Table of Contents. IntroLab < SPMLabs < Dynevor TWiki
|
|
|
- Barnard King
- 5 years ago
- Views:
Transcription
1 Table of Contents Lab 1: Introduction to SPM and data checking...1 Goals of this Lab...1 Prerequisites...1 An SPM Installation...1 SPM Defaults...2 L/R Brain Orientation...2 Memory Use for Data Processing...2 fmri and PET Analysis Threshold...2 Temporal Intervals for fmri Data...2 Start SPM...3 The SPM interface...3 Background on the data...7 Download the data:...7 Look at the data!...8 Start Matlab & SPM...8 Look at some functional data...8 Find the Susceptibility Voids...8 Look for time series artefacts...8 Review the standard deviation image...9 Compare the functional data to the anatomical data...9 ImCalc...9 Make a Binary Mask...9 Make a Mean Image...10.mat files...10 i
2 Lab 1: Introduction to SPM and data checking Goals of this Lab After this lab you will Understand the structure of the SPM package and how to best install and maintain it. 2. Be able to examine data using SPM's single and multi volume display facilities. 3. Be able to characterize the susceptibility artifacts and signal voids in functional data, as compared to similar structural data. Prerequisites A standard SPM installation The SPM single subject epoch (block) auditory fmri activation data (see below) An SPM Installation Before starting SPM, use the Windows Explorer go to the SPM directory (listed on the board). Note that SPM consists mostly of matlab.m files. There are also.c,.h C source code files, and compiled 'mex' or 'dll' files. The compiled files are custom C language extensions which can be called as if they are matlab.m files, and are used to speed up processing that would be very slow in matlab. If you install SPM yourself, you can obtain it from but be sure to obtain the latest updates (check regularly!). For SPM2, the updates are here ftp://ftp.fil.ion.ucl.ac.uk/spm/spm2_updates When you install SPM yourself, it is a good idea to create a "root" spm directory. In this root SPM directory you can put the released versions of SPM, and in a separate local folder, any local modifications. For example, our local spm directories look like /usr/local/spm /usr/local/spm/spm99 /usr/local/spm/spm99_local /usr/local/spm/spm2 /usr/local/spm/spm2_local or for Windows C:\MatlabToolboxes\spm2 C:\MatlabToolboxes\spm2_local Never change any files in a SPM release directory (e.g. spm/spm99 or spm/spm2 above), except to incorporate updates from SPM central, even then it is a good idea to backup the old copy of the file being replaced until you are sure the new file has been installed correctly and is working. Lab 1: Introduction to SPM and data checking 1
3 If you want to change some SPM code, make a copy of the.m file you want to change and put it in the local directory (e.g. "cp spm2/spm_defaults.m spm2_local/spm_defaults.m" on Unix), and then make changes to that local copy. If you don't do this, your local changes may be overwritten when you download new updates. SPM Defaults When setting up SPM for the first time it is a good idea to make sure the settings for various default variables are correct for your installation. Default settings are specified in the file spm_defaults.m. This file can be changed in the main spm2 directory where it will affect all users, or individual users can have their own copy with user specific settings. If you keep your own copy make sure the Matlab path to the copy comes BEFORE the path to the spm2 directory. L/R Brain Orientation Open the spm_defaults file by typing edit spm_defaults at the Matlab prompt. Look at the entry for defaults.analyze.flip. Does this installation of SPM expect the images to be in Neurologic (L = L) or Radiologic (L = R) convention? How can you tell? Display image sm00223_002.img. Assuming this image was made in SPM2 is there any way to tell its orientation? Now look at image sm00223_002 1.img? Is this in the same or different orientation? How would we want defaults.analyze.flip set for each of these images? Memory Use for Data Processing How can we tell spm to process the data in bigger or smaller chunks? fmri and PET Analysis Threshold Do the defaults tell spm to apply any type of threshold to the data? If so what is the threshold for fmri data? for PET data? Temporal Intervals for fmri Data IntroLab < SPMLabs < Dynevor TWiki When setting up designs for fmri, SPM divides the interscan interval into multiple intervals for finer modeling of the impulse response. How many intervals is the current interscan interval divided into? (hint: look at defaults.stats.fmri.t) What interval does SPM use as the starting time point for an event or epoch? (hint: look at defaults.stats.fmri.t0) How would the starting interval be changed to midway in the TR? Would it matter if the TR was 2 seconds or 4 seconds? SPM Defaults 2
4 Start SPM IntroLab < SPMLabs < Dynevor TWiki Open Matlab. Check that spm is on the Matlab Path by typing spm If an error occurs then you must add the spm path to the Matlab. There are two ways to set the path. You can go under the file menu and use the 'Set path...' command. Click the 'Add Folder...' button to select the spm2 folder. Click 'Close'. (Click save if you want to add the spm2 folder permanently to the Matlab path). Or you can use the command "addpath" with the path given on the board to add the SPM path to the Matlab path. addpath c:\course\spm\spm2 Here is an example of a simple m file that does this for you. You could save the following 2 lines in a file called spm2.m, and put the file in Matlab's local folder. Typing spm2 would start the fmri section of spm. % Save and name spm2.m addpath('c:\path_to_my_spm2_folder\spm2') spm('fmri') The SPM interface Now that you have run the 'spm' command, you should get a window like this: Click on the FMRI button. You now get the standard SPM interface. In the top left of the screen you have the SPM buttons window: Start SPM 3
5 Below that you have the SPM input (or Interactive) window: Start SPM 4
6 The large window on the right of the screen is the SPM graphics window. In the example below we are displaying some results: Start SPM 5
7 Start SPM 6
8 When SPM requires that you select some files, you will get the SPM file get window: Background on the data In this lab you will look at single subject fmri data from a block auditory activation experiment. 96 acquisitions were made (RT=7s), in blocks of 6, giving 16 42s blocks. The condition for successive blocks alternated between rest and auditory stimulation, starting with rest. Auditory stimulation was with bi syllabic words presented binaurally at a rate of 60 per minute. The functional data starts at acquisition 4, image fm00223_004. Due to T1 effects it is advisable to discard the first few scans (there were no "dummy" lead in scans). These whole brain BOLD/EPI images were acquired on a modified 2T Siemens MAGNETOM Vision system. Each acquisition consisted of 64 contiguous slices (64x64x64 3mm x 3mm x 3mm voxels). Acquisition took 6.05s, with the scan to scan repeat time (RT) set arbitrarily to 7s. Download the data: The data may already on your PC. If not, copy the MoEApilot data to your hard drive. Look for instructions on the board for where to put the data. Background on the data 7
