Topology-Preserving STAPLE
|
|
|
- Michael Porter
- 6 years ago
- Views:
Transcription
1 Topology-Preserving STAPLE John A. Bogovic, Pierre-Louis Bazin, Jerry L. Prince Johns Hopkins University, Baltimore, MD Abstract Methodology for fusing multiple segmentations to produce an improved result has been useful in computational anatomical studies. Although obtaining segmentations of anatomy having a particular topology are essential to studies using diffeomorphic deformation based analyses, no methods of label fusion presented to date have incorporated information regarding the topology of the anatomy. In this paper, we introduce Topology STAPLE, a novel method that statistically fuses multiple rater segmentations into a topologically correct segmentation. We evaluate the method on both simulated data and real delineations of the cerebellum produced by human raters. 1. Introduction The computational study of anatomy [8, 17] has flourished since the advent of high-resolution magnetic resonance imaging (MRI). The variability in the size and shape of different anatomical structures has been studied in both normal and diseased populations [15, 13]. These studies rely on the accurate and precise delineations of the structures of interest. Delineations are often done by human raters, but have also been produced by automatic and semiautomatic segmentation algorithms. The delineation of anatomy is vital to methods involving deformation-based morphometry (DBM) [1], statistical shape analysis [6], and atlas construction using diffeomorphic mappings [10]. DBM type analyses can localize differences in the size and shape within the structure of interest. Specifically, DBM methods reveal these local differences by studying the deformations that map each instance of that anatomy to a common, stereotactic space. Statistical shape analyses also study the differences in shape, but do so by studying the differences in shapes that result after a coarse registration. Diffeomorphic mappings have been widely sought after in the field of registration and atlas construction because they have the property that a one-to-one correspondence exists between points in the original and deformed space. This is desirable because it gives a one-toone mapping from one subject s anatomy to any other subject s anatomy. One can equivalently say that the anatomy of every person is topologically equivalent. Naturally, the stated assumption may be invalid for structures that are complex and highly-varying between individuals (e.g., vascular structures), in which case these methods should be avoided. Nevertheless, when applicable, DBM and statistical shape analysis also operate under this supposition. The pervasiveness of this assumption in many analysis methods underscores the importance of ensuring that the delineations of anatomy produced by human raters or automated methods have a particular topology. In fact, the one-to-one mapping that is assumed does not exist when the objects under consideration differ topologically. Automated segmentation methods that enforce topology have been proposed [12, 3], but human raters can only be encouraged to produce smooth results as part of a manual protocol. To our knowledge, no manual protocols for the delineation of structures mention object topology. In fact, topological defects are likely to be observed when analyzing human drawn segmentations due to the challenges of delineating highly complex and convoluted structures in three dimensions, especially given the requirement of working with one twodimensional slice at a time. A number of methods have been introduced to estimate both rater performance and the true underlying parcellation [18, 14]. It has been shown that the delineation produced by combining the results of raters is more reliable than a single rater s result [14]. Recently, multiple atlas registration methods have proven to be an efficient tool for segmentation, when many atlases are combined [11]. While these methods usually include a mechanism for encouraging smoothness in the resulting segmentation, such as a Markov random field, none explicitly enforce a particular topology. In fact, enforcing a particular topology yields smooth labels inside of objects, without oversmoothing object boundaries and is especially important when dealing with objects with boundaries of high curvature. In this paper, we describe Topology STAPLE, a method for estimating rater performances and the true, topologically correct, underlying segmentation. The details of our method are described in section 2, and include a brief summary of the STAPLE algorithm [18] on which /10/$ IEEE 1
2 it is based. Our method is then tested on simple, simulated data to demonstrate the effectiveness of the algorithm, the results of which are described in section 3. We then applied our method to a set of 22 real, hand-drawn delineations of the human cerebellum to illustrate its relevance and applicability in the context of a research or clinical study. 2. Methods In this section we describe our methodology, which follows the framework established in the STAPLE algorithm to produce a maximum-likelihood estimate of the rater performance parameters. It is a special case of the expectationmaximization (EM) algorithm, and estimates the fixed but unknown true segmentation (the hidden data) as well. Our methodology is nearly identical in formulation and similar in execution, adding an important topological constraint on the unknown true segmentation. We first introduce our notation. We denote the rater data as D i = D ir, where i {1, 2...N} indexes the voxels and r {1, 2...R} indexes the raters, where R is the number of raters. In this paper, theory is described only for binary labelings, implying that D ir {0, 1}, although the method can be generalized as demonstrated in section 3.3. The performance of the r th rater is parameterized by the true positive and true negative rates, denoted θ r11 and θ r00, respectively. The perfect rater will therefore be parameterized by θ r00 = θ r11 = 1, (i.e., the rater s true positive and true negative rates equal one). This subscript notation was chosen to emphasize that these parameters may be interpreted as the conditional probabilties: θ r11 = p(d ir = 1 T i = 1) and θ r00 = p(d ir = 0 T i = 0) where T i denotes the true label at voxel i. The probabilities of error are therefore θ r10 = 1 θ r00 = p(d ir = 1 T i = 0) and θ r01 = 1 θ r11 = p(d ir = 0 T i = 1). The performance of all raters are summarized by the 2R vector Θ = θ rss, where s {0, 1}. Maximum-likelihood estimation of the rater parameters is performed using the expectation-maximization (EM) algorithm. The E-Step is performed as in STAPLE and estimates the underlying, true segmentation. Next, this segmentation is projected onto the topologically correct space using a topology correction algorithm [2]. Finally, the ML estimates of the rater performance are computed in the M- Step. This procedure is described in detail below E-Step The E-Step of the EM iterations estimates the probability that the object is present at each voxel, and is computed by taking the conditional expectation of the complete data log likelihood, p(θ, T ), where T = T i gives the true label at each voxel i. After some manipulation, weight variables for the iteration k, W (k) si can be computed using: W (k) si = p(t i = s D i, Θ (k) ) (1) = p(t i = s) r:d ir=s θ(k) rss s p(t i = s). (2) r:d ir=s θ(k) rss the details of which can be found in [18] or [14] and where s {0, 1} as above. The result is a measure of the conditional probability that the object is present at each voxel i given the rater data D i and the estimated rater performances Θ. These probability maps are likely to be topologically defective, as the conditional probabilities are measured locally. A Markov random field (MRF) can be applied to encourage homogeneity, but does not guarantee that a particular topology results. To remedy this, we apply a topology correction algorithm on the probability maps Topology Correction We posit that the topology of the true, underlying object is known a priori and enforce that topology on our estimate of the segmentation. A topology correction algorithm using a fast-marching method [2] is applied directly to the object probability map computed as part of the E-Step. The result of this correction is a new probability map that approximates the original, but for which all isocontours or isosurfaces have the desired topology. Forming a hard segmentation of an object by thresholding this corrected probability map at any isovalue will result in an object with the correct topology. Before delving into the details of the methodology, we review some basics of digital topology. Elementary topological changes to discrete objects involve the addition or removal of a point from the object. Points that do not affect the object s topology when added or removed are called simple points. In fact, whether a point is simple or not may be determined from the point s local neighborhood [4]. In this application, we seek to correct the topology of probability maps, and so make use of generalized simple points for scalar fields [5]. For a scalar field, a point x is simple if N n (x) = N p (x) = 1, where N n (x) is the number of connected regions in the neighborhood of x with intensities less than or equal to the intensity at x. Likewise, N p (x) denotes the number of connected regions in the neighborhood of x with intensities strictly greater than the intensity at x. The topology correction methodology used in this work successively detects and removes non-simple points for all isovalues of the input scalar field. The process begins with an initialization with the desired topology which is propagated through successive isovalues using a fast-marching approach, the details of which can be found in section 2.2 of [2]. The points in the boundary around the initialized object are sorted by intensity and organized in a binary tree. At the k th iteration, the first point is removed from the tree and added to the object if it is simple. 2
