Surface fmri data processing using BrainVISA (March 4th 2011)
|
|
|
- Melanie Simpson
- 5 years ago
- Views:
Transcription
1 Surface fmri data processing using BrainVISA (March 4th 2011) Index 1. Introduction and general goal 2. Running the right pipeline 3. Practical details for computing surface processing 4. Visualization of surface results 4.1. Installation of the viewer 4.2. Viewer parameters 5. Where to find all this 1. Introduction and general goal In order to compute surface analysis of fmri data this is the right scheme to follow (from Bertrand Thirion): In this document, we describe how to compute the last steps of this pipeline. Thus, you are supposed to have computed previously: - The T1 BrainVISA pipeline, and - The SPM preprocessing: slice timing, realignment and anatomo-functional coregistration (which generates images with prefix a, for example alocalizer*.img/.hdr ) Note: the examples in this document are computed using data coming from the localizer database of Philippe Pinel and Antonio Moreno.
2 2. Running the right pipeline In BrainVISA: 1) Import SPM preprocessed data. 2) Project images/activations on the cortical surface (2 steps): o Creation of the fmri data projection kernels. o Projection of functional data on the meshes by using the convolution kernels. 3) Copy file "paradigm.csv" in the subject directory /fmri/acquisition1/minf/.
3 4) Adapt and run the python script "script_surface_localizer.py" (extract):
4 5) Visualize surface results (textures): contrasts, T map, z map See below. 3. Practical details for computing surface processing - Run BrainVISA. - Add the database: BrainVISA Préférences Databases Add + Update - Import preprocessed data: BrainVISA fmri Import fmri_images Importation de donnée d'irmf (images SPM/Analyze vers images NIFTI 4D) input = select only the first image a* output = with distortion correction = no The images preprocessed with slice timing, realignment and anatomo-functional coregistration must be selected. - Import the functional mean image (optional): BrainVISA Data management Importation General import input = mean_* output (by hand) = meanaab0036.img
5 - Project images (activations) onto the cortical surface (2 steps): BrainVISA --> Cortical Surface --> IRMf --> projection_to_cortical_surface - Creation of Kernels for fmri data projection (for the two sides: "left" and "right"): Select the white matter mesh. This creates files *KERNEL.dim/.ima (in directory /surface/). - Projection using Convolution Kernels (for the two sides: "left" and "right"). Select: white_mesh = select the mesh fmri_4d_data (the processed functional data) = a* fmri_surface_data = fill in the output This step creates files *Lwhite.tex and *Rwhite.tex (in directory /surface/functional/). - Project functional images on the meshes: BrainVISA fmri Analysis_Pipeline cortical_surface_analysis 4D-NIFTI Raw Functional Projection using Kernels Select : side = Both fct_images = a* fct_mean = mean image This step creates files R_a*.tex et L_a*.tex (in directory /fmri/acquisition1/localizer_1/) - Copy file "paradigm.csv" into the subject directory /fmri/acquisition1/minf/ - Adapt and run the python script "script_surface_localizer.py": >> python script_surface_localizer.py Do this twice (for the right side and for the left side). Lines to be adapted: (lines 24-31) DBPath = "/neurospin/unicog/protocols/irmf/maindatabaselocalizers_pinelmoreno_2008/tests_surf ace_fmri/subjects" Subjects = ["AB070075"] Acquisitions = ["acquisition1"] Sessions = ["localizer_1"] model_id = "localizer_model" side = 'right' #side = 'left' - Finally, update the database: BrainVISA Data management Update/Actualisation des bases de données update the database
6 4. Visualization of surface results For the visualization of the surface results, this new BrainVISA viewer may be used: SurfaceActivationsViewer.py. It displays the (inflated) brain with the curvature of the white matter mesh and the activation contrasts/t map This viewer allows to: - Change the inflation of the brain - Change palettes of both textures at your will (contrast of the curvature or threshold of the activations) 4.1. Installation of the viewer For installing this viewer before next BrainVISA release (which will include it), copy file SurfaceActivationsViewer.py in directory ~/.brainvisa/processes/.
7 4.2. Viewer parameters activations_texture: (.tex/.gii) contrast, T map, z map input_mesh: white matter mesh compute_inflated_mesh: True/False in order to compute the inflated brain mesh (if it has not been computed previously) inflated_mesh: inflated mesh (automatically filled) curvature_texture: (.tex/.gii) curvature (automatically filled) iterations: (500 by default) inflation number of iterations save_inflate_sequence: True/False in order to compute/display the different steps of the inflated brain
8 5. Where to find all this All the tools described in this document are available in the intranet of NeuroSpin: /neurospin/unicog/resources/brainvisa_scripts/surface/ And they are downloadable from the Unicog wiki: Acknowledgements Thank you very much to Bertrand Thirion, Denis Rivière and Antoinette Jobert for their help to implement these tools and to write this document.
Analysis of fmri data within Brainvisa Example with the Saccades database
 Analysis of fmri data within Brainvisa Example with the Saccades database 18/11/2009 Note : All the sentences in italic correspond to informations relative to the specific dataset under study TP participants
Analysis of fmri data within Brainvisa Example with the Saccades database 18/11/2009 Note : All the sentences in italic correspond to informations relative to the specific dataset under study TP participants
Instructions for FA normalization (April 15th 2010)
 Instructions for FA normalization (April 15th 2010) Index 1. Introduction and general goal 2. Environment 3. Structure 4. Preparation of the data before running the batch 5. Steps to run the batches 6.
Instructions for FA normalization (April 15th 2010) Index 1. Introduction and general goal 2. Environment 3. Structure 4. Preparation of the data before running the batch 5. Steps to run the batches 6.
