Controlling for mul2ple comparisons in imaging analysis. Where we re going. Where we re going 8/15/16
|
|
|
- Derick Knight
- 6 years ago
- Views:
Transcription
1 Controlling for mul2ple comparisons in imaging analysis Wednesday, Lecture 2 Jeane?e Mumford University of Wisconsin - Madison Where we re going Review of hypothesis tes2ng introduce mul2ple tes2ng problem Levels of inference (voxel/cluster/peak/set) Types of error rate control (none/fwer/fdr) Family- wise error control approaches (parametric/ nonparametric) FDR Rela2ng all of this to SPM output Where we re going Review of hypothesis tes2ng introduce mul2ple tes2ng problem Levels of inference (voxel/cluster/peak/set) Types of error rate control (none/fwer/fdr) Family- wise error control approaches (parametric/ nonparametric) FDR Rela2ng all of this to SPM output 1
2 Review of hypothesis tes2ng What is H0? What is HA? What are the steps of carrying out a hypothesis test? Review of hypothesis tes2ng What is H0? What is HA? What are the steps of carrying out a hypothesis test? Steps of hypothesis tes2ng
3 Steps of hypothesis tes2ng Steps of hypothesis tes2ng Steps of hypothesis tes2ng What do we compare this area to (p- value)?
4 What does the p- value mean? p = 0.01 What does the p- value mean? p = 0.01 If the null distribu2on is true What does the p- value mean? p = 0.01 If the null distribu2on is true The probability of observing my sta2s2c (or something more extreme than it) is
5 What does the p- value threshold imply? We choose 0.05 Less than 0.05 and we reject the null hypothesis Greater than 0.05 and we fail to reject the null hypothesis What does the p- value threshold imply? We choose 0.05 Less than 0.05 and we reject the null hypothesis Greater than 0.05 and we fail to reject the null hypothesis What does the p- value threshold imply? We choose 0.05 Less than 0.05 and we reject the null hypothesis Greater than 0.05 and we fail to reject the null hypothesis 5
6 Type I error Assuming the null is true, the probability that we reject the null Type I error Assuming the null is true, the probability that we reject the null 5% of the 2me, we ll have a false posi2ve Interpreta2on 1100 total voxels 100 voxels have β=δ 80% power - > 80 voxels detected 1000 voxels have β=0 5% type I error - > 50 false posi2ves Declared ac2ve Declared inac2ve Total Non- ac2ve Ac2ve Total 6
7 Interpreta2on 1100 total voxels 100 voxels have β=δ 80% power - > 80 voxels detected 1000 voxels have β=0 5% type I error - > 50 false posi2ves What we know (test results) Non- ac2ve Ac2ve Total Declared ac2ve Declared inac2ve Total Interpreta2on 1100 total voxels 100 voxels have β=δ 80% power - > 80 voxels detected 1000 voxels have β=0 5% type I error - > 50 false posi2ves What we don t know (truth) Non- ac2ve Ac2ve Total Declared ac2ve Declared inac2ve Total Interpreta2on 1100 total voxels 100 voxels have signal (null is false) 80% power - > 80 voxels detected 1000 voxels have no signal (null) 5% type I error - > 50 false posi2ves Declared ac2ve Declared inac2ve Total Non- ac2ve 1000 Ac2ve 100 Total
8 Interpreta2on 1100 total voxels 100 voxels have signal (null is false) 80% power - > 80 voxels detected 1000 voxels have no signal (null) 5% type I error - > 50 false posi2ves Declared ac2ve Declared inac2ve Total Non- ac2ve 1000 Ac2ve Total 1100 Interpreta2on 1100 total voxels 100 voxels have signal (null is false) 80% power - > 80 voxels detected 1000 voxels have no signal (null) 5% type I error - > 50 false posi2ves Declared ac2ve Declared inac2ve Total Non- ac2ve 1000 Ac2ve 80 (Power) 20 (Type II err.) 100 Total 1100 Interpreta2on 1100 total voxels 100 voxels have signal (null is false) 80% power - > 80 voxels detected 1000 voxels have no signal (null) 5% type I error - > 50 false posi2ves Declared ac2ve Declared inac2ve Total Non- ac2ve Ac2ve 80 (Power) 20 (Type II err.) 100 Total
9 Interpreta2on 1100 total voxels 100 voxels have signal (null is false) 80% power - > 80 voxels detected 1000 voxels have no signal (null) 5% type I error - > 50 false posi2ves Declared ac2ve Declared inac2ve Total Non- ac2ve 50 (Type I err.) 950 (Correct) 1000 Ac2ve 80 (Power) 20 (Type II err.) 100 Total 1100 Interpreta2on 1100 total voxels 100 voxels have signal (null is false) 80% power - > 80 voxels detected 1000 voxels have no signal (null) 5% type I error - > 50 false posi2ves Declared ac2ve Declared inac2ve Total Non- ac2ve Ac2ve Total Interpreta2on 1100 total voxels 100 voxels have signal (null is false) 80% power - > 80 voxels detected 1000 voxels have no signal (null) focus is on 5% type I error - > 50 false posi2ves controlling this number Declared ac2ve Declared inac2ve Total Non- ac2ve Ac2ve Total
10 Implica2on of type I error If you run enough tests, you ll find something that is significant This doesn t mean it is truly significant If you run 20 tests with a 5% threshold on type I errors, you expect to have at least 1 significant test This would be a false posi2ve Hypothesis Tes2ng in fmri Mass Univariate Modeling Fit a separate model for each voxel Look at images of sta2s2cs Apply Threshold Assessing Sta2s2c Images What threshold will show us signal? High Threshold Med. Threshold Low Threshold t > 5.5 t > 3.5 t > 0.5 Good Specificity Poor Power (risk of false negatives) Poor Specificity (risk of false positives) Good Power 10
11 Where we re going Review of hypothesis tes2ng introduce mul2ple tes2ng problem Levels of inference (voxel/cluster/peak/set) Types of error rate control (none/fwer/fdr) Family- wise error control approaches (parametric/ nonparametric) FDR Rela2ng all of this to SPM output Voxel level Levels of inference Cluster level Peak level Set level Voxel- level Inference Retain voxels above α- level threshold u α Gives best spa2al specificity The null hyp. at a single voxel can be rejected Statistic values space 11
12 Voxel- level Inference Retain voxels above α- level threshold u α Gives best spa2al specificity The null hyp. at a single voxel can be rejected u α space Voxel- level Inference Retain voxels above α- level threshold u α Gives best spa2al specificity The null hyp. at a single voxel can be rejected u α space Significant Voxels No significant Voxels Cluster- level Inference Two step- process Define clusters by arbitrary threshold u clus u clus space 12
13 Cluster- level Inference Two step- process Define clusters by arbitrary threshold u clus Retain clusters larger than α- level threshold k α u clus space Cluster not significant k α k α Cluster significant Cluster- level Inference Typically be?er sensi2vity Worse spa2al specificity The null hyp. of en2re cluster is rejected Only means that one or more of voxels in cluster ac2ve u clus space Cluster not significant k α k α Cluster significant Peak level inference Again start with a cluster forming threshold Instead of cluster size, focus on peak height Similarly to cluster level inference, significance applies to a set of voxels The peak and its neighbors u clus space 13
14 Peak level inference Again start with a cluster forming threshold Instead of cluster size, focus on peak height Similarly to cluster level inference, significance applies to a set of voxels The peak and its neighbors Z 4 u clus Z 1 Z 2 Z 3 Z 5 space Peak level inference Again start with a cluster forming threshold Instead of cluster size, focus on peak height Similarly to cluster level inference, significance applies to a set of voxels The peak and its neighbors Z 4 u peak Z u 1 clus Z 2 Z 3 Z 5 space Peak level inference Again start with a cluster forming threshold Instead of cluster size, focus on peak height Similarly to cluster level inference, significance applies to a set of voxels The peak and its neighbors Z 4 u peak Z u 1 clus Z 2 Z 3 Z 5 space 14
15 Set level inference Is there any ac2va2on anywhere in the brain? Omnibus hypothesis test of all voxels, simultaneously If significant, we only know there s ac2va2on somewhere in the brain Voxel level Levels of inference Cluster level Peak level Set level Ques2ons for you Why do some approaches require 2 thresholds? What thresholding strategy do people typically use? 15
16 Where we re going Review of hypothesis tes2ng introduce mul2ple tes2ng problem Levels of inference (voxel/cluster/peak/set) Types of error rate control (none/fwer/fdr) Family- wise error control approaches (parametric/ nonparametric) FDR Rela2ng all of this to SPM output What error rate should we control? Per comparison error rate? Family wise error rate? False discovery rate? Different types of error rates PCER Per comparison error rate Controlling each voxel at 5% Expect 5% of null voxels will be (mistakenly) deemed ac2ve FWER Family wise error rate Controls the probability of any false posi2ves Run 20 NULL group analyses (on 20 data sets) and only 1 analysis will have a significant finding 16
17 Different types of error rates PCER Per comparison error rate Controlling each voxel at 5% Expect 5% of null voxels will be (mistakenly) deemed ac2ve FWER Family wise error rate Controls the probability of any false posi2ves Run 20 NULL group analyses (on 20 data sets) and only 1 analysis will have a significant finding Different types of error rates FDR False discovery rate Of the voxels you deemed significant, what percentage were null FWER vs FDR FWER P(# true null declared ac2ve > 1) FDR E (# of true null declared ac2ve / # voxels declared ac2ve) Declared ac2ve Declared inac2ve Total Non- ac2ve Ac2ve Total