3 If the point x has been labeled as non-simple, its value is set to g(x) = min y Gk 1 N(x) g(y) where N(x) denotes the neighbors of x and G k 1 denotes the object at the k 1 th iteration. The correction can be done in two ways, upward or downward. The former is initialized in regions of high probability and raises areas of low probability, whereas the latter flattens regions of high probability and is initialized in regions of low probability. Figure 1 shows a simple 1D example of topology correction on scalar functions. Figure 1. An example of topology correction applied to a 1D scalar function. Displayed in black are the original scalar functions and in blue, the corrected function for both a) upward correction and b) downward correction Our method applies both topology correction schemes and chooses the result for which the sum of squared differences across all voxels is minimized. Formally, if V u and V d denote the upward and downward topology corrected probability maps, we choose our final probability map V = V i using: V = arg min (W i Y i ) 2. (3) Y {V u,v d } In this way, the resulting probability map is the closest projection (with respect to Euclidean distance) of the original on the space of topologically correct probability maps. An experiment is shown in section 3.1 that highlights the importance of approaching the problem in this way M-Step Finally, the rater performance parameter Θ that maximizes the conditional expectation of the complete data loglikelihood is computed. The details of the derivation can again be found in [18] or [14], and result in the following update equation i i:d θ (k+1) rs s = W (k) ir=s si i W. (4) (k) si Our use of the topology corrected probability map results in higher performance parameter estimates for raters that produced topologically correct objects and lower estimates for those whose labels were topologically defective. This translates into a higher weighting for the raters with correct topology in the subsequent E-Step, resulting in a new probability map that is closer to the correct topology Overview To summarize, Topology STAPLE carries out expectation-maximization (EM) iterations with 1. E-Step - estimate probability of object at each voxel 2. Project the resulting probability map onto the subspace of topologically correct probabilities 3. M-Step - estimate rater performance given the topologically correct probability map 4. Check for convergence Convergence is detected by measuring the normalized trace of Θ between subsequent iterations. The normalized trace averages the true positive and true negative fractions across raters and is computed by 1 R 2R r=1 tr(θ r). Since the topology corrected probability maps and hard segmentations are of interest in this study, the E-Step and topology correction were performed once more after the final rater performances were computed Measuring Topology The topological characteristics of the raters and the rater fusion results are described using the number of object parts, the number of object cavities, the number of object handles, and the Euler characteristic. The number of object parts (P ) is the number of connected components of the object. The number of object cavities (C) is the number of connected components of the background completely enclosed within the object. A torus or coffee mug are examples of objects with one object handle (H). The Euler characteristic of defined by χ = V E + F where V is the number of vertices, E is the number of edges, and F is the number of faces of polygonization of the object [7]. It is also related to the number of object parts, cavities and handles by: χ = 2P + 2C 2H. In most of the studies described here, the true object is presumed to have spherical topology (that is, P = 1, C = 0, H = 0, and χ = 2). Three types of topological defects can occur in three dimensions. The object may have multiple disjoint pieces, resulting in P > 1. The object may contain cavites, resulting in C > 0. Finally, the object may possess handles, resulting in H > 0. The Euler characteristic is reported along with these three quantities whenever the topology of objects is reported. 3. Experiments This proposed method was applied to both simulated and real data to gauge its efficacy and effectiveness. For 3
4 all experiments, the reported topology measures used 6- connectivity for the object, and 18-connectivity for the background, though similar results were obtained using other connectivity rules. The STAPLE algorithm was initialized with θ rjj = r, j. The algorithm was determined to have converged when the difference of the normalized trace (see Section 2.4) between iterations was less than Simulations Topology STAPLE and STAPLE were run on simple examples to demonstrate the efficacy of the topology correction methodology. A shape resembling a gray matter gyral crown and sulcal fundus was created to mimic a situation in which errors in topology are likely to occur. Five synthetic raters were created by perturbing the boundary of the shape. Figure 2 shows the true segmentation, one example rater, (c) the STAPLE hard segmentation, and (d) the Topology STAPLE hard segmentation. From these results, it is evident that our method does indeed enforce the correct topology on the resulting hard segmentation. These results demonstrate the ability of Topology STA- PLE to produce topologically correct segmentations in the presence of rater error. Notice that object was cut rather than the handle filled. This is because our algorithm chooses the topology correction method that results in a probability map closer to the computed conditional probabilities, as described in section 2.2. Finally, we note that it is possible to recover nonspherical topologies using our method, and demonstrate this ability with another simple example. This synthetic example resembles the white matter of the spinal cord and is topologically equivalent to a torus, or doughnut. The key to obtaining non-spherical topologies is to initialize the topology correction step with a mask of the desired topology. Figure 3 shows the truth model with toroidal topology,, a sample rater in white and the initialization in red. Note that the initialization is of the correct topology and is close to the true segmentation, in the sense that the cavity of the initialization encloses the cavity in the truth model. (c) Figure 3. Simulation experiment - tous/spine a) True label configuration with a toroidal topology b) Sample Rater (white) and initialization (red) d) Topology STAPLE hard segmentation Again, the segmentation produced from Topology STA- PLE has the correct topology, whereas most raters and therefore, the STAPLE segmentations are topologically defective Real Data (c) (d) Figure 2. Simulation experiment - sphere/gyrus a) True label configuration b) Sample Rater c) STAPLE hard segmentation d) Topology STAPLE hard segmentation Both Topology STAPLE and STAPLE were applied to real labelings produced by human raters. Four raters labeled the human cerebellum for 22 subjects from MP-RAGE MRI scans. The subject pool consisted of both control subjects and patients diagnosed with cerebellar degeneration. These four delineations for each subject were inputs to both STA- PLE and Topology STAPLE. Table 1 shows several topological properties for all human raters, as well as for the resulting STAPLE and Topology STAPLE segmentations. The measures of topology described in section 2.5 were computed for each of the rater segmentations and for the STAPLE and Topology STAPLE segmentations. We note that none of the input raters have spherical topology as evidenced by the measures shown below. While in some cases, the STAPLE segmentation improves upon the rater results, as evidenced by fewer object parts and cavities, many topological defects remain. Topology STAPLE produces a segmentation with the correct topology for all cases. Figure 4 shows a coronal slice of the segmentation one subject from all four raters, STAPLE, and Topology STA- PLE. It is evident the segmentation from all four raters contained a topological error in the form of a cavity in the middle of the cerebellum. As a result, the labeling produced from the maximum a posteriori label probability obtained from STAPLE contains this hole as well. By enforcing our prior knowledge that the cerebellum has spherical topology, Topology STAPLE produces a segmentation estimate free of any cavities or handles. 4
5 Table 1. Topological Characteristics of Raters, STAPLE (ST) results, and Topology STAPLE (TST) results Object Handles Object Cavities Object Parts Euler Char. Sub ST TST ST TST ST TST ST TST Finally, an experiment was carried out for which four raters annotated a volume with four object labels plus the background label. The objects under consideration were the anterior lobe, middle lobe, caudal lobe, and the corpus medullare of the cerebellum. We seek an estimate of the true label configuration for which all individual objects have spherical topology. This was done using a confusion matrix parameterization which generalizes the singleobject, true-positive, true-negative parameterization. Likewise, equations 2 and 4 were generalized to multiple objects as shown in [14]. Topology correction was done individually on the probability maps for each object. Figure 5 shows a surface rendering of volumetric labels obtained using Topology STAPLE. Figure 5 and (c) shows a close up of part of the surface obtained from STAPLE and Topology STAPLE results respectively. A handle is visible in the former, but not in the latter. (c) (d) (e) (f) Figure 4. Experiment with real rater delineations of the cerebellum a) Rater 1, Rater 2, (c) Rater 3, (d) Rater 4 delineations, e) STAPLE hard segmentation f) Topology STAPLE hard segmentation. The topology-corrected segmentation filled a number of cavities and handles that were ignored by STAPLE. (c) Figure 5. Multiple-label Topology STAPLE experiment. a) Topology STAPLE result using four objects, A close up of a handle in one object of a STAPLE segmentation, (c) The same portion of the surface obtained from the Topology STAPLE segmentation Multi Label Data We emphasize that by topology correcting the probability maps of each object individually, the resulting scalar maps are no longer memberships or probabilities, as the sum of the memberships are in general not equal to one. Furthermore, a topologically correct hard segmentation is not guaranteed when it is obtained by a max-membership rule, even if all of the memberships are topologically correct. Therefore, though closer to the correct topology than any individual rater or a STAPLE segmentation, the objects obtained using the above strategy will not have spherical topology in general, a challenge that will be addressed in future work. 5
