Demo 1: Free text search of ENCODE data
|
|
|
- Georgia Hood
- 5 years ago
- Views:
Transcription
1 Demo 1: Free text search of ENCODE data 1. Go to h(ps:// 2. Enter skin into the search box on the upper right hand corner 3. All matches on the website will be shown 4. Select Experiments
2 Demo 1: Free text search of ENCODE data (cont.) 5. View all assays that contain a match to skin 6. To limit to ENCODE data, select the Project ENCODE
3 Demo 2: Browsing and filtering of ENCODE data 1. Go to h(ps:// 2. Under the Data menu bar, select Search 3. All publicly available assays are shown. The categories on the ler are metadata describing the assays. They can be used to filter your results.
4 5. View all ENCODE assays that are filtered by selectng skin of body. Note that the results are not straight text matches. For example, fibroblast of arm, and keratnocytes are included in the results list.
5 Demo 3: Combine search & filter of ENCODE data 1. Go to h(ps:// 2. Enter skin into the search box on the upper right hand corner 3. All matches on the website will be shown 4. Select Experiments
6 5. View all ENCODE assays that contain a match to skin 6. If looking for RNA- seq data from adult samples, select RNA- seq under Assay and adult under Life Stage.
7 Demo 4: Region Search 1. Use the drop down menu from Data to select Search by Region 2. Begin typing your gene name, id, or coordinates in the box. We will search for BRCA1. After selecting your gene or coordinate, click on the Search button.
8 Demo 4: Region Search (congnued) 3. You now see all the experiments on the site that have BED files with coordinates that intersect that region. Since this is BRCA1, let s look to see what is happening with MCF- 7. It is not in the immediate biosample type list, select see more
9 Demo 4: Region Search (congnued) 4. You have narrowed your search enough to have both the Visualize and Download bu(ons available
10 Demo 5: Visualize data 1. Get the list of assays illustrated in Demo Select Visualize bu(on on the upper right corner. This will automatcally create a trackhub to visualize the results 3. A drop down menu with show available genome assemblies. Select GRCh38
11 Demo 5: Visualize data (congnued) 4. The data is now visualized via a trackhub in the UCSC Genome Browser.
12 Demo 5: Visualize data (congnued) 5. The track hub is listed as Hub (search) 6. Hover over tracks to view file- related metadata. 7. Click on ler hand grouping to configure tracks and view more metadata, including file download links and links back to the ENCODE Portal.
13 8. ENCODE metadata for the track hub
14 Demo 6: Batch download data 1. Get the list of assays illustrated in Demo Select Download bu(on on the upper right corner. 3. InstrucTons on the command to batch download files is displayed. Click on Download again. This will download a file called files.txt
15 Demo 6: Batch download data (congnued) 4. Open files.txt. This includes a list of links to all the files for those experiments. The first link contains the metadata. You can select the subset of files by using the available data facet. 5. Transfer this file to your server and use xargs n 1 curl O L < files.txt
16 Exercises using the ENCODE Portal Exercise 1: ChIP- seq assays How many ChIP- seq assays are available against H3K27me3 in mouse How many unique antbodies are used? Of the antbodies used, which one(s) have been fully characterized to current ENCODE standards? Exercise 2: Controls What is the accession for control experiment for ENCSR778SIU? How many assays use this control? Exercise 3: Recently released data What was the total number of assays released in June 2015? How many of each kind? Exercise 4: Files What are the accession(s) to the fastq files for biological replicate 1 in assay ENCSR000AFI? What is the accession of the alignment (bam) file made from them? Exercise 5: Protein factors Which assays have been performed against IGF2BP1?
17 REST API exercises using the ENCODE Portal Pre- requisites: download or install these tools to help you get started A JSON pre(y- printer plugin for your web browser, such as JSONView (for Chrome or Firefox) or JSON Forma(er (for Safari) A few python modules pip install requests pip install json pip install jsonschema Help document: h(ps:// api/ Sample script: h(ps://github.com/encode- DCC/submission_sample_scripts/blob/master/get.py Try the exercises listed on the previous page. They can be performed programmatcally.
Demo 1: Free text search of ENCODE data
 Demo 1: Free text search of ENCODE data 1. Go to h(ps://www.encodeproject.org 2. Enter skin into the search box on the upper right hand corner 3. All matches on the website will be shown 4. Select Experiments
Demo 1: Free text search of ENCODE data 1. Go to h(ps://www.encodeproject.org 2. Enter skin into the search box on the upper right hand corner 3. All matches on the website will be shown 4. Select Experiments
How To: Run the ENCODE histone ChIP- seq analysis pipeline on DNAnexus
 How To: Run the ENCODE histone ChIP- seq analysis pipeline on DNAnexus Overview: In this exercise, we will run the ENCODE Uniform Processing ChIP- seq Pipeline on a small test dataset containing reads
How To: Run the ENCODE histone ChIP- seq analysis pipeline on DNAnexus Overview: In this exercise, we will run the ENCODE Uniform Processing ChIP- seq Pipeline on a small test dataset containing reads
ChIP-seq hands-on practical using Galaxy
 ChIP-seq hands-on practical using Galaxy In this exercise we will cover some of the basic NGS analysis steps for ChIP-seq using the Galaxy framework: Quality control Mapping of reads using Bowtie2 Peak-calling
ChIP-seq hands-on practical using Galaxy In this exercise we will cover some of the basic NGS analysis steps for ChIP-seq using the Galaxy framework: Quality control Mapping of reads using Bowtie2 Peak-calling
ChIP-seq hands-on practical using Galaxy
 ChIP-seq hands-on practical using Galaxy In this exercise we will cover some of the basic NGS analysis steps for ChIP-seq using the Galaxy framework: Quality control Mapping of reads using Bowtie2 Peak-calling
ChIP-seq hands-on practical using Galaxy In this exercise we will cover some of the basic NGS analysis steps for ChIP-seq using the Galaxy framework: Quality control Mapping of reads using Bowtie2 Peak-calling
Genomic Analysis with Genome Browsers.
 Genomic Analysis with Genome Browsers http://barc.wi.mit.edu/hot_topics/ 1 Outline Genome browsers overview UCSC Genome Browser Navigating: View your list of regions in the browser Available tracks (eg.
Genomic Analysis with Genome Browsers http://barc.wi.mit.edu/hot_topics/ 1 Outline Genome browsers overview UCSC Genome Browser Navigating: View your list of regions in the browser Available tracks (eg.
Getting Started. April Strand Life Sciences, Inc All rights reserved.