9 Look at the data! IntroLab < SPMLabs < Dynevor TWiki Start Matlab & SPM The first thing you should after starting Matlab is to change the working directory to a reasonable place. In the compute lab, the working directory may default to a directory where you do not have write permissions. Use the '...' button at the top of the Matlab window, or use the 'cd' command to change to the 'c:\temp\spmcourse' directory. Look at some functional data Using the 'Display' button view a randomly selected image from among the functional data, in the fm00223 directory. You should always do this to check orientation (and any possible catastrophic problems.) 1. What is the voxel size? 2. What are the image dimension? 3. What are typical gray matter values? From to 4. Bilinear interpolation is default. Select Nearest Neighbor interpolation. Explore the image 5. Select Sinc interpolation. Explore the image. 6. Which one do you prefer? Why? Find the Susceptibility Voids Due to the differences in magnetic susceptibility of air and tissue, there are often distortions and signal voices near air tissue interfaces. Find some of these signal voids in the function data. 1. Use 'Display' to view a single functional image. 2. Position the cursor middle of the brain superior inferior, so that the axial image shows a full oval brain slice. Now slowly move down (inferiorly) while watching the axial slice. Watch the for a darkening of the medial orbital frontal region. Clicking further down, find the signal voids above the ear canals in the temporal lobes. Look for time series artefacts Almost all FMRI time series have significant artefacts due to the scanner or subject movement. It is important to check for these before we run the analysis. To do this we will use the tsdiffana utility from the CBU SPM tools: cbu.cam.ac.uk/imaging/common/downloads/spmutils/tsdiffana.tar.gz Download this archive to a temporary directory say c:\temp. Double click to open the archive, and extract into the temporary directory. You should now have some new.m files in this directory including a file tsdiffana.m. Add this directory to your path, for example with: addpath c:\temp end The " end" flag just makes sure that c:\temp goes at the end of the matlab path, so any files in the c:\temp directory can't interfere with SPM files. Look at the data! 8
10 To run the utility, go to the matlab (>>) prompt, and enter tsdiffana Then select all the raw functional images from the dataset (fm00223_*.img). You will see the time series diagnostics in the SPM graphics window. In the top panel you will see the scan to scan variance. For the first point, this the mean squared difference (MSD) in signal intensity between the first and second scan; for the second point it is the MSD for scans 2 and 3, and so on. This tells you where there has been global shift in signal between scans. What do you notice about the scan to scan variance time course? There are more details on the tsdiffana utility on the CBU web site: cbu.cam.ac.uk/imaging/common/diagnostics.shtml Review the standard deviation image The tsdiffana utility also writes out some standard deviation images for the whole time series. These images allow you to see where there may have been artefacts that are larger in particular parts of the image. 1. Click on 'Display' 2. Select the 'std_*.img' in the FMRI data directory Click around the image. What artefacts do you see? What should we do about these artefacts? Compare the functional data to the anatomical data In the sm00223 directory you'll find the structual MRI data. 1. Use 'Check Reg' to "check" the "registration" between the sm00223_002 image and one of the fm00223 functionals. 1. Unlike 'Display', 'Check Reg' lets you view upto 12 volumes. Select the anatomical and then the functional. 1. We haven't registered these two images yet; is this clear from CheckReg? ImCalc The ImCalc function of spm can be used to manipulate images in a variety of ways. In this section you will learn to make a binary image mask and to average images. Make a Binary Mask IntroLab < SPMLabs < Dynevor TWiki 1. Use the spm cd menu to move to a directory you can write to. 2. Click the imcalc button and choose the sm00223_002.img. Review the standard deviation image 9
11 3. Enter a filename for the output image. You do not need to include the.img extension. 4. Enter i1 > 0 as the expression to evaluate. 5. Look at the image output using Display. What went wrong? Make a new binary image that fixes this problem. 6. Advanced users: How would you apply the mask to the original image in a single step? Make a Mean Image 1. Click the imcalc button and select the first three EPI images (fm00223_004.img through fm00223_006.img) 2. Given the mean output image a filename say mean_4_epi (note you do not need the.img extension). 3. Enter the expression (i1+i2+i2)/3. 4. Check this worked by clicking Display, and selecting the output image (mean_4_epi.img in your current working directory). If you can't remember where your current directory is, type pwd in the matlab command window..mat files IntroLab < SPMLabs < Dynevor TWiki.mat files encode orientation information for images in SPM, and are a major source of confusion. 1. Go to the matlab console window, and type ls mean_4_epi* 2. You should see that there are two files, mean_4_epi.img, and mean_4_epi.hdr. The img file contains the actual image data, and the hdr file contains various bits of information about the image including dimensions etc. 3. If you haven't already done this for the previous exercise, click Display, and select the mean_4_epi.img file. 4. Note the Dir Cos matrix at the bottom right of the graphics window; it should be a 3x3 identity matrix (just zeros with ones on the diagonal). 5. Go to the transformation parameters box at the bottom left of the graphics window (see below for image) 6. Click in the Pitch box, and set the pitch to 0.6, and press return. Notice that the image pitches forward (in fact by 0.6 radians). 7. Click on the Reorient images... button in the bottom left hand corner. Select mean_4_epi.img (the image you are currently working with). The 0.6 pitch value has now been cleared, and you will see the Dir Cos matrix has changed reflecting the pitch transformation applied to the image 8. Click on the World space drop down menu box near the bottom mid right of the graphics window (sse below for image). 9. Try toggling between World space and Voxel space. Note that the transformation is not applied in voxel space, but it is in world space. 10. Click on Display and select mean_4_epi.img again. The transformation you saved is still being applied. 11. At the matlab console, type ls mean_4_epi* again. You will see there is a new file mean_4_epi.mat. THis file contains the reorientation that you applied to the image. 12. At the matlab console type delete mean_4_epi.mat. 13. Click on Display and select mean_4_epi.img again. The transformation has been lost. Make a Mean Image 10
12 MatthewBrett 22 Mar 2005 DarrenGitelman 22 Mar 2005 I Attachment Action Size Date Who Comment 22 Mar 2005 spm_buttons.png manage 19.2 K 23:39 MatthewBrett SPM buttons window 22 Mar 2005 spm_input.png manage 14.6 K 23:40 MatthewBrett SPM input window 22 Mar 2005 spm_graphics.png manage 44.8 K 23:40 MatthewBrett SPM graphics window Make a Mean Image 11
13 22 Mar 2005 spm_get.png manage 16.2 K 23:41 22 Mar 2005 spm_welcome.png manage 17.2 K 23:46 29 Mar 2005 orth_trans_params.png manage 4.0 K 19:14 29 Mar 2005 orth_info_box.png manage 5.3 K 19:22 MatthewBrett SPM get window MatthewBrett SPM welcome window MatthewBrett MatthewBrett Transformation parameters input box Information box for SPM image display Make a Mean Image 12
SPM99 fmri Data Analysis Workbook
 SPM99 fmri Data Analysis Workbook This file is a description of the steps needed to use SPM99 analyze a fmri data set from a single subject using a simple on/off activation paradigm. There are two parts
SPM99 fmri Data Analysis Workbook This file is a description of the steps needed to use SPM99 analyze a fmri data set from a single subject using a simple on/off activation paradigm. There are two parts
GLM for fmri data analysis Lab Exercise 1
 GLM for fmri data analysis Lab Exercise 1 March 15, 2013 Medical Image Processing Lab Medical Image Processing Lab GLM for fmri data analysis Outline 1 Getting Started 2 AUDIO 1 st level Preprocessing
GLM for fmri data analysis Lab Exercise 1 March 15, 2013 Medical Image Processing Lab Medical Image Processing Lab GLM for fmri data analysis Outline 1 Getting Started 2 AUDIO 1 st level Preprocessing