GLM for fmri data analysis Lab Exercise 1
 GLM for fmri data analysis Lab Exercise 1 March 15, 2013 Medical Image Processing Lab Medical Image Processing Lab GLM for fmri data analysis Outline 1 Getting Started 2 AUDIO 1 st level Preprocessing
GLM for fmri data analysis Lab Exercise 1 March 15, 2013 Medical Image Processing Lab Medical Image Processing Lab GLM for fmri data analysis Outline 1 Getting Started 2 AUDIO 1 st level Preprocessing
Generic batch description (last update April 12th 2010)
 Generic batch description (last update April 12th 2010) Index 1. Introduction and general goal 2. Environment 3. Structure 4. Preparation of the data before running the batch 5. Steps to run the batches
Generic batch description (last update April 12th 2010) Index 1. Introduction and general goal 2. Environment 3. Structure 4. Preparation of the data before running the batch 5. Steps to run the batches
fmri pre-processing Juergen Dukart
 fmri pre-processing Juergen Dukart Outline Why do we need pre-processing? fmri pre-processing Slice time correction Realignment Unwarping Coregistration Spatial normalisation Smoothing Overview fmri time-series
fmri pre-processing Juergen Dukart Outline Why do we need pre-processing? fmri pre-processing Slice time correction Realignment Unwarping Coregistration Spatial normalisation Smoothing Overview fmri time-series
Instructions for template creation and T1 normalization using SPM8 "new segmentation" and "Dartel" (September 27th 2010)
 Instructions for template creation and T1 normalization using SPM8 "new segmentation" and "Dartel" (September 27th 2010) Index 1. Introduction and general goal 2. Environment 3. Structure 4. Preparation
Instructions for template creation and T1 normalization using SPM8 "new segmentation" and "Dartel" (September 27th 2010) Index 1. Introduction and general goal 2. Environment 3. Structure 4. Preparation
Programming within BrainVISA project
 Programming within BrainVISA project What is the BrainVISA project? Complete development environement RedMine based forge Integration of various programming languages Multiplatform programming Documentation
Programming within BrainVISA project What is the BrainVISA project? Complete development environement RedMine based forge Integration of various programming languages Multiplatform programming Documentation
the PyHRF package P. Ciuciu1,2 and T. Vincent1,2 Methods meeting at Neurospin 1: CEA/NeuroSpin/LNAO
 Joint detection-estimation of brain activity from fmri time series: the PyHRF package Methods meeting at Neurospin P. Ciuciu1,2 and T. Vincent1,2 philippe.ciuciu@cea.fr 1: CEA/NeuroSpin/LNAO www.lnao.fr
Joint detection-estimation of brain activity from fmri time series: the PyHRF package Methods meeting at Neurospin P. Ciuciu1,2 and T. Vincent1,2 philippe.ciuciu@cea.fr 1: CEA/NeuroSpin/LNAO www.lnao.fr
fmri Preprocessing & Noise Modeling
 Translational Neuromodeling Unit fmri Preprocessing & Noise Modeling Lars Kasper September 25 th / October 17 th, 2015 MR-Technology Group & Translational Neuromodeling Unit An SPM Tutorial Institute for
Translational Neuromodeling Unit fmri Preprocessing & Noise Modeling Lars Kasper September 25 th / October 17 th, 2015 MR-Technology Group & Translational Neuromodeling Unit An SPM Tutorial Institute for
Functional MRI in Clinical Research and Practice Preprocessing
 Functional MRI in Clinical Research and Practice Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization
Functional MRI in Clinical Research and Practice Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization
SPM8 for Basic and Clinical Investigators. Preprocessing. fmri Preprocessing
 SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
SPM8 for Basic and Clinical Investigators. Preprocessing
 SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
Basic fmri Design and Analysis. Preprocessing
 Basic fmri Design and Analysis Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial filtering
Basic fmri Design and Analysis Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial filtering
fmri Basics: Spatial Pre-processing Workshop
 fmri Basics: Spatial Pre-processing Workshop Starting a VNC session: Most of your fmri analysis will be done on the central Linux machines accessed via a VNC (Virtual Network Computing) server. This is
fmri Basics: Spatial Pre-processing Workshop Starting a VNC session: Most of your fmri analysis will be done on the central Linux machines accessed via a VNC (Virtual Network Computing) server. This is
The organization of the human cerebral cortex estimated by intrinsic functional connectivity
 1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
SPM Introduction. SPM : Overview. SPM: Preprocessing SPM! SPM: Preprocessing. Scott Peltier. FMRI Laboratory University of Michigan
 SPM Introduction Scott Peltier FMRI Laboratory University of Michigan! Slides adapted from T. Nichols SPM! SPM : Overview Library of MATLAB and C functions Graphical user interface Four main components:
SPM Introduction Scott Peltier FMRI Laboratory University of Michigan! Slides adapted from T. Nichols SPM! SPM : Overview Library of MATLAB and C functions Graphical user interface Four main components:
SPM Introduction SPM! Scott Peltier. FMRI Laboratory University of Michigan. Software to perform computation, manipulation and display of imaging data
 SPM Introduction Scott Peltier FMRI Laboratory University of Michigan Slides adapted from T. Nichols SPM! Software to perform computation, manipulation and display of imaging data 1 1 SPM : Overview Library
SPM Introduction Scott Peltier FMRI Laboratory University of Michigan Slides adapted from T. Nichols SPM! Software to perform computation, manipulation and display of imaging data 1 1 SPM : Overview Library