 Getting Started April 2015 Strand Life Sciences, Inc. 2015. All rights reserved. Contents Aim... 3 Demo Project and User Interface... 3 Downloading Annotations... 4 Project and Experiment Creation... 6
Getting Started April 2015 Strand Life Sciences, Inc. 2015. All rights reserved. Contents Aim... 3 Demo Project and User Interface... 3 Downloading Annotations... 4 Project and Experiment Creation... 6
RNA-Seq in Galaxy: Tuxedo protocol. Igor Makunin, UQ RCC, QCIF
 RNA-Seq in Galaxy: Tuxedo protocol Igor Makunin, UQ RCC, QCIF Acknowledgments Genomics Virtual Lab: gvl.org.au Galaxy for tutorials: galaxy-tut.genome.edu.au Galaxy Australia: galaxy-aust.genome.edu.au
RNA-Seq in Galaxy: Tuxedo protocol Igor Makunin, UQ RCC, QCIF Acknowledgments Genomics Virtual Lab: gvl.org.au Galaxy for tutorials: galaxy-tut.genome.edu.au Galaxy Australia: galaxy-aust.genome.edu.au
Analyzing ChIP- Seq Data in Galaxy
 Analyzing ChIP- Seq Data in Galaxy Lauren Mills RISS ABSTRACT Step- by- step guide to basic ChIP- Seq analysis using the Galaxy platform. Table of Contents Introduction... 3 Links to helpful information...
Analyzing ChIP- Seq Data in Galaxy Lauren Mills RISS ABSTRACT Step- by- step guide to basic ChIP- Seq analysis using the Galaxy platform. Table of Contents Introduction... 3 Links to helpful information...
W ASHU E PI G ENOME B ROWSER
 W ASHU E PI G ENOME B ROWSER Keystone Symposium on DNA and RNA Methylation January 23 rd, 2018 Fairmont Hotel Vancouver, Vancouver, British Columbia, Canada Presenter: Renee Sears and Josh Jang Tutorial
W ASHU E PI G ENOME B ROWSER Keystone Symposium on DNA and RNA Methylation January 23 rd, 2018 Fairmont Hotel Vancouver, Vancouver, British Columbia, Canada Presenter: Renee Sears and Josh Jang Tutorial
W ASHU E PI G ENOME B ROWSER
 Roadmap Epigenomics Workshop W ASHU E PI G ENOME B ROWSER SOT 2016 Satellite Meeting March 17 th, 2016 Ernest N. Morial Convention Center, New Orleans, LA Presenter: Ting Wang Tutorial Overview: WashU
Roadmap Epigenomics Workshop W ASHU E PI G ENOME B ROWSER SOT 2016 Satellite Meeting March 17 th, 2016 Ernest N. Morial Convention Center, New Orleans, LA Presenter: Ting Wang Tutorial Overview: WashU
NGS Data Visualization and Exploration Using IGV
 1 What is Galaxy Galaxy for Bioinformaticians Galaxy for Experimental Biologists Using Galaxy for NGS Analysis NGS Data Visualization and Exploration Using IGV 2 What is Galaxy Galaxy for Bioinformaticians
1 What is Galaxy Galaxy for Bioinformaticians Galaxy for Experimental Biologists Using Galaxy for NGS Analysis NGS Data Visualization and Exploration Using IGV 2 What is Galaxy Galaxy for Bioinformaticians
ChIP-Seq Tutorial on Galaxy
 1 Introduction ChIP-Seq Tutorial on Galaxy 2 December 2010 (modified April 6, 2017) Rory Stark The aim of this practical is to give you some experience handling ChIP-Seq data. We will be working with data
1 Introduction ChIP-Seq Tutorial on Galaxy 2 December 2010 (modified April 6, 2017) Rory Stark The aim of this practical is to give you some experience handling ChIP-Seq data. We will be working with data
The UCSC Gene Sorter, Table Browser & Custom Tracks
 The UCSC Gene Sorter, Table Browser & Custom Tracks Advanced searching and discovery using the UCSC Table Browser and Custom Tracks Osvaldo Graña Bioinformatics Unit, CNIO 1 Table Browser and Custom Tracks
The UCSC Gene Sorter, Table Browser & Custom Tracks Advanced searching and discovery using the UCSC Table Browser and Custom Tracks Osvaldo Graña Bioinformatics Unit, CNIO 1 Table Browser and Custom Tracks
Instructions for downloading paid media from BSO.org and playing paid media in the BSO Media Center Revised as of 12/23/2011
 Instructions for downloading paid media from BSO.org and playing paid media in the BSO Media Center Revised as of 12/23/2011 DOWNLOADING MEDIA 1. Purchase Media Once you have completed your purchase, you
Instructions for downloading paid media from BSO.org and playing paid media in the BSO Media Center Revised as of 12/23/2011 DOWNLOADING MEDIA 1. Purchase Media Once you have completed your purchase, you
modencode Galaxy: Uniform ChIP-Seq Processing Tools for modencode and ENCODE Data
 modencode Galaxy: Uniform ChIP-Seq Processing Tools for modencode and ENCODE Data Quang M Trinh Ontario Institute for Cancer Research qtrinh@oicr.on.ca Outline Model Organism ENCyclopedia Of DNA Elements
modencode Galaxy: Uniform ChIP-Seq Processing Tools for modencode and ENCODE Data Quang M Trinh Ontario Institute for Cancer Research qtrinh@oicr.on.ca Outline Model Organism ENCyclopedia Of DNA Elements
RNA-Seq Analysis With the Tuxedo Suite
 June 2016 RNA-Seq Analysis With the Tuxedo Suite Dena Leshkowitz Introduction In this exercise we will learn how to analyse RNA-Seq data using the Tuxedo Suite tools: Tophat, Cuffmerge, Cufflinks and Cuffdiff.
June 2016 RNA-Seq Analysis With the Tuxedo Suite Dena Leshkowitz Introduction In this exercise we will learn how to analyse RNA-Seq data using the Tuxedo Suite tools: Tophat, Cuffmerge, Cufflinks and Cuffdiff.
