Group (Level 2) fmri Data Analysis - Lab 4
|
|
|
- Allen Parsons
- 6 years ago
- Views:
Transcription
1 Group (Level 2) fmri Data Analysis - Lab 4 Index Goals of this Lab Before Getting Started The Chosen Ten Checking Data Quality Create a Mean Anatomical of the Group Group Analysis: One-Sample T-Test Examine Results in SPM Visualization of Results in MRIcron Region of Interest Analyses Non-parametric model estimation Goals of this lab 1. Learn how to evaluate aspects of data quality that affect group analyses. 2. Learn how to run group analyses on the 10 subjects included in the course dataset. 3. Learn how to correct your results for multiple comparisons at both the voxel and cluster levels. 4. Learn how to visualize and explore your results. 5. Learn how to define regions of interest (ROI) and use them in a group analysis. Before Getting Started Today's lab will require use of several 3rd-party extensions to spm12 analysis in MATLAB. These are available in a folder called extensions that should be present in the folder containing the data for your 10 subjects. Before you get started, add that folder - along with all of its subfolders - to your MATLAB search path. Once you are finished, check to make sure it worked by running the following in the MATLAB command window (for each MATLAB should print the path to the file - if not, try adding the path again): which spm which files which peak_nii which nee_roi
2 The Chosen Ten By now you will have estimated (at least) two distinct models of the fmri timeseries for the subject you just happened to choose to examine during the first SPM lab on preprocessing. Now, briefly close your eyes and imagine that, magically, the same models were estimated for the remaining nine subjects in the dataset. Now, open your eyes and behold the precooked folder containing a normalized anatomical image (wanat_hires.nii) and folders for the two models estimated in the single-subject lab. IMPORTANT: Use the precooked folders to estimate your group level analyses for EVERY subject, including the one you've already done some processing on. This will ensure that everyone is starting with the same set of data.
3 Checking Data Quality There are at at least two questions you need to answer about your individual subject data before you begin a group analysis: Is everyone in the same stereotaxic space? Use check reg in SPM to verify that all 10 sets of images are in the atlas space. Before you do that, note that for this lab there will be many instances in which you will have to select a set of 10 images from each one of The Chosen Ten. To learn how to DRAMATICALLY speed up the process of selecting multiple files, read this tutorial before moving on. (Seriously, it's mandatory.) Once you're ready, examine 10 wanat_hires.nii's and see how the anatomy lines up (if the orthogonal viewers are too small, view the first 5 and then the second 5 images). Are there any subjects who seem out of registration with the others? Is anyone missing functional data? For this, use check reg to compare the 10 "mask.img" file located in your single subject analysis folders. (Do this just for the 2x2 model, but know that it is generally good practice to do this for
4 every model that you estimate.) SPM automatically generates these images to show the User (that's you!) which voxels were included in their analysis. This is called masking, in which only a subset of all voxels in the image are singled out for computation. When SPM decides which voxels to include and which to exclude (which it does by default), the masking is known as "implicit" since it happens without the User having to "explicitly" specify a mask. You can learn more about "implicit" versus "explicit" masking in the SPM manual, or here. This is very important to know, for the following reason: By default, SPM will only include a voxel in your group analysis if and only if EVERY individual subject has data at that voxel. If 9/10 have it, it will be excluded. With that in mind, check out your group's mask images, playing close attention to ventral regions of the frontal and temporal lobe that are notoriously susceptible to signal distortion and signal loss. Are there any subjects who seem to have more signal loss than others? Create a Mean Anatomical for the Group Let's create an anatomical reference image using the ImCalc tool:
5 1. Go to Options: Data Matrix. Click Yes-read images into data matrix. This will allow ImCalc to read all of the images you are about to specify into matrix X. 2. Now go to Input Images and select the 10 wanat_hires.nii which are subjects' normalized anatomical images. 3. In Output Filename type wanat_hires_mean.img. In Output Directory specify the directory into which you want the mean anatomical to be written. In Expression write mean(x). This expression will create an average of the anatomical images. 4. Run the batch. 5. Use the Check Reg utility to compare the mean anatomical to the template image. Does it look like the group is, on average, in good registration with the template? Now take a quick look at this tutorial to see how to do what you just did in less than 10 seconds using the function FILES and a second function you'll learn about called BSPM_IMCALC. Group Analysis: One-Sample T-Test Start by making an output folder for each of the analyses you plan on running. If I were you, I'd make a folder called "groupstats" in your data directory (use "mkdir groupstats" in the MATLAB command window to do this). Then, enter that folder with "cd groupstats" and make two additional folders called "2x2" and "Parametric". Finally, enter the 2x2 folder with "cd 2x2" and make folders
6 called "Why-How" and "Face-Hand". These correspond to con_0001.img and con_0002.img in your individual subject folders (you'll use the precooked models, even for your own subject). Now, let's use the Batch Editor to specify and estimate a one-sample t-test on the first of these images: 1. Open the Batch Editor 2. Open a Stats: Factorial Design Specification module 3. For Directory, select the analysis directory you created (i.e., the Why-How directory). 4. Click on "Design" and observe that you can select from a variety of statistical models; the one we need, "One Sample t test", is already selected. Expand it, then for Scans, select the ten (10) con_0001 contrast images. Tip: use the files command to create a list of all contrast images that you can copy and paste into the 'Scans' dialog when you click edit:
7 5. Add an Estimation job using the Batch Editor (as we did on Monday). Specify a dependency with the SPM.mat created from the first job. 6. Add a Contrast Manager job. Define two contrasts. First, "Why-How" requires a single contrast weight: +1. Then, "How-Why" requires the inverse: Save this Job.
8 8. Repeat the above for the Face-Hand contrast. 9. Save that job, too. 10. Load both jobs using "Load" from the Batch Editor menubar. The Module List should look like this: 11. When you're ready, run the jobs. The SPM figure window should look like this:
9 Examine Results in SPM The time has come to peer inside the group mind of The Chosen Ten. Using the Results button in the SPM menu, load in the SPM.mat file for the 2x2 model and select the Why > How contrast for viewing. For now, threshold it with:
10 1. Do not apply masking. 2. P-value adjusment should be "none" 3. Threshold should be left at Extent threshold at 0 Once you see the "glass brain", use the overlay menu (using 'sections') to use the mean anatomical as an underlay for the thresholded statistical map. Use the "whole brain" option under p-values to produce a report showing the peaks wihtin each identified cluster of activity. Below, you can see this table. The circled part at the bottom (FWEc) shows the minimum extent (which we've so far been leaving at 0) required for a cluster to considered "significant" at a family-wise error rate of.05. This is one of the forms of cluster-level correction for multiple comparisons that SPM offers.
11 To apply this correction, use the drop down menu to change the thresholding you're using, as shown below:
12 Enter all the same values except the last one, where it asks for the "extent threshold". Instead of entering "0", enter the value necessary for cluster-level correction. Does the resulting thresholding statistic map look less noisy to you? Now, let's change the threshold again but this time apply a voxel level correction for multiple comparisons. To do this, use the change threshold procedure as above. When you get "P-value adjusment", choose the option for family-wise error (FWE). Then, stick with the default threshold of.05 (which in this case refers to the FWE corrected threshold), and do not apply an extent threshold (that is, leave the last input at 0). Did this change the number of "supra" threshold voxels? At this point, this walkthrough challenges you to do the following: 1. Use SPM to load the results for the Face > Hand contrast. 2. Save a CSV file that reports those regions that survive an uncorrected voxel-wise threshold of.001 and a family-wise error corrected cluster-level threshold of.05. Which regions survived? If you did things correctly, you should see bilateral activation in some noname region of the brain that no one has every heard of. (Just in case it wasn't clear, that last sentence was meant to be ironic.) As shown below, SPM offers a number of different options for saving all or part of your results as image files. Given that we'll shortly be discussing region of interest analyses (ROIs), use the "Save current cluster" option shown below to save an image of just the right amygdala (for this, you'll need to have the crosshair positioned on a voxel in the
13 cluster). Use the Display utility to take a look at the "functional" ROI you've just created:
14 Visualization of Results in MRIcron To visualize the data, we'll use MRICron. MRIcron has already been installed on your machine and can be found here. You can launch MRIcron from Windows by doube-clicking the mricron icon. Once it is open, click on Open Templates and load in the chi2better.nii.gz. This is a skullstripped version of a canonical template in MNI space.