User s Guide Neuroimage Processing ToolKit (NPTK) Version.1.7 (beta) fmri Registration Software Pipeline for Functional Localization
 User s Guide Neuroimage Processing ToolKit (NPTK) Version.1.7 (beta) fmri Registration Software Pipeline for Functional Localization Software Written by Ali Gholipour SIP Lab, UTD, 2005-2007 Revision 1.7
User s Guide Neuroimage Processing ToolKit (NPTK) Version.1.7 (beta) fmri Registration Software Pipeline for Functional Localization Software Written by Ali Gholipour SIP Lab, UTD, 2005-2007 Revision 1.7
EPI Data Are Acquired Serially. EPI Data Are Acquired Serially 10/23/2011. Functional Connectivity Preprocessing. fmri Preprocessing
 Functional Connectivity Preprocessing Geometric distortion Head motion Geometric distortion Head motion EPI Data Are Acquired Serially EPI Data Are Acquired Serially descending 1 EPI Data Are Acquired
Functional Connectivity Preprocessing Geometric distortion Head motion Geometric distortion Head motion EPI Data Are Acquired Serially EPI Data Are Acquired Serially descending 1 EPI Data Are Acquired
Functional MRI data preprocessing. Cyril Pernet, PhD
 Functional MRI data preprocessing Cyril Pernet, PhD Data have been acquired, what s s next? time No matter the design, multiple volumes (made from multiple slices) have been acquired in time. Before getting
Functional MRI data preprocessing Cyril Pernet, PhD Data have been acquired, what s s next? time No matter the design, multiple volumes (made from multiple slices) have been acquired in time. Before getting
Introduction to fmri. Pre-processing
 Introduction to fmri Pre-processing Tibor Auer Department of Psychology Research Fellow in MRI Data Types Anatomical data: T 1 -weighted, 3D, 1/subject or session - (ME)MPRAGE/FLASH sequence, undistorted
Introduction to fmri Pre-processing Tibor Auer Department of Psychology Research Fellow in MRI Data Types Anatomical data: T 1 -weighted, 3D, 1/subject or session - (ME)MPRAGE/FLASH sequence, undistorted
Fmri Spatial Processing
 Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
Automated MR Image Analysis Pipelines
 Automated MR Image Analysis Pipelines Andy Simmons Centre for Neuroimaging Sciences, Kings College London Institute of Psychiatry. NIHR Biomedical Research Centre for Mental Health at IoP & SLAM. Neuroimaging
Automated MR Image Analysis Pipelines Andy Simmons Centre for Neuroimaging Sciences, Kings College London Institute of Psychiatry. NIHR Biomedical Research Centre for Mental Health at IoP & SLAM. Neuroimaging
Preprocessing of fmri data (basic)
 Preprocessing of fmri data (basic) Practical session SPM Course 2016, Zurich Andreea Diaconescu, Maya Schneebeli, Jakob Heinzle, Lars Kasper, and Jakob Sieerkus Translational Neuroodeling Unit (TNU) Institute
Preprocessing of fmri data (basic) Practical session SPM Course 2016, Zurich Andreea Diaconescu, Maya Schneebeli, Jakob Heinzle, Lars Kasper, and Jakob Sieerkus Translational Neuroodeling Unit (TNU) Institute
Surface-based versus volume-based fmri group analysis: a case study
 Surface-based versus volume-based fmri group analysis: a case study Alan Tucholka, Merlin Keller, Jean-Baptiste Poline, Alexis Roche, Bertrand Thirion To cite this version: Alan Tucholka, Merlin Keller,
Surface-based versus volume-based fmri group analysis: a case study Alan Tucholka, Merlin Keller, Jean-Baptiste Poline, Alexis Roche, Bertrand Thirion To cite this version: Alan Tucholka, Merlin Keller,
Anatomically Informed Convolution Kernels for the Projection of fmri Data on the Cortical Surface
 Anatomically Informed Convolution Kernels for the Projection of fmri Data on the Cortical Surface Grégory Operto 1,Rémy Bulot 1,Jean-LucAnton 2, and Olivier Coulon 1 1 Laboratoire LSIS, UMR 6168, CNRS,
Anatomically Informed Convolution Kernels for the Projection of fmri Data on the Cortical Surface Grégory Operto 1,Rémy Bulot 1,Jean-LucAnton 2, and Olivier Coulon 1 1 Laboratoire LSIS, UMR 6168, CNRS,
SPM Course! Single Subject Analysis
 SPM Course! Single Subject Analysis Practical Session Dr. Jakob Heinzle & Dr. Frederike Petzschner & Dr. Lionel Rigoux Hands up: Who has programming experience with Matlab? Who has analyzed an fmri experiment
SPM Course! Single Subject Analysis Practical Session Dr. Jakob Heinzle & Dr. Frederike Petzschner & Dr. Lionel Rigoux Hands up: Who has programming experience with Matlab? Who has analyzed an fmri experiment
An empirical comparison of surface-based and volume-based group studies in neuroimaging
 An empirical comparison of surface-based and volume-based group studies in neuroimaging Alan Tucholka, Virgile Fritsch, Jean-Baptiste Poline, Bertrand Thirion To cite this version: Alan Tucholka, Virgile
An empirical comparison of surface-based and volume-based group studies in neuroimaging Alan Tucholka, Virgile Fritsch, Jean-Baptiste Poline, Bertrand Thirion To cite this version: Alan Tucholka, Virgile
Preprocessing of fmri data
 Preprocessing of fmri data Pierre Bellec CRIUGM, DIRO, UdM Flowchart of the NIAK fmri preprocessing pipeline fmri run 1 fmri run N individual datasets CIVET NUC, segmentation, spatial normalization slice
Preprocessing of fmri data Pierre Bellec CRIUGM, DIRO, UdM Flowchart of the NIAK fmri preprocessing pipeline fmri run 1 fmri run N individual datasets CIVET NUC, segmentation, spatial normalization slice
NA-MIC National Alliance for Medical Image Computing fmri Data Analysis
 NA-MIC fmri Data Analysis Sonia Pujol, Ph.D. Wendy Plesniak, Ph.D. Randy Gollub, M.D., Ph.D. Acknowledgments NIH U54EB005149 Neuroimage Analysis Center NIH P41RR013218 FIRST Biomedical Informatics Research
NA-MIC fmri Data Analysis Sonia Pujol, Ph.D. Wendy Plesniak, Ph.D. Randy Gollub, M.D., Ph.D. Acknowledgments NIH U54EB005149 Neuroimage Analysis Center NIH P41RR013218 FIRST Biomedical Informatics Research
Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question.