Protocol: peak-calling for ChIP-seq data / segmentation analysis for histone modification data
 Protocol: peak-calling for ChIP-seq data / segmentation analysis for histone modification data Table of Contents Protocol: peak-calling for ChIP-seq data / segmentation analysis for histone modification
Protocol: peak-calling for ChIP-seq data / segmentation analysis for histone modification data Table of Contents Protocol: peak-calling for ChIP-seq data / segmentation analysis for histone modification
Genome Browser. Background and Strategy
 Genome Browser Background and Strategy Contents What is a genome browser? Purpose of a genome browser Examples Structure Extra Features Contents What is a genome browser? Purpose of a genome browser Examples
Genome Browser Background and Strategy Contents What is a genome browser? Purpose of a genome browser Examples Structure Extra Features Contents What is a genome browser? Purpose of a genome browser Examples
Easy visualization of the read coverage using the CoverageView package
 Easy visualization of the read coverage using the CoverageView package Ernesto Lowy European Bioinformatics Institute EMBL June 13, 2018 > options(width=40) > library(coverageview) 1 Introduction This
Easy visualization of the read coverage using the CoverageView package Ernesto Lowy European Bioinformatics Institute EMBL June 13, 2018 > options(width=40) > library(coverageview) 1 Introduction This
Analysis of ChIP-seq data
 Before we start: 1. Log into tak (step 0 on the exercises) 2. Go to your lab space and create a folder for the class (see separate hand out) 3. Connect to your lab space through the wihtdata network and
Before we start: 1. Log into tak (step 0 on the exercises) 2. Go to your lab space and create a folder for the class (see separate hand out) 3. Connect to your lab space through the wihtdata network and
Colorado State University Bioinformatics Algorithms Assignment 6: Analysis of High- Throughput Biological Data Hamidreza Chitsaz, Ali Sharifi- Zarchi
 Colorado State University Bioinformatics Algorithms Assignment 6: Analysis of High- Throughput Biological Data Hamidreza Chitsaz, Ali Sharifi- Zarchi Although a little- bit long, this is an easy exercise
Colorado State University Bioinformatics Algorithms Assignment 6: Analysis of High- Throughput Biological Data Hamidreza Chitsaz, Ali Sharifi- Zarchi Although a little- bit long, this is an easy exercise
ChIP-seq practical: peak detection and peak annotation. Mali Salmon-Divon Remco Loos Myrto Kostadima
 ChIP-seq practical: peak detection and peak annotation Mali Salmon-Divon Remco Loos Myrto Kostadima March 2012 Introduction The goal of this hands-on session is to perform some basic tasks in the analysis
ChIP-seq practical: peak detection and peak annotation Mali Salmon-Divon Remco Loos Myrto Kostadima March 2012 Introduction The goal of this hands-on session is to perform some basic tasks in the analysis
Student Evaluation of Teaching (SET)
 Testing, Evaluation and Research Services Student Evaluation of Teaching (SET) A guide to retrieving your faculty longitudinal report online. FOR ADDITIONAL INFORMATION, PLEASE CONTACT THE SET HELPDESK
Testing, Evaluation and Research Services Student Evaluation of Teaching (SET) A guide to retrieving your faculty longitudinal report online. FOR ADDITIONAL INFORMATION, PLEASE CONTACT THE SET HELPDESK
Tiling Assembly for Annotation-independent Novel Gene Discovery
 Tiling Assembly for Annotation-independent Novel Gene Discovery By Jennifer Lopez and Kenneth Watanabe Last edited on September 7, 2015 by Kenneth Watanabe The following procedure explains how to run the
Tiling Assembly for Annotation-independent Novel Gene Discovery By Jennifer Lopez and Kenneth Watanabe Last edited on September 7, 2015 by Kenneth Watanabe The following procedure explains how to run the
Getting Started with the WorkKeys Reports Portal
 Contents Topic Page Working with the Report 2 Log In & Access the Reports Portal 3 Running a Report 5 Print/Export the Report 7 Supported Browsers 7 Reports Portal: 1 of 7 Working with the Report Navigation
Contents Topic Page Working with the Report 2 Log In & Access the Reports Portal 3 Running a Report 5 Print/Export the Report 7 Supported Browsers 7 Reports Portal: 1 of 7 Working with the Report Navigation
Student Evaluation of Teaching (SET)
 Testing, Evaluation and Research Services Student Evaluation of Teaching (SET) A guide to retrieving your Chair administrative reports online. FOR ADDITIONAL INFORMATION, PLEASE CONTACT THE SET HELPDESK
Testing, Evaluation and Research Services Student Evaluation of Teaching (SET) A guide to retrieving your Chair administrative reports online. FOR ADDITIONAL INFORMATION, PLEASE CONTACT THE SET HELPDESK
Today's outline. Resources. Genome browser components. Genome browsers: Discovering biology through genomics. Genome browser tutorial materials
 Today's outline Genome browsers: Discovering biology through genomics BaRC Hot Topics April 2013 George Bell, Ph.D. http://jura.wi.mit.edu/bio/education/hot_topics/ Genome browser introduction Popular
Today's outline Genome browsers: Discovering biology through genomics BaRC Hot Topics April 2013 George Bell, Ph.D. http://jura.wi.mit.edu/bio/education/hot_topics/ Genome browser introduction Popular
From genomic regions to biology
 Before we start: 1. Log into tak (step 0 on the exercises) 2. Go to your lab space and create a folder for the class (see separate hand out) 3. Connect to your lab space through the wihtdata network and
Before we start: 1. Log into tak (step 0 on the exercises) 2. Go to your lab space and create a folder for the class (see separate hand out) 3. Connect to your lab space through the wihtdata network and
A short Introduction to UCSC Genome Browser
 A short Introduction to UCSC Genome Browser Elodie Girard, Nicolas Servant Institut Curie/INSERM U900 Bioinformatics, Biostatistics, Epidemiology and computational Systems Biology of Cancer 1 Why using
A short Introduction to UCSC Genome Browser Elodie Girard, Nicolas Servant Institut Curie/INSERM U900 Bioinformatics, Biostatistics, Epidemiology and computational Systems Biology of Cancer 1 Why using
Galaxy workshop at the Winter School Igor Makunin
 Galaxy workshop at the Winter School 2016 Igor Makunin i.makunin@uq.edu.au Winter school, UQ, July 6, 2016 Plan Overview of the Genomics Virtual Lab Introduce Galaxy, a web based platform for analysis
Galaxy workshop at the Winter School 2016 Igor Makunin i.makunin@uq.edu.au Winter school, UQ, July 6, 2016 Plan Overview of the Genomics Virtual Lab Introduce Galaxy, a web based platform for analysis
ChIP-seq Analysis. BaRC Hot Topics - March 21 st 2017 Bioinformatics and Research Computing Whitehead Institute.