18 FWER vs FDR FWER P(# true null declared ac2ve > 1) FDR E (# of true null declared ac2ve / # voxels declared ac2ve) Declared ac2ve Declared inac2ve Total Non- ac2ve Ac2ve Total False Discovery Rate Illustra2on: Noise Signal Signal+Noise Control of Per Comparison Rate at 10% 11.3% 11.3% 12.5% 10.8% 11.5% 10.0% 10.7% 11.2% 10.2% 9.5% Percentage of Null Pixels that are False Positives Control of Familywise Error Rate at 10% Occurrence of Familywise Error FWE Control of False Discovery Rate at 10% 6.7% 10.4% 14.9% 9.3% 16.2% 13.8% 14.0% 10.5% 12.2% 8.7% Percentage of Activated Pixels that are False Positives 18
19 Control of Per Comparison Rate at 10% 11.3% 11.3% 12.5% 10.8% 11.5% 10.0% 10.7% 11.2% 10.2% 9.5% Percentage of Null Pixels that are False Positives Control of Familywise Error Rate at 10% Occurrence of Familywise Error FWE Control of False Discovery Rate at 10% 6.7% 10.4% 14.9% 9.3% 16.2% 13.8% 14.0% 10.5% 12.2% 8.7% Percentage of Activated Pixels that are False Positives Control of Per Comparison Rate at 10% 11.3% 11.3% 12.5% 10.8% 11.5% 10.0% 10.7% 11.2% 10.2% 9.5% Percentage of Null Pixels that are False Positives Control of Familywise Error Rate at 10% Occurrence of Familywise Error FWE Control of False Discovery Rate at 10% 6.7% 10.4% 14.9% 9.3% 16.2% 13.8% 14.0% 10.5% 12.2% 8.7% Percentage of Activated Pixels that are False Positives Considera2ons with mul2ple comparisons What sta2s2c you re working with Voxel wise? Cluster wise? What error rate you re controlling Per comparison error rate Family wise error rate False discovery rate 19
20 Correlated data Images typically have correlated voxels # of false posi2ves = 0.05 x (# of independent tests) Extreme example Data are smoothed so much all voxels are iden2cal Only 1 out of 20 data sets would have a false posi2ve Correlated data Images typically have correlated voxels # of false posi2ves = 0.05 x (# of independent tests) Extreme example Data are smoothed so much all voxels are iden2cal Only 1 out of 20 data sets would have a false posi2ve Correlated data Coun2ng false posi2ves becomes tricky, since you don t know the number of independent things 20
21 When data are not correlated P- values computed from simulated null data When data are not correlated Thresholded p< > 4.7% are false posi2ves Correlated data Coun2ng false posi2ves becomes tricky, since you don t know the number of independent things 21
22 Same demo, with smoothed data Thresholded p- value map - > 4.2% are FP Where we re going Review of hypothesis tes2ng introduce mul2ple tes2ng problem Levels of inference (voxel/cluster/peak/set) Types of error rate control (none/fwer/fdr) Family- wise error control approaches (parametric/ nonparametric) FDR Rela2ng all of this to SPM output FWER FWER P(# true null declared ac2ve > 1) FDR E (# of true null declared ac2ve / # voxels declared ac2ve) Declared ac2ve Declared inac2ve Total Non- ac2ve Ac2ve Total
23 FWER Correc2on - Bonferroni Based on the Bonferroni inequality nx P (E 1 or E 2 or...e n ) apple P (E i ) i=1 If P (Y i passes H 0 ) apple /n then nx P (some Y i passes H 0 ) apple P (Y i passes H 0 ) apple i=1 For 100,000 voxels = 0.05/100, 000 = FWER Correc2on - Bonferroni Based on the Bonferroni inequality nx P (E 1 or E 2 or...e n ) apple P (E i ) i=1 If P (Y i passes H 0 ) apple /n then nx P (some Y i passes H 0 ) apple P (Y i passes H 0 ) apple i=1 For 100,000 voxels = 0.05/100, 000 = FWER Correc2on - Bonferroni Based on the Bonferroni inequality nx P (E 1 or E 2 or...e n ) apple P (E i ) i=1 If P (Y i passes H 0 ) apple /n then nx P (some Y i passes H 0 ) apple P (Y i passes H 0 ) apple i=1 For 100,000 voxels = 0.05/100, 000 =
24 FWER Correc2on - Bonferroni Based on the Bonferroni inequality nx P (E 1 or E 2 or...e n ) apple P (E i ) i=1 If P (Y i passes H 0 ) apple /n then nx P (some Y i passes H 0 ) apple P (Y i passes H 0 ) apple i=1 For 100,000 voxels = 0.05/100, 000 = FWER Correc2on - Bonferroni Can be too conserva2ve Bonferroni assumes all tests are independent fmri data tend to be spa2ally correlated # of independent tests < # voxels Smooth data How will the Bonferroni correc2on work with smoothed data? Will false posi2ve rate increase or decrease? 24
25 Ques2ons Why doesn t Bonferroni work well with our imaging data? Why does smoothness make mul2ple comparison correc2on more tricky? FWER Random Field theory Parametric approach to controlling false posi2ves Parametric = there s an equa2on that will spit out the p- value Beyond the scope of this course Tends to be as conserva2ve as Bonferroni FWER Random Field theory Parametric approach to controlling false posi2ves Parametric = there s an equa2on that will spit out the p- value Beyond the scope of this course Tends to be as conserva2ve as Bonferroni 25
26 FWER Random Field theory Parametric approach to controlling false posi2ves Parametric = there s an equa2on that will spit out the p- value Voxelwise version tends to be as conserva2ve as Bonferroni FWER with max sta2s2c FWER & distribu2on of maximum FWER = P(FWE) = P(One or more voxels u H o ) = P(Max voxel u H o ) 100(1- α)%ile of max dist n controls FWER FWER = P(Max voxel u α H o ) α u α α FWER with max sta2s2c FWER & distribu2on of maximum FWER = P(FWE) = P(One or more voxels u H o ) = P(Max voxel u H o ) 100(1- α)%ile of max dist n controls FWER FWER = P(Max voxel u α H o ) α u α α 26
27 FWER with max sta2s2c FWER & distribu2on of maximum FWER = P(FWE) = P(One or more voxels u H o ) = P(Max voxel u H o ) 100(1- α)%ile of max dist n controls FWER FWER = P(Max voxel u α H o ) α u α α FWER with max sta2s2c FWER & distribu2on of maximum FWER = P(FWE) = P(One or more voxels u H o ) = P(Max voxel u H o ) 100(1- α)%ile of max dist n controls FWER FWER = P(Max voxel u α H o ) α u α α FWER with max sta2s2c FWER & distribu2on of maximum FWER = P(FWE) = P(One or more voxels u H o ) = P(Max voxel u H o ) 100(1- α)%ile of max dist n controls FWER FWER = P(Max voxel u α H o ) α u α α 27
28 FWER with max sta2s2c FWER & distribu2on of maximum FWER = P(FWE) = P(One or more voxels u H o ) = P(Max voxel u H o ) 100(1- α)%ile of max dist n controls FWER FWER = P(Max voxel u α H o ) α FWER with max sta2s2c FWER & distribu2on of maximum FWER = P(FWE) = P(One or more voxels u H o ) = P(Max voxel u H o ) 100(1- α)%ile of max dist n controls FWER FWER = P(Max voxel u α H o ) α u α FWER with max sta2s2c FWER & distribu2on of maximum FWER = P(FWE) = P(One or more voxels u H o ) = P(Max voxel u H o ) 100(1- α)%ile of max dist n controls FWER FWER = P(Max voxel u α H o ) α u α α 28
29 FWER MTP Solu2ons: Random Field Theory Euler Characteris2c χ u Topological Measure #blobs - #holes At high thresholds, just counts blobs No holes Never more than 1 blob Random Field FWER = P(Max voxel u H o ) = P(One or more blobs H o ) P(χ u 1 H o ) E(χ u H o ) Threshold Suprathreshold Sets Distribution details Math is hairy! Nichols and Hayasaka 2003 Cao and Worsley 2001 What you need to know Depends on smoothness of your image Must quantify smoothness and it is important to report when using RFT General idea E(χ u ) Mathy stuff *Volume/Smoothness We know what the volume is What is smoothness? 29
30 Smoothness How smooth are the data? Measured by FWHM=[FWHM x, FWHM y, FWHM z ] Starting with white noise smooth with a gaussian How large does the variance of that gaussian need to be such that the smoothness matches your data? RESEL RESolution Element RESEL=FWHM x x FWHM y x FWHM z RESEL count If your voxels were the size of a RESEL, how many are required to fill your volume? 10 voxels, 2.5 voxel FWHM smoothness 4 RESELS voxels FWHM= 2.5 voxels RESEL count=4 30
31 Note about RESEL count Not the number of independent tests Not the magic bullet for a better Bonferroni Re-expression of volume in terms of smoothness We need it, since it is necessary to calculate our p-values Revisit distribution E(χ u ) Mathy stuff *Volume/Smoothness Smoothness is defined in RESELs E(χ u ) is our p-value How does a p-value change as volume increases? How does a p-value change as smoothness increases? RFT adapts For larger volumes it is more strict Multiple comparison problem is worse For smoother data it is less strict Multiple comparison problem is less severe 31
32 Shortcomings of RFT Requires estimating a lot of parameters Random field must be sufficiently smooth If you don t spatially smooth the data enough, RFT doesn t work well I ll cover the Eklund paper later on today! Bonferroni and RFT u RF = 9.87 u Bonf = sig. vox. t 11 Sta2s2c, RF & Bonf. Threshold RFT Voxelwise RFT is rarely used in prac2ce Too conserva2ve Cluster wise RFT is very common We ll learn about cluster stats with permuta2on tes2ng 32
33 FYI If you re using RFT, you probably shouldn t lower the cluster forming threshold Assump2ons could break down If you really want to lower it, switch to nonparameteric approaches SnPM Randomise Ques2ons for you Why do we use the max sta2s2c for mul2ple comparison correc2on? Was this a voxelwise or clusterwise approach? Parametric vs Nonparametric Parametric Assume distribu2on shape Typically 1 or more parameters must be es2mated Nonparametric No assump2on on distribu2on shape Use data to construct distribu2on Related to bootstrap and jackknife, BUT not the same!!! 33