 (1) conn - Functional connectivity toolbox v1.0 Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question. 2) in "Basic" enter "1" subject, "6" seconds
(1) conn - Functional connectivity toolbox v1.0 Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question. 2) in "Basic" enter "1" subject, "6" seconds
Analysis of group fmri datasets: Voxel-based techniques and beyond
 Jirfni 2009, Marseille Analysis of group fmri datasets: Voxel-based techniques and beyond Bertrand Thirion INRIA Saclay Ile de France, PARIETAL team, Neurospin bertrand.thirion@inria.fr 1/80 Outline Background
Jirfni 2009, Marseille Analysis of group fmri datasets: Voxel-based techniques and beyond Bertrand Thirion INRIA Saclay Ile de France, PARIETAL team, Neurospin bertrand.thirion@inria.fr 1/80 Outline Background
ASAP_2.0 (Automatic Software for ASL Processing) USER S MANUAL
 ASAP_2.0 (Automatic Software for ASL Processing) USER S MANUAL ASAP was developed as part of the COST Action "Arterial Spin Labelling Initiative in Dementia (AID)" by: Department of Neuroimaging, Institute
ASAP_2.0 (Automatic Software for ASL Processing) USER S MANUAL ASAP was developed as part of the COST Action "Arterial Spin Labelling Initiative in Dementia (AID)" by: Department of Neuroimaging, Institute
Event-related design efficiency and How to plot fmri time series. Sepideh Sadaghiani NeuroSpin Methods Meeting 15. Sept. 2008
 Event-related design efficiency and How to plot fmri time series Sepideh Sadaghiani NeuroSpin Methods Meeting 15. Sept. 2008 Event-related averaging or FIR? event-related fmri ability to average responses
Event-related design efficiency and How to plot fmri time series Sepideh Sadaghiani NeuroSpin Methods Meeting 15. Sept. 2008 Event-related averaging or FIR? event-related fmri ability to average responses
CONN fmri functional connectivity toolbox
 CONN fmri functional connectivity toolbox Gabrieli Lab. McGovern Institute for Brain Research Massachusetts Institute of Technology http://www.nitrc.org/projects/conn Susan Whitfield-Gabrieli Alfonso Nieto-Castanon
CONN fmri functional connectivity toolbox Gabrieli Lab. McGovern Institute for Brain Research Massachusetts Institute of Technology http://www.nitrc.org/projects/conn Susan Whitfield-Gabrieli Alfonso Nieto-Castanon
User s Guide Neuroimage Processing ToolKit (NPTK) Version 2.0 fmri Registration Software Pipeline for Functional Localization
 User s Guide Neuroimage Processing ToolKit (NPTK) Version 2.0 fmri Registration Software Pipeline for Functional Localization Software Written by Ali Gholipour SIP Lab, UTD, 2005-2010 Revision 2.0 February
User s Guide Neuroimage Processing ToolKit (NPTK) Version 2.0 fmri Registration Software Pipeline for Functional Localization Software Written by Ali Gholipour SIP Lab, UTD, 2005-2010 Revision 2.0 February
Tutorial MICCAI'04. fmri data analysis: state of the art and future challenges. Jean-Baptiste Poline Philippe Ciuciu Alexis Roche
 Tutorial MICCAI'04 fmri data analysis: state of the art and future challenges Jean-Baptiste Poline Philippe Ciuciu Alexis Roche CEA/SHFJ, Orsay (France) Goal of this tutorial and plan I. Talk 1: will orientate
Tutorial MICCAI'04 fmri data analysis: state of the art and future challenges Jean-Baptiste Poline Philippe Ciuciu Alexis Roche CEA/SHFJ, Orsay (France) Goal of this tutorial and plan I. Talk 1: will orientate
Function-Structure Integration in FreeSurfer
 Function-Structure Integration in FreeSurfer Outline Function-Structure Integration Function-Structure Registration in FreeSurfer fmri Analysis Preprocessing First-Level Analysis Higher-Level (Group) Analysis
Function-Structure Integration in FreeSurfer Outline Function-Structure Integration Function-Structure Registration in FreeSurfer fmri Analysis Preprocessing First-Level Analysis Higher-Level (Group) Analysis
OHBA M/EEG Analysis Workshop. Mark Woolrich Diego Vidaurre Andrew Quinn Romesh Abeysuriya Robert Becker
 OHBA M/EEG Analysis Workshop Mark Woolrich Diego Vidaurre Andrew Quinn Romesh Abeysuriya Robert Becker Workshop Schedule Tuesday Session 1: Preprocessing, manual and automatic pipelines Session 2: Task
OHBA M/EEG Analysis Workshop Mark Woolrich Diego Vidaurre Andrew Quinn Romesh Abeysuriya Robert Becker Workshop Schedule Tuesday Session 1: Preprocessing, manual and automatic pipelines Session 2: Task
seam Documentation Release 0.0 Scott Burns
 seam Documentation Release 0.0 Scott Burns April 08, 2014 Contents 1 Contents 3 1.1 Installation................................................ 3 1.2 Philosophy................................................
seam Documentation Release 0.0 Scott Burns April 08, 2014 Contents 1 Contents 3 1.1 Installation................................................ 3 1.2 Philosophy................................................
TractoR and Other Software
 TractoR and Other Software Jon Clayden DIBS Teaching Seminar, 11 Dec 2015 Photo by José Martín Ramírez Carrasco https://www.behance.net/martini_rc TractoR A set of R packages Additional
TractoR and Other Software Jon Clayden DIBS Teaching Seminar, 11 Dec 2015 Photo by José Martín Ramírez Carrasco https://www.behance.net/martini_rc TractoR A set of R packages Additional
2. Creating Field Maps Using the Field Map GUI (Version 2.0) in SPM5
 1. Introduction This manual describes how to use the Field Map Toolbox Version 2.0 for creating unwrapped field maps that can be used to do geometric distortion correction of EPI images in SPM5. 1. 1.
1. Introduction This manual describes how to use the Field Map Toolbox Version 2.0 for creating unwrapped field maps that can be used to do geometric distortion correction of EPI images in SPM5. 1. 1.
7/15/2016 ARE YOUR ANALYSES TOO WHY IS YOUR ANALYSIS PARAMETRIC? PARAMETRIC? That s not Normal!
 ARE YOUR ANALYSES TOO PARAMETRIC? That s not Normal! Martin M Monti http://montilab.psych.ucla.edu WHY IS YOUR ANALYSIS PARAMETRIC? i. Optimal power (defined as the probability to detect a real difference)
ARE YOUR ANALYSES TOO PARAMETRIC? That s not Normal! Martin M Monti http://montilab.psych.ucla.edu WHY IS YOUR ANALYSIS PARAMETRIC? i. Optimal power (defined as the probability to detect a real difference)
BrainSuite Lab Exercises. presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014
 BrainSuite Lab Exercises presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 1. Opening and Displaying an MRI Start BrainSuite Drag and drop the T1 image from the native space
BrainSuite Lab Exercises presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 1. Opening and Displaying an MRI Start BrainSuite Drag and drop the T1 image from the native space
Surface-Based Structural Group Analysis of fmri Data
 Surface-Based Structural Group Analysis of fmri Data Grégory Operto 1,Cédric Clouchoux 1,Rémy Bulot 1,Jean-LucAnton 2, and Olivier Coulon 1 1 Laboratoire LSIS, UMR CNRS 6168, Marseille, France gregory.operto@univmed.fr
Surface-Based Structural Group Analysis of fmri Data Grégory Operto 1,Cédric Clouchoux 1,Rémy Bulot 1,Jean-LucAnton 2, and Olivier Coulon 1 1 Laboratoire LSIS, UMR CNRS 6168, Marseille, France gregory.operto@univmed.fr
Basic Introduction to Data Analysis. Block Design Demonstration. Robert Savoy
 Basic Introduction to Data Analysis Block Design Demonstration Robert Savoy Sample Block Design Experiment Demonstration Use of Visual and Motor Task Separability of Responses Combined Visual and Motor
Basic Introduction to Data Analysis Block Design Demonstration Robert Savoy Sample Block Design Experiment Demonstration Use of Visual and Motor Task Separability of Responses Combined Visual and Motor
Image Processing for fmri John Ashburner. Wellcome Trust Centre for Neuroimaging, 12 Queen Square, London, UK.