15 Next, click on Overlay -> Add and open the spm_t map located in your group analysis of the Why > How contrast in the 2x2 model. Once loaded, we can change the color map and the upper and lower bounds of the color map:
16 Let's start with the "Multislice" feature. Go to Window -> Multislice Under View, you can change which slices you want to look at (Slices...), how much the slices visually overlap (Overslice), and various other features. When you re happy with the figure, you can go to File -> Save as bitmap... to cherish forever.
17 Not satisfied? Try the Window -> Render option that is much more flexible than SPM: The Azimuth and Elevation change the view. Make sure to check the option Overlay -> Search -> Infront/below BG surface so that deep activations are not mysteriously brought to the surface. You
18 can control how much activations are brought to the surface under Overlay -> Search Depth. Finally, we can peel back chunks of the brain to observe activations lurking in inner structures. Click on View -> Cutout and then click the Show cutout checkbox. The lower and upper limits of X, Y, and Z will determine what parts of the brain are cutout. Feel free to play with those values and perform different cuts. Brain surgery made simple... When you re satisfied with your cuts, you can click OK in the cutout window. Finally, click the
19 double red arrow at the top of the render window to generate a high resolution rendering. Note, that doing this multiple times may cause MRIcron to crash. It appears to be good for one high resolution rendering per use of the program. Now, you can click on File -> Save as bitmap... for a nice figure. Region of Interest Analyses You've already seen above how you can easily make functional ROIs from activation maps in SPM. In this tutorial, you'll see how to make ROIs based on the peak coordinates reported in previous neuroimaging studies. Finally, let s extract data from ROIs. First, gather the contrast images for the Face > Hand contrast. You can use the function files again, or you can find them neatly stored in the Group Level SPM.mat file under the field SPM.xY.P. Next, we ll extract ROI averaged data for each subject. P = SPM.xY.P; % alternatively, set P to the images you gathered using files ROI = 'ROI_Sphere9_-50_9_30_L_IFG.nii'; % Or you can use a different ROI file roidata = nee_roi(p,roi); roidata This returns an n x 1 vector with the contrast estimate averaged across the ROI for each subject. You can perform a t-test on it: [h,p,ci,stats] = ttest(roidata) Next, you can do the same analysis on the functional ROI you created above. That analysis
20 should be highly significant. But we already knew that since the functional ROI was defined as a region that showed a significant effect (and reporting it as if it were another independent test in a paper is one form of double-dipping). However, extracting data from a functional ROI can sometimes be useful to serve as a correlate (e.g. with behavior or another ROI). Non-parametric model estimation (SnPM) With the Statistical nonparametric Mapping (SnPM) toolbox you can estimate your models using a non-parametric permutation-based approach. Non-parametric model estimation has the advantage that it does not have any assumptions about the distribution of your outcome measure (i.e., your voxel values) and is therefore also appropriate for non-normally distributed residuals. You can start the SnPM toolbox by typing snpm_ui, or by selecting a design from the batch editor: In the sample below we will run the same one-sample t-test as we ran previously, but now using SnPM. For this, create a new folder to store your model and results. Next, select MultiSub: One Sample T test on diffs/contrasts. Specify your analysis directory, and the contrast images of the contrast you want to analyze:
21 You can select your first level parametric contrast images for this second level non-parametric group level analysis. For this example, we will select each second contrast image (Face-Hand) of the parametric 1st level analysis of all 10 subjects (i.e, con_0002.img). SnPM has a couple of options that you have to set. As you know by now, SPM (and SnPM as well) provide a lot of good information about the options in the batch editor in the bottom part of the batch editor. Below, we will briefly go over the most important options. For more information see the text boxes in the SnPM batch window: Number of permutations: The number of permutations that is being conducted can be exhaustive, but this normally takes a very long time to complete. This is why you can select to run a random subset of the possible permutations. The default is You can reduce the number of permutations for
22 testing. Generally, for publication you should set the number of permutations to 10,000 or more. The more permutations, the more precise the p-value. For now, let's set the number of permutations to: (Note that for a one-sample t-test the non-parametric sign flip test is used instead of permutation tests). Variance smoothing: This is particularly useful if you have a small sample (i.e., n<20). In this case SnPM regularizes your variance image. In general, the smoothing kernel that you select for your variance maps should not exceed the smoothing kernel that you selected for your data (i.e., 8mm in our case). For now, let's set the variance smoothing to: [8 8 8]. Additional background information about variance smoothing by Thomas Nichols can be found HERE. Cluster inference: By default cluster inference is not performed because it takes a long time to run. If you want to easily switch between different cluster size inferences it is recommended to set cluster inference to: Yes (slow, may create huge SnPM_ST.mat file). However, for now we are going to use cluster inference with a predefined statistical threshold. For this, select Yes, set cluster-forming threshold now (fast) and set it to Now that we have created a model, we are going to add add compute to our batch (this is the equivalent of estimate in the parametric analysis). To add compute to your batch select it from the batch menu: SPM --> Tools --> SnPM --> Compute. Click on dependency and select the design that you just created. Finally, add inference to your batch by selecting it from the batch menu: SPM --> Tools --> SnPM --> Inference. Inference is the equivalent for results in SPM. Set the SnPM.mat results file to dependency and select: Compute: SnPM.mat results file. For type of thresholding you can select either voxel wise or cluster wise inference. Both are valid options, but for now we are going to select Cluster-Level Inference. We will leave the Cluster-Forming Threshold at NaN, because we have already set it to p<0.001 when we were defining our model. Now, keep the default value for all other options, save your batch, and run it! Results: Your results overview should look like this (click to enlarge):
23 If you still have time, compare these results to the results from the parametric One-Sample T-Test of the second contrast (Face-Hand). Do they differ a lot? If so, what could be an explanation? Note that SnPM does not offer all the tools in the results window that SPM offers. However, SnPM stores the pseudo-t image in the file "snpmt-.img". You can open this file in MRICron and use the T-threshold that you will find in your SnPM output, under "Cluster-defining thresh".