34 Where we re going Review of hypothesis tes2ng introduce mul2ple tes2ng problem Levels of inference (voxel/cluster/peak/set) Types of error rate control (none/fwer/fdr) Family- wise error control approaches (parametric/ nonparametric) FDR Rela2ng all of this to SPM output Permuta2on test Generally can be used when the true distribu2on shape is unknown Data don t follow a normal distribu2on Generally doesn t control for mul2ple comparisons Using in conjunc2on with the max sta2s2c tackles 2 problems Not knowing the structure of the distribu2on Control FWER Permuta2on test Generally can be used when the true distribu2on shape is unknown Data don t follow a normal distribu2on Generally doesn t control for mul2ple comparisons Using in conjunc2on with the max sta2s2c tackles 2 problems Not knowing the structure of the distribu2on Control FWER 34
35 Permuta2on test Generally can be used when the true distribu2on shape is unknown Data don t follow a normal distribu2on Generally doesn t control for mul2ple comparisons Using in conjunc2on with the max sta2s2c tackles 2 problems Not knowing the structure of the distribu2on Control FWER Permuta2on test Without using max sta2s2c So we understand how it generally works With max sta2s2c So we understand how to control FWER Permuta2on test Parametric methods Assume distribu2on of sta2s2c under null hypothesis Nonparametric methods Use data to find distribu2on of sta2s2c under null hypothesis Any sta2s2c! 5% Parametric Null Distribu2on 5% Nonparametric Null Distribu2on 35
36 Permuta2on Test Toy Example Data from voxel in visual s2m. experiment A: Ac2ve, flashing checkerboard B: Baseline, fixa2on 6 blocks, ABABAB Just consider block averages... A B A B A B Null hypothesis H o No experimental effect, A & B labels arbitrary Sta2s2c Mean difference Permuta2on Test Toy Example Under H o Consider all equivalent relabelings AAABBB ABABAB BAAABB BABBAA AABABB ABABBA BAABAB BBAAAB AABBAB ABBAAB BAABBA BBAABA AABBBA ABBABA BABAAB BBABAA ABAABB ABBBAA BABABA BBBAAA Permuta2on Test Toy Example Under H o Consider all equivalent relabelings Compute all possible sta2s2c values AAABBB 4.82 ABABAB 9.45 BAAABB BABBAA AABABB ABABBA 6.97 BAABAB 1.10 BBAAAB 3.15 AABBAB ABBAAB 1.38 BAABBA BBAABA 0.67 AABBBA ABBABA BABAAB BBABAA 3.25 ABAABB 6.86 ABBBAA 1.48 BABABA BBBAAA
37 Permuta2on Test Toy Example Under H o Consider all equivalent relabelings Compute all possible sta2s2c values Find 95%ile of permuta2on distribu2on AAABBB 4.82 ABABAB 9.45 BAAABB BABBAA AABABB ABABBA 6.97 BAABAB 1.10 BBAAAB 3.15 AABBAB ABBAAB 1.38 BAABBA BBAABA 0.67 AABBBA ABBABA BABAAB BBABAA 3.25 ABAABB 6.86 ABBBAA 1.48 BABABA BBBAAA Permuta2on Test Toy Example Under H o Consider all equivalent relabelings Compute all possible sta2s2c values Find 95%ile of permuta2on distribu2on AAABBB 4.82 ABABAB 9.45 BAAABB BABBAA AABABB ABABBA 6.97 BAABAB 1.10 BBAAAB 3.15 AABBAB ABBAAB 1.38 BAABBA BBAABA 0.67 AABBBA ABBABA BABAAB BBABAA 3.25 ABAABB 6.86 ABBBAA 1.48 BABABA BBBAAA Permuta2on Test Toy Example Under H o Consider all equivalent relabelings Compute all possible sta2s2c values Find 95%ile of permuta2on distribu2on
38 Small Sample Sizes Permutation test doesn t work well with small sample sizes Possible p-values for previous example: 0.05, 0.1, 0.15, 0.2, etc Tends to be conservative for small sample sizes Permuta2on Test & Exchangeability Exchangeability is fundamental Def: Distribu2on of the data unperturbed by permuta2on Under H 0, exchangeability jus2fies permu2ng data Allows us to build permuta2on distribu2on Permuta2on Test & Exchangeability Subjects are exchangeable Under Ho, each subject s A/B labels can be flipped fmri scans are not exchangeable under Ho If no signal, can we permute over 2me? No, permu2ng disrupts order, temporal autocorrela2on 38
39 Permuta2on Test & Exchangeability Two sample t test Compare subjects in group 1 to subjects in group 2 Randomly assign group labels in permuta2ons One sample t test Randomly flip sign of values for some subjects Ques2ons for you What is permuted for a 1- sample t- test? What is permuted for a 2- sample t- test? What is permuted for a correla2on? Why are small sample sizes problema2c for permuta2on tes2ng? Controlling FWER: Permuta2on Test Parametric methods Assume distribu2on of max sta2s2c under null hypothesis Nonparametric methods Use data to find distribu2on of max sta2s2c under null hypothesis Again, any max sta2s2c! 5% Parametric Null Max Distribu2on 5% Nonparametric Null Max Distribu2on 39
40 Permuta2on Test Other Sta2s2cs Collect max distribu2on To find threshold that controls FWER Consider smoothed variance t sta2s2c To regularize low- df variance es2mate Max sta2s2c for imaging data 1. Compute your sta2s2cs map for original data 2. Shuffle labels and compute sta2s2cs map 3. Save the largest sta2s2c over the whole brain 4. Repeat steps 2-3 many 2mes ( ,000) 5. Use distribu2on of stats over permuta2ons to compute threshold 6. Apply threshold to map from step 1 Max sta2s2c for imaging data 1. Compute your sta2s2cs map for original data 2. Shuffle labels and compute sta2s2cs map 3. Save the largest sta2s2c over the whole brain 4. Repeat steps 2-3 many 2mes ( ,000) 5. Use distribu2on of stats over permuta2ons to compute threshold 6. Apply threshold to map from step 1 40
41 Max sta2s2c for imaging data 1. Compute your sta2s2cs map for original data 2. Shuffle labels and compute sta2s2cs map 3. Save the largest sta2s2c over the whole brain 4. Repeat steps 2-3 many 2mes ( ,000) 5. Use distribu2on of stats over permuta2ons to compute threshold 6. Apply threshold to map from step 1 Max sta2s2c for imaging data 1. Compute your sta2s2cs map for original data 2. Shuffle labels and compute sta2s2cs map 3. Save the largest sta2s2c over the whole brain 4. Repeat steps 2-3 many 2mes ( ,000) 5. Use distribu2on of stats over permuta2ons to compute threshold 6. Apply threshold to map from step 1 Max sta2s2c for imaging data 1. Compute your sta2s2cs map for original data 2. Shuffle labels and compute sta2s2cs map 3. Save the largest sta2s2c over the whole brain 4. Repeat steps 2-3 many 2mes ( ,000) 5. Use distribu2on of stats over permuta2ons to compute threshold 6. Apply threshold to map from step 1 41
42 Max sta2s2c for imaging data 1. Compute your sta2s2cs map for original data 2. Shuffle labels and compute sta2s2cs map 3. Save the largest sta2s2c over the whole brain 4. Repeat steps 2-3 many 2mes ( ,000) 5. Use distribu2on of stats over permuta2ons to compute threshold 6. Apply threshold to map from step 1 Permuta2on Test Smoothed Variance t Collect max distribu2on To find threshold that controls FWER Consider smoothed variance t sta2s2c mean difference variance t-statistic Permuta2on Test Smoothed Variance t Collect max distribu2on To find threshold that controls FWER Consider smoothed variance t sta2s2c mean difference smoothed variance Smoothed Variance t-statistic 42
43 Permuta2on Test Example fmri Study of Working Memory 12 subjects, block design Marshuetz et al (2000) Item Recogni2on Ac2ve: View five le?ers, 2s pause, view probe le?er, respond Baseline: View XXXXX, 2s pause, view Y or N, respond Second Level RFX Difference image, A- B constructed for each subject One sample t test UBKDA Active... Baseline XXXXX D yes... N no Permuta2on Test Example Permute! 2 12 = 4,096 ways to flip 12 A/B labels For each, note maximum of t image. Permuta2on Distribu2on Maximum t Maximum Intensity Projec2on Thresholded t u Perm = sig. vox. t 11 Sta2s2c, Nonparametric Threshold u RF = 9.87 u Bonf = sig. vox. t 11 Sta2s2c, RF & Bonf. Threshold 378 sig. vox. Smoothed Variance t Sta2s2c, Nonparametric Threshold RFT threshold is conservative (not smooth enough, d.f. too small) Permutation test is more efficient than Bonferroni since it accounts for smoothness Smooth variance is more efficient for small d.f. 43
44 u Perm = sig. vox. t 11 Sta2s2c, Nonparametric Threshold u RF = 9.87 u Bonf = sig. vox. t 11 Sta2s2c, RF & Bonf. Threshold 378 sig. vox. Smoothed Variance t Sta2s2c, Nonparametric Threshold RFT threshold is conservative (not smooth enough, d.f. too small) Permutation test is more efficient than Bonferroni since it accounts for smoothness Smooth variance is more efficient for small d.f. u Perm = sig. vox. t 11 Sta2s2c, Nonparametric Threshold u RF = 9.87 u Bonf = sig. vox. t 11 Sta2s2c, RF & Bonf. Threshold 378 sig. vox. Smoothed Variance t Sta2s2c, Nonparametric Threshold RFT threshold is conservative (not smooth enough, d.f. too small) Permutation test is more efficient than Bonferroni since it accounts for smoothness Smooth variance is more efficient for small d.f. u Perm = sig. vox. t 11 Sta2s2c, Nonparametric Threshold u RF = 9.87 u Bonf = sig. vox. t 11 Sta2s2c, RF & Bonf. Threshold 378 sig. vox. Smoothed Variance t Sta2s2c, Nonparametric Threshold RFT threshold is conservative (not smooth enough, d.f. too small) Permutation test is more efficient than Bonferroni since it accounts for smoothness Smooth variance is more efficient for small d.f. 44