6 4. Conclusion This paper presents a novel method, Topology STAPLE, for estimating the hidden, topologically correct, true segmentation of an object and rater performances given rater delineations. Topology STAPLE was demonstrated to produce a topologically correct label estimate on synthetic data sets as well as on real cerebellar delineations from human raters. We have shown that none of the human raters, nor the STAPLE estimate applied on these results produced the correct topology. We verified on these data that our method produce a specified topology in practice. Our method was also applied on a dataset with multiple object labels, by applying topology correction to each label individually and updating probabilities accordingly. Future work will study estimating the global, closest projection onto the topologically correct space. This could be done using graph-based correction methods [9, 16], for example, or by further development of the current fast marching correction method. Past work in the topology of multiple objects has employed topology templates, or initializations of the correct topology that are deformed according to membership functions. We will investigate the use of such templates and alternative techniques into a multiobject, topology-preserving version of STAPLE. With this tool we will be able to guarantee topological equivalence between segmentations obtained by combining multiple atlas-based labelings. References [1] J. Ashburner, C. Hutton, R. Frackowiak, I. Johnsrude, C. Price, and K. Friston. Identifying global anatomical differences: deformation-based morphometry. Hum. Brain Mapp., 6(5-6): , [2] P.-l. Bazin and D. L. Pham. Topology Correction of Segmented Medical Images using a Fast Marching Algorithm. Comput Methods Programs Biomed, 88(2): , [3] P.-L. Bazin and D. L. Pham. Topology-preserving tissue classification of magnetic resonance brain images. IEEE transactions on medical imaging, 26(4):487 96, April [4] G. Bertrand. Simple points, topological numbers and geodesic neighborhood in cubic grids. Pattern Recognition Letters, 15(10): , [5] M. Couprie, F. N. Bezerra, and G. Bertrand. Topological operators for grayscale image processing. Journal of Electronic Imaging, 10(4):1003, [6] P. Golland, W. E. L. Grimson, and R. Kikinis. Statistical Shape Analysis Using Fixed Topology Skeletons: Corpus Callosum Study. In Proc. IPMI, LNCS, 1613: , [7] R. C. Gonzalez and R. E. Woods. Digital Image Processing. Addison-Wesley, [8] U. Grenader and M. I. Miller. Computational anatomy: an emerging discipline. Quarterly of Applied Mathematics, 56(4): , [9] X. Han, C. Xu, U. Braga-neto, and J. L. Prince. Topology Correction in Brain Cortex Segmentation Using a Multiscale, Graph-Based Algorithm. IEEE Trans. Med. Imag, 21(2): , [10] S. Joshi, B. Davis, M. Jomier, and G. Gerig. Unbiased diffeomorphic atlas construction for computational anatomy. NeuroImage, 23 Suppl 1:S151 60, [11] T. Langerak, U. van Der Heide, I. Lips, A. Kotte, M. van Vulpen, and J. Pluim. Label fusion using performance estimation with iterative label selection IEEE ISBI, pages , June [12] J.-F. Mangin, V. Frouin, I. Bloch, J. Regis, and J. Lopez-Krahe. From 3D magnetic resonance images to structural representations of the cortex topography using topology preserving deformations. Journal of Mathematical Imaging and Vision, 5(4): , [13] D. Mietchen and C. Gaser. Computational morphometry for detecting changes in brain structure due to development, aging, learning, disease and evolution. Frontiers in neuroinformatics, 3(25), [14] T. Rohlfing, D. B. Russakoff, and C. R. Maurer. Performance-Based Classifier Combination in Atlas-Based Image Segmentation Using Expectation- Maximization Parameter Estimation. IEEE Trans. Med. Imaging, 23(8): , [15] A. I. Scher, Y. Xu, E. S. C. Korf, L. R. White, P. Scheltens, A. W. Toga, P. M. Thompson, S. W. Hartley, M. P. Witter, D. J. Valentino, and L. J. Launer. Hippocampal shape analysis in Alzheimer s disease: a population-based study. NeuroImage, 36(1):8 18, [16] D. W. Shattuck and R. M. Leahy. Automated graphbased analysis and correction of cortical volume topology. IEEE Trans. Med. Imag, 20(11): , [17] P. M. Thompson and A. W. Toga. A framework for computational anatomy. Computing and Visualization in Science, 5(1):13 34, July [18] S. K. Warfield, K. H. Zou, and W. M. Wells. Simultaneous Truth and Performance Level Estimation (STAPLE): An Algorithm for the Validation of Image Segmentation. IEEE Trans. Med. Imaging, 23(7): ,
Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm
 Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm Lin Chen and Gudrun Wagenknecht Central Institute for Electronics, Research Center Jülich, Jülich, Germany Email: l.chen@fz-juelich.de
Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm Lin Chen and Gudrun Wagenknecht Central Institute for Electronics, Research Center Jülich, Jülich, Germany Email: l.chen@fz-juelich.de
ABSTRACT 1. INTRODUCTION 2. METHODS
 Finding Seeds for Segmentation Using Statistical Fusion Fangxu Xing *a, Andrew J. Asman b, Jerry L. Prince a,c, Bennett A. Landman b,c,d a Department of Electrical and Computer Engineering, Johns Hopkins
Finding Seeds for Segmentation Using Statistical Fusion Fangxu Xing *a, Andrew J. Asman b, Jerry L. Prince a,c, Bennett A. Landman b,c,d a Department of Electrical and Computer Engineering, Johns Hopkins
Topology Preserving Brain Tissue Segmentation Using Graph Cuts
 Topology Preserving Brain Tissue Segmentation Using Graph Cuts Xinyang Liu 1, Pierre-Louis Bazin 2, Aaron Carass 3, and Jerry Prince 3 1 Brigham and Women s Hospital, Boston, MA 1 xinyang@bwh.harvard.edu
Topology Preserving Brain Tissue Segmentation Using Graph Cuts Xinyang Liu 1, Pierre-Louis Bazin 2, Aaron Carass 3, and Jerry Prince 3 1 Brigham and Women s Hospital, Boston, MA 1 xinyang@bwh.harvard.edu
A Genus Bound for Digital Image Boundaries
 A Genus Bound for Digital Image Boundaries Lowell Abrams and Donniell E. Fishkind March 9, 2005 Abstract Shattuck and Leahy [4] conjectured and Abrams, Fishkind, and Priebe [1],[2] proved that the boundary
A Genus Bound for Digital Image Boundaries Lowell Abrams and Donniell E. Fishkind March 9, 2005 Abstract Shattuck and Leahy [4] conjectured and Abrams, Fishkind, and Priebe [1],[2] proved that the boundary
Open Topology: A Toolkit for Brain Isosurface Correction
 Open Topology: A Toolkit for Brain Isosurface Correction Sylvain Jaume 1, Patrice Rondao 2, and Benoît Macq 2 1 National Institute of Research in Computer Science and Control, INRIA, France, sylvain@mit.edu,
Open Topology: A Toolkit for Brain Isosurface Correction Sylvain Jaume 1, Patrice Rondao 2, and Benoît Macq 2 1 National Institute of Research in Computer Science and Control, INRIA, France, sylvain@mit.edu,
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge
 Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry
 Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry Nivedita Agarwal, MD Nivedita.agarwal@apss.tn.it Nivedita.agarwal@unitn.it Volume and surface morphometry Brain volume White matter
Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry Nivedita Agarwal, MD Nivedita.agarwal@apss.tn.it Nivedita.agarwal@unitn.it Volume and surface morphometry Brain volume White matter
Methods for data preprocessing
 Methods for data preprocessing John Ashburner Wellcome Trust Centre for Neuroimaging, 12 Queen Square, London, UK. Overview Voxel-Based Morphometry Morphometry in general Volumetrics VBM preprocessing
Methods for data preprocessing John Ashburner Wellcome Trust Centre for Neuroimaging, 12 Queen Square, London, UK. Overview Voxel-Based Morphometry Morphometry in general Volumetrics VBM preprocessing
Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE)
 Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE) Mahnaz Maddah, Kelly H. Zou, William M. Wells, Ron Kikinis, and Simon K. Warfield
Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE) Mahnaz Maddah, Kelly H. Zou, William M. Wells, Ron Kikinis, and Simon K. Warfield
Prototype of Silver Corpus Merging Framework
 www.visceral.eu Prototype of Silver Corpus Merging Framework Deliverable number D3.3 Dissemination level Public Delivery data 30.4.2014 Status Authors Final Markus Krenn, Allan Hanbury, Georg Langs This
www.visceral.eu Prototype of Silver Corpus Merging Framework Deliverable number D3.3 Dissemination level Public Delivery data 30.4.2014 Status Authors Final Markus Krenn, Allan Hanbury, Georg Langs This
An Introduction To Automatic Tissue Classification Of Brain MRI. Colm Elliott Mar 2014
 An Introduction To Automatic Tissue Classification Of Brain MRI Colm Elliott Mar 2014 Tissue Classification Tissue classification is part of many processing pipelines. We often want to classify each voxel
An Introduction To Automatic Tissue Classification Of Brain MRI Colm Elliott Mar 2014 Tissue Classification Tissue classification is part of many processing pipelines. We often want to classify each voxel
An Intensity Consistent Approach to the Cross Sectional Analysis of Deformation Tensor Derived Maps of Brain Shape
 An Intensity Consistent Approach to the Cross Sectional Analysis of Deformation Tensor Derived Maps of Brain Shape C. Studholme, V. Cardenas, A. Maudsley, and M. Weiner U.C.S.F., Dept of Radiology, VAMC
An Intensity Consistent Approach to the Cross Sectional Analysis of Deformation Tensor Derived Maps of Brain Shape C. Studholme, V. Cardenas, A. Maudsley, and M. Weiner U.C.S.F., Dept of Radiology, VAMC
The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy
 The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy Sokratis K. Makrogiannis, PhD From post-doctoral research at SBIA lab, Department of Radiology,
The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy Sokratis K. Makrogiannis, PhD From post-doctoral research at SBIA lab, Department of Radiology,
Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow
 Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow Abstract. Finding meaningful 1-1 correspondences between hippocampal (HP) surfaces is an important but difficult
Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow Abstract. Finding meaningful 1-1 correspondences between hippocampal (HP) surfaces is an important but difficult
Where are we now? Structural MRI processing and analysis
 Where are we now? Structural MRI processing and analysis Pierre-Louis Bazin bazin@cbs.mpg.de Leipzig, Germany Structural MRI processing: why bother? Just use the standards? SPM FreeSurfer FSL However:
Where are we now? Structural MRI processing and analysis Pierre-Louis Bazin bazin@cbs.mpg.de Leipzig, Germany Structural MRI processing: why bother? Just use the standards? SPM FreeSurfer FSL However:
Automatic Registration-Based Segmentation for Neonatal Brains Using ANTs and Atropos
 Automatic Registration-Based Segmentation for Neonatal Brains Using ANTs and Atropos Jue Wu and Brian Avants Penn Image Computing and Science Lab, University of Pennsylvania, Philadelphia, USA Abstract.