 ChIP-seq Analysis BaRC Hot Topics - March 21 st 2017 Bioinformatics and Research Computing Whitehead Institute http://barc.wi.mit.edu/hot_topics/ Outline ChIP-seq overview Experimental design Quality control/preprocessing
ChIP-seq Analysis BaRC Hot Topics - March 21 st 2017 Bioinformatics and Research Computing Whitehead Institute http://barc.wi.mit.edu/hot_topics/ Outline ChIP-seq overview Experimental design Quality control/preprocessing
Using Tools for API Development and Testing
 This chapter contains the following sections: Using the API Inspector, page 1 Using the Managed Object Browser, page 3 Testing the API, page 6 Using the API Inspector Viewing an API Interchange in the
This chapter contains the following sections: Using the API Inspector, page 1 Using the Managed Object Browser, page 3 Testing the API, page 6 Using the API Inspector Viewing an API Interchange in the
CDIS Biomedical Data Commons
 CDIS Biomedical Data Commons Computational Life Science Seminar Series October 18, 2017 Michael Fitzsimons Center for Data Intensive Science Agenda What is a Data Commons? Data Commons at CDIS NCI GDC
CDIS Biomedical Data Commons Computational Life Science Seminar Series October 18, 2017 Michael Fitzsimons Center for Data Intensive Science Agenda What is a Data Commons? Data Commons at CDIS NCI GDC
Sequence Alignment. GBIO0002 Archana Bhardwaj University of Liege
 Sequence Alignment GBIO0002 Archana Bhardwaj University of Liege 1 What is Sequence Alignment? A sequence alignment is a way of arranging the sequences of DNA, RNA, or protein to identify regions of similarity.
Sequence Alignment GBIO0002 Archana Bhardwaj University of Liege 1 What is Sequence Alignment? A sequence alignment is a way of arranging the sequences of DNA, RNA, or protein to identify regions of similarity.
Services Performed. The following checklist confirms the steps of the RNA-Seq Service that were performed on your samples.
 Services Performed The following checklist confirms the steps of the RNA-Seq Service that were performed on your samples. SERVICE Sample Received Sample Quality Evaluated Sample Prepared for Sequencing
Services Performed The following checklist confirms the steps of the RNA-Seq Service that were performed on your samples. SERVICE Sample Received Sample Quality Evaluated Sample Prepared for Sequencing
Jersey City Free Public Library WIFI Hotspot
 1. Windows 2000, XP, 7 and Vista Users: a. Select the wireless icon in the system tray. or or or b. Select the SSID of the library you are currently located: JCPL- c. Launch a web browser (Internet Explorer,
1. Windows 2000, XP, 7 and Vista Users: a. Select the wireless icon in the system tray. or or or b. Select the SSID of the library you are currently located: JCPL- c. Launch a web browser (Internet Explorer,
epigenomegateway.wustl.edu
 Everything can be found at epigenomegateway.wustl.edu REFERENCES 1. Zhou X, et al., Nature Methods 8, 989-990 (2011) 2. Zhou X & Wang T, Current Protocols in Bioinformatics Unit 10.10 (2012) 3. Zhou X,
Everything can be found at epigenomegateway.wustl.edu REFERENCES 1. Zhou X, et al., Nature Methods 8, 989-990 (2011) 2. Zhou X & Wang T, Current Protocols in Bioinformatics Unit 10.10 (2012) 3. Zhou X,
Quantification. Part I, using Excel
 Quantification In this exercise we will work with RNA-seq data from a study by Serin et al (2017). RNA-seq was performed on Arabidopsis seeds matured at standard temperature (ST, 22 C day/18 C night) or
Quantification In this exercise we will work with RNA-seq data from a study by Serin et al (2017). RNA-seq was performed on Arabidopsis seeds matured at standard temperature (ST, 22 C day/18 C night) or
Introduction to Genome Browsers
 Introduction to Genome Browsers Rolando Garcia-Milian, MLS, AHIP (Rolando.milian@ufl.edu) Department of Biomedical and Health Information Services Health Sciences Center Libraries, University of Florida
Introduction to Genome Browsers Rolando Garcia-Milian, MLS, AHIP (Rolando.milian@ufl.edu) Department of Biomedical and Health Information Services Health Sciences Center Libraries, University of Florida
ChIP-seq (NGS) Data Formats
 ChIP-seq (NGS) Data Formats Biological samples Sequence reads SRA/SRF, FASTQ Quality control SAM/BAM/Pileup?? Mapping Assembly... DE Analysis Variant Detection Peak Calling...? Counts, RPKM VCF BED/narrowPeak/
ChIP-seq (NGS) Data Formats Biological samples Sequence reads SRA/SRF, FASTQ Quality control SAM/BAM/Pileup?? Mapping Assembly... DE Analysis Variant Detection Peak Calling...? Counts, RPKM VCF BED/narrowPeak/
Tutorial. RNA-Seq Analysis of Breast Cancer Data. Sample to Insight. November 21, 2017
 RNA-Seq Analysis of Breast Cancer Data November 21, 2017 Sample to Insight QIAGEN Aarhus Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.qiagenbioinformatics.com AdvancedGenomicsSupport@qiagen.com
RNA-Seq Analysis of Breast Cancer Data November 21, 2017 Sample to Insight QIAGEN Aarhus Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.qiagenbioinformatics.com AdvancedGenomicsSupport@qiagen.com
UCSC Genome Browser Pittsburgh Workshop -- Practical Exercises
 UCSC Genome Browser Pittsburgh Workshop -- Practical Exercises We will be using human assembly hg19. These problems will take you through a variety of resources at the UCSC Genome Browser. You will learn
UCSC Genome Browser Pittsburgh Workshop -- Practical Exercises We will be using human assembly hg19. These problems will take you through a variety of resources at the UCSC Genome Browser. You will learn
Browser Exercises - I. Alignments and Comparative genomics
 Browser Exercises - I Alignments and Comparative genomics 1. Navigating to the Genome Browser (GBrowse) Note: For this exercise use http://www.tritrypdb.org a. Navigate to the Genome Browser (GBrowse)
Browser Exercises - I Alignments and Comparative genomics 1. Navigating to the Genome Browser (GBrowse) Note: For this exercise use http://www.tritrypdb.org a. Navigate to the Genome Browser (GBrowse)
mirnet Tutorial Starting with expression data
 mirnet Tutorial Starting with expression data Computer and Browser Requirements A modern web browser with Java Script enabled Chrome, Safari, Firefox, and Internet Explorer 9+ For best performance and
mirnet Tutorial Starting with expression data Computer and Browser Requirements A modern web browser with Java Script enabled Chrome, Safari, Firefox, and Internet Explorer 9+ For best performance and
GETTING STARTED. A Step-by-Step Guide to Using MarketSight
 GETTING STARTED A Step-by-Step Guide to Using MarketSight Analyze any dataset Run crosstabs Test statistical significance Create charts and dashboards Share results online Introduction MarketSight is a
GETTING STARTED A Step-by-Step Guide to Using MarketSight Analyze any dataset Run crosstabs Test statistical significance Create charts and dashboards Share results online Introduction MarketSight is a
Integrated Genome browser (IGB) installation
 Integrated Genome browser (IGB) installation Navigate to the IGB download page http://bioviz.org/igb/download.html You will see three icons for download: The three icons correspond to different memory
Integrated Genome browser (IGB) installation Navigate to the IGB download page http://bioviz.org/igb/download.html You will see three icons for download: The three icons correspond to different memory
The European Variation Archive
 The European Variation Archive Webinar: A database of all types of genomic variation data from all species Hannah McLaren www.ebi.ac.uk/eva eva-helpdesk@ebi.ac.uk Learning objectives Establish the key
The European Variation Archive Webinar: A database of all types of genomic variation data from all species Hannah McLaren www.ebi.ac.uk/eva eva-helpdesk@ebi.ac.uk Learning objectives Establish the key
Web page:
 mirdeep* manual 2013-01-10 Jiyuan An, John Lai, Melanie Lehman, Colleen Nelson: Australian Prostate Cancer Research Center (APCRC-Q) and Institute of Health and Biomedical Innovation (IHBI), Queensland
mirdeep* manual 2013-01-10 Jiyuan An, John Lai, Melanie Lehman, Colleen Nelson: Australian Prostate Cancer Research Center (APCRC-Q) and Institute of Health and Biomedical Innovation (IHBI), Queensland
But before understanding the Selenium WebDriver concept, we need to know about the Selenium first.