45 Permuta2on test cluster sta2s2c Two step- process Define clusters by arbitrary threshold u clus u clus space Permuta2on test cluster sta2s2c Two step- process Define clusters by arbitrary threshold u clus Retain clusters larger than α- level threshold k α u clus space Cluster not significant k α k α Cluster significant Permuta2on test cluster sta2s2cs Cluster size Simply count how many voxels are in the sta2s2c Cluster mass Sum up the sta2s2cs in the cluster 45
46 Permuta2on test cluster sta2s2cs 1. Find clusters with original data 2. Permute labels 3. Compute sta2s2cs 4. Apply cluster- forming threshold 5. Compute cluster sta2s2cs 6. Save largest (cluster size or mass) 7. Repeat steps 2-3 many 2mes ( ,000) 8. Use distribu2on from step 7 to find cluster (size or mass) threshold Permuta2on test cluster sta2s2cs 1. Find clusters with original data 2. Permute labels 3. Compute sta2s2cs 4. Apply cluster- forming threshold 5. Compute cluster sta2s2cs 6. Save largest (cluster size or mass) 7. Repeat steps 2-3 many 2mes ( ,000) 8. Use distribu2on from step 7 to find cluster (size or mass) threshold Permuta2on test cluster sta2s2cs 1. Find clusters with original data 2. Permute labels 3. Compute sta2s2cs 4. Apply cluster- forming threshold 5. Compute cluster sta2s2cs 6. Save largest (cluster size or mass) 7. Repeat steps 2-3 many 2mes ( ,000) 8. Use distribu2on from step 7 to find cluster (size or mass) threshold 46
47 Permuta2on test cluster sta2s2cs 1. Find clusters with original data 2. Permute labels 3. Compute sta2s2cs 4. Apply cluster- forming threshold 5. Compute cluster sta2s2cs 6. Save largest (cluster size or mass) 7. Repeat steps 2-3 many 2mes ( ,000) 8. Use distribu2on from step 7 to find cluster (size or mass) threshold Permuta2on test cluster sta2s2cs 1. Find clusters with original data 2. Permute labels 3. Compute sta2s2cs 4. Apply cluster- forming threshold 5. Compute cluster sta2s2cs 6. Save largest (cluster size or mass) 7. Repeat steps 2-3 many 2mes ( ,000) 8. Use distribu2on from step 7 to find cluster (size or mass) threshold Permuta2on test cluster sta2s2cs 1. Find clusters with original data 2. Permute labels 3. Compute sta2s2cs 4. Apply cluster- forming threshold 5. Compute cluster sta2s2cs 6. Save largest (cluster size or mass) 7. Repeat steps 2-3 many 2mes ( ,000) 8. Use distribu2on from step 7 to find cluster (size or mass) threshold 47
48 Permuta2on test cluster sta2s2cs 1. Find clusters with original data 2. Permute labels 3. Compute sta2s2cs 4. Apply cluster- forming threshold 5. Compute cluster sta2s2cs 6. Save largest (cluster size or mass) 7. Repeat steps 2-3 many 2mes ( ,000) 8. Use distribu2on from step 7 to find cluster (size or mass) threshold Permuta2on test cluster sta2s2cs 1. Find clusters with original data 2. Permute labels 3. Compute sta2s2cs 4. Apply cluster- forming threshold 5. Compute cluster sta2s2cs 6. Save largest (cluster size or mass) 7. Repeat steps 2-3 many 2mes ( ,000) 8. Use distribu2on from step 7 to find cluster (size or mass) threshold & apply to step 1 Ques2ons for you Why don t permuta2on tests, alone, fix mul2ple comparisons? What did we need to use to address mul2ple comparisons? How are the voxelwise and clusterwise permuta2on tests set up? 48
49 Where we re going Review of hypothesis tes2ng introduce mul2ple tes2ng problem Levels of inference (voxel/cluster/peak/set) Types of error rate control (none/fwer/fdr) Family- wise error control approaches (parametric/ nonparametric) FDR Rela2ng all of this to SPM output FWER vs FDR FWER P(# true null declared ac2ve > 1) FDR E (# of true null declared ac2ve / # voxels declared ac2ve) Declared ac2ve Declared inac2ve Total Non- ac2ve Ac2ve Total Controlling FDR Tends to be less conserva2ve than controlling FWER What rate is appropriate? Imagers use 5%...out of habit FDR people I ve met outside of imaging ozen use higher values Decide before you threshold your data Don t choose what makes your data look good 49
50 /15/16 Benjamini & Hochberg Procedure Select desired limit α on FDR Order p- values, p (1) p (2)... p (v) Let r be largest i such that p (i) i/v α Reject all hypotheses corresponding to p (1),..., p (r). p-value p (i) 0 i/v 1 Benjamini & Hochberg Procedure Select desired limit α on FDR Order p- values, p (1) p (2)... p (v) Let r be largest i such that p (i) i/v α Reject all hypotheses corresponding to p (1),..., p (r). p-value 0 p (i) i/v α 1 i/v Benjamini & Hochberg Procedure Select desired limit α on FDR Order p- values, p (1) p (2)... p (v) Let r be largest i such that p (i) i/v α Reject all hypotheses corresponding to p (1),..., p (r). p-value 0 p (i) i/v α i/v 1 50
51 FDR Example FWER Perm. Thresh. = voxels FDR Threshold = ,073 voxels Where we re going Review of hypothesis tes2ng introduce mul2ple tes2ng problem Levels of inference (voxel/cluster/peak/set) Types of error rate control (none/fwer/fdr) Family- wise error control approaches (parametric/ nonparametric) FDR Rela2ng all of this to SPM output Guess what? Now you have the knowledge needed to understand a huge/daun2ng table SPM spits out! Let s do it 51
52 SPM output SPM output Which level of inference is missing? SPM output what exci2ng conclusion can we make? 52
53 SPM output Recall: FWE correc2on shown earlier was super conserva2ve compared to FDR. Why does this look different? SPM output What do you think K E is? What sta2s2c does the p- value correspond to? SPM output The uncorrected stat doesn t take the search volume into account 53
54 SPM output See the note at the bo?om? SPM output Do any clusters have more than one peak? SPM output Last, but not least, you ll use this in lab. This is used to threshold clusters so you can look at only the significant ones 54
55 SPM output Compare this threshold to the FWE p- values for cluster stats That s it! Ques2ons? 55
Controlling for multiple comparisons in imaging analysis. Wednesday, Lecture 2 Jeanette Mumford University of Wisconsin - Madison
 Controlling for multiple comparisons in imaging analysis Wednesday, Lecture 2 Jeanette Mumford University of Wisconsin - Madison Motivation Run 100 hypothesis tests on null data using p
Controlling for multiple comparisons in imaging analysis Wednesday, Lecture 2 Jeanette Mumford University of Wisconsin - Madison Motivation Run 100 hypothesis tests on null data using p
Multiple Testing and Thresholding
 Multiple Testing and Thresholding NITP, 2010 Thanks for the slides Tom Nichols! Overview Multiple Testing Problem Which of my 100,000 voxels are active? Two methods for controlling false positives Familywise
Multiple Testing and Thresholding NITP, 2010 Thanks for the slides Tom Nichols! Overview Multiple Testing Problem Which of my 100,000 voxels are active? Two methods for controlling false positives Familywise
Controlling for mul-ple comparisons in imaging analysis. Wednesday, Lecture 2 Jeane:e Mumford University of Wisconsin - Madison
 Controlling for mul-ple comparisons in imaging analysis Wednesday, Lecture 2 Jeane:e Mumford University of Wisconsin - Madison Where we re going Review of hypothesis tes-ng introduce mul-ple tes-ng problem
Controlling for mul-ple comparisons in imaging analysis Wednesday, Lecture 2 Jeane:e Mumford University of Wisconsin - Madison Where we re going Review of hypothesis tes-ng introduce mul-ple tes-ng problem
Multiple Testing and Thresholding
 Multiple Testing and Thresholding UCLA Advanced NeuroImaging Summer School, 2007 Thanks for the slides Tom Nichols! Overview Multiple Testing Problem Which of my 100,000 voxels are active? Two methods
Multiple Testing and Thresholding UCLA Advanced NeuroImaging Summer School, 2007 Thanks for the slides Tom Nichols! Overview Multiple Testing Problem Which of my 100,000 voxels are active? Two methods
Multiple Testing and Thresholding
 Multiple Testing and Thresholding NITP, 2009 Thanks for the slides Tom Nichols! Overview Multiple Testing Problem Which of my 100,000 voxels are active? Two methods for controlling false positives Familywise
Multiple Testing and Thresholding NITP, 2009 Thanks for the slides Tom Nichols! Overview Multiple Testing Problem Which of my 100,000 voxels are active? Two methods for controlling false positives Familywise
Contents. comparison. Multiple comparison problem. Recap & Introduction Inference & multiple. «Take home» message. DISCOS SPM course, CRC, Liège, 2009
 DISCOS SPM course, CRC, Liège, 2009 Contents Multiple comparison problem Recap & Introduction Inference & multiple comparison «Take home» message C. Phillips, Centre de Recherches du Cyclotron, ULg, Belgium
DISCOS SPM course, CRC, Liège, 2009 Contents Multiple comparison problem Recap & Introduction Inference & multiple comparison «Take home» message C. Phillips, Centre de Recherches du Cyclotron, ULg, Belgium
New and best-practice approaches to thresholding
 New and best-practice approaches to thresholding Thomas Nichols, Ph.D. Department of Statistics & Warwick Manufacturing Group University of Warwick FIL SPM Course 17 May, 2012 1 Overview Why threshold?
New and best-practice approaches to thresholding Thomas Nichols, Ph.D. Department of Statistics & Warwick Manufacturing Group University of Warwick FIL SPM Course 17 May, 2012 1 Overview Why threshold?