Automatic Registration-Based Segmentation for Neonatal Brains Using ANTs and Atropos Jue Wu and Brian Avants Penn Image Computing and Science Lab, University of Pennsylvania, Philadelphia, USA Abstract.
CHAPTER 2. Morphometry on rodent brains. A.E.H. Scheenstra J. Dijkstra L. van der Weerd
 CHAPTER 2 Morphometry on rodent brains A.E.H. Scheenstra J. Dijkstra L. van der Weerd This chapter was adapted from: Volumetry and other quantitative measurements to assess the rodent brain, In vivo NMR
CHAPTER 2 Morphometry on rodent brains A.E.H. Scheenstra J. Dijkstra L. van der Weerd This chapter was adapted from: Volumetry and other quantitative measurements to assess the rodent brain, In vivo NMR
Neuroimage Processing
 Neuroimage Processing Instructor: Moo K. Chung mkchung@wisc.edu Lecture 2-3. General Linear Models (GLM) Voxel-based Morphometry (VBM) September 11, 2009 What is GLM The general linear model (GLM) is a
Neuroimage Processing Instructor: Moo K. Chung mkchung@wisc.edu Lecture 2-3. General Linear Models (GLM) Voxel-based Morphometry (VBM) September 11, 2009 What is GLM The general linear model (GLM) is a
Johns Hopkins University, 3400 N. Charles St., Baltimore, MD, USA ABSTRACT 1. INTRODUCTION
 Simultaneous Truth and Performance Level Estimation with Incomplete, Over-complete, and Ancillary Data Bennett A. Landman *a,c, John A. Bogovic b, Jerry L. Prince a,b a Biomedical Engineering, b Electrical
Simultaneous Truth and Performance Level Estimation with Incomplete, Over-complete, and Ancillary Data Bennett A. Landman *a,c, John A. Bogovic b, Jerry L. Prince a,b a Biomedical Engineering, b Electrical
PROSTATE CANCER DETECTION USING LABEL IMAGE CONSTRAINED MULTIATLAS SELECTION
 PROSTATE CANCER DETECTION USING LABEL IMAGE CONSTRAINED MULTIATLAS SELECTION Ms. Vaibhavi Nandkumar Jagtap 1, Mr. Santosh D. Kale 2 1 PG Scholar, 2 Assistant Professor, Department of Electronics and Telecommunication,
PROSTATE CANCER DETECTION USING LABEL IMAGE CONSTRAINED MULTIATLAS SELECTION Ms. Vaibhavi Nandkumar Jagtap 1, Mr. Santosh D. Kale 2 1 PG Scholar, 2 Assistant Professor, Department of Electronics and Telecommunication,
The organization of the human cerebral cortex estimated by intrinsic functional connectivity
 1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques
 Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques Sea Chen Department of Biomedical Engineering Advisors: Dr. Charles A. Bouman and Dr. Mark J. Lowe S. Chen Final Exam October
Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques Sea Chen Department of Biomedical Engineering Advisors: Dr. Charles A. Bouman and Dr. Mark J. Lowe S. Chen Final Exam October
Simultaneous Cortical Surface Labeling and Sulcal Curve Extraction
 Simultaneous Cortical Surface Labeling and Sulcal Curve Extraction Zhen Yang a, Aaron Carass a, Chen Chen a, Jerry L Prince a,b a Electrical and Computer Engineering, b Biomedical Engineering, Johns Hopkins
Simultaneous Cortical Surface Labeling and Sulcal Curve Extraction Zhen Yang a, Aaron Carass a, Chen Chen a, Jerry L Prince a,b a Electrical and Computer Engineering, b Biomedical Engineering, Johns Hopkins
Norbert Schuff VA Medical Center and UCSF
 Norbert Schuff Medical Center and UCSF Norbert.schuff@ucsf.edu Medical Imaging Informatics N.Schuff Course # 170.03 Slide 1/67 Objective Learn the principle segmentation techniques Understand the role
Norbert Schuff Medical Center and UCSF Norbert.schuff@ucsf.edu Medical Imaging Informatics N.Schuff Course # 170.03 Slide 1/67 Objective Learn the principle segmentation techniques Understand the role
Morphological Analysis of Brain Structures Using Spatial Normalization
 Morphological Analysis of Brain Structures Using Spatial Normalization C. Davatzikos 1, M. Vaillant 1, S. Resnick 2, J.L. Prince 3;1, S. Letovsky 1, and R.N. Bryan 1 1 Department of Radiology, Johns Hopkins
Morphological Analysis of Brain Structures Using Spatial Normalization C. Davatzikos 1, M. Vaillant 1, S. Resnick 2, J.L. Prince 3;1, S. Letovsky 1, and R.N. Bryan 1 1 Department of Radiology, Johns Hopkins
City, University of London Institutional Repository
 City Research Online City, University of London Institutional Repository Citation: Doan, H., Slabaugh, G.G., Unal, G.B. & Fang, T. (2006). Semi-Automatic 3-D Segmentation of Anatomical Structures of Brain
City Research Online City, University of London Institutional Repository Citation: Doan, H., Slabaugh, G.G., Unal, G.B. & Fang, T. (2006). Semi-Automatic 3-D Segmentation of Anatomical Structures of Brain
ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION
 ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION Abstract: MIP Project Report Spring 2013 Gaurav Mittal 201232644 This is a detailed report about the course project, which was to implement
ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION Abstract: MIP Project Report Spring 2013 Gaurav Mittal 201232644 This is a detailed report about the course project, which was to implement
Efficient population registration of 3D data
 Efficient population registration of 3D data Lilla Zöllei 1, Erik Learned-Miller 2, Eric Grimson 1, William Wells 1,3 1 Computer Science and Artificial Intelligence Lab, MIT; 2 Dept. of Computer Science,
Efficient population registration of 3D data Lilla Zöllei 1, Erik Learned-Miller 2, Eric Grimson 1, William Wells 1,3 1 Computer Science and Artificial Intelligence Lab, MIT; 2 Dept. of Computer Science,
Automatic Generation of Training Data for Brain Tissue Classification from MRI
 MICCAI-2002 1 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. Cocosco, Alex P. Zijdenbos, and Alan C. Evans McConnell Brain Imaging Centre, Montreal Neurological
MICCAI-2002 1 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. Cocosco, Alex P. Zijdenbos, and Alan C. Evans McConnell Brain Imaging Centre, Montreal Neurological
Distance Transforms in Multi Channel MR Image Registration
 Distance Transforms in Multi Channel MR Image Registration Min Chen 1, Aaron Carass 1, John Bogovic 1, Pierre-Louis Bazin 2 and Jerry L. Prince 1 1 Image Analysis and Communications Laboratory, 2 The Laboratory
Distance Transforms in Multi Channel MR Image Registration Min Chen 1, Aaron Carass 1, John Bogovic 1, Pierre-Louis Bazin 2 and Jerry L. Prince 1 1 Image Analysis and Communications Laboratory, 2 The Laboratory
Preprocessing II: Between Subjects John Ashburner
 Preprocessing II: Between Subjects John Ashburner Pre-processing Overview Statistics or whatever fmri time-series Anatomical MRI Template Smoothed Estimate Spatial Norm Motion Correct Smooth Coregister
Preprocessing II: Between Subjects John Ashburner Pre-processing Overview Statistics or whatever fmri time-series Anatomical MRI Template Smoothed Estimate Spatial Norm Motion Correct Smooth Coregister
MR IMAGE SEGMENTATION
 MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
GLIRT: Groupwise and Longitudinal Image Registration Toolbox
 Software Release (1.0.1) Last updated: March. 30, 2011. GLIRT: Groupwise and Longitudinal Image Registration Toolbox Guorong Wu 1, Qian Wang 1,2, Hongjun Jia 1, and Dinggang Shen 1 1 Image Display, Enhancement,
Software Release (1.0.1) Last updated: March. 30, 2011. GLIRT: Groupwise and Longitudinal Image Registration Toolbox Guorong Wu 1, Qian Wang 1,2, Hongjun Jia 1, and Dinggang Shen 1 1 Image Display, Enhancement,
Subvoxel Segmentation and Representation of Brain Cortex Using Fuzzy Clustering and Gradient Vector Diffusion
 Subvoxel Segmentation and Representation of Brain Cortex Using Fuzzy Clustering and Gradient Vector Diffusion Ming-Ching Chang Xiaodong Tao GE Global Research Center {changm, taox} @ research.ge.com SPIE
Subvoxel Segmentation and Representation of Brain Cortex Using Fuzzy Clustering and Gradient Vector Diffusion Ming-Ching Chang Xiaodong Tao GE Global Research Center {changm, taox} @ research.ge.com SPIE
A multi-atlas approach for prostate segmentation in MR images