 Iage Processing for fmri John Ashburner Wellcoe Trust Centre for Neuroiaging, 12 Queen Square, London, UK. Contents * Preliinaries * Rigid-Body and Affine Transforations * Optiisation and Objective Functions
Iage Processing for fmri John Ashburner Wellcoe Trust Centre for Neuroiaging, 12 Queen Square, London, UK. Contents * Preliinaries * Rigid-Body and Affine Transforations * Optiisation and Objective Functions
White Pixel Artifact. Caused by a noise spike during acquisition Spike in K-space <--> sinusoid in image space
 White Pixel Artifact Caused by a noise spike during acquisition Spike in K-space sinusoid in image space Susceptibility Artifacts Off-resonance artifacts caused by adjacent regions with different
White Pixel Artifact Caused by a noise spike during acquisition Spike in K-space sinusoid in image space Susceptibility Artifacts Off-resonance artifacts caused by adjacent regions with different
SPM99 fmri Data Analysis Workbook
 SPM99 fmri Data Analysis Workbook This file is a description of the steps needed to use SPM99 analyze a fmri data set from a single subject using a simple on/off activation paradigm. There are two parts
SPM99 fmri Data Analysis Workbook This file is a description of the steps needed to use SPM99 analyze a fmri data set from a single subject using a simple on/off activation paradigm. There are two parts
An independent component analysis based tool for exploring functional connections in the brain
 An independent component analysis based tool for exploring functional connections in the brain S. M. Rolfe a, L. Finney b, R. F. Tungaraza b, J. Guan b, L.G. Shapiro b, J. F. Brinkely b, A. Poliakov c,
An independent component analysis based tool for exploring functional connections in the brain S. M. Rolfe a, L. Finney b, R. F. Tungaraza b, J. Guan b, L.G. Shapiro b, J. F. Brinkely b, A. Poliakov c,
TractoR and Other Software
 TractoR and Other Software Jon Clayden DIBS Teaching Seminar, 15 Nov 2016 Photo by José Martín Ramírez Carrasco https://www.behance.net/martini_rc TractoR A platform for multimodal
TractoR and Other Software Jon Clayden DIBS Teaching Seminar, 15 Nov 2016 Photo by José Martín Ramírez Carrasco https://www.behance.net/martini_rc TractoR A platform for multimodal
Anatomic parcellation based on DTI data with FSL taking the example of SMA/preSMA
 Groupe de Travail IRMf/MEG 02/2014 Anatomic parcellation based on DTI data with FSL taking the example of SMA/preSMA Magdalena Wutte, Lucile Brun & Boris Burle SMA/preSMA Parcellation - what for? - connectivity
Groupe de Travail IRMf/MEG 02/2014 Anatomic parcellation based on DTI data with FSL taking the example of SMA/preSMA Magdalena Wutte, Lucile Brun & Boris Burle SMA/preSMA Parcellation - what for? - connectivity
n o r d i c B r a i n E x Tutorial DSC Module
 m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DSC Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DSC Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
Brain Explorer for Connectomic Analysis (BECA) Software Manual
 Brain Explorer for Connectomic Analysis (BECA) Software Manual Table of Contents 1. Launch the application... 2 2. Tractography Visualization... 3 3. Gray Matter Visualization... 5 4. Fiber classification
Brain Explorer for Connectomic Analysis (BECA) Software Manual Table of Contents 1. Launch the application... 2 2. Tractography Visualization... 3 3. Gray Matter Visualization... 5 4. Fiber classification
Math in image processing
 Math in image processing Math in image processing Nyquist theorem Math in image processing Discrete Fourier Transformation Math in image processing Image enhancement: scaling Math in image processing Image
Math in image processing Math in image processing Nyquist theorem Math in image processing Discrete Fourier Transformation Math in image processing Image enhancement: scaling Math in image processing Image
FSL Workshop Session 3 David Smith & John Clithero
 FSL Workshop 12.09.08 Session 3 David Smith & John Clithero What is MELODIC? Probabilistic ICA Improves upon standard ICA Allows for inference Avoids over-fitting Three stage process ( ppca ) 1.) Dimension
FSL Workshop 12.09.08 Session 3 David Smith & John Clithero What is MELODIC? Probabilistic ICA Improves upon standard ICA Allows for inference Avoids over-fitting Three stage process ( ppca ) 1.) Dimension
0.1. Setting up the system path to allow use of BIAC XML headers (BXH). Depending on the computer(s), you may only have to do this once.