 As per the today s scenario, companies not only desire to test software adequately, but they also want to get the work done as quickly and thoroughly as possible. To accomplish this goal, organizations
As per the today s scenario, companies not only desire to test software adequately, but they also want to get the work done as quickly and thoroughly as possible. To accomplish this goal, organizations
Supplementary Figure 1. Fast read-mapping algorithm of BrowserGenome.
 Supplementary Figure 1 Fast read-mapping algorithm of BrowserGenome. (a) Indexing strategy: The genome sequence of interest is divided into non-overlapping 12-mers. A Hook table is generated that contains
Supplementary Figure 1 Fast read-mapping algorithm of BrowserGenome. (a) Indexing strategy: The genome sequence of interest is divided into non-overlapping 12-mers. A Hook table is generated that contains
SAS STUDIO. JUNE 2014 PRESENTER: MARY HARDING Education SAS Canada. Copyr i g ht 2014, SAS Ins titut e Inc. All rights res er ve d.
 JUNE 2014 PRESENTER: MARY HARDING Education SAS Canada NEW SAS PROGRAMMING ENVIRONMENT Available Consistent Assistive AVAILABLE THROUGH ALL MODERN WEB BROWSERS Available Consistent Assistive ONE INTERFACE
JUNE 2014 PRESENTER: MARY HARDING Education SAS Canada NEW SAS PROGRAMMING ENVIRONMENT Available Consistent Assistive AVAILABLE THROUGH ALL MODERN WEB BROWSERS Available Consistent Assistive ONE INTERFACE
HymenopteraMine Documentation
 HymenopteraMine Documentation Release 1.0 Aditi Tayal, Deepak Unni, Colin Diesh, Chris Elsik, Darren Hagen Apr 06, 2017 Contents 1 Welcome to HymenopteraMine 3 1.1 Overview of HymenopteraMine.....................................
HymenopteraMine Documentation Release 1.0 Aditi Tayal, Deepak Unni, Colin Diesh, Chris Elsik, Darren Hagen Apr 06, 2017 Contents 1 Welcome to HymenopteraMine 3 1.1 Overview of HymenopteraMine.....................................
replace my_user_id in the commands with your actual user ID
 Exercise 1. Alignment with TOPHAT Part 1. Prepare the working directory. 1. Find out the name of the computer that has been reserved for you (https://cbsu.tc.cornell.edu/ww/machines.aspx?i=57 ). Everyone
Exercise 1. Alignment with TOPHAT Part 1. Prepare the working directory. 1. Find out the name of the computer that has been reserved for you (https://cbsu.tc.cornell.edu/ww/machines.aspx?i=57 ). Everyone
Sequence Analysis Pipeline
 Sequence Analysis Pipeline Transcript fragments 1. PREPROCESSING 2. ASSEMBLY (today) Removal of contaminants, vector, adaptors, etc Put overlapping sequence together and calculate bigger sequences 3. Analysis/Annotation
Sequence Analysis Pipeline Transcript fragments 1. PREPROCESSING 2. ASSEMBLY (today) Removal of contaminants, vector, adaptors, etc Put overlapping sequence together and calculate bigger sequences 3. Analysis/Annotation
Index Introduction Setting up an account Searching and accessing Download Advanced features
 ESGF Earth System Grid Federation Tutorial Index Introduction Setting up an account Searching and accessing Download Advanced features Index Introduction IT Challenges of Climate Change Research ESGF Introduction
ESGF Earth System Grid Federation Tutorial Index Introduction Setting up an account Searching and accessing Download Advanced features Index Introduction IT Challenges of Climate Change Research ESGF Introduction
Transcript quantification using Salmon and differential expression analysis using bayseq
 Introduction to expression analysis (RNA-seq) Transcript quantification using Salmon and differential expression analysis using bayseq Philippine Genome Center University of the Philippines Prepared by
Introduction to expression analysis (RNA-seq) Transcript quantification using Salmon and differential expression analysis using bayseq Philippine Genome Center University of the Philippines Prepared by
Bioinformatics Hubs on the Web
 Bioinformatics Hubs on the Web Take a class The Galter Library teaches a related class called Bioinformatics Hubs on the Web. See our Classes schedule for the next available offering. If this class is
Bioinformatics Hubs on the Web Take a class The Galter Library teaches a related class called Bioinformatics Hubs on the Web. See our Classes schedule for the next available offering. If this class is
Lecture 8. Sequence alignments
 Lecture 8 Sequence alignments DATA FORMATS bioawk bioawk is a program that extends awk s powerful processing of tabular data to processing tasks involving common bioinformatics formats like FASTA/FASTQ,
Lecture 8 Sequence alignments DATA FORMATS bioawk bioawk is a program that extends awk s powerful processing of tabular data to processing tasks involving common bioinformatics formats like FASTA/FASTQ,
Viewing CyVerse Hosted Data at UCSC
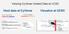 Viewing CyVerse Hosted Data at UCSC Host data at CyVerse Binary indexed files: twobitpath, bigdataurl Text files: hub.txt, genomes.txt, trackdb.txt Visualize at UCSC CyVerse s Send to Genome Browser option
Viewing CyVerse Hosted Data at UCSC Host data at CyVerse Binary indexed files: twobitpath, bigdataurl Text files: hub.txt, genomes.txt, trackdb.txt Visualize at UCSC CyVerse s Send to Genome Browser option
Useful software utilities for computational genomics. Shamith Samarajiwa CRUK Autumn School in Bioinformatics September 2017
 Useful software utilities for computational genomics Shamith Samarajiwa CRUK Autumn School in Bioinformatics September 2017 Overview Search and download genomic datasets: GEOquery, GEOsearch and GEOmetadb,
Useful software utilities for computational genomics Shamith Samarajiwa CRUK Autumn School in Bioinformatics September 2017 Overview Search and download genomic datasets: GEOquery, GEOsearch and GEOmetadb,