 A multi-atlas approach for prostate segmentation in MR images Geert Litjens, Nico Karssemeijer, and Henkjan Huisman Diagnostic Image Analysis Group, Radboud University Nijmegen Medical Centre, Nijmegen,
A multi-atlas approach for prostate segmentation in MR images Geert Litjens, Nico Karssemeijer, and Henkjan Huisman Diagnostic Image Analysis Group, Radboud University Nijmegen Medical Centre, Nijmegen,
Segmenting Glioma in Multi-Modal Images using a Generative Model for Brain Lesion Segmentation
 Segmenting Glioma in Multi-Modal Images using a Generative Model for Brain Lesion Segmentation Bjoern H. Menze 1,2, Koen Van Leemput 3, Danial Lashkari 4 Marc-André Weber 5, Nicholas Ayache 2, and Polina
Segmenting Glioma in Multi-Modal Images using a Generative Model for Brain Lesion Segmentation Bjoern H. Menze 1,2, Koen Van Leemput 3, Danial Lashkari 4 Marc-André Weber 5, Nicholas Ayache 2, and Polina
Classification of Abdominal Tissues by k-means Clustering for 3D Acoustic and Shear-Wave Modeling
 1 Classification of Abdominal Tissues by k-means Clustering for 3D Acoustic and Shear-Wave Modeling Kevin T. Looby klooby@stanford.edu I. ABSTRACT Clutter is an effect that degrades the quality of medical
1 Classification of Abdominal Tissues by k-means Clustering for 3D Acoustic and Shear-Wave Modeling Kevin T. Looby klooby@stanford.edu I. ABSTRACT Clutter is an effect that degrades the quality of medical
Regional Manifold Learning for Deformable Registration of Brain MR Images
 Regional Manifold Learning for Deformable Registration of Brain MR Images Dong Hye Ye, Jihun Hamm, Dongjin Kwon, Christos Davatzikos, and Kilian M. Pohl Department of Radiology, University of Pennsylvania,
Regional Manifold Learning for Deformable Registration of Brain MR Images Dong Hye Ye, Jihun Hamm, Dongjin Kwon, Christos Davatzikos, and Kilian M. Pohl Department of Radiology, University of Pennsylvania,
Watersheds on the Cortical Surface for Automated Sulcal Segmentation
 Watersheds on the Cortical Surface for Automated Sulcal Segmentation Maryam E. Rettmann,XiaoHan y, and Jerry L. Prince y Department of Biomedical Engineering y Department of Electrical and Computer Engineering
Watersheds on the Cortical Surface for Automated Sulcal Segmentation Maryam E. Rettmann,XiaoHan y, and Jerry L. Prince y Department of Biomedical Engineering y Department of Electrical and Computer Engineering
Optimal MAP Parameters Estimation in STAPLE - Learning from Performance Parameters versus Image Similarity Information
 Optimal MAP Parameters Estimation in STAPLE - Learning from Performance Parameters versus Image Similarity Information Subrahmanyam Gorthi 1, Alireza Akhondi-Asl 1, Jean-Philippe Thiran 2, and Simon K.
Optimal MAP Parameters Estimation in STAPLE - Learning from Performance Parameters versus Image Similarity Information Subrahmanyam Gorthi 1, Alireza Akhondi-Asl 1, Jean-Philippe Thiran 2, and Simon K.
Structural MRI analysis
 Structural MRI analysis volumetry and voxel-based morphometry cortical thickness measurements structural covariance network mapping Boris Bernhardt, PhD Department of Social Neuroscience, MPI-CBS bernhardt@cbs.mpg.de
Structural MRI analysis volumetry and voxel-based morphometry cortical thickness measurements structural covariance network mapping Boris Bernhardt, PhD Department of Social Neuroscience, MPI-CBS bernhardt@cbs.mpg.de
MARS: Multiple Atlases Robust Segmentation
 Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Low-Rank to the Rescue Atlas-based Analyses in the Presence of Pathologies
 Low-Rank to the Rescue Atlas-based Analyses in the Presence of Pathologies Xiaoxiao Liu 1, Marc Niethammer 2, Roland Kwitt 3, Matthew McCormick 1, and Stephen Aylward 1 1 Kitware Inc., USA 2 University
Low-Rank to the Rescue Atlas-based Analyses in the Presence of Pathologies Xiaoxiao Liu 1, Marc Niethammer 2, Roland Kwitt 3, Matthew McCormick 1, and Stephen Aylward 1 1 Kitware Inc., USA 2 University
Detecting Changes In Non-Isotropic Images
 Detecting Changes In Non-Isotropic Images K.J. Worsley 1, M. Andermann 1, T. Koulis 1, D. MacDonald, 2 and A.C. Evans 2 August 4, 1999 1 Department of Mathematics and Statistics, 2 Montreal Neurological
Detecting Changes In Non-Isotropic Images K.J. Worsley 1, M. Andermann 1, T. Koulis 1, D. MacDonald, 2 and A.C. Evans 2 August 4, 1999 1 Department of Mathematics and Statistics, 2 Montreal Neurological
STATISTICAL ATLAS-BASED SUB-VOXEL SEGMENTATION OF 3D BRAIN MRI
 STATISTICA ATAS-BASED SUB-VOXE SEGMENTATION OF 3D BRAIN MRI Marcel Bosc 1,2, Fabrice Heitz 1, Jean-Paul Armspach 2 (1) SIIT UMR-7005 CNRS / Strasbourg I University, 67400 Illkirch, France (2) IPB UMR-7004
STATISTICA ATAS-BASED SUB-VOXE SEGMENTATION OF 3D BRAIN MRI Marcel Bosc 1,2, Fabrice Heitz 1, Jean-Paul Armspach 2 (1) SIIT UMR-7005 CNRS / Strasbourg I University, 67400 Illkirch, France (2) IPB UMR-7004
Performance Evaluation of the TINA Medical Image Segmentation Algorithm on Brainweb Simulated Images
 Tina Memo No. 2008-003 Internal Memo Performance Evaluation of the TINA Medical Image Segmentation Algorithm on Brainweb Simulated Images P. A. Bromiley Last updated 20 / 12 / 2007 Imaging Science and
Tina Memo No. 2008-003 Internal Memo Performance Evaluation of the TINA Medical Image Segmentation Algorithm on Brainweb Simulated Images P. A. Bromiley Last updated 20 / 12 / 2007 Imaging Science and
Using a Statistical Shape Model to Extract Sulcal Curves on the Outer Cortex of the Human Brain
 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 21, NO. 5, MAY 2002 513 Using a Statistical Shape Model to Extract Sulcal Curves on the Outer Cortex of the Human Brain Xiaodong Tao, Jerry L. Prince, and Christos
IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 21, NO. 5, MAY 2002 513 Using a Statistical Shape Model to Extract Sulcal Curves on the Outer Cortex of the Human Brain Xiaodong Tao, Jerry L. Prince, and Christos
3D VISUALIZATION AND SEGMENTATION OF BRAIN MRI DATA
 3D VISUALIZATION AND SEGMENTATION OF BRAIN MRI DATA Konstantin Levinski, Alexei Sourin and Vitali Zagorodnov Nanyang Technological University, Nanyang Avenue, Singapore konstantin.levinski@gmail.com, assourin@ntu.edu.sg,
3D VISUALIZATION AND SEGMENTATION OF BRAIN MRI DATA Konstantin Levinski, Alexei Sourin and Vitali Zagorodnov Nanyang Technological University, Nanyang Avenue, Singapore konstantin.levinski@gmail.com, assourin@ntu.edu.sg,
A Fast and Accurate Method for Automated Cortical Surface Registration and Labeling
 A Fast and Accurate Method for Automated Cortical Surface Registration and Labeling Anand A Joshi 1, David W Shattuck 2 and Richard M Leahy 1 1 Signal and Image Processing Institute, University of Southern
A Fast and Accurate Method for Automated Cortical Surface Registration and Labeling Anand A Joshi 1, David W Shattuck 2 and Richard M Leahy 1 1 Signal and Image Processing Institute, University of Southern
Automatic Subcortical Segmentation Using a Contextual Model
 Automatic Subcortical Segmentation Using a Contextual Model Jonathan H. Morra 1, Zhuowen Tu 1, Liana G. Apostolova 1,2, Amity E. Green 1,2, Arthur W. Toga 1, and Paul M. Thompson 1 1 Laboratory of Neuro
Automatic Subcortical Segmentation Using a Contextual Model Jonathan H. Morra 1, Zhuowen Tu 1, Liana G. Apostolova 1,2, Amity E. Green 1,2, Arthur W. Toga 1, and Paul M. Thompson 1 1 Laboratory of Neuro
Subcortical Structure Segmentation using Probabilistic Atlas Priors
 Subcortical Structure Segmentation using Probabilistic Atlas Priors Sylvain Gouttard 1, Martin Styner 1,2, Sarang Joshi 3, Brad Davis 2, Rachel G. Smith 1, Heather Cody Hazlett 1, Guido Gerig 1,2 1 Department
Subcortical Structure Segmentation using Probabilistic Atlas Priors Sylvain Gouttard 1, Martin Styner 1,2, Sarang Joshi 3, Brad Davis 2, Rachel G. Smith 1, Heather Cody Hazlett 1, Guido Gerig 1,2 1 Department
Automated Graph-Based Analysis and Correction of Cortical Volume Topology