Statistical Methods in functional MRI. Standard Analysis. Data Processing Pipeline. Multiple Comparisons Problem. Multiple Comparisons Problem
 Statistical Methods in fnctional MRI Lectre 7: Mltiple Comparisons 04/3/13 Martin Lindqist Department of Biostatistics Johns Hopkins University Data Processing Pipeline Standard Analysis Data Acqisition
Statistical Methods in fnctional MRI Lectre 7: Mltiple Comparisons 04/3/13 Martin Lindqist Department of Biostatistics Johns Hopkins University Data Processing Pipeline Standard Analysis Data Acqisition
Power analysis. Wednesday, Lecture 3 Jeanette Mumford University of Wisconsin - Madison
 Power analysis Wednesday, Lecture 3 Jeanette Mumford University of Wisconsin - Madison Power Analysis-Why? To answer the question. How many subjects do I need for my study? How many runs per subject should
Power analysis Wednesday, Lecture 3 Jeanette Mumford University of Wisconsin - Madison Power Analysis-Why? To answer the question. How many subjects do I need for my study? How many runs per subject should
Multiple comparisons problem and solutions
 Multiple comparisons problem and solutions James M. Kilner http://sites.google.com/site/kilnerlab/home What is the multiple comparisons problem How can it be avoided Ways to correct for the multiple comparisons
Multiple comparisons problem and solutions James M. Kilner http://sites.google.com/site/kilnerlab/home What is the multiple comparisons problem How can it be avoided Ways to correct for the multiple comparisons
Extending the GLM. Outline. Mixed effects motivation Evaluating mixed effects methods Three methods. Conclusions. Overview
 Extending the GLM So far, we have considered the GLM for one run in one subject The same logic can be applied to multiple runs and multiple subjects GLM Stats For any given region, we can evaluate the
Extending the GLM So far, we have considered the GLM for one run in one subject The same logic can be applied to multiple runs and multiple subjects GLM Stats For any given region, we can evaluate the
Correction for multiple comparisons. Cyril Pernet, PhD SBIRC/SINAPSE University of Edinburgh
 Correction for multiple comparisons Cyril Pernet, PhD SBIRC/SINAPSE University of Edinburgh Overview Multiple comparisons correction procedures Levels of inferences (set, cluster, voxel) Circularity issues
Correction for multiple comparisons Cyril Pernet, PhD SBIRC/SINAPSE University of Edinburgh Overview Multiple comparisons correction procedures Levels of inferences (set, cluster, voxel) Circularity issues
Introductory Concepts for Voxel-Based Statistical Analysis
 Introductory Concepts for Voxel-Based Statistical Analysis John Kornak University of California, San Francisco Department of Radiology and Biomedical Imaging Department of Epidemiology and Biostatistics
Introductory Concepts for Voxel-Based Statistical Analysis John Kornak University of California, San Francisco Department of Radiology and Biomedical Imaging Department of Epidemiology and Biostatistics
Linear Models in Medical Imaging. John Kornak MI square February 22, 2011
 Linear Models in Medical Imaging John Kornak MI square February 22, 2011 Acknowledgement / Disclaimer Many of the slides in this lecture have been adapted from slides available in talks available on the
Linear Models in Medical Imaging John Kornak MI square February 22, 2011 Acknowledgement / Disclaimer Many of the slides in this lecture have been adapted from slides available in talks available on the
Linear Models in Medical Imaging. John Kornak MI square February 19, 2013
 Linear Models in Medical Imaging John Kornak MI square February 19, 2013 Acknowledgement / Disclaimer Many of the slides in this lecture have been adapted from slides available in talks available on the
Linear Models in Medical Imaging John Kornak MI square February 19, 2013 Acknowledgement / Disclaimer Many of the slides in this lecture have been adapted from slides available in talks available on the
Linear Models in Medical Imaging. John Kornak MI square February 23, 2010
 Linear Models in Medical Imaging John Kornak MI square February 23, 2010 Acknowledgement / Disclaimer Many of the slides in this lecture have been adapted from slides available in talks available on the
Linear Models in Medical Imaging John Kornak MI square February 23, 2010 Acknowledgement / Disclaimer Many of the slides in this lecture have been adapted from slides available in talks available on the
Group Sta*s*cs in MEG/EEG
 Group Sta*s*cs in MEG/EEG Will Woods NIF Fellow Brain and Psychological Sciences Research Centre Swinburne University of Technology A Cau*onary tale. A Cau*onary tale. A Cau*onary tale. Overview Introduc*on
Group Sta*s*cs in MEG/EEG Will Woods NIF Fellow Brain and Psychological Sciences Research Centre Swinburne University of Technology A Cau*onary tale. A Cau*onary tale. A Cau*onary tale. Overview Introduc*on
Medical Image Analysis
 Medical Image Analysis Instructor: Moo K. Chung mchung@stat.wisc.edu Lecture 10. Multiple Comparisons March 06, 2007 This lecture will show you how to construct P-value maps fmri Multiple Comparisons 4-Dimensional
Medical Image Analysis Instructor: Moo K. Chung mchung@stat.wisc.edu Lecture 10. Multiple Comparisons March 06, 2007 This lecture will show you how to construct P-value maps fmri Multiple Comparisons 4-Dimensional
Nonparametric Permutation Tests For Functional Neuroimaging: APrimer with Examples
 Human Brain Mapping 15:1 25(2001) Nonparametric Permutation Tests For Functional Neuroimaging: APrimer with Examples Thomas E. Nichols 1 and Andrew P. Holmes 2,3 * 1 Department of Biostatistics, University
Human Brain Mapping 15:1 25(2001) Nonparametric Permutation Tests For Functional Neuroimaging: APrimer with Examples Thomas E. Nichols 1 and Andrew P. Holmes 2,3 * 1 Department of Biostatistics, University
7/15/2016 ARE YOUR ANALYSES TOO WHY IS YOUR ANALYSIS PARAMETRIC? PARAMETRIC? That s not Normal!
 ARE YOUR ANALYSES TOO PARAMETRIC? That s not Normal! Martin M Monti http://montilab.psych.ucla.edu WHY IS YOUR ANALYSIS PARAMETRIC? i. Optimal power (defined as the probability to detect a real difference)
ARE YOUR ANALYSES TOO PARAMETRIC? That s not Normal! Martin M Monti http://montilab.psych.ucla.edu WHY IS YOUR ANALYSIS PARAMETRIC? i. Optimal power (defined as the probability to detect a real difference)
Linear Models in Medical Imaging. John Kornak MI square February 21, 2012
 Linear Models in Medical Imaging John Kornak MI square February 21, 2012 Acknowledgement / Disclaimer Many of the slides in this lecture have been adapted from slides available in talks available on the
Linear Models in Medical Imaging John Kornak MI square February 21, 2012 Acknowledgement / Disclaimer Many of the slides in this lecture have been adapted from slides available in talks available on the
Statistical inference on images
 7 Statistical inference on images The goal of statistical inference is to make decisions based on our data, while accounting for uncertainty due to noise in the data. From a broad perspective, statistical
7 Statistical inference on images The goal of statistical inference is to make decisions based on our data, while accounting for uncertainty due to noise in the data. From a broad perspective, statistical
Advances in FDR for fmri -p.1
 Advances in FDR for fmri Ruth Heller Department of Statistics, University of Pennsylvania Joint work with: Yoav Benjamini, Nava Rubin, Damian Stanley, Daniel Yekutieli, Yulia Golland, Rafael Malach Advances
Advances in FDR for fmri Ruth Heller Department of Statistics, University of Pennsylvania Joint work with: Yoav Benjamini, Nava Rubin, Damian Stanley, Daniel Yekutieli, Yulia Golland, Rafael Malach Advances
SnPM is an SPM toolbox developed by Andrew Holmes & Tom Nichols
 1 of 14 3/30/2005 9:24 PM SnPM A Worked fmri Example SnPM is an SPM toolbox developed by Andrew Holmes & Tom Nichols This page... introduction example data background design setup computation viewing results
1 of 14 3/30/2005 9:24 PM SnPM A Worked fmri Example SnPM is an SPM toolbox developed by Andrew Holmes & Tom Nichols This page... introduction example data background design setup computation viewing results
CHAPTER 2. Morphometry on rodent brains. A.E.H. Scheenstra J. Dijkstra L. van der Weerd
 CHAPTER 2 Morphometry on rodent brains A.E.H. Scheenstra J. Dijkstra L. van der Weerd This chapter was adapted from: Volumetry and other quantitative measurements to assess the rodent brain, In vivo NMR
CHAPTER 2 Morphometry on rodent brains A.E.H. Scheenstra J. Dijkstra L. van der Weerd This chapter was adapted from: Volumetry and other quantitative measurements to assess the rodent brain, In vivo NMR
fmri Analysis Sackler Ins2tute 2011
 fmri Analysis Sackler Ins2tute 2011 How do we get from this to this? How do we get from this to this? And what are those colored blobs we re all trying to see, anyway? Raw fmri data straight from the scanner
fmri Analysis Sackler Ins2tute 2011 How do we get from this to this? How do we get from this to this? And what are those colored blobs we re all trying to see, anyway? Raw fmri data straight from the scanner
Statistical Methods in functional MRI. False Discovery Rate. Issues with FWER. Lecture 7.2: Multiple Comparisons ( ) 04/25/13
 Statistical Methods in functional MRI Lecture 7.2: Multiple Comparisons 04/25/13 Martin Lindquist Department of iostatistics Johns Hopkins University Issues with FWER Methods that control the FWER (onferroni,
Statistical Methods in functional MRI Lecture 7.2: Multiple Comparisons 04/25/13 Martin Lindquist Department of iostatistics Johns Hopkins University Issues with FWER Methods that control the FWER (onferroni,
A Non-Parametric Approach
 Andrew P. Holmes. Ph.D., 1994. Chapter Six A Non-Parametric Approach In this chapter, a non-parametric approach to assessing functional mapping experiments is presented. A multiple comparisons randomisation
Andrew P. Holmes. Ph.D., 1994. Chapter Six A Non-Parametric Approach In this chapter, a non-parametric approach to assessing functional mapping experiments is presented. A multiple comparisons randomisation
Group (Level 2) fmri Data Analysis - Lab 4
 Group (Level 2) fmri Data Analysis - Lab 4 Index Goals of this Lab Before Getting Started The Chosen Ten Checking Data Quality Create a Mean Anatomical of the Group Group Analysis: One-Sample T-Test Examine
Group (Level 2) fmri Data Analysis - Lab 4 Index Goals of this Lab Before Getting Started The Chosen Ten Checking Data Quality Create a Mean Anatomical of the Group Group Analysis: One-Sample T-Test Examine
First-level fmri modeling
 First-level fmri modeling Monday, Lecture 3 Jeanette Mumford University of Wisconsin - Madison What do we need to remember from the last lecture? What is the general structure of a t- statistic? How about
First-level fmri modeling Monday, Lecture 3 Jeanette Mumford University of Wisconsin - Madison What do we need to remember from the last lecture? What is the general structure of a t- statistic? How about
The Effect of Correlation and Error Rate Specification on Thresholding Methods in fmri Analysis
 The Effect of Correlation and Error Rate Specification on Thresholding Methods in fmri Analysis Brent R. Logan and Daniel B. Rowe, Division of Biostatistics and Department of Biophysics Division of Biostatistics
The Effect of Correlation and Error Rate Specification on Thresholding Methods in fmri Analysis Brent R. Logan and Daniel B. Rowe, Division of Biostatistics and Department of Biophysics Division of Biostatistics
Introduction to Neuroimaging Janaina Mourao-Miranda
 Introduction to Neuroimaging Janaina Mourao-Miranda Neuroimaging techniques have changed the way neuroscientists address questions about functional anatomy, especially in relation to behavior and clinical
Introduction to Neuroimaging Janaina Mourao-Miranda Neuroimaging techniques have changed the way neuroscientists address questions about functional anatomy, especially in relation to behavior and clinical
Network statistics and thresholding
 Network statistics and thresholding Andrew Zalesky azalesky@unimelb.edu.au HBM Educational Course June 25, 2017 Network thresholding Unthresholded Moderate thresholding Severe thresholding Strong link
Network statistics and thresholding Andrew Zalesky azalesky@unimelb.edu.au HBM Educational Course June 25, 2017 Network thresholding Unthresholded Moderate thresholding Severe thresholding Strong link
Multiple Linear Regression: Global tests and Multiple Testing
 Multiple Linear Regression: Global tests and Multiple Testing Author: Nicholas G Reich, Jeff Goldsmith This material is part of the statsteachr project Made available under the Creative Commons Attribution-ShareAlike
Multiple Linear Regression: Global tests and Multiple Testing Author: Nicholas G Reich, Jeff Goldsmith This material is part of the statsteachr project Made available under the Creative Commons Attribution-ShareAlike
Introduc)on to fmri. Natalia Zaretskaya
 Introduc)on to fmri Natalia Zaretskaya Content fmri signal fmri versus neural ac)vity A classical experiment: flickering checkerboard Preprocessing Univariate analysis Single- subject analysis Group analysis
Introduc)on to fmri Natalia Zaretskaya Content fmri signal fmri versus neural ac)vity A classical experiment: flickering checkerboard Preprocessing Univariate analysis Single- subject analysis Group analysis
MS&E 226: Small Data
 MS&E 226: Small Data Lecture 14: Introduction to hypothesis testing (v2) Ramesh Johari ramesh.johari@stanford.edu 1 / 10 Hypotheses 2 / 10 Quantifying uncertainty Recall the two key goals of inference:
MS&E 226: Small Data Lecture 14: Introduction to hypothesis testing (v2) Ramesh Johari ramesh.johari@stanford.edu 1 / 10 Hypotheses 2 / 10 Quantifying uncertainty Recall the two key goals of inference:
Introduc)on to FreeSurfer h0p://surfer.nmr.mgh.harvard.edu. Jenni Pacheco.