ArrayExpress and Expression Atlas: Mining Functional Genomics data
 and Expression Atlas: Mining Functional Genomics data Gabriella Rustici, PhD Functional Genomics Team EBI-EMBL gabry@ebi.ac.uk What is functional genomics (FG)? The aim of FG is to understand the function
and Expression Atlas: Mining Functional Genomics data Gabriella Rustici, PhD Functional Genomics Team EBI-EMBL gabry@ebi.ac.uk What is functional genomics (FG)? The aim of FG is to understand the function
Automated Bioinformatics Analysis System on Chip ABASOC. version 1.1
 Automated Bioinformatics Analysis System on Chip ABASOC version 1.1 Phillip Winston Miller, Priyam Patel, Daniel L. Johnson, PhD. University of Tennessee Health Science Center Office of Research Molecular
Automated Bioinformatics Analysis System on Chip ABASOC version 1.1 Phillip Winston Miller, Priyam Patel, Daniel L. Johnson, PhD. University of Tennessee Health Science Center Office of Research Molecular
Expression Analysis with the Advanced RNA-Seq Plugin
 Expression Analysis with the Advanced RNA-Seq Plugin May 24, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com
Expression Analysis with the Advanced RNA-Seq Plugin May 24, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com
Online App Access: Quick Set-up & Instructions
 Online App Access: Quick Set-up & Instructions Welcome to The Media Audit You will be accessing data ONLINE through our Citrix app, available by visiting: app.themediaaudit.com This simple guidebook contains
Online App Access: Quick Set-up & Instructions Welcome to The Media Audit You will be accessing data ONLINE through our Citrix app, available by visiting: app.themediaaudit.com This simple guidebook contains
G-TACT User Guide Ensuring Accurate Classification of BRCA Variants Module: RUN 1
 G-TACT User Guide Ensuring Accurate Classification of BRCA Variants Module: RUN 1 Description The G-TACT User Guide provides assistance to users in all aspects of using the Ensuring Accurate Classification
G-TACT User Guide Ensuring Accurate Classification of BRCA Variants Module: RUN 1 Description The G-TACT User Guide provides assistance to users in all aspects of using the Ensuring Accurate Classification
Genome Browser. Background & Strategy. Spring 2017 Faction II
 Genome Browser Background & Strategy Spring 2017 Faction II Outline Beginning of the Last Phase Goals State of Art Applicable Genome Browsers Not So Genome Browsers Storing Data Strategy for the website
Genome Browser Background & Strategy Spring 2017 Faction II Outline Beginning of the Last Phase Goals State of Art Applicable Genome Browsers Not So Genome Browsers Storing Data Strategy for the website
ChIP-seq Analysis. BaRC Hot Topics - Feb 23 th 2016 Bioinformatics and Research Computing Whitehead Institute.
 ChIP-seq Analysis BaRC Hot Topics - Feb 23 th 2016 Bioinformatics and Research Computing Whitehead Institute http://barc.wi.mit.edu/hot_topics/ Outline ChIP-seq overview Experimental design Quality control/preprocessing
ChIP-seq Analysis BaRC Hot Topics - Feb 23 th 2016 Bioinformatics and Research Computing Whitehead Institute http://barc.wi.mit.edu/hot_topics/ Outline ChIP-seq overview Experimental design Quality control/preprocessing
Programming in the Life Sciences
 Programming in the Life Sciences In the Maastricht Science Programme Open PHACTS Community Workshop London, 26 June 2014 1 Who am I? Teacher at Dept. Bioinformatics BiGCaT, NUTRIM, FHML, UM. http://chem-bla-ics.blogspot.com/
Programming in the Life Sciences In the Maastricht Science Programme Open PHACTS Community Workshop London, 26 June 2014 1 Who am I? Teacher at Dept. Bioinformatics BiGCaT, NUTRIM, FHML, UM. http://chem-bla-ics.blogspot.com/
Bioinformatics in next generation sequencing projects
 Bioinformatics in next generation sequencing projects Rickard Sandberg Assistant Professor Department of Cell and Molecular Biology Karolinska Institutet March 2011 Once sequenced the problem becomes computational
Bioinformatics in next generation sequencing projects Rickard Sandberg Assistant Professor Department of Cell and Molecular Biology Karolinska Institutet March 2011 Once sequenced the problem becomes computational
How to block ads in the right side of Yahoo Mail
 How to block ads in the right side of Yahoo Mail Submitted by Jess on Sun, 01/04/2015-15:27 Millions of people use Yahoo! Mail and its other email infrastructure like SBCGlobal, ymail, and others. Yahoo!
How to block ads in the right side of Yahoo Mail Submitted by Jess on Sun, 01/04/2015-15:27 Millions of people use Yahoo! Mail and its other email infrastructure like SBCGlobal, ymail, and others. Yahoo!
Exercise 1. RNA-seq alignment and quantification. Part 1. Prepare the working directory. Part 2. Examine qualities of the RNA-seq data files
 Exercise 1. RNA-seq alignment and quantification Part 1. Prepare the working directory. 1. Connect to your assigned computer. If you do not know how, follow the instruction at http://cbsu.tc.cornell.edu/lab/doc/remote_access.pdf
Exercise 1. RNA-seq alignment and quantification Part 1. Prepare the working directory. 1. Connect to your assigned computer. If you do not know how, follow the instruction at http://cbsu.tc.cornell.edu/lab/doc/remote_access.pdf
ChIP- seq Analysis. BaRC Hot Topics - Feb 24 th 2015 BioinformaBcs and Research CompuBng Whitehead InsBtute. hgp://barc.wi.mit.
 ChIP- seq Analysis BaRC Hot Topics - Feb 24 th 2015 BioinformaBcs and Research CompuBng Whitehead InsBtute hgp://barc.wi.mit.edu/hot_topics/ Before we start: 1. Log into tak (step 0 on the exercises) 2.
ChIP- seq Analysis BaRC Hot Topics - Feb 24 th 2015 BioinformaBcs and Research CompuBng Whitehead InsBtute hgp://barc.wi.mit.edu/hot_topics/ Before we start: 1. Log into tak (step 0 on the exercises) 2.