 Week 3 Exercises Last week you began working with MR data, both in the form of anatomical images and functional time series. This week we will discuss some concepts related to the idea of fmri data as
Week 3 Exercises Last week you began working with MR data, both in the form of anatomical images and functional time series. This week we will discuss some concepts related to the idea of fmri data as
Correction of Partial Volume Effects in Arterial Spin Labeling MRI
 Correction of Partial Volume Effects in Arterial Spin Labeling MRI By: Tracy Ssali Supervisors: Dr. Keith St. Lawrence and Udunna Anazodo Medical Biophysics 3970Z Six Week Project April 13 th 2012 Introduction
Correction of Partial Volume Effects in Arterial Spin Labeling MRI By: Tracy Ssali Supervisors: Dr. Keith St. Lawrence and Udunna Anazodo Medical Biophysics 3970Z Six Week Project April 13 th 2012 Introduction
Surface-based Analysis: Inter-subject Registration and Smoothing
 Surface-based Analysis: Inter-subject Registration and Smoothing Outline Exploratory Spatial Analysis Coordinate Systems 3D (Volumetric) 2D (Surface-based) Inter-subject registration Volume-based Surface-based
Surface-based Analysis: Inter-subject Registration and Smoothing Outline Exploratory Spatial Analysis Coordinate Systems 3D (Volumetric) 2D (Surface-based) Inter-subject registration Volume-based Surface-based
Neuroimaging Group Pipeline Quick Start Manual
 Centre for Healthy Brain Ageing (CHeBA) Neuroimaging Group Pipeline Quick Start Manual UBO Detector UBO Detector is a cluster-based white matter hyperintensity (WMH) extraction pipeline based on k-nearest
Centre for Healthy Brain Ageing (CHeBA) Neuroimaging Group Pipeline Quick Start Manual UBO Detector UBO Detector is a cluster-based white matter hyperintensity (WMH) extraction pipeline based on k-nearest
Methods for data preprocessing
 Methods for data preprocessing John Ashburner Wellcome Trust Centre for Neuroimaging, 12 Queen Square, London, UK. Overview Voxel-Based Morphometry Morphometry in general Volumetrics VBM preprocessing
Methods for data preprocessing John Ashburner Wellcome Trust Centre for Neuroimaging, 12 Queen Square, London, UK. Overview Voxel-Based Morphometry Morphometry in general Volumetrics VBM preprocessing
6 credits. BMSC-GA Practical Magnetic Resonance Imaging II
 BMSC-GA 4428 - Practical Magnetic Resonance Imaging II 6 credits Course director: Ricardo Otazo, PhD Course description: This course is a practical introduction to image reconstruction, image analysis
BMSC-GA 4428 - Practical Magnetic Resonance Imaging II 6 credits Course director: Ricardo Otazo, PhD Course description: This course is a practical introduction to image reconstruction, image analysis
A Graph Theoretical Network Analysis Toolbox
 A Graph Theoretical Network Analysis Toolbox Reference Manual for GRETNA (v2.0) June 2017 National Key Laboratory of Cognitive Neuroscience and Learning Beijing Key Laboratory of Brain Imaging and Connectomics
A Graph Theoretical Network Analysis Toolbox Reference Manual for GRETNA (v2.0) June 2017 National Key Laboratory of Cognitive Neuroscience and Learning Beijing Key Laboratory of Brain Imaging and Connectomics
- Graphical editing of user montages for convenient data review - Import of user-defined file formats using generic reader
 Data review and processing Source montages and 3D whole-head mapping Onset of epileptic seizure with 3D whole-head maps and hemispheric comparison of density spectral arrays (DSA) Graphical display of
Data review and processing Source montages and 3D whole-head mapping Onset of epileptic seizure with 3D whole-head maps and hemispheric comparison of density spectral arrays (DSA) Graphical display of
syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets
 syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets Julien Gervais; Lisa Chuah Siemens Healthcare, Magnetic
syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets Julien Gervais; Lisa Chuah Siemens Healthcare, Magnetic
Introduction to Neuroimaging Janaina Mourao-Miranda
 Introduction to Neuroimaging Janaina Mourao-Miranda Neuroimaging techniques have changed the way neuroscientists address questions about functional anatomy, especially in relation to behavior and clinical
Introduction to Neuroimaging Janaina Mourao-Miranda Neuroimaging techniques have changed the way neuroscientists address questions about functional anatomy, especially in relation to behavior and clinical
Correction for multiple comparisons. Cyril Pernet, PhD SBIRC/SINAPSE University of Edinburgh
 Correction for multiple comparisons Cyril Pernet, PhD SBIRC/SINAPSE University of Edinburgh Overview Multiple comparisons correction procedures Levels of inferences (set, cluster, voxel) Circularity issues
Correction for multiple comparisons Cyril Pernet, PhD SBIRC/SINAPSE University of Edinburgh Overview Multiple comparisons correction procedures Levels of inferences (set, cluster, voxel) Circularity issues
Where are we now? Structural MRI processing and analysis
 Where are we now? Structural MRI processing and analysis Pierre-Louis Bazin bazin@cbs.mpg.de Leipzig, Germany Structural MRI processing: why bother? Just use the standards? SPM FreeSurfer FSL However:
Where are we now? Structural MRI processing and analysis Pierre-Louis Bazin bazin@cbs.mpg.de Leipzig, Germany Structural MRI processing: why bother? Just use the standards? SPM FreeSurfer FSL However:
Detecting White Matter Lesions in Lupus
 Slicer3 Training Compendium Detecting White Matter Lesions in Lupus Version 2.1 6/25/2009 H. Jeremy Bockholt Mark Scully -1- Learning objective This tutorial demonstrates an automated, multi-level method
Slicer3 Training Compendium Detecting White Matter Lesions in Lupus Version 2.1 6/25/2009 H. Jeremy Bockholt Mark Scully -1- Learning objective This tutorial demonstrates an automated, multi-level method
Data pre-processing framework in SPM. Bogdan Draganski
 Data pre-processing fraework in SPM Bogdan Draganski Outline Why do we need pre-processing? Overview Structural MRI pre-processing fmri pre-processing Why do we need pre-processing? What do we want? Reason
Data pre-processing fraework in SPM Bogdan Draganski Outline Why do we need pre-processing? Overview Structural MRI pre-processing fmri pre-processing Why do we need pre-processing? What do we want? Reason
Surface Projection Method for Visualizing Volumetric Data
 Surface Projection Method for Visualizing Volumetric Data by Peter Lincoln A senior thesis submitted in partial fulfillment of the requirements for the degree of Bachelor of Science With Departmental Honors
Surface Projection Method for Visualizing Volumetric Data by Peter Lincoln A senior thesis submitted in partial fulfillment of the requirements for the degree of Bachelor of Science With Departmental Honors
This exercise uses one anatomical data set (ANAT1) and two functional data sets (FUNC1 and FUNC2).