SnPM is an SPM toolbox developed by Andrew Holmes & Tom Nichols
 1 of 14 3/30/2005 9:24 PM SnPM A Worked fmri Example SnPM is an SPM toolbox developed by Andrew Holmes & Tom Nichols This page... introduction example data background design setup computation viewing results
1 of 14 3/30/2005 9:24 PM SnPM A Worked fmri Example SnPM is an SPM toolbox developed by Andrew Holmes & Tom Nichols This page... introduction example data background design setup computation viewing results
SPM Introduction. SPM : Overview. SPM: Preprocessing SPM! SPM: Preprocessing. Scott Peltier. FMRI Laboratory University of Michigan
 SPM Introduction Scott Peltier FMRI Laboratory University of Michigan! Slides adapted from T. Nichols SPM! SPM : Overview Library of MATLAB and C functions Graphical user interface Four main components:
SPM Introduction Scott Peltier FMRI Laboratory University of Michigan! Slides adapted from T. Nichols SPM! SPM : Overview Library of MATLAB and C functions Graphical user interface Four main components:
SPM Introduction SPM! Scott Peltier. FMRI Laboratory University of Michigan. Software to perform computation, manipulation and display of imaging data
 SPM Introduction Scott Peltier FMRI Laboratory University of Michigan Slides adapted from T. Nichols SPM! Software to perform computation, manipulation and display of imaging data 1 1 SPM : Overview Library
SPM Introduction Scott Peltier FMRI Laboratory University of Michigan Slides adapted from T. Nichols SPM! Software to perform computation, manipulation and display of imaging data 1 1 SPM : Overview Library
Lab 5: Multi Subject fmri
 USA SPM Short Course April 6-8, 2005 Yale University Lab 5: Multi Subject fmri Rik Henson mailto:rik.henson@mrc-cbu.cam.ac.uk MRC Cognition & Brain Sciences Unit, Cambridge, UK CONTENTS Goals Introduction
USA SPM Short Course April 6-8, 2005 Yale University Lab 5: Multi Subject fmri Rik Henson mailto:rik.henson@mrc-cbu.cam.ac.uk MRC Cognition & Brain Sciences Unit, Cambridge, UK CONTENTS Goals Introduction
Resources for Nonparametric, Power and Meta-Analysis Practical SPM Course 2015, Zurich
 Resources for Nonparametric, Power and Meta-Analysis Practical SPM Course 2015, Zurich http://www.translationalneuromodeling.org/practical-sessions/ Preliminary Get this file: http://warwick.ac.uk/tenichols/zurich.pdf
Resources for Nonparametric, Power and Meta-Analysis Practical SPM Course 2015, Zurich http://www.translationalneuromodeling.org/practical-sessions/ Preliminary Get this file: http://warwick.ac.uk/tenichols/zurich.pdf
Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question.
 (1) conn - Functional connectivity toolbox v1.0 Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question. 2) in "Basic" enter "1" subject, "6" seconds
(1) conn - Functional connectivity toolbox v1.0 Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question. 2) in "Basic" enter "1" subject, "6" seconds
Data Visualisation in SPM: An introduction
 Data Visualisation in SPM: An introduction Alexa Morcom Edinburgh SPM course, April 2015 SPMmip [-30, 3, -9] 3 Visualising results remembered vs. fixation contrast(s) < < After the results table - what
Data Visualisation in SPM: An introduction Alexa Morcom Edinburgh SPM course, April 2015 SPMmip [-30, 3, -9] 3 Visualising results remembered vs. fixation contrast(s) < < After the results table - what
fmri Basics: Single Subject Analysis
 fmri Basics: Single Subject Analysis This session is intended to give an overview of the basic process of setting up a general linear model for a single subject. This stage of the analysis is also variously
fmri Basics: Single Subject Analysis This session is intended to give an overview of the basic process of setting up a general linear model for a single subject. This stage of the analysis is also variously
Documentation for imcalc (SPM 5/8/12) Robert J Ellis
 (_) _ '_ ` _ \ / / _` / Image calculations and transformations (using SPM) (_ (_ ( This software version: 09-Nov-2017 _ _ _ _ \ \,_ _ \ (C) Robert J Ellis (http://tools.robjellis.net) Documentation for
(_) _ '_ ` _ \ / / _` / Image calculations and transformations (using SPM) (_ (_ ( This software version: 09-Nov-2017 _ _ _ _ \ \,_ _ \ (C) Robert J Ellis (http://tools.robjellis.net) Documentation for
Version. Getting Started: An fmri-cpca Tutorial
 Version 11 Getting Started: An fmri-cpca Tutorial 2 Table of Contents Table of Contents... 2 Introduction... 3 Definition of fmri-cpca Data... 3 Purpose of this tutorial... 3 Prerequisites... 4 Used Terms
Version 11 Getting Started: An fmri-cpca Tutorial 2 Table of Contents Table of Contents... 2 Introduction... 3 Definition of fmri-cpca Data... 3 Purpose of this tutorial... 3 Prerequisites... 4 Used Terms
GLM for fmri data analysis Lab Exercise 1
 GLM for fmri data analysis Lab Exercise 1 March 15, 2013 Medical Image Processing Lab Medical Image Processing Lab GLM for fmri data analysis Outline 1 Getting Started 2 AUDIO 1 st level Preprocessing
GLM for fmri data analysis Lab Exercise 1 March 15, 2013 Medical Image Processing Lab Medical Image Processing Lab GLM for fmri data analysis Outline 1 Getting Started 2 AUDIO 1 st level Preprocessing
Data Visualisation in SPM: An introduction
 Data Visualisation in SPM: An introduction Alexa Morcom Edinburgh SPM course, April 2010 Centre for Cognitive & Neural Systems/ Department of Psychology University of Edinburgh Visualising results remembered
Data Visualisation in SPM: An introduction Alexa Morcom Edinburgh SPM course, April 2010 Centre for Cognitive & Neural Systems/ Department of Psychology University of Edinburgh Visualising results remembered
Normalization for clinical data
 Normalization for clinical data Christopher Rorden, Leonardo Bonilha, Julius Fridriksson, Benjamin Bender, Hans-Otto Karnath (2012) Agespecific CT and MRI templates for spatial normalization. NeuroImage
Normalization for clinical data Christopher Rorden, Leonardo Bonilha, Julius Fridriksson, Benjamin Bender, Hans-Otto Karnath (2012) Agespecific CT and MRI templates for spatial normalization. NeuroImage
Correction for multiple comparisons. Cyril Pernet, PhD SBIRC/SINAPSE University of Edinburgh
 Correction for multiple comparisons Cyril Pernet, PhD SBIRC/SINAPSE University of Edinburgh Overview Multiple comparisons correction procedures Levels of inferences (set, cluster, voxel) Circularity issues
Correction for multiple comparisons Cyril Pernet, PhD SBIRC/SINAPSE University of Edinburgh Overview Multiple comparisons correction procedures Levels of inferences (set, cluster, voxel) Circularity issues
Analysis of fmri data within Brainvisa Example with the Saccades database
 Analysis of fmri data within Brainvisa Example with the Saccades database 18/11/2009 Note : All the sentences in italic correspond to informations relative to the specific dataset under study TP participants
Analysis of fmri data within Brainvisa Example with the Saccades database 18/11/2009 Note : All the sentences in italic correspond to informations relative to the specific dataset under study TP participants
fmri Preprocessing & Noise Modeling
 Translational Neuromodeling Unit fmri Preprocessing & Noise Modeling Lars Kasper September 25 th / October 17 th, 2015 MR-Technology Group & Translational Neuromodeling Unit An SPM Tutorial Institute for
Translational Neuromodeling Unit fmri Preprocessing & Noise Modeling Lars Kasper September 25 th / October 17 th, 2015 MR-Technology Group & Translational Neuromodeling Unit An SPM Tutorial Institute for
SPM Course! Single Subject Analysis
 SPM Course! Single Subject Analysis Practical Session Dr. Jakob Heinzle & Dr. Frederike Petzschner & Dr. Lionel Rigoux Hands up: Who has programming experience with Matlab? Who has analyzed an fmri experiment
SPM Course! Single Subject Analysis Practical Session Dr. Jakob Heinzle & Dr. Frederike Petzschner & Dr. Lionel Rigoux Hands up: Who has programming experience with Matlab? Who has analyzed an fmri experiment
CONN fmri functional connectivity toolbox
 CONN fmri functional connectivity toolbox Gabrieli Lab. McGovern Institute for Brain Research Massachusetts Institute of Technology http://www.nitrc.org/projects/conn Susan Whitfield-Gabrieli Alfonso Nieto-Castanon
CONN fmri functional connectivity toolbox Gabrieli Lab. McGovern Institute for Brain Research Massachusetts Institute of Technology http://www.nitrc.org/projects/conn Susan Whitfield-Gabrieli Alfonso Nieto-Castanon
Quality Checking an fmri Group Result (art_groupcheck)
 Quality Checking an fmri Group Result (art_groupcheck) Paul Mazaika, Feb. 24, 2009 A statistical parameter map of fmri group analyses relies on the assumptions of the General Linear Model (GLM). The assumptions
Quality Checking an fmri Group Result (art_groupcheck) Paul Mazaika, Feb. 24, 2009 A statistical parameter map of fmri group analyses relies on the assumptions of the General Linear Model (GLM). The assumptions
Table of Contents. IntroLab < SPMLabs < Dynevor TWiki
 Table of Contents Lab 1: Introduction to SPM and data checking...1 Goals of this Lab...1 Prerequisites...1 An SPM Installation...1 SPM Defaults...2 L/R Brain Orientation...2 Memory Use for Data Processing...2
Table of Contents Lab 1: Introduction to SPM and data checking...1 Goals of this Lab...1 Prerequisites...1 An SPM Installation...1 SPM Defaults...2 L/R Brain Orientation...2 Memory Use for Data Processing...2
BPM e Toolbox Biological Parametric Mapping - Extended
 BPM e Toolbox Biological Parametric Mapping - Extended The BPM Random Regressors Toolbox is provided as an overlay on the WFU Biological Parametric Mapping Toolbox release 1.5d as modified by the robust
BPM e Toolbox Biological Parametric Mapping - Extended The BPM Random Regressors Toolbox is provided as an overlay on the WFU Biological Parametric Mapping Toolbox release 1.5d as modified by the robust
New and best-practice approaches to thresholding
 New and best-practice approaches to thresholding Thomas Nichols, Ph.D. Department of Statistics & Warwick Manufacturing Group University of Warwick FIL SPM Course 17 May, 2012 1 Overview Why threshold?