 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 20, NO. 11, NOVEMBER 2001 1167 Automated Graph-Based Analysis and Correction of Cortical Volume Topology David W. Shattuck* and Richard M. Leahy Abstract The
IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 20, NO. 11, NOVEMBER 2001 1167 Automated Graph-Based Analysis and Correction of Cortical Volume Topology David W. Shattuck* and Richard M. Leahy Abstract The
Multivariate Tensor-based Brain Anatomical Surface Morphometry via Holomorphic One-Forms
 Multivariate Tensor-based Brain Anatomical Surface Morphometry via Holomorphic One-Forms Yalin Wang 1,2, Tony F. Chan 2, Arthur W. Toga 1, and Paul M. Thompson 1 1 Lab. of Neuro Imaging, UCLA School of
Multivariate Tensor-based Brain Anatomical Surface Morphometry via Holomorphic One-Forms Yalin Wang 1,2, Tony F. Chan 2, Arthur W. Toga 1, and Paul M. Thompson 1 1 Lab. of Neuro Imaging, UCLA School of
Population Modeling in Neuroscience Using Computer Vision and Machine Learning to learn from Brain Images. Overview. Image Registration
 Population Modeling in Neuroscience Using Computer Vision and Machine Learning to learn from Brain Images Overview 1. Part 1: Theory 1. 2. Learning 2. Part 2: Applications ernst.schwartz@meduniwien.ac.at
Population Modeling in Neuroscience Using Computer Vision and Machine Learning to learn from Brain Images Overview 1. Part 1: Theory 1. 2. Learning 2. Part 2: Applications ernst.schwartz@meduniwien.ac.at
Attribute Similarity and Mutual-Saliency Weighting for Registration and Label Fusion
 Attribute Similarity and Mutual-Saliency Weighting for Registration and Label Fusion Yangming Ou, Jimit Doshi, Guray Erus, and Christos Davatzikos Section of Biomedical Image Analysis (SBIA) Department
Attribute Similarity and Mutual-Saliency Weighting for Registration and Label Fusion Yangming Ou, Jimit Doshi, Guray Erus, and Christos Davatzikos Section of Biomedical Image Analysis (SBIA) Department
Hybrid Approach for MRI Human Head Scans Classification using HTT based SFTA Texture Feature Extraction Technique
 Volume 118 No. 17 2018, 691-701 ISSN: 1311-8080 (printed version); ISSN: 1314-3395 (on-line version) url: http://www.ijpam.eu ijpam.eu Hybrid Approach for MRI Human Head Scans Classification using HTT
Volume 118 No. 17 2018, 691-701 ISSN: 1311-8080 (printed version); ISSN: 1314-3395 (on-line version) url: http://www.ijpam.eu ijpam.eu Hybrid Approach for MRI Human Head Scans Classification using HTT
Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00
 Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00 Marcel Prastawa 1 and Guido Gerig 1 Abstract July 17, 2008 1 Scientific Computing and Imaging
Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00 Marcel Prastawa 1 and Guido Gerig 1 Abstract July 17, 2008 1 Scientific Computing and Imaging
NIH Public Access Author Manuscript Proc Soc Photo Opt Instrum Eng. Author manuscript; available in PMC 2014 October 07.
 NIH Public Access Author Manuscript Published in final edited form as: Proc Soc Photo Opt Instrum Eng. 2014 March 21; 9034: 903442. doi:10.1117/12.2042915. MRI Brain Tumor Segmentation and Necrosis Detection
NIH Public Access Author Manuscript Published in final edited form as: Proc Soc Photo Opt Instrum Eng. 2014 March 21; 9034: 903442. doi:10.1117/12.2042915. MRI Brain Tumor Segmentation and Necrosis Detection
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit
 Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Random projection for non-gaussian mixture models
 Random projection for non-gaussian mixture models Győző Gidófalvi Department of Computer Science and Engineering University of California, San Diego La Jolla, CA 92037 gyozo@cs.ucsd.edu Abstract Recently,
Random projection for non-gaussian mixture models Győző Gidófalvi Department of Computer Science and Engineering University of California, San Diego La Jolla, CA 92037 gyozo@cs.ucsd.edu Abstract Recently,
Comparison of Vessel Segmentations Using STAPLE
 Comparison of Vessel Segmentations Using STAPLE Julien Jomier, Vincent LeDigarcher, and Stephen R. Aylward Computer-Aided Diagnosis and Display Lab, The University of North Carolina at Chapel Hill, Department
Comparison of Vessel Segmentations Using STAPLE Julien Jomier, Vincent LeDigarcher, and Stephen R. Aylward Computer-Aided Diagnosis and Display Lab, The University of North Carolina at Chapel Hill, Department
Brain Warping Via Landmark Points and Curves with a Level Set Representation
 Brain Warping Via Landmark Points and Curves with a Level Set Representation Andrew Y. Wang, Alex D. Leow, 2 Hillary D. Protas, Arthur W. Toga, Paul M. Thompson UCLA Laboratory of Neuro Imaging, Los Angeles,
Brain Warping Via Landmark Points and Curves with a Level Set Representation Andrew Y. Wang, Alex D. Leow, 2 Hillary D. Protas, Arthur W. Toga, Paul M. Thompson UCLA Laboratory of Neuro Imaging, Los Angeles,
Segmenting Glioma in Multi-Modal Images using a Generative-Discriminative Model for Brain Lesion Segmentation
 Segmenting Glioma in Multi-Modal Images using a Generative-Discriminative Model for Brain Lesion Segmentation Bjoern H. Menze 1,2, Ezequiel Geremia 2, Nicholas Ayache 2, and Gabor Szekely 1 1 Computer
Segmenting Glioma in Multi-Modal Images using a Generative-Discriminative Model for Brain Lesion Segmentation Bjoern H. Menze 1,2, Ezequiel Geremia 2, Nicholas Ayache 2, and Gabor Szekely 1 1 Computer
Semi-automated Basal Ganglia Segmentation Using Large Deformation Diffeomorphic Metric Mapping
 Semi-automated Basal Ganglia Segmentation Using Large Deformation Diffeomorphic Metric Mapping Ali Khan 1, Elizabeth Aylward 2, Patrick Barta 3, Michael Miller 4,andM.FaisalBeg 1 1 School of Engineering
Semi-automated Basal Ganglia Segmentation Using Large Deformation Diffeomorphic Metric Mapping Ali Khan 1, Elizabeth Aylward 2, Patrick Barta 3, Michael Miller 4,andM.FaisalBeg 1 1 School of Engineering
Scalar Visualization
 Scalar Visualization 5-1 Motivation Visualizing scalar data is frequently encountered in science, engineering, and medicine, but also in daily life. Recalling from earlier, scalar datasets, or scalar fields,
Scalar Visualization 5-1 Motivation Visualizing scalar data is frequently encountered in science, engineering, and medicine, but also in daily life. Recalling from earlier, scalar datasets, or scalar fields,
weighted minimal surface model for surface reconstruction from scattered points, curves, and/or pieces of surfaces.
 weighted minimal surface model for surface reconstruction from scattered points, curves, and/or pieces of surfaces. joint work with (S. Osher, R. Fedkiw and M. Kang) Desired properties for surface reconstruction:
weighted minimal surface model for surface reconstruction from scattered points, curves, and/or pieces of surfaces. joint work with (S. Osher, R. Fedkiw and M. Kang) Desired properties for surface reconstruction:
Learning-based Neuroimage Registration
 Learning-based Neuroimage Registration Leonid Teverovskiy and Yanxi Liu 1 October 2004 CMU-CALD-04-108, CMU-RI-TR-04-59 School of Computer Science Carnegie Mellon University Pittsburgh, PA 15213 Abstract
Learning-based Neuroimage Registration Leonid Teverovskiy and Yanxi Liu 1 October 2004 CMU-CALD-04-108, CMU-RI-TR-04-59 School of Computer Science Carnegie Mellon University Pittsburgh, PA 15213 Abstract
Active Contours Under Topology Control Genus Preserving Level Sets. Abstract
 Active Contours Under Topology Control Genus Preserving Level Sets Florent Ségonne Willow - Certis Laboratory - ENS / INRIA / ENPC Paris, France Abstract We present a novel framework to exert topology
Active Contours Under Topology Control Genus Preserving Level Sets Florent Ségonne Willow - Certis Laboratory - ENS / INRIA / ENPC Paris, France Abstract We present a novel framework to exert topology
RECENT advances in medical imaging of the brain allow
 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 18, NO. 6, JUNE 1999 467 Reconstruction of the Human Cerebral Cortex from Magnetic Resonance Images Chenyang Xu, Dzung L. Pham, Maryam E. Rettmann, Daphne N.
IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 18, NO. 6, JUNE 1999 467 Reconstruction of the Human Cerebral Cortex from Magnetic Resonance Images Chenyang Xu, Dzung L. Pham, Maryam E. Rettmann, Daphne N.
Session scaling; global mean scaling; block effect; mean intensity scaling
 Types of Scaling Session scaling; global mean scaling; block effect; mean intensity scaling Purpose remove intensity differences between runs (i.e., the mean of the whole time series). whole time series
Types of Scaling Session scaling; global mean scaling; block effect; mean intensity scaling Purpose remove intensity differences between runs (i.e., the mean of the whole time series). whole time series
Anatomically Informed Convolution Kernels for the Projection of fmri Data on the Cortical Surface
 Anatomically Informed Convolution Kernels for the Projection of fmri Data on the Cortical Surface Grégory Operto 1,Rémy Bulot 1,Jean-LucAnton 2, and Olivier Coulon 1 1 Laboratoire LSIS, UMR 6168, CNRS,
Anatomically Informed Convolution Kernels for the Projection of fmri Data on the Cortical Surface Grégory Operto 1,Rémy Bulot 1,Jean-LucAnton 2, and Olivier Coulon 1 1 Laboratoire LSIS, UMR 6168, CNRS,
LOCUS: LOcal Cooperative Unified Segmentation of MRI Brain Scans
 Author manuscript, published in "MICCAI07-10th International Conference on Medical Image Computing and Computer Assisted Intervention, Brisbane : Australia (2007)" DOI : 10.1007/978-3-540-75757-3_27 LOCUS:
Author manuscript, published in "MICCAI07-10th International Conference on Medical Image Computing and Computer Assisted Intervention, Brisbane : Australia (2007)" DOI : 10.1007/978-3-540-75757-3_27 LOCUS:
A Generative Model for Image Segmentation Based on Label Fusion Mert R. Sabuncu*, B. T. Thomas Yeo, Koen Van Leemput, Bruce Fischl, and Polina Golland
 1714 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 29, NO. 10, OCTOBER 2010 A Generative Model for Image Segmentation Based on Label Fusion Mert R. Sabuncu*, B. T. Thomas Yeo, Koen Van Leemput, Bruce Fischl,
1714 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 29, NO. 10, OCTOBER 2010 A Generative Model for Image Segmentation Based on Label Fusion Mert R. Sabuncu*, B. T. Thomas Yeo, Koen Van Leemput, Bruce Fischl,
A MORPHOLOGY-BASED FILTER STRUCTURE FOR EDGE-ENHANCING SMOOTHING
 Proceedings of the 1994 IEEE International Conference on Image Processing (ICIP-94), pp. 530-534. (Austin, Texas, 13-16 November 1994.) A MORPHOLOGY-BASED FILTER STRUCTURE FOR EDGE-ENHANCING SMOOTHING
Proceedings of the 1994 IEEE International Conference on Image Processing (ICIP-94), pp. 530-534. (Austin, Texas, 13-16 November 1994.) A MORPHOLOGY-BASED FILTER STRUCTURE FOR EDGE-ENHANCING SMOOTHING
Methodological progress in image registration for ventilation estimation, segmentation propagation and multi-modal fusion
 Methodological progress in image registration for ventilation estimation, segmentation propagation and multi-modal fusion Mattias P. Heinrich Julia A. Schnabel, Mark Jenkinson, Sir Michael Brady 2 Clinical
Methodological progress in image registration for ventilation estimation, segmentation propagation and multi-modal fusion Mattias P. Heinrich Julia A. Schnabel, Mark Jenkinson, Sir Michael Brady 2 Clinical
Research Proposal: Computational Geometry with Applications on Medical Images
 Research Proposal: Computational Geometry with Applications on Medical Images MEI-HENG YUEH yueh@nctu.edu.tw National Chiao Tung University 1 Introduction My research mainly focuses on the issues of computational
Research Proposal: Computational Geometry with Applications on Medical Images MEI-HENG YUEH yueh@nctu.edu.tw National Chiao Tung University 1 Introduction My research mainly focuses on the issues of computational
Segmentation and Modeling of the Spinal Cord for Reality-based Surgical Simulator
 Segmentation and Modeling of the Spinal Cord for Reality-based Surgical Simulator Li X.C.,, Chui C. K.,, and Ong S. H.,* Dept. of Electrical and Computer Engineering Dept. of Mechanical Engineering, National
Segmentation and Modeling of the Spinal Cord for Reality-based Surgical Simulator Li X.C.,, Chui C. K.,, and Ong S. H.,* Dept. of Electrical and Computer Engineering Dept. of Mechanical Engineering, National
NIH Public Access Author Manuscript Proc SPIE. Author manuscript; available in PMC 2013 December 30.
 NIH Public Access Author Manuscript Published in final edited form as: Proc SPIE. 2013 March 12; 8669:. doi:10.1117/12.2006682. Longitudinal Intensity Normalization of Magnetic Resonance Images using Patches
NIH Public Access Author Manuscript Published in final edited form as: Proc SPIE. 2013 March 12; 8669:. doi:10.1117/12.2006682. Longitudinal Intensity Normalization of Magnetic Resonance Images using Patches
Transformation Model and Constraints Cause Bias in Statistics on Deformation Fields
 Transformation Model and Constraints Cause Bias in Statistics on Deformation Fields Torsten Rohlfing Neuroscience Program, SRI International, Menlo Park, CA, USA torsten@synapse.sri.com http://www.stanford.edu/
Transformation Model and Constraints Cause Bias in Statistics on Deformation Fields Torsten Rohlfing Neuroscience Program, SRI International, Menlo Park, CA, USA torsten@synapse.sri.com http://www.stanford.edu/
Computational Neuroanatomy
 Computational Neuroanatomy John Ashburner john@fil.ion.ucl.ac.uk Smoothing Motion Correction Between Modality Co-registration Spatial Normalisation Segmentation Morphometry Overview fmri time-series kernel
Computational Neuroanatomy John Ashburner john@fil.ion.ucl.ac.uk Smoothing Motion Correction Between Modality Co-registration Spatial Normalisation Segmentation Morphometry Overview fmri time-series kernel
Comparison of Vessel Segmentations using STAPLE
 Comparison of Vessel Segmentations using STAPLE Julien Jomier, Vincent LeDigarcher, and Stephen R. Aylward Computer-Aided Diagnosis and Display Lab The University of North Carolina at Chapel Hill, Department
Comparison of Vessel Segmentations using STAPLE Julien Jomier, Vincent LeDigarcher, and Stephen R. Aylward Computer-Aided Diagnosis and Display Lab The University of North Carolina at Chapel Hill, Department
Automatic Generation of Training Data for Brain Tissue Classification from MRI
 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. COCOSCO, Alex P. ZIJDENBOS, and Alan C. EVANS http://www.bic.mni.mcgill.ca/users/crisco/ McConnell Brain Imaging
Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. COCOSCO, Alex P. ZIJDENBOS, and Alan C. EVANS http://www.bic.mni.mcgill.ca/users/crisco/ McConnell Brain Imaging
Discriminative, Semantic Segmentation of Brain Tissue in MR Images
 Discriminative, Semantic Segmentation of Brain Tissue in MR Images Zhao Yi 1, Antonio Criminisi 2, Jamie Shotton 2, and Andrew Blake 2 1 University of California, Los Angeles, USA. zyi@ucla.edu. 2 Microsoft
Discriminative, Semantic Segmentation of Brain Tissue in MR Images Zhao Yi 1, Antonio Criminisi 2, Jamie Shotton 2, and Andrew Blake 2 1 University of California, Los Angeles, USA. zyi@ucla.edu. 2 Microsoft
Measuring longitudinal brain changes in humans and small animal models. Christos Davatzikos
 Measuring longitudinal brain changes in humans and small animal models Christos Davatzikos Section of Biomedical Image Analysis University of Pennsylvania (Radiology) http://www.rad.upenn.edu/sbia Computational
Measuring longitudinal brain changes in humans and small animal models Christos Davatzikos Section of Biomedical Image Analysis University of Pennsylvania (Radiology) http://www.rad.upenn.edu/sbia Computational
Automated Cerebellar Lobule Segmentation using Graph Cuts