Maximizing Public Data Sources for Sequencing and GWAS
 Maximizing Public Data Sources for Sequencing and GWAS February 4, 2014 G Bryce Christensen Director of Services Questions during the presentation Use the Questions pane in your GoToWebinar window Agenda
Maximizing Public Data Sources for Sequencing and GWAS February 4, 2014 G Bryce Christensen Director of Services Questions during the presentation Use the Questions pane in your GoToWebinar window Agenda
Chromatin signature discovery via histone modification profile alignments Jianrong Wang, Victoria V. Lunyak and I. King Jordan
 Supplementary Information for: Chromatin signature discovery via histone modification profile alignments Jianrong Wang, Victoria V. Lunyak and I. King Jordan Contents Instructions for installing and running
Supplementary Information for: Chromatin signature discovery via histone modification profile alignments Jianrong Wang, Victoria V. Lunyak and I. King Jordan Contents Instructions for installing and running
Creating a Bookmark/Link for the Portal(my.cuw.edu)
 Here are the links to each guide on this PDF Internet Explorer EDGE Firefox Chrome Safari Items to Note Always remember to click the log out button on the Portal when you are finished with your session.
Here are the links to each guide on this PDF Internet Explorer EDGE Firefox Chrome Safari Items to Note Always remember to click the log out button on the Portal when you are finished with your session.
PRINTING IN ESCRIBE...2
 PRINTING IN ESCRIBE...2 PRINTING FROM MOZILLA FIREFOX...2 ALLOWING POPUPS IN MOZILLA FIREFOX...3 PRINTING FROM GOOGLE CHROME...4 ALLOWING POPUPS IN GOOGLE CHROME...5 PRINTING FROM APPLE SAFARI...6 ALLOWING
PRINTING IN ESCRIBE...2 PRINTING FROM MOZILLA FIREFOX...2 ALLOWING POPUPS IN MOZILLA FIREFOX...3 PRINTING FROM GOOGLE CHROME...4 ALLOWING POPUPS IN GOOGLE CHROME...5 PRINTING FROM APPLE SAFARI...6 ALLOWING
EPITENSOR MANUAL. Version 0.9. Yun Zhu
 EPITENSOR MANUAL Version 0.9 Yun Zhu May 4, 2015 1. Overview EpiTensor is a program that can construct 3-D interaction maps in the genome from 1-D epigenomes. It takes aligned reads in bam format as input
EPITENSOR MANUAL Version 0.9 Yun Zhu May 4, 2015 1. Overview EpiTensor is a program that can construct 3-D interaction maps in the genome from 1-D epigenomes. It takes aligned reads in bam format as input
Tutorial: How to Download Your 23andme.com Raw Data & How to Resolve issues with PayPal
 Tutorial: How to Download Your 23andme.com Raw Data & How to Resolve issues with PayPal I. Access 23andme.com account II. Sign In III. Browse Raw Data IV. a, Download Raw Data b, Download Raw Data (cont.)
Tutorial: How to Download Your 23andme.com Raw Data & How to Resolve issues with PayPal I. Access 23andme.com account II. Sign In III. Browse Raw Data IV. a, Download Raw Data b, Download Raw Data (cont.)
How to store and visualize RNA-seq data
 How to store and visualize RNA-seq data Gabriella Rustici Functional Genomics Group gabry@ebi.ac.uk EBI is an Outstation of the European Molecular Biology Laboratory. Talk summary How do we archive RNA-seq
How to store and visualize RNA-seq data Gabriella Rustici Functional Genomics Group gabry@ebi.ac.uk EBI is an Outstation of the European Molecular Biology Laboratory. Talk summary How do we archive RNA-seq
ChIP-seq Analysis Practical
 ChIP-seq Analysis Practical Vladimir Teif (vteif@essex.ac.uk) An updated version of this document will be available at http://generegulation.info/index.php/teaching In this practical we will learn how
ChIP-seq Analysis Practical Vladimir Teif (vteif@essex.ac.uk) An updated version of this document will be available at http://generegulation.info/index.php/teaching In this practical we will learn how
HIPPIE User Manual. (v0.0.2-beta, 2015/4/26, Yih-Chii Hwang, yihhwang [at] mail.med.upenn.edu)
![HIPPIE User Manual. (v0.0.2-beta, 2015/4/26, Yih-Chii Hwang, yihhwang [at] mail.med.upenn.edu) HIPPIE User Manual. (v0.0.2-beta, 2015/4/26, Yih-Chii Hwang, yihhwang [at] mail.med.upenn.edu)](/thumbs/71/65752105.jpg) HIPPIE User Manual (v0.0.2-beta, 2015/4/26, Yih-Chii Hwang, yihhwang [at] mail.med.upenn.edu) OVERVIEW OF HIPPIE o Flowchart of HIPPIE o Requirements PREPARE DIRECTORY STRUCTURE FOR HIPPIE EXECUTION o
HIPPIE User Manual (v0.0.2-beta, 2015/4/26, Yih-Chii Hwang, yihhwang [at] mail.med.upenn.edu) OVERVIEW OF HIPPIE o Flowchart of HIPPIE o Requirements PREPARE DIRECTORY STRUCTURE FOR HIPPIE EXECUTION o
Agilent Genomic Workbench Lite Edition 6.5
 Agilent Genomic Workbench Lite Edition 6.5 SureSelect Quality Analyzer User Guide For Research Use Only. Not for use in diagnostic procedures. Agilent Technologies Notices Agilent Technologies, Inc. 2010
Agilent Genomic Workbench Lite Edition 6.5 SureSelect Quality Analyzer User Guide For Research Use Only. Not for use in diagnostic procedures. Agilent Technologies Notices Agilent Technologies, Inc. 2010
Genome Browsers - The UCSC Genome Browser
 Genome Browsers - The UCSC Genome Browser Background The UCSC Genome Browser is a well-curated site that provides users with a view of gene or sequence information in genomic context for a specific species,
Genome Browsers - The UCSC Genome Browser Background The UCSC Genome Browser is a well-curated site that provides users with a view of gene or sequence information in genomic context for a specific species,
ITMO Ecole de Bioinformatique Hands-on session: smallrna-seq N. Servant 21 rd November 2013
 ITMO Ecole de Bioinformatique Hands-on session: smallrna-seq N. Servant 21 rd November 2013 1. Data and objectives We will use the data from GEO (GSE35368, Toedling, Servant et al. 2011). Two samples were
ITMO Ecole de Bioinformatique Hands-on session: smallrna-seq N. Servant 21 rd November 2013 1. Data and objectives We will use the data from GEO (GSE35368, Toedling, Servant et al. 2011). Two samples were
How to Access Live Video
 CHAPTER 5 This chapter discusses how live video works with in the Video Portal. Figure 5-1 shows the Live Video Lists. From the initial Featured Playlist tab, you can glance at the lineup of spotlighted
CHAPTER 5 This chapter discusses how live video works with in the Video Portal. Figure 5-1 shows the Live Video Lists. From the initial Featured Playlist tab, you can glance at the lineup of spotlighted
Mapping RNA sequence data (Part 1: using pathogen portal s RNAseq pipeline) Exercise 6
 Mapping RNA sequence data (Part 1: using pathogen portal s RNAseq pipeline) Exercise 6 The goal of this exercise is to retrieve an RNA-seq dataset in FASTQ format and run it through an RNA-sequence analysis
Mapping RNA sequence data (Part 1: using pathogen portal s RNAseq pipeline) Exercise 6 The goal of this exercise is to retrieve an RNA-seq dataset in FASTQ format and run it through an RNA-sequence analysis
YouTube: Instructor Guide
 YouTube: Instructor Guide Before you begin, you WILL need an @mocs.utc.edu email to setup a YouTube channel. This is not the same as your Outlook email. It is the same type of email that students have.