New and best-practice approaches to thresholding Thomas Nichols, Ph.D. Department of Statistics & Warwick Manufacturing Group University of Warwick FIL SPM Course 17 May, 2012 1 Overview Why threshold?
7/15/2016 ARE YOUR ANALYSES TOO WHY IS YOUR ANALYSIS PARAMETRIC? PARAMETRIC? That s not Normal!
 ARE YOUR ANALYSES TOO PARAMETRIC? That s not Normal! Martin M Monti http://montilab.psych.ucla.edu WHY IS YOUR ANALYSIS PARAMETRIC? i. Optimal power (defined as the probability to detect a real difference)
ARE YOUR ANALYSES TOO PARAMETRIC? That s not Normal! Martin M Monti http://montilab.psych.ucla.edu WHY IS YOUR ANALYSIS PARAMETRIC? i. Optimal power (defined as the probability to detect a real difference)
Figure 1. Comparison of the frequency of centrality values for central London and our region of Soho studied. The comparison shows that Soho falls
 A B C Figure 1. Comparison of the frequency of centrality values for central London and our region of Soho studied. The comparison shows that Soho falls well within the distribution for London s streets.
A B C Figure 1. Comparison of the frequency of centrality values for central London and our region of Soho studied. The comparison shows that Soho falls well within the distribution for London s streets.
BrainSuite Lab Exercises. presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014
 BrainSuite Lab Exercises presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 1. Opening and Displaying an MRI Start BrainSuite Drag and drop the T1 image from the native space
BrainSuite Lab Exercises presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 1. Opening and Displaying an MRI Start BrainSuite Drag and drop the T1 image from the native space
NEURO M203 & BIOMED M263 WINTER 2014
 NEURO M203 & BIOMED M263 WINTER 2014 MRI Lab 2: Neuroimaging Connectivity Lab In today s lab we will work with sample diffusion imaging data and the group averaged fmri data collected during your scanning
NEURO M203 & BIOMED M263 WINTER 2014 MRI Lab 2: Neuroimaging Connectivity Lab In today s lab we will work with sample diffusion imaging data and the group averaged fmri data collected during your scanning
Multiple comparisons problem and solutions
 Multiple comparisons problem and solutions James M. Kilner http://sites.google.com/site/kilnerlab/home What is the multiple comparisons problem How can it be avoided Ways to correct for the multiple comparisons
Multiple comparisons problem and solutions James M. Kilner http://sites.google.com/site/kilnerlab/home What is the multiple comparisons problem How can it be avoided Ways to correct for the multiple comparisons
Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry
 Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry Nivedita Agarwal, MD Nivedita.agarwal@apss.tn.it Nivedita.agarwal@unitn.it Volume and surface morphometry Brain volume White matter
Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry Nivedita Agarwal, MD Nivedita.agarwal@apss.tn.it Nivedita.agarwal@unitn.it Volume and surface morphometry Brain volume White matter
Press Release. Introduction. Prerequisites for LORETA
 Press Release Support & Tips A guided tour through LORETA Source localization in BrainVision Analyzer 2 by Dr.-Ing. Kidist Mideksa, Scientific Consultant at Brain Products Scientific Support What do I
Press Release Support & Tips A guided tour through LORETA Source localization in BrainVision Analyzer 2 by Dr.-Ing. Kidist Mideksa, Scientific Consultant at Brain Products Scientific Support What do I
Artifact Detection and Repair: Overview and Sample Outputs
 Artifact Detection and Repair: Overview and Sample Outputs Paul Mazaika February 2007 Programs originated in Gabrieli Neuroscience Laboratory, updated and enhanced at Center for Interdisciplinary Brain
Artifact Detection and Repair: Overview and Sample Outputs Paul Mazaika February 2007 Programs originated in Gabrieli Neuroscience Laboratory, updated and enhanced at Center for Interdisciplinary Brain
Batch processing of GLMs
 Batch processing of GLMs 1 Contents Batch processing of GLMs... 1 Contents... 2 Introduction... 3 Installing the project... 4 1. Location of the script project... 4 2. Setting the project as current in
Batch processing of GLMs 1 Contents Batch processing of GLMs... 1 Contents... 2 Introduction... 3 Installing the project... 4 1. Location of the script project... 4 2. Setting the project as current in
FNC Toolbox Walk Through
 FNC Toolbox Walk Through Date: August 10 th, 2009 By: Nathan Swanson, Vince Calhoun at The Mind Research Network Email: nswanson@mrn.org Introduction The FNC Toolbox is an extension of the GIFT toolbox
FNC Toolbox Walk Through Date: August 10 th, 2009 By: Nathan Swanson, Vince Calhoun at The Mind Research Network Email: nswanson@mrn.org Introduction The FNC Toolbox is an extension of the GIFT toolbox
This exercise uses one anatomical data set (ANAT1) and two functional data sets (FUNC1 and FUNC2).