 Exploring Brain Anatomy This week s exercises will let you explore the anatomical organization of the brain to learn some of its basic properties, as well as the location of different structures. The human
Exploring Brain Anatomy This week s exercises will let you explore the anatomical organization of the brain to learn some of its basic properties, as well as the location of different structures. The human
Preprocessing II: Between Subjects John Ashburner
 Preprocessing II: Between Subjects John Ashburner Pre-processing Overview Statistics or whatever fmri time-series Anatomical MRI Template Smoothed Estimate Spatial Norm Motion Correct Smooth Coregister
Preprocessing II: Between Subjects John Ashburner Pre-processing Overview Statistics or whatever fmri time-series Anatomical MRI Template Smoothed Estimate Spatial Norm Motion Correct Smooth Coregister
ANALYSIS OF FUNCTIONAL MAGNETIC RESONANCE IMAGING DATA USING SPM99: VOXEL-BASED MORPHOMETRY DONNA ROSE ADDIS
 Donna Rose Addis, TWRI, May 2004 1 ANALYSIS OF FUNCTIONAL MAGNETIC RESONANCE IMAGING DATA USING SPM99: VOXEL-BASED MORPHOMETRY DONNA ROSE ADDIS DEPT. OF PSYCHOLOGY, UNIVERSITY OF TORONTO TORONTO WESTERN
Donna Rose Addis, TWRI, May 2004 1 ANALYSIS OF FUNCTIONAL MAGNETIC RESONANCE IMAGING DATA USING SPM99: VOXEL-BASED MORPHOMETRY DONNA ROSE ADDIS DEPT. OF PSYCHOLOGY, UNIVERSITY OF TORONTO TORONTO WESTERN
fmri Analysis Sackler Ins2tute 2011
 fmri Analysis Sackler Ins2tute 2011 How do we get from this to this? How do we get from this to this? And what are those colored blobs we re all trying to see, anyway? Raw fmri data straight from the scanner
fmri Analysis Sackler Ins2tute 2011 How do we get from this to this? How do we get from this to this? And what are those colored blobs we re all trying to see, anyway? Raw fmri data straight from the scanner
fmri Basics: Single Subject Analysis
 fmri Basics: Single Subject Analysis This session is intended to give an overview of the basic process of setting up a general linear model for a single subject. This stage of the analysis is also variously
fmri Basics: Single Subject Analysis This session is intended to give an overview of the basic process of setting up a general linear model for a single subject. This stage of the analysis is also variously
Detecting Changes In Non-Isotropic Images
 Detecting Changes In Non-Isotropic Images K.J. Worsley 1, M. Andermann 1, T. Koulis 1, D. MacDonald, 2 and A.C. Evans 2 August 4, 1999 1 Department of Mathematics and Statistics, 2 Montreal Neurological
Detecting Changes In Non-Isotropic Images K.J. Worsley 1, M. Andermann 1, T. Koulis 1, D. MacDonald, 2 and A.C. Evans 2 August 4, 1999 1 Department of Mathematics and Statistics, 2 Montreal Neurological
Playing with data from lab
 Playing with data from lab Getting data off the scanner From the Patient Browser, select the folder for the study you want (or within that study, the set of images you want), and then from the Transfer
Playing with data from lab Getting data off the scanner From the Patient Browser, select the folder for the study you want (or within that study, the set of images you want), and then from the Transfer
How to make ROI files
 How to make ROI files Ver. 1.0 2012/08/07 Introduction Functional ROI Structural ROI Hand-made ROI History Contact 2 Introduction This manual explains how to make ROI (Region Of Interest) files that you
How to make ROI files Ver. 1.0 2012/08/07 Introduction Functional ROI Structural ROI Hand-made ROI History Contact 2 Introduction This manual explains how to make ROI (Region Of Interest) files that you
MB-EPI PCASL. Release Notes for Version February 2015
 MB-EPI PCASL Release Notes for Version 1.0 20 February 2015 1 Background High-resolution arterial spin labeling (ASL) imaging is highly desirable in both neuroscience research and clinical applications
MB-EPI PCASL Release Notes for Version 1.0 20 February 2015 1 Background High-resolution arterial spin labeling (ASL) imaging is highly desirable in both neuroscience research and clinical applications
Input tab. Insert the path to the DTI Image that you want to tract.
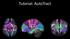 Tutorial: AutoTract Input tab Insert the path to the DTI Image that you want to tract. Input tab (optional)if you already have a WM/CSF mask you want to use to process the tracts, insert the path here.
Tutorial: AutoTract Input tab Insert the path to the DTI Image that you want to tract. Input tab (optional)if you already have a WM/CSF mask you want to use to process the tracts, insert the path here.
Basic Python 3 Programming (Theory & Practical)
 Basic Python 3 Programming (Theory & Practical) Length Delivery Method : 5 Days : Instructor-led (Classroom) Course Overview This Python 3 Programming training leads the student from the basics of writing
Basic Python 3 Programming (Theory & Practical) Length Delivery Method : 5 Days : Instructor-led (Classroom) Course Overview This Python 3 Programming training leads the student from the basics of writing
DynaConn Users Guide. Dynamic Function Connectivity Graphical User Interface. Last saved by John Esquivel Last update: 3/27/14 4:26 PM
 DynaConn Users Guide Dynamic Function Connectivity Graphical User Interface Last saved by John Esquivel Last update: 3/27/14 4:26 PM Table Of Contents Chapter 1 Introduction... 1 1.1 Introduction to this
DynaConn Users Guide Dynamic Function Connectivity Graphical User Interface Last saved by John Esquivel Last update: 3/27/14 4:26 PM Table Of Contents Chapter 1 Introduction... 1 1.1 Introduction to this
Group (Level 2) fmri Data Analysis - Lab 4
 Group (Level 2) fmri Data Analysis - Lab 4 Index Goals of this Lab Before Getting Started The Chosen Ten Checking Data Quality Create a Mean Anatomical of the Group Group Analysis: One-Sample T-Test Examine
Group (Level 2) fmri Data Analysis - Lab 4 Index Goals of this Lab Before Getting Started The Chosen Ten Checking Data Quality Create a Mean Anatomical of the Group Group Analysis: One-Sample T-Test Examine
Using the History Palette Part 2 - Create & Convert Quick Scripts
 Using the History Palette Part 2 - Create & Convert Quick Scripts By JP Kabala Quick Scripts are such a useful and intuitive new feature of the History Palette that it really is worth your while to take
Using the History Palette Part 2 - Create & Convert Quick Scripts By JP Kabala Quick Scripts are such a useful and intuitive new feature of the History Palette that it really is worth your while to take
How to create a head model
 How to create a head model This document describes the command line tools: mri2mesh: Central tool to reconstruct a head model from T1w and T2w data dwi2cond: Reconstruct conductivity tensors for brain
How to create a head model This document describes the command line tools: mri2mesh: Central tool to reconstruct a head model from T1w and T2w data dwi2cond: Reconstruct conductivity tensors for brain
Getting Started Guide
 Getting Started Guide Version 2.5 for BVQX 1.9 Rainer Goebel, Henk Jansma and Jochen Seitz Copyright 2007 Brain Innovation B.V. 2 Contents Preface...4 The Objects Tutorial...5 Scanning session information...5
Getting Started Guide Version 2.5 for BVQX 1.9 Rainer Goebel, Henk Jansma and Jochen Seitz Copyright 2007 Brain Innovation B.V. 2 Contents Preface...4 The Objects Tutorial...5 Scanning session information...5
Previously. Iteration. Date and time structures. Modularisation.