 Introduc)on to FreeSurfer h0p://surfer.nmr.mgh.harvard.edu Jenni Pacheco jpacheco@mail.utexas.edu Outline Processing Stages Command line Stream Assemble Data (mris_preproc, mri_surf2surf) Design/Contrast
Introduc)on to FreeSurfer h0p://surfer.nmr.mgh.harvard.edu Jenni Pacheco jpacheco@mail.utexas.edu Outline Processing Stages Command line Stream Assemble Data (mris_preproc, mri_surf2surf) Design/Contrast
fmri Basics: Single Subject Analysis
 fmri Basics: Single Subject Analysis This session is intended to give an overview of the basic process of setting up a general linear model for a single subject. This stage of the analysis is also variously
fmri Basics: Single Subject Analysis This session is intended to give an overview of the basic process of setting up a general linear model for a single subject. This stage of the analysis is also variously
Cluster failure: Why fmri inferences for spatial extent have inflated false positive rates
 Supporting Information Appendix Cluster failure: Why fmri inferences for spatial extent have inflated false positive rates Anders Eklund, Thomas Nichols, Hans Knutsson Methods Resting state fmri data Resting
Supporting Information Appendix Cluster failure: Why fmri inferences for spatial extent have inflated false positive rates Anders Eklund, Thomas Nichols, Hans Knutsson Methods Resting state fmri data Resting
Controlling the Familywise Error Rate in Functional Neuroimaging: A Comparative Review
 Controlling the Familywise Error Rate in Functional Neuroimaging: A Comparative Review Thomas Nichols & Satoru Hayasaka Departments of Biostatistics University of Michigan, Ann Arbor, MI 48109, U.S.A.
Controlling the Familywise Error Rate in Functional Neuroimaging: A Comparative Review Thomas Nichols & Satoru Hayasaka Departments of Biostatistics University of Michigan, Ann Arbor, MI 48109, U.S.A.
arxiv: v1 [stat.ap] 1 Jun 2016
![arxiv: v1 [stat.ap] 1 Jun 2016 arxiv: v1 [stat.ap] 1 Jun 2016](/thumbs/81/83094240.jpg) Permutation-based cluster size correction for voxel-based lesion-symptom mapping arxiv:1606.00475v1 [stat.ap] 1 Jun 2016 June 3, 2016 Daniel Mirman a,b,1 Jon-Frederick Landrigan a Spiro Kokolis a Sean
Permutation-based cluster size correction for voxel-based lesion-symptom mapping arxiv:1606.00475v1 [stat.ap] 1 Jun 2016 June 3, 2016 Daniel Mirman a,b,1 Jon-Frederick Landrigan a Spiro Kokolis a Sean
Network Analysis Integra2ve Genomics module
 Network Analysis Integra2ve Genomics module Michael Inouye Centre for Systems Genomics University of Melbourne, Australia Summer Ins@tute in Sta@s@cal Gene@cs 2016 SeaBle, USA @minouye271 inouyelab.org
Network Analysis Integra2ve Genomics module Michael Inouye Centre for Systems Genomics University of Melbourne, Australia Summer Ins@tute in Sta@s@cal Gene@cs 2016 SeaBle, USA @minouye271 inouyelab.org
Lab 5: Multi Subject fmri
 USA SPM Short Course April 6-8, 2005 Yale University Lab 5: Multi Subject fmri Rik Henson mailto:rik.henson@mrc-cbu.cam.ac.uk MRC Cognition & Brain Sciences Unit, Cambridge, UK CONTENTS Goals Introduction
USA SPM Short Course April 6-8, 2005 Yale University Lab 5: Multi Subject fmri Rik Henson mailto:rik.henson@mrc-cbu.cam.ac.uk MRC Cognition & Brain Sciences Unit, Cambridge, UK CONTENTS Goals Introduction
Pa#ern Recogni-on for Neuroimaging Toolbox
 Pa#ern Recogni-on for Neuroimaging Toolbox Pa#ern Recogni-on Methods: Basics João M. Monteiro Based on slides from Jessica Schrouff and Janaina Mourão-Miranda PRoNTo course UCL, London, UK 2017 Outline
Pa#ern Recogni-on for Neuroimaging Toolbox Pa#ern Recogni-on Methods: Basics João M. Monteiro Based on slides from Jessica Schrouff and Janaina Mourão-Miranda PRoNTo course UCL, London, UK 2017 Outline
Figure 1. Comparison of the frequency of centrality values for central London and our region of Soho studied. The comparison shows that Soho falls
 A B C Figure 1. Comparison of the frequency of centrality values for central London and our region of Soho studied. The comparison shows that Soho falls well within the distribution for London s streets.
A B C Figure 1. Comparison of the frequency of centrality values for central London and our region of Soho studied. The comparison shows that Soho falls well within the distribution for London s streets.
General Factorial Models
 In Chapter 8 in Oehlert STAT:5201 Week 9 - Lecture 1 1 / 31 It is possible to have many factors in a factorial experiment. We saw some three-way factorials earlier in the DDD book (HW 1 with 3 factors:
In Chapter 8 in Oehlert STAT:5201 Week 9 - Lecture 1 1 / 31 It is possible to have many factors in a factorial experiment. We saw some three-way factorials earlier in the DDD book (HW 1 with 3 factors:
General Factorial Models
 In Chapter 8 in Oehlert STAT:5201 Week 9 - Lecture 2 1 / 34 It is possible to have many factors in a factorial experiment. In DDD we saw an example of a 3-factor study with ball size, height, and surface
In Chapter 8 in Oehlert STAT:5201 Week 9 - Lecture 2 1 / 34 It is possible to have many factors in a factorial experiment. In DDD we saw an example of a 3-factor study with ball size, height, and surface
Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question.
 (1) conn - Functional connectivity toolbox v1.0 Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question. 2) in "Basic" enter "1" subject, "6" seconds
(1) conn - Functional connectivity toolbox v1.0 Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question. 2) in "Basic" enter "1" subject, "6" seconds
R-Square Coeff Var Root MSE y Mean
 STAT:50 Applied Statistics II Exam - Practice 00 possible points. Consider a -factor study where each of the factors has 3 levels. The factors are Diet (,,3) and Drug (A,B,C) and there are n = 3 observations
STAT:50 Applied Statistics II Exam - Practice 00 possible points. Consider a -factor study where each of the factors has 3 levels. The factors are Diet (,,3) and Drug (A,B,C) and there are n = 3 observations
9.2 Types of Errors in Hypothesis testing
 9.2 Types of Errors in Hypothesis testing 1 Mistakes we could make As I mentioned, when we take a sample we won t be 100% sure of something because we do not take a census (we only look at information
9.2 Types of Errors in Hypothesis testing 1 Mistakes we could make As I mentioned, when we take a sample we won t be 100% sure of something because we do not take a census (we only look at information
An improved theoretical P-value for SPMs based on discrete local maxima
 An improved theoretical P-value for SPMs based on discrete local maxima K.J. Worsley August 9, 25 Department of Mathematics and Statistics, McGill University, 85 Sherbrooke St. West, Montreal, Québec,
An improved theoretical P-value for SPMs based on discrete local maxima K.J. Worsley August 9, 25 Department of Mathematics and Statistics, McGill University, 85 Sherbrooke St. West, Montreal, Québec,
Efficiency and design optimization
 Efficiency and design optimization Tuesday, Lecture 3 Jeanette Mumford University of Wisconsin - Madison Thanks to Tom Liu for letting me use some of his slides! What is the best way to increase your power?
Efficiency and design optimization Tuesday, Lecture 3 Jeanette Mumford University of Wisconsin - Madison Thanks to Tom Liu for letting me use some of his slides! What is the best way to increase your power?