 Exploring Brain Anatomy This week s exercises will let you explore the anatomical organization of the brain to learn some of its basic properties, as well as the location of different structures. The human
Exploring Brain Anatomy This week s exercises will let you explore the anatomical organization of the brain to learn some of its basic properties, as well as the location of different structures. The human
How to make ROI files
 How to make ROI files Ver. 1.0 2012/08/07 Introduction Functional ROI Structural ROI Hand-made ROI History Contact 2 Introduction This manual explains how to make ROI (Region Of Interest) files that you
How to make ROI files Ver. 1.0 2012/08/07 Introduction Functional ROI Structural ROI Hand-made ROI History Contact 2 Introduction This manual explains how to make ROI (Region Of Interest) files that you
fmri/dti analysis using Dynasuite
 fmri/dti analysis using Dynasuite Contents 1 Logging in 2 Finding patient session 3 Viewing and adjusting images 4 Checking brain segmentation 5 Checking image registration 6 Seeing fmri results 7 Saving
fmri/dti analysis using Dynasuite Contents 1 Logging in 2 Finding patient session 3 Viewing and adjusting images 4 Checking brain segmentation 5 Checking image registration 6 Seeing fmri results 7 Saving
Session scaling; global mean scaling; block effect; mean intensity scaling
 Types of Scaling Session scaling; global mean scaling; block effect; mean intensity scaling Purpose remove intensity differences between runs (i.e., the mean of the whole time series). whole time series
Types of Scaling Session scaling; global mean scaling; block effect; mean intensity scaling Purpose remove intensity differences between runs (i.e., the mean of the whole time series). whole time series
The organization of the human cerebral cortex estimated by intrinsic functional connectivity
 1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
EMPIRICALLY INVESTIGATING THE STATISTICAL VALIDITY OF SPM, FSL AND AFNI FOR SINGLE SUBJECT FMRI ANALYSIS
 EMPIRICALLY INVESTIGATING THE STATISTICAL VALIDITY OF SPM, FSL AND AFNI FOR SINGLE SUBJECT FMRI ANALYSIS Anders Eklund a,b,c, Thomas Nichols d, Mats Andersson a,c, Hans Knutsson a,c a Department of Biomedical
EMPIRICALLY INVESTIGATING THE STATISTICAL VALIDITY OF SPM, FSL AND AFNI FOR SINGLE SUBJECT FMRI ANALYSIS Anders Eklund a,b,c, Thomas Nichols d, Mats Andersson a,c, Hans Knutsson a,c a Department of Biomedical
SPM99 fmri Data Analysis Workbook
 SPM99 fmri Data Analysis Workbook This file is a description of the steps needed to use SPM99 analyze a fmri data set from a single subject using a simple on/off activation paradigm. There are two parts
SPM99 fmri Data Analysis Workbook This file is a description of the steps needed to use SPM99 analyze a fmri data set from a single subject using a simple on/off activation paradigm. There are two parts
NA-MIC National Alliance for Medical Image Computing fmri Data Analysis
 NA-MIC fmri Data Analysis Sonia Pujol, Ph.D. Wendy Plesniak, Ph.D. Randy Gollub, M.D., Ph.D. Acknowledgments NIH U54EB005149 Neuroimage Analysis Center NIH P41RR013218 FIRST Biomedical Informatics Research
NA-MIC fmri Data Analysis Sonia Pujol, Ph.D. Wendy Plesniak, Ph.D. Randy Gollub, M.D., Ph.D. Acknowledgments NIH U54EB005149 Neuroimage Analysis Center NIH P41RR013218 FIRST Biomedical Informatics Research
Preprocessing of fmri Data in SPM 12 - Lab 1
 Preprocessing of fmri Data in SPM 12 - Lab 1 Index Goals of this Lab Preprocessing Overview MATLAB, SPM, Data Setup Preprocessing I: Checking Motion Correction Preprocessing II: Coregistration Preprocessing
Preprocessing of fmri Data in SPM 12 - Lab 1 Index Goals of this Lab Preprocessing Overview MATLAB, SPM, Data Setup Preprocessing I: Checking Motion Correction Preprocessing II: Coregistration Preprocessing
Spatial Filtering Methods in MEG. Part 3: Template Normalization and Group Analysis"
 Spatial Filtering Methods in MEG Part 3: Template Normalization and Group Analysis" Douglas Cheyne, PhD" Program in Neurosciences and Mental Health" Hospital for Sick Children Research Institute " &" Department
Spatial Filtering Methods in MEG Part 3: Template Normalization and Group Analysis" Douglas Cheyne, PhD" Program in Neurosciences and Mental Health" Hospital for Sick Children Research Institute " &" Department
Extending the GLM. Outline. Mixed effects motivation Evaluating mixed effects methods Three methods. Conclusions. Overview
 Extending the GLM So far, we have considered the GLM for one run in one subject The same logic can be applied to multiple runs and multiple subjects GLM Stats For any given region, we can evaluate the
Extending the GLM So far, we have considered the GLM for one run in one subject The same logic can be applied to multiple runs and multiple subjects GLM Stats For any given region, we can evaluate the
ASAP_2.0 (Automatic Software for ASL Processing) USER S MANUAL
 ASAP_2.0 (Automatic Software for ASL Processing) USER S MANUAL ASAP was developed as part of the COST Action "Arterial Spin Labelling Initiative in Dementia (AID)" by: Department of Neuroimaging, Institute
ASAP_2.0 (Automatic Software for ASL Processing) USER S MANUAL ASAP was developed as part of the COST Action "Arterial Spin Labelling Initiative in Dementia (AID)" by: Department of Neuroimaging, Institute
Learn Image Segmentation Basics with Hands-on Introduction to ITK-SNAP. RSNA 2016 Courses RCB22 and RCB54
 Learn Image Segmentation Basics with Hands-on Introduction to ITK-SNAP RSNA 2016 Courses RCB22 and RCB54 RCB22 Mon, Nov 28 10:30-12:00 PM, Room S401CD RCB54 Thu, Dec 1 2:30-4:30 PM, Room S401CD Presenters:
Learn Image Segmentation Basics with Hands-on Introduction to ITK-SNAP RSNA 2016 Courses RCB22 and RCB54 RCB22 Mon, Nov 28 10:30-12:00 PM, Room S401CD RCB54 Thu, Dec 1 2:30-4:30 PM, Room S401CD Presenters:
A User s Guide to Graphical-Model-based Multivariate Analysis
 A User s Guide to Graphical-Model-based Multivariate Analysis 1. Introduction Rong Chen December 2006 Graphical-Model-based Multivariate Analysis (GAMMA) is a Bayesian data mining software for structural
A User s Guide to Graphical-Model-based Multivariate Analysis 1. Introduction Rong Chen December 2006 Graphical-Model-based Multivariate Analysis (GAMMA) is a Bayesian data mining software for structural
VBM Tutorial. 1 Getting Started. John Ashburner. March 12, 2015
 VBM Tutorial John Ashburner March 12, 2015 1 Getting Started The data provided are a selection of T1-weighted scans from the freely available IXI dataset 1. The overall plan will be to Start up SPM. Check
VBM Tutorial John Ashburner March 12, 2015 1 Getting Started The data provided are a selection of T1-weighted scans from the freely available IXI dataset 1. The overall plan will be to Start up SPM. Check
Introduction to fmri. Pre-processing
 Introduction to fmri Pre-processing Tibor Auer Department of Psychology Research Fellow in MRI Data Types Anatomical data: T 1 -weighted, 3D, 1/subject or session - (ME)MPRAGE/FLASH sequence, undistorted
Introduction to fmri Pre-processing Tibor Auer Department of Psychology Research Fellow in MRI Data Types Anatomical data: T 1 -weighted, 3D, 1/subject or session - (ME)MPRAGE/FLASH sequence, undistorted
Function-Structure Integration in FreeSurfer
 Function-Structure Integration in FreeSurfer Outline Function-Structure Integration Function-Structure Registration in FreeSurfer fmri Analysis Preprocessing First-Level Analysis Higher-Level (Group) Analysis
Function-Structure Integration in FreeSurfer Outline Function-Structure Integration Function-Structure Registration in FreeSurfer fmri Analysis Preprocessing First-Level Analysis Higher-Level (Group) Analysis