 2017 2018 Previously Iteration. Date and time structures. Modularisation. Today File handling. Reading and writing files. In order for a program to work with a file, the program must create a file object
2017 2018 Previously Iteration. Date and time structures. Modularisation. Today File handling. Reading and writing files. In order for a program to work with a file, the program must create a file object
BrainVoyager TM. Getting Started Guide. Version 3.0. for BV 21
 BrainVoyager TM Getting Started Guide Version 3.0 for BV 21 Rainer Goebel, Henk Jansma, Caroline Benjamins, Judith Eck, Hester Breman and Armin Heinecke Copyright 2018 Brain Innovation B.V. Contents About
BrainVoyager TM Getting Started Guide Version 3.0 for BV 21 Rainer Goebel, Henk Jansma, Caroline Benjamins, Judith Eck, Hester Breman and Armin Heinecke Copyright 2018 Brain Innovation B.V. Contents About
Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm
 Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm Lin Chen and Gudrun Wagenknecht Central Institute for Electronics, Research Center Jülich, Jülich, Germany Email: l.chen@fz-juelich.de
Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm Lin Chen and Gudrun Wagenknecht Central Institute for Electronics, Research Center Jülich, Jülich, Germany Email: l.chen@fz-juelich.de
Artifact Detection and Repair: Overview and Sample Outputs
 Artifact Detection and Repair: Overview and Sample Outputs Paul Mazaika February 2007 Programs originated in Gabrieli Neuroscience Laboratory, updated and enhanced at Center for Interdisciplinary Brain
Artifact Detection and Repair: Overview and Sample Outputs Paul Mazaika February 2007 Programs originated in Gabrieli Neuroscience Laboratory, updated and enhanced at Center for Interdisciplinary Brain
TMS NEURONAVIGATOR (ZEBRIS)
 TMS NEURONAVIGATOR (ZEBRIS) Getting Started Guide Copyright 2015: Brain Innovation bv Table of contents About This Document...6 Preface...6 Hardware and Software Requirements...6 Contact...6 Tutorial
TMS NEURONAVIGATOR (ZEBRIS) Getting Started Guide Copyright 2015: Brain Innovation bv Table of contents About This Document...6 Preface...6 Hardware and Software Requirements...6 Contact...6 Tutorial
Supplementary methods
 Supplementary methods This section provides additional technical details on the sample, the applied imaging and analysis steps and methods. Structural imaging Trained radiographers placed all participants
Supplementary methods This section provides additional technical details on the sample, the applied imaging and analysis steps and methods. Structural imaging Trained radiographers placed all participants
NEURO M203 & BIOMED M263 WINTER 2014
 NEURO M203 & BIOMED M263 WINTER 2014 MRI Lab 2: Neuroimaging Connectivity Lab In today s lab we will work with sample diffusion imaging data and the group averaged fmri data collected during your scanning
NEURO M203 & BIOMED M263 WINTER 2014 MRI Lab 2: Neuroimaging Connectivity Lab In today s lab we will work with sample diffusion imaging data and the group averaged fmri data collected during your scanning
The alignment process uses two alignment methods: (1) the mrvista function, rxalign, for a rough alignment, and then (2) KNK's alignment for
 Alignment (Allegra) Download example alignment script here. Anatomical images are collected at much higher resolution (isotropic 1 mm voxels) than functional images (isotropic 2 mm voxels; sometimes 2.5).
Alignment (Allegra) Download example alignment script here. Anatomical images are collected at much higher resolution (isotropic 1 mm voxels) than functional images (isotropic 2 mm voxels; sometimes 2.5).
Tutorial BOLD Module
 m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial BOLD Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial BOLD Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
MSI 2D Viewer Software Guide
 MSI 2D Viewer Software Guide Page:1 DISCLAIMER We have used reasonable effort to include accurate and up-to-date information in this manual; it does not, however, make any warranties, conditions or representations
MSI 2D Viewer Software Guide Page:1 DISCLAIMER We have used reasonable effort to include accurate and up-to-date information in this manual; it does not, however, make any warranties, conditions or representations
Walk Through of CERR Capabilities. CERR: Introduction. Outline. Getting CERR: Control Panel. Documentation Support Community
 Walk Through of CERR Capabilities Aditya P. Apte, Ph.D. Department of Medical Physics Memorial Sloan Kettering Cancer Center New York aptea@mskcc.org AAPM 2015, July 15, 2015 CERR: Computational Environment
Walk Through of CERR Capabilities Aditya P. Apte, Ph.D. Department of Medical Physics Memorial Sloan Kettering Cancer Center New York aptea@mskcc.org AAPM 2015, July 15, 2015 CERR: Computational Environment
Introduction to Visualization on Stampede
 Introduction to Visualization on Stampede Aaron Birkland Cornell CAC With contributions from TACC visualization training materials Parallel Computing on Stampede June 11, 2013 From data to Insight Data
Introduction to Visualization on Stampede Aaron Birkland Cornell CAC With contributions from TACC visualization training materials Parallel Computing on Stampede June 11, 2013 From data to Insight Data
n o r d i c B r a i n E x Tutorial DTI Module
 m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DTI Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DTI Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