Search Engines. Informa1on Retrieval in Prac1ce. Annota1ons by Michael L. Nelson
 Search Engines Informa1on Retrieval in Prac1ce Annota1ons by Michael L. Nelson All slides Addison Wesley, 2008 Evalua1on Evalua1on is key to building effec$ve and efficient search engines measurement usually
Search Engines Informa1on Retrieval in Prac1ce Annota1ons by Michael L. Nelson All slides Addison Wesley, 2008 Evalua1on Evalua1on is key to building effec$ve and efficient search engines measurement usually
Bias in Resampling-Based Thresholding of Statistical Maps in fmri
 Bias in Resampling-Based Thresholding of Statistical Maps in fmri Ola Friman and Carl-Fredrik Westin Laboratory of Mathematics in Imaging, Department of Radiology Brigham and Women s Hospital, Harvard
Bias in Resampling-Based Thresholding of Statistical Maps in fmri Ola Friman and Carl-Fredrik Westin Laboratory of Mathematics in Imaging, Department of Radiology Brigham and Women s Hospital, Harvard
Sta$s$cs & Experimental Design with R. Barbara Kitchenham Keele University
 Sta$s$cs & Experimental Design with R Barbara Kitchenham Keele University 1 Comparing two or more groups Part 5 2 Aim To cover standard approaches for independent and dependent groups For two groups Student
Sta$s$cs & Experimental Design with R Barbara Kitchenham Keele University 1 Comparing two or more groups Part 5 2 Aim To cover standard approaches for independent and dependent groups For two groups Student
Informa(on Retrieval
 Introduc)on to Informa)on Retrieval CS3245 Informa(on Retrieval Lecture 7: Scoring, Term Weigh9ng and the Vector Space Model 7 Last Time: Index Compression Collec9on and vocabulary sta9s9cs: Heaps and
Introduc)on to Informa)on Retrieval CS3245 Informa(on Retrieval Lecture 7: Scoring, Term Weigh9ng and the Vector Space Model 7 Last Time: Index Compression Collec9on and vocabulary sta9s9cs: Heaps and
Bayesian Spherical Wavelet Shrinkage: Applications to Shape Analysis
 Bayesian Spherical Wavelet Shrinkage: Applications to Shape Analysis Xavier Le Faucheur a, Brani Vidakovic b and Allen Tannenbaum a a School of Electrical and Computer Engineering, b Department of Biomedical
Bayesian Spherical Wavelet Shrinkage: Applications to Shape Analysis Xavier Le Faucheur a, Brani Vidakovic b and Allen Tannenbaum a a School of Electrical and Computer Engineering, b Department of Biomedical
Journal of Articles in Support of The Null Hypothesis
 Data Preprocessing Martin M. Monti, PhD UCLA Psychology NITP 2016 Typical (task-based) fmri analysis sequence Image Pre-processing Single Subject Analysis Group Analysis Journal of Articles in Support
Data Preprocessing Martin M. Monti, PhD UCLA Psychology NITP 2016 Typical (task-based) fmri analysis sequence Image Pre-processing Single Subject Analysis Group Analysis Journal of Articles in Support
Fields. J-B. Poline A.P. Holmes K.J. Worsley K.J. Friston. 1 Introduction 2. 2 Testing for the intensity of an activation in SPMs 3
 Statistical Inference and the Theory of Random Fields J-B. Poline A.P. Holmes K.J. Worsley K.J. Friston Contents 1 Introduction 2 2 Testing for the intensity of an activation in SPMs 3 2.1 Theory : : :
Statistical Inference and the Theory of Random Fields J-B. Poline A.P. Holmes K.J. Worsley K.J. Friston Contents 1 Introduction 2 2 Testing for the intensity of an activation in SPMs 3 2.1 Theory : : :
Multiple Comparisons of Treatments vs. a Control (Simulation)
 Chapter 585 Multiple Comparisons of Treatments vs. a Control (Simulation) Introduction This procedure uses simulation to analyze the power and significance level of two multiple-comparison procedures that
Chapter 585 Multiple Comparisons of Treatments vs. a Control (Simulation) Introduction This procedure uses simulation to analyze the power and significance level of two multiple-comparison procedures that
Clustering. Barna Saha
 Clustering Barna Saha The Problem of Clustering Given a set of points, with a no;on of distance between points, group the points into some number of clusters, so that members of a cluster are close to
Clustering Barna Saha The Problem of Clustering Given a set of points, with a no;on of distance between points, group the points into some number of clusters, so that members of a cluster are close to
Machine Learning Crash Course: Part I
 Machine Learning Crash Course: Part I Ariel Kleiner August 21, 2012 Machine learning exists at the intersec
Machine Learning Crash Course: Part I Ariel Kleiner August 21, 2012 Machine learning exists at the intersec
STA 4273H: Sta-s-cal Machine Learning
 STA 4273H: Sta-s-cal Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! h0p://www.cs.toronto.edu/~rsalakhu/ Lecture 3 Parametric Distribu>ons We want model the probability
STA 4273H: Sta-s-cal Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! h0p://www.cs.toronto.edu/~rsalakhu/ Lecture 3 Parametric Distribu>ons We want model the probability
Predictive Analysis: Evaluation and Experimentation. Heejun Kim
 Predictive Analysis: Evaluation and Experimentation Heejun Kim June 19, 2018 Evaluation and Experimentation Evaluation Metrics Cross-Validation Significance Tests Evaluation Predictive analysis: training
Predictive Analysis: Evaluation and Experimentation Heejun Kim June 19, 2018 Evaluation and Experimentation Evaluation Metrics Cross-Validation Significance Tests Evaluation Predictive analysis: training
Pattern Recognition for Neuroimaging Data
 Pattern Recognition for Neuroimaging Data Edinburgh, SPM course April 2013 C. Phillips, Cyclotron Research Centre, ULg, Belgium http://www.cyclotron.ulg.ac.be Overview Introduction Univariate & multivariate
Pattern Recognition for Neuroimaging Data Edinburgh, SPM course April 2013 C. Phillips, Cyclotron Research Centre, ULg, Belgium http://www.cyclotron.ulg.ac.be Overview Introduction Univariate & multivariate
Discovering Visual Hierarchy through Unsupervised Learning Haider Razvi
 Discovering Visual Hierarchy through Unsupervised Learning Haider Razvi hrazvi@stanford.edu 1 Introduction: We present a method for discovering visual hierarchy in a set of images. Automatically grouping
Discovering Visual Hierarchy through Unsupervised Learning Haider Razvi hrazvi@stanford.edu 1 Introduction: We present a method for discovering visual hierarchy in a set of images. Automatically grouping
Chapter 6: Linear Model Selection and Regularization
 Chapter 6: Linear Model Selection and Regularization As p (the number of predictors) comes close to or exceeds n (the sample size) standard linear regression is faced with problems. The variance of the
Chapter 6: Linear Model Selection and Regularization As p (the number of predictors) comes close to or exceeds n (the sample size) standard linear regression is faced with problems. The variance of the
Evaluation. Evaluate what? For really large amounts of data... A: Use a validation set.
 Evaluate what? Evaluation Charles Sutton Data Mining and Exploration Spring 2012 Do you want to evaluate a classifier or a learning algorithm? Do you want to predict accuracy or predict which one is better?
Evaluate what? Evaluation Charles Sutton Data Mining and Exploration Spring 2012 Do you want to evaluate a classifier or a learning algorithm? Do you want to predict accuracy or predict which one is better?
EMPIRICALLY INVESTIGATING THE STATISTICAL VALIDITY OF SPM, FSL AND AFNI FOR SINGLE SUBJECT FMRI ANALYSIS
 EMPIRICALLY INVESTIGATING THE STATISTICAL VALIDITY OF SPM, FSL AND AFNI FOR SINGLE SUBJECT FMRI ANALYSIS Anders Eklund a,b,c, Thomas Nichols d, Mats Andersson a,c, Hans Knutsson a,c a Department of Biomedical
EMPIRICALLY INVESTIGATING THE STATISTICAL VALIDITY OF SPM, FSL AND AFNI FOR SINGLE SUBJECT FMRI ANALYSIS Anders Eklund a,b,c, Thomas Nichols d, Mats Andersson a,c, Hans Knutsson a,c a Department of Biomedical
Supplementary methods
 Supplementary methods This section provides additional technical details on the sample, the applied imaging and analysis steps and methods. Structural imaging Trained radiographers placed all participants
Supplementary methods This section provides additional technical details on the sample, the applied imaging and analysis steps and methods. Structural imaging Trained radiographers placed all participants
Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques
 Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques Sea Chen Department of Biomedical Engineering Advisors: Dr. Charles A. Bouman and Dr. Mark J. Lowe S. Chen Final Exam October
Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques Sea Chen Department of Biomedical Engineering Advisors: Dr. Charles A. Bouman and Dr. Mark J. Lowe S. Chen Final Exam October
Notes on Simulations in SAS Studio
 Notes on Simulations in SAS Studio If you are not careful about simulations in SAS Studio, you can run into problems. In particular, SAS Studio has a limited amount of memory that you can use to write
Notes on Simulations in SAS Studio If you are not careful about simulations in SAS Studio, you can run into problems. In particular, SAS Studio has a limited amount of memory that you can use to write
Coarse-to-fine image registration
 Today we will look at a few important topics in scale space in computer vision, in particular, coarseto-fine approaches, and the SIFT feature descriptor. I will present only the main ideas here to give
Today we will look at a few important topics in scale space in computer vision, in particular, coarseto-fine approaches, and the SIFT feature descriptor. I will present only the main ideas here to give
Pair-Wise Multiple Comparisons (Simulation)
 Chapter 580 Pair-Wise Multiple Comparisons (Simulation) Introduction This procedure uses simulation analyze the power and significance level of three pair-wise multiple-comparison procedures: Tukey-Kramer,
Chapter 580 Pair-Wise Multiple Comparisons (Simulation) Introduction This procedure uses simulation analyze the power and significance level of three pair-wise multiple-comparison procedures: Tukey-Kramer,
Functional MRI in Clinical Research and Practice Preprocessing
 Functional MRI in Clinical Research and Practice Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization
Functional MRI in Clinical Research and Practice Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization
Data Visualisation in SPM: An introduction
 Data Visualisation in SPM: An introduction Alexa Morcom Edinburgh SPM course, April 2010 Centre for Cognitive & Neural Systems/ Department of Psychology University of Edinburgh Visualising results remembered
Data Visualisation in SPM: An introduction Alexa Morcom Edinburgh SPM course, April 2010 Centre for Cognitive & Neural Systems/ Department of Psychology University of Edinburgh Visualising results remembered
Resources for Nonparametric, Power and Meta-Analysis Practical SPM Course 2015, Zurich
 Resources for Nonparametric, Power and Meta-Analysis Practical SPM Course 2015, Zurich http://www.translationalneuromodeling.org/practical-sessions/ Preliminary Get this file: http://warwick.ac.uk/tenichols/zurich.pdf
Resources for Nonparametric, Power and Meta-Analysis Practical SPM Course 2015, Zurich http://www.translationalneuromodeling.org/practical-sessions/ Preliminary Get this file: http://warwick.ac.uk/tenichols/zurich.pdf
Data Visualisation in SPM: An introduction
 Data Visualisation in SPM: An introduction Alexa Morcom Edinburgh SPM course, April 2015 SPMmip [-30, 3, -9] 3 Visualising results remembered vs. fixation contrast(s) < < After the results table - what
Data Visualisation in SPM: An introduction Alexa Morcom Edinburgh SPM course, April 2015 SPMmip [-30, 3, -9] 3 Visualising results remembered vs. fixation contrast(s) < < After the results table - what
Quality Checking an fmri Group Result (art_groupcheck)
 Quality Checking an fmri Group Result (art_groupcheck) Paul Mazaika, Feb. 24, 2009 A statistical parameter map of fmri group analyses relies on the assumptions of the General Linear Model (GLM). The assumptions
Quality Checking an fmri Group Result (art_groupcheck) Paul Mazaika, Feb. 24, 2009 A statistical parameter map of fmri group analyses relies on the assumptions of the General Linear Model (GLM). The assumptions
We can see that some anatomical details are lost after aligning and averaging brains, especially on the cortical level.