ANALYSIS OF FUNCTIONAL MAGNETIC RESONANCE IMAGING DATA USING SPM99: VOXEL-BASED MORPHOMETRY DONNA ROSE ADDIS
 Donna Rose Addis, TWRI, May 2004 1 ANALYSIS OF FUNCTIONAL MAGNETIC RESONANCE IMAGING DATA USING SPM99: VOXEL-BASED MORPHOMETRY DONNA ROSE ADDIS DEPT. OF PSYCHOLOGY, UNIVERSITY OF TORONTO TORONTO WESTERN
Donna Rose Addis, TWRI, May 2004 1 ANALYSIS OF FUNCTIONAL MAGNETIC RESONANCE IMAGING DATA USING SPM99: VOXEL-BASED MORPHOMETRY DONNA ROSE ADDIS DEPT. OF PSYCHOLOGY, UNIVERSITY OF TORONTO TORONTO WESTERN
Cluster failure: Why fmri inferences for spatial extent have inflated false positive rates
 Supporting Information Appendix Cluster failure: Why fmri inferences for spatial extent have inflated false positive rates Anders Eklund, Thomas Nichols, Hans Knutsson Methods Resting state fmri data Resting
Supporting Information Appendix Cluster failure: Why fmri inferences for spatial extent have inflated false positive rates Anders Eklund, Thomas Nichols, Hans Knutsson Methods Resting state fmri data Resting
Functional MRI in Clinical Research and Practice Preprocessing
 Functional MRI in Clinical Research and Practice Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization
Functional MRI in Clinical Research and Practice Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization
Supplementary methods
 Supplementary methods This section provides additional technical details on the sample, the applied imaging and analysis steps and methods. Structural imaging Trained radiographers placed all participants
Supplementary methods This section provides additional technical details on the sample, the applied imaging and analysis steps and methods. Structural imaging Trained radiographers placed all participants
Controlling for multiple comparisons in imaging analysis. Wednesday, Lecture 2 Jeanette Mumford University of Wisconsin - Madison
 Controlling for multiple comparisons in imaging analysis Wednesday, Lecture 2 Jeanette Mumford University of Wisconsin - Madison Motivation Run 100 hypothesis tests on null data using p
Controlling for multiple comparisons in imaging analysis Wednesday, Lecture 2 Jeanette Mumford University of Wisconsin - Madison Motivation Run 100 hypothesis tests on null data using p
Hi everyone. I hope everyone had a good Fourth of July. Today we're going to be covering graph search. Now, whenever we bring up graph algorithms, we
 Hi everyone. I hope everyone had a good Fourth of July. Today we're going to be covering graph search. Now, whenever we bring up graph algorithms, we have to talk about the way in which we represent the
Hi everyone. I hope everyone had a good Fourth of July. Today we're going to be covering graph search. Now, whenever we bring up graph algorithms, we have to talk about the way in which we represent the
SPM8 for Basic and Clinical Investigators. Preprocessing. fmri Preprocessing
 SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
PRANA. Project Review and Analysis. March (Pre-release and still under development!)
 PRANA Project Review and Analysis March 2010- (Pre-release and still under development!) A.A. Maudsley Contents: PRANA... 1 1. Introduction... 2 1.1. Examples... 2 2. Subject/Study Selection and Filter...
PRANA Project Review and Analysis March 2010- (Pre-release and still under development!) A.A. Maudsley Contents: PRANA... 1 1. Introduction... 2 1.1. Examples... 2 2. Subject/Study Selection and Filter...
Group Sta*s*cs in MEG/EEG
 Group Sta*s*cs in MEG/EEG Will Woods NIF Fellow Brain and Psychological Sciences Research Centre Swinburne University of Technology A Cau*onary tale. A Cau*onary tale. A Cau*onary tale. Overview Introduc*on
Group Sta*s*cs in MEG/EEG Will Woods NIF Fellow Brain and Psychological Sciences Research Centre Swinburne University of Technology A Cau*onary tale. A Cau*onary tale. A Cau*onary tale. Overview Introduc*on
Statistical Analysis of Neuroimaging Data. Phebe Kemmer BIOS 516 Sept 24, 2015
 Statistical Analysis of Neuroimaging Data Phebe Kemmer BIOS 516 Sept 24, 2015 Review from last time Structural Imaging modalities MRI, CAT, DTI (diffusion tensor imaging) Functional Imaging modalities
Statistical Analysis of Neuroimaging Data Phebe Kemmer BIOS 516 Sept 24, 2015 Review from last time Structural Imaging modalities MRI, CAT, DTI (diffusion tensor imaging) Functional Imaging modalities
We can see that some anatomical details are lost after aligning and averaging brains, especially on the cortical level.
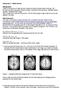 Homework 3 - Model answer Background: Listen to the lecture on Data formats (Voxel and affine transformation matrices, nifti formats) and group analysis (Group Analysis, Anatomical Normalization, Multiple
Homework 3 - Model answer Background: Listen to the lecture on Data formats (Voxel and affine transformation matrices, nifti formats) and group analysis (Group Analysis, Anatomical Normalization, Multiple
EMSegment Tutorial. How to Define and Fine-Tune Automatic Brain Compartment Segmentation and the Detection of White Matter Hyperintensities
 EMSegment Tutorial How to Define and Fine-Tune Automatic Brain Compartment Segmentation and the Detection of White Matter Hyperintensities This documentation serves as a tutorial to learn to customize
EMSegment Tutorial How to Define and Fine-Tune Automatic Brain Compartment Segmentation and the Detection of White Matter Hyperintensities This documentation serves as a tutorial to learn to customize
Machine Learning Practical NITP Summer Course Pamela K. Douglas UCLA Semel Institute
 Machine Learning Practical NITP Summer Course 2013 Pamela K. Douglas UCLA Semel Institute Email: pamelita@g.ucla.edu Topics Covered Part I: WEKA Basics J Part II: MONK Data Set & Feature Selection (from
Machine Learning Practical NITP Summer Course 2013 Pamela K. Douglas UCLA Semel Institute Email: pamelita@g.ucla.edu Topics Covered Part I: WEKA Basics J Part II: MONK Data Set & Feature Selection (from
Medical Image Analysis
 Medical Image Analysis Instructor: Moo K. Chung mchung@stat.wisc.edu Lecture 10. Multiple Comparisons March 06, 2007 This lecture will show you how to construct P-value maps fmri Multiple Comparisons 4-Dimensional
Medical Image Analysis Instructor: Moo K. Chung mchung@stat.wisc.edu Lecture 10. Multiple Comparisons March 06, 2007 This lecture will show you how to construct P-value maps fmri Multiple Comparisons 4-Dimensional
Basic fmri Design and Analysis. Preprocessing
 Basic fmri Design and Analysis Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial filtering
Basic fmri Design and Analysis Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial filtering
Multi-voxel pattern analysis: Decoding Mental States from fmri Activity Patterns
 Multi-voxel pattern analysis: Decoding Mental States from fmri Activity Patterns Artwork by Leon Zernitsky Jesse Rissman NITP Summer Program 2012 Part 1 of 2 Goals of Multi-voxel Pattern Analysis Decoding
Multi-voxel pattern analysis: Decoding Mental States from fmri Activity Patterns Artwork by Leon Zernitsky Jesse Rissman NITP Summer Program 2012 Part 1 of 2 Goals of Multi-voxel Pattern Analysis Decoding
FSL Workshop Session 3 David Smith & John Clithero
 FSL Workshop 12.09.08 Session 3 David Smith & John Clithero What is MELODIC? Probabilistic ICA Improves upon standard ICA Allows for inference Avoids over-fitting Three stage process ( ppca ) 1.) Dimension
FSL Workshop 12.09.08 Session 3 David Smith & John Clithero What is MELODIC? Probabilistic ICA Improves upon standard ICA Allows for inference Avoids over-fitting Three stage process ( ppca ) 1.) Dimension