Basic fmri Design and Analysis. Preprocessing
 Basic fmri Design and Analysis Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial filtering
Basic fmri Design and Analysis Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial filtering
White Pixel Artifact. Caused by a noise spike during acquisition Spike in K-space <--> sinusoid in image space
 White Pixel Artifact Caused by a noise spike during acquisition Spike in K-space sinusoid in image space Susceptibility Artifacts Off-resonance artifacts caused by adjacent regions with different
White Pixel Artifact Caused by a noise spike during acquisition Spike in K-space sinusoid in image space Susceptibility Artifacts Off-resonance artifacts caused by adjacent regions with different
SPM Introduction. SPM : Overview. SPM: Preprocessing SPM! SPM: Preprocessing. Scott Peltier. FMRI Laboratory University of Michigan
 SPM Introduction Scott Peltier FMRI Laboratory University of Michigan! Slides adapted from T. Nichols SPM! SPM : Overview Library of MATLAB and C functions Graphical user interface Four main components:
SPM Introduction Scott Peltier FMRI Laboratory University of Michigan! Slides adapted from T. Nichols SPM! SPM : Overview Library of MATLAB and C functions Graphical user interface Four main components:
ASAP_2.0 (Automatic Software for ASL Processing) USER S MANUAL
 ASAP_2.0 (Automatic Software for ASL Processing) USER S MANUAL ASAP was developed as part of the COST Action "Arterial Spin Labelling Initiative in Dementia (AID)" by: Department of Neuroimaging, Institute
ASAP_2.0 (Automatic Software for ASL Processing) USER S MANUAL ASAP was developed as part of the COST Action "Arterial Spin Labelling Initiative in Dementia (AID)" by: Department of Neuroimaging, Institute
SPM Introduction SPM! Scott Peltier. FMRI Laboratory University of Michigan. Software to perform computation, manipulation and display of imaging data
 SPM Introduction Scott Peltier FMRI Laboratory University of Michigan Slides adapted from T. Nichols SPM! Software to perform computation, manipulation and display of imaging data 1 1 SPM : Overview Library
SPM Introduction Scott Peltier FMRI Laboratory University of Michigan Slides adapted from T. Nichols SPM! Software to perform computation, manipulation and display of imaging data 1 1 SPM : Overview Library
Version. Getting Started: An fmri-cpca Tutorial
 Version 11 Getting Started: An fmri-cpca Tutorial 2 Table of Contents Table of Contents... 2 Introduction... 3 Definition of fmri-cpca Data... 3 Purpose of this tutorial... 3 Prerequisites... 4 Used Terms
Version 11 Getting Started: An fmri-cpca Tutorial 2 Table of Contents Table of Contents... 2 Introduction... 3 Definition of fmri-cpca Data... 3 Purpose of this tutorial... 3 Prerequisites... 4 Used Terms
HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Group (Level 2) fmri Data Analysis - Lab 4
 Group (Level 2) fmri Data Analysis - Lab 4 Index Goals of this Lab Before Getting Started The Chosen Ten Checking Data Quality Create a Mean Anatomical of the Group Group Analysis: One-Sample T-Test Examine
Group (Level 2) fmri Data Analysis - Lab 4 Index Goals of this Lab Before Getting Started The Chosen Ten Checking Data Quality Create a Mean Anatomical of the Group Group Analysis: One-Sample T-Test Examine
Introduction to fmri. Pre-processing
 Introduction to fmri Pre-processing Tibor Auer Department of Psychology Research Fellow in MRI Data Types Anatomical data: T 1 -weighted, 3D, 1/subject or session - (ME)MPRAGE/FLASH sequence, undistorted
Introduction to fmri Pre-processing Tibor Auer Department of Psychology Research Fellow in MRI Data Types Anatomical data: T 1 -weighted, 3D, 1/subject or session - (ME)MPRAGE/FLASH sequence, undistorted
Analysis of fmri data within Brainvisa Example with the Saccades database
 Analysis of fmri data within Brainvisa Example with the Saccades database 18/11/2009 Note : All the sentences in italic correspond to informations relative to the specific dataset under study TP participants
Analysis of fmri data within Brainvisa Example with the Saccades database 18/11/2009 Note : All the sentences in italic correspond to informations relative to the specific dataset under study TP participants
SPM8 for Basic and Clinical Investigators. Preprocessing. fmri Preprocessing
 SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
This exercise uses one anatomical data set (ANAT1) and two functional data sets (FUNC1 and FUNC2).
 Exploring Brain Anatomy This week s exercises will let you explore the anatomical organization of the brain to learn some of its basic properties, as well as the location of different structures. The human
Exploring Brain Anatomy This week s exercises will let you explore the anatomical organization of the brain to learn some of its basic properties, as well as the location of different structures. The human
This Time. fmri Data analysis
 This Time Reslice example Spatial Normalization Noise in fmri Methods for estimating and correcting for physiologic noise SPM Example Spatial Normalization: Remind ourselves what a typical functional image
This Time Reslice example Spatial Normalization Noise in fmri Methods for estimating and correcting for physiologic noise SPM Example Spatial Normalization: Remind ourselves what a typical functional image
Functional MRI in Clinical Research and Practice Preprocessing
 Functional MRI in Clinical Research and Practice Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization
Functional MRI in Clinical Research and Practice Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization
Last Time. This Time. Thru-plane dephasing: worse at long TE. Local susceptibility gradients: thru-plane dephasing
 Motion Correction Last Time Mutual Information Optimiation Decoupling Translation & Rotation Interpolation SPM Example (Least Squares & MI) A Simple Derivation This Time Reslice example SPM Example : Remind
Motion Correction Last Time Mutual Information Optimiation Decoupling Translation & Rotation Interpolation SPM Example (Least Squares & MI) A Simple Derivation This Time Reslice example SPM Example : Remind
Preprocessing of fmri Data in SPM 12 - Lab 1
 Preprocessing of fmri Data in SPM 12 - Lab 1 Index Goals of this Lab Preprocessing Overview MATLAB, SPM, Data Setup Preprocessing I: Checking Motion Correction Preprocessing II: Coregistration Preprocessing
Preprocessing of fmri Data in SPM 12 - Lab 1 Index Goals of this Lab Preprocessing Overview MATLAB, SPM, Data Setup Preprocessing I: Checking Motion Correction Preprocessing II: Coregistration Preprocessing
EPI Data Are Acquired Serially. EPI Data Are Acquired Serially 10/23/2011. Functional Connectivity Preprocessing. fmri Preprocessing
 Functional Connectivity Preprocessing Geometric distortion Head motion Geometric distortion Head motion EPI Data Are Acquired Serially EPI Data Are Acquired Serially descending 1 EPI Data Are Acquired
Functional Connectivity Preprocessing Geometric distortion Head motion Geometric distortion Head motion EPI Data Are Acquired Serially EPI Data Are Acquired Serially descending 1 EPI Data Are Acquired
fmri Basics: Spatial Pre-processing Workshop
 fmri Basics: Spatial Pre-processing Workshop Starting a VNC session: Most of your fmri analysis will be done on the central Linux machines accessed via a VNC (Virtual Network Computing) server. This is
fmri Basics: Spatial Pre-processing Workshop Starting a VNC session: Most of your fmri analysis will be done on the central Linux machines accessed via a VNC (Virtual Network Computing) server. This is
SPM8 for Basic and Clinical Investigators. Preprocessing
 SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
2. Creating Field Maps Using the Field Map GUI (Version 2.0) in SPM5
 1. Introduction This manual describes how to use the Field Map Toolbox Version 2.0 for creating unwrapped field maps that can be used to do geometric distortion correction of EPI images in SPM5. 1. 1.