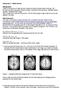 Homework 3 - Model answer Background: Listen to the lecture on Data formats (Voxel and affine transformation matrices, nifti formats) and group analysis (Group Analysis, Anatomical Normalization, Multiple
Homework 3 - Model answer Background: Listen to the lecture on Data formats (Voxel and affine transformation matrices, nifti formats) and group analysis (Group Analysis, Anatomical Normalization, Multiple
About the Course. Reading List. Assignments and Examina5on
 Uppsala University Department of Linguis5cs and Philology About the Course Introduc5on to machine learning Focus on methods used in NLP Decision trees and nearest neighbor methods Linear models for classifica5on
Uppsala University Department of Linguis5cs and Philology About the Course Introduc5on to machine learning Focus on methods used in NLP Decision trees and nearest neighbor methods Linear models for classifica5on
Informa(on Retrieval
 Introduc)on to Informa)on Retrieval CS3245 Informa(on Retrieval Lecture 7: Scoring, Term Weigh9ng and the Vector Space Model 7 Last Time: Index Construc9on Sort- based indexing Blocked Sort- Based Indexing
Introduc)on to Informa)on Retrieval CS3245 Informa(on Retrieval Lecture 7: Scoring, Term Weigh9ng and the Vector Space Model 7 Last Time: Index Construc9on Sort- based indexing Blocked Sort- Based Indexing
Sta$s$cs & Experimental Design with R. Barbara Kitchenham Keele University
 Sta$s$cs & Experimental Design with R Barbara Kitchenham Keele University 1 Analysis of Variance Mul$ple groups with Normally distributed data 2 Experimental Design LIST Factors you may be able to control
Sta$s$cs & Experimental Design with R Barbara Kitchenham Keele University 1 Analysis of Variance Mul$ple groups with Normally distributed data 2 Experimental Design LIST Factors you may be able to control
Lecture 13: Tracking mo3on features op3cal flow
 Lecture 13: Tracking mo3on features op3cal flow Professor Fei- Fei Li Stanford Vision Lab Lecture 13-1! What we will learn today? Introduc3on Op3cal flow Feature tracking Applica3ons (Problem Set 3 (Q1))
Lecture 13: Tracking mo3on features op3cal flow Professor Fei- Fei Li Stanford Vision Lab Lecture 13-1! What we will learn today? Introduc3on Op3cal flow Feature tracking Applica3ons (Problem Set 3 (Q1))
FMRI Pre-Processing and Model- Based Statistics
 FMRI Pre-Processing and Model- Based Statistics Brief intro to FMRI experiments and analysis FMRI pre-stats image processing Simple Single-Subject Statistics Multi-Level FMRI Analysis Advanced FMRI Analysis
FMRI Pre-Processing and Model- Based Statistics Brief intro to FMRI experiments and analysis FMRI pre-stats image processing Simple Single-Subject Statistics Multi-Level FMRI Analysis Advanced FMRI Analysis
Zurich SPM Course Voxel-Based Morphometry. Ged Ridgway (Oxford & UCL) With thanks to John Ashburner and the FIL Methods Group
 Zurich SPM Course 2015 Voxel-Based Morphometry Ged Ridgway (Oxford & UCL) With thanks to John Ashburner and the FIL Methods Group Examples applications of VBM Many scientifically or clinically interesting
Zurich SPM Course 2015 Voxel-Based Morphometry Ged Ridgway (Oxford & UCL) With thanks to John Ashburner and the FIL Methods Group Examples applications of VBM Many scientifically or clinically interesting
Introduc)on to Computer Networks
 Introduc)on to Computer Networks COSC 4377 Lecture 9 Spring 2012 February 15, 2012 Announcements HW4 due today Start working on HW5 In- class student presenta)ons TA office hours this week TR 1030a 100p
Introduc)on to Computer Networks COSC 4377 Lecture 9 Spring 2012 February 15, 2012 Announcements HW4 due today Start working on HW5 In- class student presenta)ons TA office hours this week TR 1030a 100p
Surface-based Analysis: Inter-subject Registration and Smoothing
 Surface-based Analysis: Inter-subject Registration and Smoothing Outline Exploratory Spatial Analysis Coordinate Systems 3D (Volumetric) 2D (Surface-based) Inter-subject registration Volume-based Surface-based
Surface-based Analysis: Inter-subject Registration and Smoothing Outline Exploratory Spatial Analysis Coordinate Systems 3D (Volumetric) 2D (Surface-based) Inter-subject registration Volume-based Surface-based
Introduction to fmri. Pre-processing
 Introduction to fmri Pre-processing Tibor Auer Department of Psychology Research Fellow in MRI Data Types Anatomical data: T 1 -weighted, 3D, 1/subject or session - (ME)MPRAGE/FLASH sequence, undistorted
Introduction to fmri Pre-processing Tibor Auer Department of Psychology Research Fellow in MRI Data Types Anatomical data: T 1 -weighted, 3D, 1/subject or session - (ME)MPRAGE/FLASH sequence, undistorted
TOPOLOGICAL INFERENCE FOR EEG AND MEG 1. BY JAMES M. KILNER AND KARL J. FRISTON University College London
 The Annals of Applied Statistics 2010, Vol. 4, No. 3, 1272 1290 DOI: 10.1214/10-AOAS337 Institute of Mathematical Statistics, 2010 TOPOLOGICAL INFERENCE FOR EEG AND MEG 1 BY JAMES M. KILNER AND KARL J.
The Annals of Applied Statistics 2010, Vol. 4, No. 3, 1272 1290 DOI: 10.1214/10-AOAS337 Institute of Mathematical Statistics, 2010 TOPOLOGICAL INFERENCE FOR EEG AND MEG 1 BY JAMES M. KILNER AND KARL J.
EPI Data Are Acquired Serially. EPI Data Are Acquired Serially 10/23/2011. Functional Connectivity Preprocessing. fmri Preprocessing
 Functional Connectivity Preprocessing Geometric distortion Head motion Geometric distortion Head motion EPI Data Are Acquired Serially EPI Data Are Acquired Serially descending 1 EPI Data Are Acquired
Functional Connectivity Preprocessing Geometric distortion Head motion Geometric distortion Head motion EPI Data Are Acquired Serially EPI Data Are Acquired Serially descending 1 EPI Data Are Acquired
Lecture 14: Tracking mo3on features op3cal flow
 Lecture 14: Tracking mo3on features op3cal flow Dr. Juan Carlos Niebles Stanford AI Lab Professor Fei- Fei Li Stanford Vision Lab Lecture 14-1 What we will learn today? Introduc3on Op3cal flow Feature
Lecture 14: Tracking mo3on features op3cal flow Dr. Juan Carlos Niebles Stanford AI Lab Professor Fei- Fei Li Stanford Vision Lab Lecture 14-1 What we will learn today? Introduc3on Op3cal flow Feature
SPM8 for Basic and Clinical Investigators. Preprocessing. fmri Preprocessing
 SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
Bootstrapping Methods
 Bootstrapping Methods example of a Monte Carlo method these are one Monte Carlo statistical method some Bayesian statistical methods are Monte Carlo we can also simulate models using Monte Carlo methods
Bootstrapping Methods example of a Monte Carlo method these are one Monte Carlo statistical method some Bayesian statistical methods are Monte Carlo we can also simulate models using Monte Carlo methods
Flash Reliability in Produc4on: The Importance of Measurement and Analysis in Improving System Reliability
 Flash Reliability in Produc4on: The Importance of Measurement and Analysis in Improving System Reliability Bianca Schroeder University of Toronto (Currently on sabbatical at Microsoft Research Redmond)
Flash Reliability in Produc4on: The Importance of Measurement and Analysis in Improving System Reliability Bianca Schroeder University of Toronto (Currently on sabbatical at Microsoft Research Redmond)
Quiz Section Week 3 April 12, Functions Reading from and writing to files Complexity
 Quiz Section Week 3 April 12, 2016 Functions Reading from and writing to files Complexity Hypothesis testing: how interesting is my data? Test statistic: a number that describes how interesting your data
Quiz Section Week 3 April 12, 2016 Functions Reading from and writing to files Complexity Hypothesis testing: how interesting is my data? Test statistic: a number that describes how interesting your data
Machine Learning and Data Mining. Clustering (1): Basics. Kalev Kask
 Machine Learning and Data Mining Clustering (1): Basics Kalev Kask Unsupervised learning Supervised learning Predict target value ( y ) given features ( x ) Unsupervised learning Understand patterns of
Machine Learning and Data Mining Clustering (1): Basics Kalev Kask Unsupervised learning Supervised learning Predict target value ( y ) given features ( x ) Unsupervised learning Understand patterns of
Lab 9. Julia Janicki. Introduction
 Lab 9 Julia Janicki Introduction My goal for this project is to map a general land cover in the area of Alexandria in Egypt using supervised classification, specifically the Maximum Likelihood and Support
Lab 9 Julia Janicki Introduction My goal for this project is to map a general land cover in the area of Alexandria in Egypt using supervised classification, specifically the Maximum Likelihood and Support
Introduction to hypothesis testing
 Introduction to hypothesis testing Mark Johnson Macquarie University Sydney, Australia February 27, 2017 1 / 38 Outline Introduction Hypothesis tests and confidence intervals Classical hypothesis tests
Introduction to hypothesis testing Mark Johnson Macquarie University Sydney, Australia February 27, 2017 1 / 38 Outline Introduction Hypothesis tests and confidence intervals Classical hypothesis tests
Apply. A. Michelle Lawing Ecosystem Science and Management Texas A&M University College Sta,on, TX
 Apply A. Michelle Lawing Ecosystem Science and Management Texas A&M University College Sta,on, TX 77843 alawing@tamu.edu Schedule for today My presenta,on Review New stuff Mixed, Fixed, and Random Models
Apply A. Michelle Lawing Ecosystem Science and Management Texas A&M University College Sta,on, TX 77843 alawing@tamu.edu Schedule for today My presenta,on Review New stuff Mixed, Fixed, and Random Models