fmri pre-processing Juergen Dukart
 fmri pre-processing Juergen Dukart Outline Why do we need pre-processing? fmri pre-processing Slice time correction Realignment Unwarping Coregistration Spatial normalisation Smoothing Overview fmri time-series
fmri pre-processing Juergen Dukart Outline Why do we need pre-processing? fmri pre-processing Slice time correction Realignment Unwarping Coregistration Spatial normalisation Smoothing Overview fmri time-series
I.e. Sex differences in child appetitive traits and Eating in the Absence of Hunger:
 Supplementary Materials I. Evidence of sex differences on eating behavior in children I.e. Sex differences in child appetitive traits and Eating in the Absence of Hunger: Table 2. Parent Report for Child
Supplementary Materials I. Evidence of sex differences on eating behavior in children I.e. Sex differences in child appetitive traits and Eating in the Absence of Hunger: Table 2. Parent Report for Child
BrainMask. Quick Start
 BrainMask Quick Start Segmentation of the brain from three-dimensional MR images is a crucial preprocessing step in morphological and volumetric brain studies. BrainMask software implements a fully automatic
BrainMask Quick Start Segmentation of the brain from three-dimensional MR images is a crucial preprocessing step in morphological and volumetric brain studies. BrainMask software implements a fully automatic
Brain Images. Download the data. Comparing Images. 13 th February Version 1.4. For GIMIAS-1.4. Contact:
 Brain Images Comparing Images 13 th February 2012 Version 1.4 For GIMIAS-1.4 Contact: cistib.developers@gmail.com The aim of this exercise is to show different visualization and fusion functionalities
Brain Images Comparing Images 13 th February 2012 Version 1.4 For GIMIAS-1.4 Contact: cistib.developers@gmail.com The aim of this exercise is to show different visualization and fusion functionalities
CS 229 Final Project Report Learning to Decode Cognitive States of Rat using Functional Magnetic Resonance Imaging Time Series
 CS 229 Final Project Report Learning to Decode Cognitive States of Rat using Functional Magnetic Resonance Imaging Time Series Jingyuan Chen //Department of Electrical Engineering, cjy2010@stanford.edu//
CS 229 Final Project Report Learning to Decode Cognitive States of Rat using Functional Magnetic Resonance Imaging Time Series Jingyuan Chen //Department of Electrical Engineering, cjy2010@stanford.edu//
BrainMask. Quick Start
 BrainMask Quick Start Segmentation of the brain from three-dimensional MR images is a crucial pre-processing step in morphological and volumetric brain studies. BrainMask software implements a fully automatic
BrainMask Quick Start Segmentation of the brain from three-dimensional MR images is a crucial pre-processing step in morphological and volumetric brain studies. BrainMask software implements a fully automatic
Crop Counting and Metrics Tutorial
 Crop Counting and Metrics Tutorial The ENVI Crop Science platform contains remote sensing analytic tools for precision agriculture and agronomy. In this tutorial you will go through a typical workflow
Crop Counting and Metrics Tutorial The ENVI Crop Science platform contains remote sensing analytic tools for precision agriculture and agronomy. In this tutorial you will go through a typical workflow
Event-related design efficiency and How to plot fmri time series. Sepideh Sadaghiani NeuroSpin Methods Meeting 15. Sept. 2008
 Event-related design efficiency and How to plot fmri time series Sepideh Sadaghiani NeuroSpin Methods Meeting 15. Sept. 2008 Event-related averaging or FIR? event-related fmri ability to average responses
Event-related design efficiency and How to plot fmri time series Sepideh Sadaghiani NeuroSpin Methods Meeting 15. Sept. 2008 Event-related averaging or FIR? event-related fmri ability to average responses
Caret5 Tutorial Segmentation, Flattening, and Registration
 Caret5 Tutorial Segmentation, Flattening, and Registration John Harwell, Donna Dierker, Heather A. Drury, and David C. Van Essen Version 5.5 September 2006 Copyright 1995-2006 Washington University Washington
Caret5 Tutorial Segmentation, Flattening, and Registration John Harwell, Donna Dierker, Heather A. Drury, and David C. Van Essen Version 5.5 September 2006 Copyright 1995-2006 Washington University Washington
Playing with data from lab
 Playing with data from lab Getting data off the scanner From the Patient Browser, select the folder for the study you want (or within that study, the set of images you want), and then from the Transfer
Playing with data from lab Getting data off the scanner From the Patient Browser, select the folder for the study you want (or within that study, the set of images you want), and then from the Transfer
Supplementary Information. Anne Maass, Hartmut Schütze, Oliver Speck, Andrew Yonelinas, Claus Tempelmann,
 Supplementary Information Laminar activity in the hippocampus and entorhinal cortex related to novelty and episodic encoding Anne Maass, Hartmut Schütze, Oliver Speck, Andrew Yonelinas, Claus Tempelmann,
Supplementary Information Laminar activity in the hippocampus and entorhinal cortex related to novelty and episodic encoding Anne Maass, Hartmut Schütze, Oliver Speck, Andrew Yonelinas, Claus Tempelmann,
Fmri Spatial Processing
 Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
arxiv: v1 [stat.ap] 1 Jun 2016
![arxiv: v1 [stat.ap] 1 Jun 2016 arxiv: v1 [stat.ap] 1 Jun 2016](/thumbs/81/83094240.jpg) Permutation-based cluster size correction for voxel-based lesion-symptom mapping arxiv:1606.00475v1 [stat.ap] 1 Jun 2016 June 3, 2016 Daniel Mirman a,b,1 Jon-Frederick Landrigan a Spiro Kokolis a Sean
Permutation-based cluster size correction for voxel-based lesion-symptom mapping arxiv:1606.00475v1 [stat.ap] 1 Jun 2016 June 3, 2016 Daniel Mirman a,b,1 Jon-Frederick Landrigan a Spiro Kokolis a Sean
Power analysis. Wednesday, Lecture 3 Jeanette Mumford University of Wisconsin - Madison
 Power analysis Wednesday, Lecture 3 Jeanette Mumford University of Wisconsin - Madison Power Analysis-Why? To answer the question. How many subjects do I need for my study? How many runs per subject should
Power analysis Wednesday, Lecture 3 Jeanette Mumford University of Wisconsin - Madison Power Analysis-Why? To answer the question. How many subjects do I need for my study? How many runs per subject should
DynaConn Users Guide. Dynamic Function Connectivity Graphical User Interface. Last saved by John Esquivel Last update: 3/27/14 4:26 PM
 DynaConn Users Guide Dynamic Function Connectivity Graphical User Interface Last saved by John Esquivel Last update: 3/27/14 4:26 PM Table Of Contents Chapter 1 Introduction... 1 1.1 Introduction to this
DynaConn Users Guide Dynamic Function Connectivity Graphical User Interface Last saved by John Esquivel Last update: 3/27/14 4:26 PM Table Of Contents Chapter 1 Introduction... 1 1.1 Introduction to this
Group Level Imputation of Statistic Maps, Version 1.0. User Manual by Kenny Vaden, Ph.D. musc. edu)
 Updated: March 29, 2012 1 Group Level Imputation of Statistic Maps, Version 1.0 User Manual by Kenny Vaden, Ph.D. (vaden @ musc. edu) Notice for use in Academic Work: If our toolkit is used in an academic
Updated: March 29, 2012 1 Group Level Imputation of Statistic Maps, Version 1.0 User Manual by Kenny Vaden, Ph.D. (vaden @ musc. edu) Notice for use in Academic Work: If our toolkit is used in an academic
Zurich SPM Course Voxel-Based Morphometry. Ged Ridgway (Oxford & UCL) With thanks to John Ashburner and the FIL Methods Group