1. Introduction This manual describes how to use the Field Map Toolbox Version 2.0 for creating unwrapped field maps that can be used to do geometric distortion correction of EPI images in SPM5. 1. 1.
Playing with data from lab
 Playing with data from lab Getting data off the scanner From the Patient Browser, select the folder for the study you want (or within that study, the set of images you want), and then from the Transfer
Playing with data from lab Getting data off the scanner From the Patient Browser, select the folder for the study you want (or within that study, the set of images you want), and then from the Transfer
HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2006
 MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2006 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2006 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
SPM Course! Single Subject Analysis
 SPM Course! Single Subject Analysis Practical Session Dr. Jakob Heinzle & Dr. Frederike Petzschner & Dr. Lionel Rigoux Hands up: Who has programming experience with Matlab? Who has analyzed an fmri experiment
SPM Course! Single Subject Analysis Practical Session Dr. Jakob Heinzle & Dr. Frederike Petzschner & Dr. Lionel Rigoux Hands up: Who has programming experience with Matlab? Who has analyzed an fmri experiment
0.1. Setting up the system path to allow use of BIAC XML headers (BXH). Depending on the computer(s), you may only have to do this once.
 Week 3 Exercises Last week you began working with MR data, both in the form of anatomical images and functional time series. This week we will discuss some concepts related to the idea of fmri data as
Week 3 Exercises Last week you began working with MR data, both in the form of anatomical images and functional time series. This week we will discuss some concepts related to the idea of fmri data as
fmri Basics: Single Subject Analysis
 fmri Basics: Single Subject Analysis This session is intended to give an overview of the basic process of setting up a general linear model for a single subject. This stage of the analysis is also variously
fmri Basics: Single Subject Analysis This session is intended to give an overview of the basic process of setting up a general linear model for a single subject. This stage of the analysis is also variously
Measuring baseline whole-brain perfusion on GE 3.0T using arterial spin labeling (ASL) MRI
 Measuring baseline whole-brain perfusion on GE 3.0T using arterial spin labeling (ASL) MRI Revision date: 09/15/2008 Overview This document describes the procedure for measuring baseline whole-brain perfusion
Measuring baseline whole-brain perfusion on GE 3.0T using arterial spin labeling (ASL) MRI Revision date: 09/15/2008 Overview This document describes the procedure for measuring baseline whole-brain perfusion
Functional MRI data preprocessing. Cyril Pernet, PhD
 Functional MRI data preprocessing Cyril Pernet, PhD Data have been acquired, what s s next? time No matter the design, multiple volumes (made from multiple slices) have been acquired in time. Before getting
Functional MRI data preprocessing Cyril Pernet, PhD Data have been acquired, what s s next? time No matter the design, multiple volumes (made from multiple slices) have been acquired in time. Before getting
CS/NEUR125 Brains, Minds, and Machines. Due: Wednesday, April 5
 CS/NEUR125 Brains, Minds, and Machines Lab 8: Using fmri to Discover Language Areas in the Brain Due: Wednesday, April 5 In this lab, you will analyze fmri data from an experiment that was designed to
CS/NEUR125 Brains, Minds, and Machines Lab 8: Using fmri to Discover Language Areas in the Brain Due: Wednesday, April 5 In this lab, you will analyze fmri data from an experiment that was designed to
fmri pre-processing Juergen Dukart
 fmri pre-processing Juergen Dukart Outline Why do we need pre-processing? fmri pre-processing Slice time correction Realignment Unwarping Coregistration Spatial normalisation Smoothing Overview fmri time-series
fmri pre-processing Juergen Dukart Outline Why do we need pre-processing? fmri pre-processing Slice time correction Realignment Unwarping Coregistration Spatial normalisation Smoothing Overview fmri time-series
The alignment process uses two alignment methods: (1) the mrvista function, rxalign, for a rough alignment, and then (2) KNK's alignment for
 Alignment (Allegra) Download example alignment script here. Anatomical images are collected at much higher resolution (isotropic 1 mm voxels) than functional images (isotropic 2 mm voxels; sometimes 2.5).
Alignment (Allegra) Download example alignment script here. Anatomical images are collected at much higher resolution (isotropic 1 mm voxels) than functional images (isotropic 2 mm voxels; sometimes 2.5).
Fmri Spatial Processing
 Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
FMRI Pre-Processing and Model- Based Statistics
 FMRI Pre-Processing and Model- Based Statistics Brief intro to FMRI experiments and analysis FMRI pre-stats image processing Simple Single-Subject Statistics Multi-Level FMRI Analysis Advanced FMRI Analysis
FMRI Pre-Processing and Model- Based Statistics Brief intro to FMRI experiments and analysis FMRI pre-stats image processing Simple Single-Subject Statistics Multi-Level FMRI Analysis Advanced FMRI Analysis
FILE MAINTENANCE COMMANDS
 Birla Institute of Technology & Science, Pilani Computer Programming (CS F111) Lab-2 ----------------------------------------------------------------------------------------------------------------------
Birla Institute of Technology & Science, Pilani Computer Programming (CS F111) Lab-2 ----------------------------------------------------------------------------------------------------------------------
fmri/dti analysis using Dynasuite
 fmri/dti analysis using Dynasuite Contents 1 Logging in 2 Finding patient session 3 Viewing and adjusting images 4 Checking brain segmentation 5 Checking image registration 6 Seeing fmri results 7 Saving
fmri/dti analysis using Dynasuite Contents 1 Logging in 2 Finding patient session 3 Viewing and adjusting images 4 Checking brain segmentation 5 Checking image registration 6 Seeing fmri results 7 Saving
syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets
 syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets Julien Gervais; Lisa Chuah Siemens Healthcare, Magnetic
syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets Julien Gervais; Lisa Chuah Siemens Healthcare, Magnetic
CHAPTER 9: Magnetic Susceptibility Effects in High Field MRI
 Figure 1. In the brain, the gray matter has substantially more blood vessels and capillaries than white matter. The magnified image on the right displays the rich vasculature in gray matter forming porous,
Figure 1. In the brain, the gray matter has substantially more blood vessels and capillaries than white matter. The magnified image on the right displays the rich vasculature in gray matter forming porous,
You should see something like this, called the prompt :
 CSE 1030 Lab 1 Basic Use of the Command Line PLEASE NOTE this lab will not be graded and does not count towards your final grade. However, all of these techniques are considered testable in a labtest.