 Automated Cerebellar Lobule Segmentation using Graph Cuts Zhen Yang 1, John A. Bogovic 2, Chuyang Ye 1, Aaron Carass 1, Sarah Ying 3, and Jerry L. Prince 1 1 Johns Hopkins University, Baltimore, USA 2
Automated Cerebellar Lobule Segmentation using Graph Cuts Zhen Yang 1, John A. Bogovic 2, Chuyang Ye 1, Aaron Carass 1, Sarah Ying 3, and Jerry L. Prince 1 1 Johns Hopkins University, Baltimore, USA 2
Biometrics Technology: Image Processing & Pattern Recognition (by Dr. Dickson Tong)
 Biometrics Technology: Image Processing & Pattern Recognition (by Dr. Dickson Tong) References: [1] http://homepages.inf.ed.ac.uk/rbf/hipr2/index.htm [2] http://www.cs.wisc.edu/~dyer/cs540/notes/vision.html
Biometrics Technology: Image Processing & Pattern Recognition (by Dr. Dickson Tong) References: [1] http://homepages.inf.ed.ac.uk/rbf/hipr2/index.htm [2] http://www.cs.wisc.edu/~dyer/cs540/notes/vision.html
Robust Statistical Fusion of Image Labels
 Re-Submitted to IEEE TMI April 12, 2011 1 Robust Statistical Fusion of Image Labels Bennett A. Landman, Member IEEE, Andrew J. Asman, Student Member, IEEE, Andrew G. Scoggins, John A. Bogovic, Student
Re-Submitted to IEEE TMI April 12, 2011 1 Robust Statistical Fusion of Image Labels Bennett A. Landman, Member IEEE, Andrew J. Asman, Student Member, IEEE, Andrew G. Scoggins, John A. Bogovic, Student
CS 231A Computer Vision (Winter 2014) Problem Set 3
 CS 231A Computer Vision (Winter 2014) Problem Set 3 Due: Feb. 18 th, 2015 (11:59pm) 1 Single Object Recognition Via SIFT (45 points) In his 2004 SIFT paper, David Lowe demonstrates impressive object recognition
CS 231A Computer Vision (Winter 2014) Problem Set 3 Due: Feb. 18 th, 2015 (11:59pm) 1 Single Object Recognition Via SIFT (45 points) In his 2004 SIFT paper, David Lowe demonstrates impressive object recognition
Resting state network estimation in individual subjects
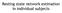 Resting state network estimation in individual subjects Data 3T NIL(21,17,10), Havard-MGH(692) Young adult fmri BOLD Method Machine learning algorithm MLP DR LDA Network image Correlation Spatial Temporal
Resting state network estimation in individual subjects Data 3T NIL(21,17,10), Havard-MGH(692) Young adult fmri BOLD Method Machine learning algorithm MLP DR LDA Network image Correlation Spatial Temporal
A Novel Contrast for DTI Visualization for Thalamus Delineation
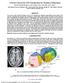 A Novel Contrast for DTI Visualization for Thalamus Delineation Xian Fan a, Meredith Thompson a,b, John A. Bogovic a, Pierre-Louis Bazin c, Jerry L. Prince a,c a Johns Hopkins University, Baltimore, MD,
A Novel Contrast for DTI Visualization for Thalamus Delineation Xian Fan a, Meredith Thompson a,b, John A. Bogovic a, Pierre-Louis Bazin c, Jerry L. Prince a,c a Johns Hopkins University, Baltimore, MD,
Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity
 Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity Pahal Dalal 1, Feng Shi 2, Dinggang Shen 2, and Song Wang 1 1 Department of Computer Science and Engineering, University of South
Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity Pahal Dalal 1, Feng Shi 2, Dinggang Shen 2, and Song Wang 1 1 Department of Computer Science and Engineering, University of South
NIH Public Access Author Manuscript Conf Proc IEEE Eng Med Biol Soc. Author manuscript; available in PMC 2009 December 11.
 NIH Public Access Author Manuscript Published in final edited form as: Conf Proc IEEE Eng Med Biol Soc. 2004 ; 2: 1521 1524. doi:10.1109/iembs.2004.1403466. An Elliptic PDE Approach for Shape Characterization
NIH Public Access Author Manuscript Published in final edited form as: Conf Proc IEEE Eng Med Biol Soc. 2004 ; 2: 1521 1524. doi:10.1109/iembs.2004.1403466. An Elliptic PDE Approach for Shape Characterization
This exercise uses one anatomical data set (ANAT1) and two functional data sets (FUNC1 and FUNC2).
 Exploring Brain Anatomy This week s exercises will let you explore the anatomical organization of the brain to learn some of its basic properties, as well as the location of different structures. The human
Exploring Brain Anatomy This week s exercises will let you explore the anatomical organization of the brain to learn some of its basic properties, as well as the location of different structures. The human
Volume visualization. Volume visualization. Volume visualization methods. Sources of volume visualization. Sources of volume visualization
 Volume visualization Volume visualization Volumes are special cases of scalar data: regular 3D grids of scalars, typically interpreted as density values. Each data value is assumed to describe a cubic
Volume visualization Volume visualization Volumes are special cases of scalar data: regular 3D grids of scalars, typically interpreted as density values. Each data value is assumed to describe a cubic
Chapter 3 Set Redundancy in Magnetic Resonance Brain Images
 16 Chapter 3 Set Redundancy in Magnetic Resonance Brain Images 3.1 MRI (magnetic resonance imaging) MRI is a technique of measuring physical structure within the human anatomy. Our proposed research focuses
16 Chapter 3 Set Redundancy in Magnetic Resonance Brain Images 3.1 MRI (magnetic resonance imaging) MRI is a technique of measuring physical structure within the human anatomy. Our proposed research focuses
3D Surface Reconstruction of the Brain based on Level Set Method
 3D Surface Reconstruction of the Brain based on Level Set Method Shijun Tang, Bill P. Buckles, and Kamesh Namuduri Department of Computer Science & Engineering Department of Electrical Engineering University
3D Surface Reconstruction of the Brain based on Level Set Method Shijun Tang, Bill P. Buckles, and Kamesh Namuduri Department of Computer Science & Engineering Department of Electrical Engineering University
Histograms. h(r k ) = n k. p(r k )= n k /NM. Histogram: number of times intensity level rk appears in the image
 Histograms h(r k ) = n k Histogram: number of times intensity level rk appears in the image p(r k )= n k /NM normalized histogram also a probability of occurence 1 Histogram of Image Intensities Create
Histograms h(r k ) = n k Histogram: number of times intensity level rk appears in the image p(r k )= n k /NM normalized histogram also a probability of occurence 1 Histogram of Image Intensities Create
Brain Portion Peeling from T2 Axial MRI Head Scans using Clustering and Morphological Operation
 159 Brain Portion Peeling from T2 Axial MRI Head Scans using Clustering and Morphological Operation K. Somasundaram Image Processing Lab Dept. of Computer Science and Applications Gandhigram Rural Institute
159 Brain Portion Peeling from T2 Axial MRI Head Scans using Clustering and Morphological Operation K. Somasundaram Image Processing Lab Dept. of Computer Science and Applications Gandhigram Rural Institute
Ensemble registration: Combining groupwise registration and segmentation
 PURWANI, COOTES, TWINING: ENSEMBLE REGISTRATION 1 Ensemble registration: Combining groupwise registration and segmentation Sri Purwani 1,2 sri.purwani@postgrad.manchester.ac.uk Tim Cootes 1 t.cootes@manchester.ac.uk
PURWANI, COOTES, TWINING: ENSEMBLE REGISTRATION 1 Ensemble registration: Combining groupwise registration and segmentation Sri Purwani 1,2 sri.purwani@postgrad.manchester.ac.uk Tim Cootes 1 t.cootes@manchester.ac.uk
Statistical Shape Analysis of Anatomical Structures. Polina Golland
 Statistical Shape Analysis of Anatomical Structures by Polina Golland B.A., Technion, Israel (1993) M.Sc., Technion, Israel (1995) Submitted to the Department of Electrical Engineering and Computer Science
Statistical Shape Analysis of Anatomical Structures by Polina Golland B.A., Technion, Israel (1993) M.Sc., Technion, Israel (1995) Submitted to the Department of Electrical Engineering and Computer Science