 Zurich SPM Course 2015 Voxel-Based Morphometry Ged Ridgway (Oxford & UCL) With thanks to John Ashburner and the FIL Methods Group Examples applications of VBM Many scientifically or clinically interesting
Zurich SPM Course 2015 Voxel-Based Morphometry Ged Ridgway (Oxford & UCL) With thanks to John Ashburner and the FIL Methods Group Examples applications of VBM Many scientifically or clinically interesting
MR IMAGE SEGMENTATION
 MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
Surface-based Analysis: Inter-subject Registration and Smoothing
 Surface-based Analysis: Inter-subject Registration and Smoothing Outline Exploratory Spatial Analysis Coordinate Systems 3D (Volumetric) 2D (Surface-based) Inter-subject registration Volume-based Surface-based
Surface-based Analysis: Inter-subject Registration and Smoothing Outline Exploratory Spatial Analysis Coordinate Systems 3D (Volumetric) 2D (Surface-based) Inter-subject registration Volume-based Surface-based
Pattern Recognition for Neuroimaging Toolbox: PRoNTo
 Click to edit Master title style Pattern Recognition for Neuroimaging Toolbox: PRoNTo Jessica Schrouff PRNI 2018 June 14 th NUS, Singapore Click Outline to edit Master title style PRoNTo s goals and history
Click to edit Master title style Pattern Recognition for Neuroimaging Toolbox: PRoNTo Jessica Schrouff PRNI 2018 June 14 th NUS, Singapore Click Outline to edit Master title style PRoNTo s goals and history
Artifact detection and repair in fmri
 Artifact detection and repair in fmri Paul K. Mazaika, Ph.D. Center for Interdisciplinary Brain Sciences Research (CIBSR) Division of Interdisciplinary Behavioral Sciences Stanford University School of
Artifact detection and repair in fmri Paul K. Mazaika, Ph.D. Center for Interdisciplinary Brain Sciences Research (CIBSR) Division of Interdisciplinary Behavioral Sciences Stanford University School of
SIVIC Scripting Tutorial
 SIVIC Scripting Tutorial HMTRC Workshop - March 23-24, 2017 Department of Radiology and Biomedical Imaging, UCSF Supported by NIBIB P41EB013598 Goal: The purpose of this tutorial is to introduce you to
SIVIC Scripting Tutorial HMTRC Workshop - March 23-24, 2017 Department of Radiology and Biomedical Imaging, UCSF Supported by NIBIB P41EB013598 Goal: The purpose of this tutorial is to introduce you to
An independent component analysis based tool for exploring functional connections in the brain
 An independent component analysis based tool for exploring functional connections in the brain S. M. Rolfe a, L. Finney b, R. F. Tungaraza b, J. Guan b, L.G. Shapiro b, J. F. Brinkely b, A. Poliakov c,
An independent component analysis based tool for exploring functional connections in the brain S. M. Rolfe a, L. Finney b, R. F. Tungaraza b, J. Guan b, L.G. Shapiro b, J. F. Brinkely b, A. Poliakov c,
EPI Data Are Acquired Serially. EPI Data Are Acquired Serially 10/23/2011. Functional Connectivity Preprocessing. fmri Preprocessing
 Functional Connectivity Preprocessing Geometric distortion Head motion Geometric distortion Head motion EPI Data Are Acquired Serially EPI Data Are Acquired Serially descending 1 EPI Data Are Acquired
Functional Connectivity Preprocessing Geometric distortion Head motion Geometric distortion Head motion EPI Data Are Acquired Serially EPI Data Are Acquired Serially descending 1 EPI Data Are Acquired
A giant leap for particle size analysis
 6.7 A giant leap for particle size analysis With the new and improved detection and splitting options SPIP version 6.7 has taken a giant leap towards easy and robust particle analysis in SPM and SEM images.
6.7 A giant leap for particle size analysis With the new and improved detection and splitting options SPIP version 6.7 has taken a giant leap towards easy and robust particle analysis in SPM and SEM images.
Supplementary Information. Task-induced brain state manipulation improves prediction of individual traits. Greene et al.
 Supplementary Information Task-induced brain state manipulation improves prediction of individual traits Greene et al. Supplementary Note 1 Analyses of effects of gf measurement technique PNC CPM results
Supplementary Information Task-induced brain state manipulation improves prediction of individual traits Greene et al. Supplementary Note 1 Analyses of effects of gf measurement technique PNC CPM results
fmri Basics: Spatial Pre-processing Workshop
 fmri Basics: Spatial Pre-processing Workshop Starting a VNC session: Most of your fmri analysis will be done on the central Linux machines accessed via a VNC (Virtual Network Computing) server. This is
fmri Basics: Spatial Pre-processing Workshop Starting a VNC session: Most of your fmri analysis will be done on the central Linux machines accessed via a VNC (Virtual Network Computing) server. This is
9/1/09 1 DOCUMENTATION FOR. Multi-Level Kernel Density Analysis (MKDA): TOR WAGER S META-ANALYTIC PROCEDURES FOR NEUROIMAGING FINDINGS
 9/1/09 1 DOCUMENTATION FOR Multi-Level Kernel Density Analysis (MKDA): TOR WAGER S META-ANALYTIC PROCEDURES FOR NEUROIMAGING FINDINGS Documentation by: Lisa Feldman Barrett Boston College Kristen Lindquist
9/1/09 1 DOCUMENTATION FOR Multi-Level Kernel Density Analysis (MKDA): TOR WAGER S META-ANALYTIC PROCEDURES FOR NEUROIMAGING FINDINGS Documentation by: Lisa Feldman Barrett Boston College Kristen Lindquist
SIVIC Scripting Tutorial
 SIVIC Scripting Tutorial HMTRC Workshop - March 23-24, 2017 Department of Radiology and Biomedical Imaging, UCSF Supported by NIBIB P41EB013598 Goal: The purpose of this tutorial is to introduce you to
SIVIC Scripting Tutorial HMTRC Workshop - March 23-24, 2017 Department of Radiology and Biomedical Imaging, UCSF Supported by NIBIB P41EB013598 Goal: The purpose of this tutorial is to introduce you to
Files Used in this Tutorial
 Generate Point Clouds and DSM Tutorial This tutorial shows how to generate point clouds and a digital surface model (DSM) from IKONOS satellite stereo imagery. You will view the resulting point clouds
Generate Point Clouds and DSM Tutorial This tutorial shows how to generate point clouds and a digital surface model (DSM) from IKONOS satellite stereo imagery. You will view the resulting point clouds
Structural MRI of Amygdala Tutorial: Observation, Segmentation, Quantification
 Structural MRI of Amygdala Tutorial: Observation, Segmentation, Quantification The FMRIB Software Library (FSL) is a powerful tool that allows users to observe the human brain in various planes and dimensions,
Structural MRI of Amygdala Tutorial: Observation, Segmentation, Quantification The FMRIB Software Library (FSL) is a powerful tool that allows users to observe the human brain in various planes and dimensions,
HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Voxel-Based Morphometry & DARTEL. Ged Ridgway, London With thanks to John Ashburner and the FIL Methods Group
 Zurich SPM Course 2012 Voxel-Based Morphometry & DARTEL Ged Ridgway, London With thanks to John Ashburner and the FIL Methods Group Aims of computational neuroanatomy * Many interesting and clinically
Zurich SPM Course 2012 Voxel-Based Morphometry & DARTEL Ged Ridgway, London With thanks to John Ashburner and the FIL Methods Group Aims of computational neuroanatomy * Many interesting and clinically
Multiple Testing and Thresholding
 Multiple Testing and Thresholding NITP, 2010 Thanks for the slides Tom Nichols! Overview Multiple Testing Problem Which of my 100,000 voxels are active? Two methods for controlling false positives Familywise
Multiple Testing and Thresholding NITP, 2010 Thanks for the slides Tom Nichols! Overview Multiple Testing Problem Which of my 100,000 voxels are active? Two methods for controlling false positives Familywise