CSE 1030 Lab 1 Basic Use of the Command Line PLEASE NOTE this lab will not be graded and does not count towards your final grade. However, all of these techniques are considered testable in a labtest.
ANALYSIS OF FUNCTIONAL MAGNETIC RESONANCE IMAGING DATA USING SPM99: VOXEL-BASED MORPHOMETRY DONNA ROSE ADDIS
 Donna Rose Addis, TWRI, May 2004 1 ANALYSIS OF FUNCTIONAL MAGNETIC RESONANCE IMAGING DATA USING SPM99: VOXEL-BASED MORPHOMETRY DONNA ROSE ADDIS DEPT. OF PSYCHOLOGY, UNIVERSITY OF TORONTO TORONTO WESTERN
Donna Rose Addis, TWRI, May 2004 1 ANALYSIS OF FUNCTIONAL MAGNETIC RESONANCE IMAGING DATA USING SPM99: VOXEL-BASED MORPHOMETRY DONNA ROSE ADDIS DEPT. OF PSYCHOLOGY, UNIVERSITY OF TORONTO TORONTO WESTERN
All have parameters: Voxel dimensions: 90 * 108 * 78, Voxel size: 1.5mm * 1.5mm * 1.5mm, Data type: float, Orientation: Axial (S=>I).
 Week 5 Exercises This week we will discuss some concepts related to the idea of fmri data as sets of numbers that can be manipulated computationally. We will also introduce the concept of "statistical
Week 5 Exercises This week we will discuss some concepts related to the idea of fmri data as sets of numbers that can be manipulated computationally. We will also introduce the concept of "statistical
Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question.
 (1) conn - Functional connectivity toolbox v1.0 Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question. 2) in "Basic" enter "1" subject, "6" seconds
(1) conn - Functional connectivity toolbox v1.0 Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question. 2) in "Basic" enter "1" subject, "6" seconds
Measuring baseline whole-brain perfusion on GE 3.0T using arterial spin labeling (ASL) MRI
 Measuring baseline whole-brain perfusion on GE 3.0T using arterial spin labeling (ASL) MRI Revision date: 11/20/2006 Overview This document describes the procedure for measuring baseline whole-brain perfusion
Measuring baseline whole-brain perfusion on GE 3.0T using arterial spin labeling (ASL) MRI Revision date: 11/20/2006 Overview This document describes the procedure for measuring baseline whole-brain perfusion
The organization of the human cerebral cortex estimated by intrinsic functional connectivity
 1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
fmri Preprocessing & Noise Modeling
 Translational Neuromodeling Unit fmri Preprocessing & Noise Modeling Lars Kasper September 25 th / October 17 th, 2015 MR-Technology Group & Translational Neuromodeling Unit An SPM Tutorial Institute for
Translational Neuromodeling Unit fmri Preprocessing & Noise Modeling Lars Kasper September 25 th / October 17 th, 2015 MR-Technology Group & Translational Neuromodeling Unit An SPM Tutorial Institute for
Artifact detection and repair in fmri
 Artifact detection and repair in fmri Paul K. Mazaika, Ph.D. Center for Interdisciplinary Brain Sciences Research (CIBSR) Division of Interdisciplinary Behavioral Sciences Stanford University School of
Artifact detection and repair in fmri Paul K. Mazaika, Ph.D. Center for Interdisciplinary Brain Sciences Research (CIBSR) Division of Interdisciplinary Behavioral Sciences Stanford University School of
HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
An independent component analysis based tool for exploring functional connections in the brain
 An independent component analysis based tool for exploring functional connections in the brain S. M. Rolfe a, L. Finney b, R. F. Tungaraza b, J. Guan b, L.G. Shapiro b, J. F. Brinkely b, A. Poliakov c,
An independent component analysis based tool for exploring functional connections in the brain S. M. Rolfe a, L. Finney b, R. F. Tungaraza b, J. Guan b, L.G. Shapiro b, J. F. Brinkely b, A. Poliakov c,
Characterization and Correction of Interpolation Effects in the Realignment of fmri Time Series
 NeuroImage 11, 49 57 (2000) doi:10.1006/nimg.1999.0515, available online at http://www.idealibrary.com on Characterization and Correction of Interpolation Effects in the Realignment of fmri Time Series
NeuroImage 11, 49 57 (2000) doi:10.1006/nimg.1999.0515, available online at http://www.idealibrary.com on Characterization and Correction of Interpolation Effects in the Realignment of fmri Time Series
Functional MRI. Jerry Allison, Ph. D. Medical College of Georgia
 Functional MRI Jerry Allison, Ph. D. Medical College of Georgia BOLD Imaging Technique Blood Oxygen Level Dependent contrast can be used to map brain function Right Hand Motor Task Outline fmri BOLD Contrast
Functional MRI Jerry Allison, Ph. D. Medical College of Georgia BOLD Imaging Technique Blood Oxygen Level Dependent contrast can be used to map brain function Right Hand Motor Task Outline fmri BOLD Contrast
HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Supplementary Figure 1
 Supplementary Figure 1 BOLD and CBV functional maps showing EPI versus line-scanning FLASH fmri. A. Colored BOLD and CBV functional maps are shown in the highlighted window (green frame) of the raw EPI
Supplementary Figure 1 BOLD and CBV functional maps showing EPI versus line-scanning FLASH fmri. A. Colored BOLD and CBV functional maps are shown in the highlighted window (green frame) of the raw EPI
FNC Toolbox Walk Through
 FNC Toolbox Walk Through Date: August 10 th, 2009 By: Nathan Swanson, Vince Calhoun at The Mind Research Network Email: nswanson@mrn.org Introduction The FNC Toolbox is an extension of the GIFT toolbox
FNC Toolbox Walk Through Date: August 10 th, 2009 By: Nathan Swanson, Vince Calhoun at The Mind Research Network Email: nswanson@mrn.org Introduction The FNC Toolbox is an extension of the GIFT toolbox
Basic principles of MR image analysis. Basic principles of MR image analysis. Basic principles of MR image analysis
 Basic principles of MR image analysis Basic principles of MR image analysis Julien Milles Leiden University Medical Center Terminology of fmri Brain extraction Registration Linear registration Non-linear
Basic principles of MR image analysis Basic principles of MR image analysis Julien Milles Leiden University Medical Center Terminology of fmri Brain extraction Registration Linear registration Non-linear