YouTube: Instructor Guide Before you begin, you WILL need an @mocs.utc.edu email to setup a YouTube channel. This is not the same as your Outlook email. It is the same type of email that students have.
Genetics 211 Genomics Winter 2014 Problem Set 4
 Genomics - Part 1 due Friday, 2/21/2014 by 9:00am Part 2 due Friday, 3/7/2014 by 9:00am For this problem set, we re going to use real data from a high-throughput sequencing project to look for differential
Genomics - Part 1 due Friday, 2/21/2014 by 9:00am Part 2 due Friday, 3/7/2014 by 9:00am For this problem set, we re going to use real data from a high-throughput sequencing project to look for differential
Installation Guide for Python
 GPDI 513 Beginner s Guide to the Python Programming Language Installation Guide for Python Linux Operating System If you are using a Linux computer, open the terminal and type the following commands in
GPDI 513 Beginner s Guide to the Python Programming Language Installation Guide for Python Linux Operating System If you are using a Linux computer, open the terminal and type the following commands in
Reference guided RNA-seq data analysis using BioHPC Lab computers
 Reference guided RNA-seq data analysis using BioHPC Lab computers This document assumes that you already know some basics of how to use a Linux computer. Some of the command lines in this document are
Reference guided RNA-seq data analysis using BioHPC Lab computers This document assumes that you already know some basics of how to use a Linux computer. Some of the command lines in this document are
New Viewer Functionality PRINT FUNCTIONALITY
 HTML5 Viewer using Google Chrome This document provides information on using the new HTLM5 viewer with Google Chrome. Note that examples and screenshots in this document have been provided from the esearch
HTML5 Viewer using Google Chrome This document provides information on using the new HTLM5 viewer with Google Chrome. Note that examples and screenshots in this document have been provided from the esearch
WebEx Cloud Quick Reference Guide
 The purpose of this document is to be used as a quick reference guide only. To provide basis instructions on how to setup meetings within in WebEx. For detailed instructions see Lynda.com WebEx Cloud allows
The purpose of this document is to be used as a quick reference guide only. To provide basis instructions on how to setup meetings within in WebEx. For detailed instructions see Lynda.com WebEx Cloud allows
Advanced UCSC Browser Functions
 Advanced UCSC Browser Functions Dr. Thomas Randall tarandal@email.unc.edu bioinformatics.unc.edu UCSC Browser: genome.ucsc.edu Overview Custom Tracks adding your own datasets Utilities custom tools for
Advanced UCSC Browser Functions Dr. Thomas Randall tarandal@email.unc.edu bioinformatics.unc.edu UCSC Browser: genome.ucsc.edu Overview Custom Tracks adding your own datasets Utilities custom tools for
Tutorial 1: Exploring the UCSC Genome Browser
 Last updated: May 12, 2011 Tutorial 1: Exploring the UCSC Genome Browser Open the homepage of the UCSC Genome Browser at: http://genome.ucsc.edu/ In the blue bar at the top, click on the Genomes link.
Last updated: May 12, 2011 Tutorial 1: Exploring the UCSC Genome Browser Open the homepage of the UCSC Genome Browser at: http://genome.ucsc.edu/ In the blue bar at the top, click on the Genomes link.
Supplementary Material. Cell type-specific termination of transcription by transposable element sequences
 Supplementary Material Cell type-specific termination of transcription by transposable element sequences Andrew B. Conley and I. King Jordan Controls for TTS identification using PET A series of controls
Supplementary Material Cell type-specific termination of transcription by transposable element sequences Andrew B. Conley and I. King Jordan Controls for TTS identification using PET A series of controls
2) NCBI BLAST tutorial This is a users guide written by the education department at NCBI.
 Web resources -- Tour. page 1 of 8 This is a guided tour. Any homework is separate. In fact, this exercise is used for multiple classes and is publicly available to everyone. The entire tour will take
Web resources -- Tour. page 1 of 8 This is a guided tour. Any homework is separate. In fact, this exercise is used for multiple classes and is publicly available to everyone. The entire tour will take
High-throughout sequencing and using short-read aligners. Simon Anders
 High-throughout sequencing and using short-read aligners Simon Anders High-throughput sequencing (HTS) Sequencing millions of short DNA fragments in parallel. a.k.a.: next-generation sequencing (NGS) massively-parallel
High-throughout sequencing and using short-read aligners Simon Anders High-throughput sequencing (HTS) Sequencing millions of short DNA fragments in parallel. a.k.a.: next-generation sequencing (NGS) massively-parallel
Integrative Genomics Viewer. Prat Thiru
 Integrative Genomics Viewer Prat Thiru 1 Overview User Interface Basics Browsing the Data Data Formats IGV Tools Demo Outline Based on ISMB 2010 Tutorial by Robinson and Thorvaldsdottir 2 Why IGV? IGV
Integrative Genomics Viewer Prat Thiru 1 Overview User Interface Basics Browsing the Data Data Formats IGV Tools Demo Outline Based on ISMB 2010 Tutorial by Robinson and Thorvaldsdottir 2 Why IGV? IGV
Analyzing Variant Call results using EuPathDB Galaxy, Part II
 Analyzing Variant Call results using EuPathDB Galaxy, Part II In this exercise, we will work in groups to examine the results from the SNP analysis workflow that we started yesterday. The first step is
Analyzing Variant Call results using EuPathDB Galaxy, Part II In this exercise, we will work in groups to examine the results from the SNP analysis workflow that we started yesterday. The first step is

 Searching the ENCODE Portal Asia- Pacific Bioinforma
Searching the ENCODE Portal Asia- Pacific Bioinforma