling and Simulation of Human Movement BME 599 Laboratory Developers: Jeff Reinbolt, Hoa Hoang, B.J. Fregley, Kate Saul Holzbaur, Darryl Thelen,
Lines of Symmetry. Grade 3. Amy Hahn. Education 334: MW 8 9:20 a.m.
 Lines of Symmetry Grade 3 Amy Hahn Education 334: MW 8 9:20 a.m. GRADE 3 V. SPATIAL SENSE, GEOMETRY AND MEASUREMENT A. Spatial Sense Understand the concept of reflection symmetry as applied to geometric
Lines of Symmetry Grade 3 Amy Hahn Education 334: MW 8 9:20 a.m. GRADE 3 V. SPATIAL SENSE, GEOMETRY AND MEASUREMENT A. Spatial Sense Understand the concept of reflection symmetry as applied to geometric
Building Vector Layers
 Building Vector Layers in QGIS Introduction: Spatially referenced data can be separated into two categories, raster and vector data. This week, we focus on the building of vector features. Vector shapefiles
Building Vector Layers in QGIS Introduction: Spatially referenced data can be separated into two categories, raster and vector data. This week, we focus on the building of vector features. Vector shapefiles
FSL Workshop Session 3 David Smith & John Clithero
 FSL Workshop 12.09.08 Session 3 David Smith & John Clithero What is MELODIC? Probabilistic ICA Improves upon standard ICA Allows for inference Avoids over-fitting Three stage process ( ppca ) 1.) Dimension
FSL Workshop 12.09.08 Session 3 David Smith & John Clithero What is MELODIC? Probabilistic ICA Improves upon standard ICA Allows for inference Avoids over-fitting Three stage process ( ppca ) 1.) Dimension
A guide to XiT. Main user interface of XiT
 A guide to XiT XiT stands for Xdimensional image analysis Toolbox. It is a tool written in Matlab (R2015b, The MathWorks, Inc., Natick, Massachusetts, United States), which enables performing multidimensional
A guide to XiT XiT stands for Xdimensional image analysis Toolbox. It is a tool written in Matlab (R2015b, The MathWorks, Inc., Natick, Massachusetts, United States), which enables performing multidimensional
2 Lab 2: LabVIEW and Control System Building Blocks
 2 Lab 2: LabVIEW and Control System Building Blocks 2.1 Introduction Controllers are built from mechanical or electrical building blocks. Most controllers are implemented in a program using sensors to
2 Lab 2: LabVIEW and Control System Building Blocks 2.1 Introduction Controllers are built from mechanical or electrical building blocks. Most controllers are implemented in a program using sensors to
Bonus Ch. 1. Subdivisional Modeling. Understanding Sub-Ds
 Bonus Ch. 1 Subdivisional Modeling Throughout this book, you ve used the modo toolset to create various objects. Some objects included the use of subdivisional surfaces, and some did not. But I ve yet
Bonus Ch. 1 Subdivisional Modeling Throughout this book, you ve used the modo toolset to create various objects. Some objects included the use of subdivisional surfaces, and some did not. But I ve yet
L3 Series Software for Material Testing
 L3 Series Software for Material Testing Performing a Test The Better Solution Table of Contents Page 4.0 performing an L3 Test 4 4.1 L3 Test Modes of Operation 4 4.1.1 Standard Operating Mode 4 4.1.2
L3 Series Software for Material Testing Performing a Test The Better Solution Table of Contents Page 4.0 performing an L3 Test 4 4.1 L3 Test Modes of Operation 4 4.1.1 Standard Operating Mode 4 4.1.2
GUIDE TO VIEW3D. Introduction to View3D
 View3D Guide Introduction to View3D... 1 Starting Hampson-Russell Software... 2 Starting View3D... 4 A Brief Summary of the View3D Process... 8 Loading the Seismic and Horizon Data... 8 Selection Errors...
View3D Guide Introduction to View3D... 1 Starting Hampson-Russell Software... 2 Starting View3D... 4 A Brief Summary of the View3D Process... 8 Loading the Seismic and Horizon Data... 8 Selection Errors...