Supplementary methods
 Supplementary methods This section provides additional technical details on the sample, the applied imaging and analysis steps and methods. Structural imaging Trained radiographers placed all participants
Supplementary methods This section provides additional technical details on the sample, the applied imaging and analysis steps and methods. Structural imaging Trained radiographers placed all participants
SnPM is an SPM toolbox developed by Andrew Holmes & Tom Nichols
 1 of 14 3/30/2005 9:24 PM SnPM A Worked fmri Example SnPM is an SPM toolbox developed by Andrew Holmes & Tom Nichols This page... introduction example data background design setup computation viewing results
1 of 14 3/30/2005 9:24 PM SnPM A Worked fmri Example SnPM is an SPM toolbox developed by Andrew Holmes & Tom Nichols This page... introduction example data background design setup computation viewing results
BDP: BrainSuite Diffusion Pipeline. Chitresh Bhushan
 BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
icatvision Quick Reference
 icatvision Quick Reference Navigating the i-cat Interface This guide shows how to: View reconstructed images Use main features and tools to optimize an image. REMINDER Images are displayed as if you are
icatvision Quick Reference Navigating the i-cat Interface This guide shows how to: View reconstructed images Use main features and tools to optimize an image. REMINDER Images are displayed as if you are
Statistical Analysis of Neuroimaging Data. Phebe Kemmer BIOS 516 Sept 24, 2015
 Statistical Analysis of Neuroimaging Data Phebe Kemmer BIOS 516 Sept 24, 2015 Review from last time Structural Imaging modalities MRI, CAT, DTI (diffusion tensor imaging) Functional Imaging modalities
Statistical Analysis of Neuroimaging Data Phebe Kemmer BIOS 516 Sept 24, 2015 Review from last time Structural Imaging modalities MRI, CAT, DTI (diffusion tensor imaging) Functional Imaging modalities
The BERGEN Plug-in for EEGLAB
 The BERGEN Plug-in for EEGLAB July 2009, Version 1.0 What is the Bergen Plug-in for EEGLAB? The Bergen plug-in is a set of Matlab tools developed at the fmri group, University of Bergen, Norway, which
The BERGEN Plug-in for EEGLAB July 2009, Version 1.0 What is the Bergen Plug-in for EEGLAB? The Bergen plug-in is a set of Matlab tools developed at the fmri group, University of Bergen, Norway, which
QIBA PET Amyloid BC March 11, Agenda
 QIBA PET Amyloid BC March 11, 2016 - Agenda 1. QIBA Round 6 Funding a. Deadlines b. What projects can be funded, what cannot c. Discussion of projects Mechanical phantom and DRO Paul & John? Any Profile
QIBA PET Amyloid BC March 11, 2016 - Agenda 1. QIBA Round 6 Funding a. Deadlines b. What projects can be funded, what cannot c. Discussion of projects Mechanical phantom and DRO Paul & John? Any Profile
Function-Structure Integration in FreeSurfer
 Function-Structure Integration in FreeSurfer Outline Function-Structure Integration Function-Structure Registration in FreeSurfer fmri Analysis Preprocessing First-Level Analysis Higher-Level (Group) Analysis
Function-Structure Integration in FreeSurfer Outline Function-Structure Integration Function-Structure Registration in FreeSurfer fmri Analysis Preprocessing First-Level Analysis Higher-Level (Group) Analysis
CSCI 161: Introduction to Programming I Lab 1b: Hello, World (Eclipse, Java)
 Goals - to learn how to compile and execute a Java program - to modify a program to enhance it Overview This activity will introduce you to the Java programming language. You will type in the Java program
Goals - to learn how to compile and execute a Java program - to modify a program to enhance it Overview This activity will introduce you to the Java programming language. You will type in the Java program
Quality Checking an fmri Group Result (art_groupcheck)
 Quality Checking an fmri Group Result (art_groupcheck) Paul Mazaika, Feb. 24, 2009 A statistical parameter map of fmri group analyses relies on the assumptions of the General Linear Model (GLM). The assumptions
Quality Checking an fmri Group Result (art_groupcheck) Paul Mazaika, Feb. 24, 2009 A statistical parameter map of fmri group analyses relies on the assumptions of the General Linear Model (GLM). The assumptions
Journal of Articles in Support of The Null Hypothesis
 Data Preprocessing Martin M. Monti, PhD UCLA Psychology NITP 2016 Typical (task-based) fmri analysis sequence Image Pre-processing Single Subject Analysis Group Analysis Journal of Articles in Support
Data Preprocessing Martin M. Monti, PhD UCLA Psychology NITP 2016 Typical (task-based) fmri analysis sequence Image Pre-processing Single Subject Analysis Group Analysis Journal of Articles in Support
Brain Extraction, Registration & EPI Distortion Correction
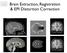 Brain Extraction, Registration & EPI Distortion Correction What use is Registration? Some common uses of registration: Combining across individuals in group studies: including fmri & diffusion Quantifying
Brain Extraction, Registration & EPI Distortion Correction What use is Registration? Some common uses of registration: Combining across individuals in group studies: including fmri & diffusion Quantifying
SIVIC Scripting Tutorial
 SIVIC Scripting Tutorial HMTRC Workshop - March 23-24, 2017 Department of Radiology and Biomedical Imaging, UCSF Supported by NIBIB P41EB013598 Goal: The purpose of this tutorial is to introduce you to
SIVIC Scripting Tutorial HMTRC Workshop - March 23-24, 2017 Department of Radiology and Biomedical Imaging, UCSF Supported by NIBIB P41EB013598 Goal: The purpose of this tutorial is to introduce you to
BDP: BrainSuite Diffusion Pipeline. Chitresh Bhushan
 BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
Image Acquisition Systems
 Image Acquisition Systems Goals and Terminology Conventional Radiography Axial Tomography Computer Axial Tomography (CAT) Magnetic Resonance Imaging (MRI) PET, SPECT Ultrasound Microscopy Imaging ITCS
Image Acquisition Systems Goals and Terminology Conventional Radiography Axial Tomography Computer Axial Tomography (CAT) Magnetic Resonance Imaging (MRI) PET, SPECT Ultrasound Microscopy Imaging ITCS
Computational Neuroanatomy
 Computational Neuroanatomy John Ashburner john@fil.ion.ucl.ac.uk Smoothing Motion Correction Between Modality Co-registration Spatial Normalisation Segmentation Morphometry Overview fmri time-series kernel
Computational Neuroanatomy John Ashburner john@fil.ion.ucl.ac.uk Smoothing Motion Correction Between Modality Co-registration Spatial Normalisation Segmentation Morphometry Overview fmri time-series kernel
Correction of Partial Volume Effects in Arterial Spin Labeling MRI
 Correction of Partial Volume Effects in Arterial Spin Labeling MRI By: Tracy Ssali Supervisors: Dr. Keith St. Lawrence and Udunna Anazodo Medical Biophysics 3970Z Six Week Project April 13 th 2012 Introduction
Correction of Partial Volume Effects in Arterial Spin Labeling MRI By: Tracy Ssali Supervisors: Dr. Keith St. Lawrence and Udunna Anazodo Medical Biophysics 3970Z Six Week Project April 13 th 2012 Introduction
Diffusion Mapping with FireVoxel Quick Start Guide
 Diffusion Mapping with FireVoxel Quick Start Guide Medical image analysis tool developed by Artem Mikheev and Henry Rusinek Radiology Department, NYU School of Medicine Original version prepared by Jinyu
Diffusion Mapping with FireVoxel Quick Start Guide Medical image analysis tool developed by Artem Mikheev and Henry Rusinek Radiology Department, NYU School of Medicine Original version prepared by Jinyu
User s Guide Neuroimage Processing ToolKit (NPTK) Version.1.7 (beta) fmri Registration Software Pipeline for Functional Localization
 User s Guide Neuroimage Processing ToolKit (NPTK) Version.1.7 (beta) fmri Registration Software Pipeline for Functional Localization Software Written by Ali Gholipour SIP Lab, UTD, 2005-2007 Revision 1.7
User s Guide Neuroimage Processing ToolKit (NPTK) Version.1.7 (beta) fmri Registration Software Pipeline for Functional Localization Software Written by Ali Gholipour SIP Lab, UTD, 2005-2007 Revision 1.7
ACQUIRING AND PROCESSING SUSCEPTIBILITY WEIGHTED IMAGING (SWI) DATA ON GE 3.0T
 ACQUIRING AND PROCESSING SUSCEPTIBILITY WEIGHTED IMAGING (SWI) DATA ON GE 3.0T Revision date: 12/13/2010 Overview Susceptibility Weighted Imaging (SWI) is a relatively new data acquisition and processing
ACQUIRING AND PROCESSING SUSCEPTIBILITY WEIGHTED IMAGING (SWI) DATA ON GE 3.0T Revision date: 12/13/2010 Overview Susceptibility Weighted Imaging (SWI) is a relatively new data acquisition and processing
Normalization for clinical data
 Normalization for clinical data Christopher Rorden, Leonardo Bonilha, Julius Fridriksson, Benjamin Bender, Hans-Otto Karnath (2012) Agespecific CT and MRI templates for spatial normalization. NeuroImage
Normalization for clinical data Christopher Rorden, Leonardo Bonilha, Julius Fridriksson, Benjamin Bender, Hans-Otto Karnath (2012) Agespecific CT and MRI templates for spatial normalization. NeuroImage
Documentation for imcalc (SPM 5/8/12) Robert J Ellis
 (_) _ '_ ` _ \ / / _` / Image calculations and transformations (using SPM) (_ (_ ( This software version: 09-Nov-2017 _ _ _ _ \ \,_ _ \ (C) Robert J Ellis (http://tools.robjellis.net) Documentation for
(_) _ '_ ` _ \ / / _` / Image calculations and transformations (using SPM) (_ (_ ( This software version: 09-Nov-2017 _ _ _ _ \ \,_ _ \ (C) Robert J Ellis (http://tools.robjellis.net) Documentation for
Tutorial 1: Unix Basics
 Tutorial 1: Unix Basics To log in to your ece account, enter your ece username and password in the space provided in the login screen. Note that when you type your password, nothing will show up in the
Tutorial 1: Unix Basics To log in to your ece account, enter your ece username and password in the space provided in the login screen. Note that when you type your password, nothing will show up in the
Tutorial BOLD Module
 m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial BOLD Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial BOLD Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
Tools for NIfTI and ANALYZE image (Click LINK HERE for the latest update)
 Tools for NIfTI and ANALYZE image (Click LINK HERE for the latest update) Description: If you are confused by identifying which one is the Left-hand-side / Right-hand-side of an ANALYZE image, the UseANALYZE.pdf
Tools for NIfTI and ANALYZE image (Click LINK HERE for the latest update) Description: If you are confused by identifying which one is the Left-hand-side / Right-hand-side of an ANALYZE image, the UseANALYZE.pdf
Diffusion MRI Acquisition. Karla Miller FMRIB Centre, University of Oxford
 Diffusion MRI Acquisition Karla Miller FMRIB Centre, University of Oxford karla@fmrib.ox.ac.uk Diffusion Imaging How is diffusion weighting achieved? How is the image acquired? What are the limitations,
Diffusion MRI Acquisition Karla Miller FMRIB Centre, University of Oxford karla@fmrib.ox.ac.uk Diffusion Imaging How is diffusion weighting achieved? How is the image acquired? What are the limitations,
MRI Segmentation MIDAS, 2007, 2010
 MRI Segmentation MIDAS, 2007, 2010 Lawrence O. Hall, Dmitry Goldgof, Yuhua Gu, Prodip Hore Dept. of Computer Science & Engineering University of South Florida CONTENTS: 1. Introduction... 1 2. Installing
MRI Segmentation MIDAS, 2007, 2010 Lawrence O. Hall, Dmitry Goldgof, Yuhua Gu, Prodip Hore Dept. of Computer Science & Engineering University of South Florida CONTENTS: 1. Introduction... 1 2. Installing
Lecture 4 Image Enhancement in Spatial Domain
 Digital Image Processing Lecture 4 Image Enhancement in Spatial Domain Fall 2010 2 domains Spatial Domain : (image plane) Techniques are based on direct manipulation of pixels in an image Frequency Domain
Digital Image Processing Lecture 4 Image Enhancement in Spatial Domain Fall 2010 2 domains Spatial Domain : (image plane) Techniques are based on direct manipulation of pixels in an image Frequency Domain
Stat 5411 Lab 1 Fall Assignment: Turn in copies of bike.sas and bike.lst from your SAS run. Turn this in next Friday with your assignment.
 Stat 5411 Lab 1 Fall 2009 Assignment: Turn in copies of bike.sas and bike.lst from your SAS run. Turn this in next Friday with your assignment. To run SAS on UMD's UNIX machine ub, you can either (1) create
Stat 5411 Lab 1 Fall 2009 Assignment: Turn in copies of bike.sas and bike.lst from your SAS run. Turn this in next Friday with your assignment. To run SAS on UMD's UNIX machine ub, you can either (1) create
Transforming Datasets to Talairach-Tournoux Coordinates
 -1- Transforming Datasets to Talairach-Tournoux Coordinates The original purpose of AFNI was to perform the transformation of datasets to Talairach-Tournoux (stereotaxic) coordinates The transformation
-1- Transforming Datasets to Talairach-Tournoux Coordinates The original purpose of AFNI was to perform the transformation of datasets to Talairach-Tournoux (stereotaxic) coordinates The transformation
Brain Images. Download the data. Comparing Images. 13 th February Version 1.4. For GIMIAS-1.4. Contact:
 Brain Images Comparing Images 13 th February 2012 Version 1.4 For GIMIAS-1.4 Contact: cistib.developers@gmail.com The aim of this exercise is to show different visualization and fusion functionalities
Brain Images Comparing Images 13 th February 2012 Version 1.4 For GIMIAS-1.4 Contact: cistib.developers@gmail.com The aim of this exercise is to show different visualization and fusion functionalities
Transform Introduction page 96 Spatial Transforms page 97
 Transform Introduction page 96 Spatial Transforms page 97 Pad page 97 Subregion page 101 Resize page 104 Shift page 109 1. Correcting Wraparound Using the Shift Tool page 109 Flip page 116 2. Flipping
Transform Introduction page 96 Spatial Transforms page 97 Pad page 97 Subregion page 101 Resize page 104 Shift page 109 1. Correcting Wraparound Using the Shift Tool page 109 Flip page 116 2. Flipping
Part I. Introduction to Linux
 Part I Introduction to Linux 7 Chapter 1 Linux operating system Goal-of-the-Day Familiarisation with basic Linux commands and creation of data plots. 1.1 What is Linux? All astronomical data processing
Part I Introduction to Linux 7 Chapter 1 Linux operating system Goal-of-the-Day Familiarisation with basic Linux commands and creation of data plots. 1.1 What is Linux? All astronomical data processing
Exercise #5b - Geometric Correction of Image Data
 Exercise #5b - Geometric Correction of Image Data 5.6 Geocoding or Registration of geometrically uncorrected image data 5.7 Resampling 5.8 The Ukrainian coordinate system 5.9 Selecting Ground Control Points
Exercise #5b - Geometric Correction of Image Data 5.6 Geocoding or Registration of geometrically uncorrected image data 5.7 Resampling 5.8 The Ukrainian coordinate system 5.9 Selecting Ground Control Points
A Study of Medical Image Analysis System
 Indian Journal of Science and Technology, Vol 8(25), DOI: 10.17485/ijst/2015/v8i25/80492, October 2015 ISSN (Print) : 0974-6846 ISSN (Online) : 0974-5645 A Study of Medical Image Analysis System Kim Tae-Eun
Indian Journal of Science and Technology, Vol 8(25), DOI: 10.17485/ijst/2015/v8i25/80492, October 2015 ISSN (Print) : 0974-6846 ISSN (Online) : 0974-5645 A Study of Medical Image Analysis System Kim Tae-Eun
Lab 2. Hanz Cuevas Velásquez, Bob Fisher Advanced Vision School of Informatics, University of Edinburgh Week 3, 2018
 Lab 2 Hanz Cuevas Velásquez, Bob Fisher Advanced Vision School of Informatics, University of Edinburgh Week 3, 2018 This lab will focus on learning simple image transformations and the Canny edge detector.
Lab 2 Hanz Cuevas Velásquez, Bob Fisher Advanced Vision School of Informatics, University of Edinburgh Week 3, 2018 This lab will focus on learning simple image transformations and the Canny edge detector.
fmri Image Preprocessing
 fmri Image Preprocessing Rick Hoge, Ph.D. Laboratoire de neuroimagerie vasculaire (LINeV) Centre de recherche de l institut universitaire de gériatrie de Montréal, Université de Montréal Outline Motion
fmri Image Preprocessing Rick Hoge, Ph.D. Laboratoire de neuroimagerie vasculaire (LINeV) Centre de recherche de l institut universitaire de gériatrie de Montréal, Université de Montréal Outline Motion
LAB 1 INTRODUCTION TO LINUX ENVIRONMENT AND C COMPILER
 LAB 1 INTRODUCTION TO LINUX ENVIRONMENT AND C COMPILER School of Computer and Communication Engineering Universiti Malaysia Perlis 1 1. GETTING STARTED: 1.1 Steps to Create New Folder: 1.1.1 Click Places
LAB 1 INTRODUCTION TO LINUX ENVIRONMENT AND C COMPILER School of Computer and Communication Engineering Universiti Malaysia Perlis 1 1. GETTING STARTED: 1.1 Steps to Create New Folder: 1.1.1 Click Places
Getting Started With UNIX Lab Exercises
 Getting Started With UNIX Lab Exercises This is the lab exercise handout for the Getting Started with UNIX tutorial. The exercises provide hands-on experience with the topics discussed in the tutorial.
Getting Started With UNIX Lab Exercises This is the lab exercise handout for the Getting Started with UNIX tutorial. The exercises provide hands-on experience with the topics discussed in the tutorial.
Source Reconstruction in MEG & EEG
 Source Reconstruction in MEG & EEG ~ From Brain-Waves to Neural Sources ~ Workshop Karolinska Institutet June 16 th 2017 Program for today Intro Overview of a source reconstruction pipeline Overview of
Source Reconstruction in MEG & EEG ~ From Brain-Waves to Neural Sources ~ Workshop Karolinska Institutet June 16 th 2017 Program for today Intro Overview of a source reconstruction pipeline Overview of
Slicer3 Tutorial: Registration Library Case 14. Intra-subject Brain PET-MRI fusion
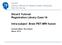 NA-MIC Slicer3 Tutorial: Registration Library Case 14 Intra-subject Brain PET-MRI fusion Dominik Meier, Ron Kikinis March 2010 Overview 1. Introduction 2. Prerequisites 3. Modules Used takes how long to
NA-MIC Slicer3 Tutorial: Registration Library Case 14 Intra-subject Brain PET-MRI fusion Dominik Meier, Ron Kikinis March 2010 Overview 1. Introduction 2. Prerequisites 3. Modules Used takes how long to
Section 9. Human Anatomy and Physiology
 Section 9. Human Anatomy and Physiology 9.1 MR Neuroimaging 9.2 Electroencephalography Overview As stated throughout, electrophysiology is the key tool in current systems neuroscience. However, single-
Section 9. Human Anatomy and Physiology 9.1 MR Neuroimaging 9.2 Electroencephalography Overview As stated throughout, electrophysiology is the key tool in current systems neuroscience. However, single-
Role of Parallel Imaging in High Field Functional MRI
 Role of Parallel Imaging in High Field Functional MRI Douglas C. Noll & Bradley P. Sutton Department of Biomedical Engineering, University of Michigan Supported by NIH Grant DA15410 & The Whitaker Foundation
Role of Parallel Imaging in High Field Functional MRI Douglas C. Noll & Bradley P. Sutton Department of Biomedical Engineering, University of Michigan Supported by NIH Grant DA15410 & The Whitaker Foundation
UNIX Tutorial One
 1.1 Listing files and directories ls (list) When you first login, your current working directory is your home directory. Your home directory has the same name as your user-name, for example, ee91ab, and
1.1 Listing files and directories ls (list) When you first login, your current working directory is your home directory. Your home directory has the same name as your user-name, for example, ee91ab, and
Chapter 3 Set Redundancy in Magnetic Resonance Brain Images
 16 Chapter 3 Set Redundancy in Magnetic Resonance Brain Images 3.1 MRI (magnetic resonance imaging) MRI is a technique of measuring physical structure within the human anatomy. Our proposed research focuses
16 Chapter 3 Set Redundancy in Magnetic Resonance Brain Images 3.1 MRI (magnetic resonance imaging) MRI is a technique of measuring physical structure within the human anatomy. Our proposed research focuses
MSI 2D Viewer Software Guide
 MSI 2D Viewer Software Guide Page:1 DISCLAIMER We have used reasonable effort to include accurate and up-to-date information in this manual; it does not, however, make any warranties, conditions or representations
MSI 2D Viewer Software Guide Page:1 DISCLAIMER We have used reasonable effort to include accurate and up-to-date information in this manual; it does not, however, make any warranties, conditions or representations
