Comparison of Several Classes of Algorithms for Cytoplasm to Nucleus Translocation
|
|
|
- Eugenia Hunter
- 5 years ago
- Views:
Transcription
1 Abstract Comparison of Several Classes of Algorithms for Cytoplasm to Nucleus Translocation Ilya Ravkin and Vladimir Temov, Vitra Bioscience, Inc. Cellular assays assessed by image analysis provide a new type of information but also present several new sources of variability compared to traditional screening. Understanding of these sources and of the behavior of image analysis algorithms under different conditions is important for obtaining reliable data from imaging assays. We have developed several new and implemented several commonly used measures of cytoplasm to nucleus translocation. The quality of these measures is analyzed with respect to user parameters, number of analyzed cells, uniformity of illumination, accuracy of focusing, and depth of field. Slide: 1
2 Classes of Algorithms in this Study. Algorithms for Image Analysis of Cytoplasm to Nucleus Translocation Algorithms based on 2D distributions of signal stain and counter stain, e.g. Slope algorithms 9 Algorithms based on measuring signal stain in cellular compartments: nucleus, cytoplasm, or part of cytoplasm (ring) Algorithms based on measuring by segmenting the image into cellular compartments 8 Algorithms based on measuring without image segmentation into cellular compartments Slide: 2
3 Preprocessing Common to all Algorithms. Before running specific algorithms for cytoplasm to nucleus translocation we perform several preprocessing steps: 1. Rejection of artifacts. This removes areas of fluorescent debris determined primarily by saturation of intensity. 2. Shading correction is available, but was turned off in this study because we used intensity normalization (see below). 3. Background removal. This operation removes background by morphological opening. Saturated areas are not changed. 4. Intensity normalization. This is a complex operation based on partitioning the image into components; it consists of the following parts: - Detection of markers of components - Detection of watersheds, which serve as separating lines between components - Adjustment of intensity in individual components Steps 1-3 are rather common in processing of microscope images. Step 4 will be described in some detail below. Slide: 3
4 Partitioning into Components. Markers. Partitioning into components serves a dual purpose. The first is that partitioning is needed for individual cell analysis; the second as a step in intensity equalization. There are two main parts in the partitioning method: finding of markers and finding of separation lines. Markers are found by the following algorithm. A fixed value (marker contrast) is subtracted with saturation from the image of nuclear counterstain and the resulting image is reconstructed 7 within the image of nuclear counterstain. This image is then subtracted from the counterstain image and the result is converted to binary image. The components of this image are the markers. A further restriction may be imposed on markers only markers that have at least one pixel in the original intensity image above a given threshold (marker brightness) are retained for the second step. Depending on magnification and noise level the image of nuclear counterstain may be smoothed prior to this algorithm. This method of determining markers can handle cells of different size and shape. Other methods, e.g. based on top-hat transform 7 may be used too. Left - original image of cells expressing GFP-fused p65 protein. Middle - smoothed nuclear counterstain image shown in grayscale with overlaid contours of markers (cyan) and contours of background areas (white). Right - partitioning of the image induced by the markers. Slide: 4
5 Partitioning into Components. Watersheds of Combined Intensity Images. Separation lines are defined as the watershed 4,5,6 of the inverted image of the linear combination of the counterstain image and the signal stain image. The reason to use linear combination rather than just the nuclear counterstain image is that cells are often nonsymmetrical and unevenly spaced. Separation lines from nuclear stain image may cut through the middle of cells. The use of signal stain produces more accurate separation lines. Coefficients of the linear combination may be varied depending on the peculiarities of staining and of image acquisition (for such nuclear stains as DRAQ5, which give a low level staining of the whole cell, the use of signal stain is not needed). C1 Background C10 C5 C11 C2 C7 Background C6 C9 Background C3 C8 C4 C12 C13 C16 C17 C15 C14 C18 C19 Background C20 Left - image partitioning using only nuclear stain image (blue). Right image partitioning using linear combination of nuclear and signal stains. Coefficient of linear combination for the nuclear image = 0.3, for the signal stain = 0.7 The result of partitioning is the labeling of the background and of the components. Components contain preferably individual cells, but may contain clusters of cells; they do not necessarily correspond to cell shape. Slide: 5
6 Equalization of Intensities. Equalization of intensities is not strictly required by the algorithms described below. Identical results can be achieved by adaptively choosing parameters (specifically, thresholds) for individual cells or by normalizing the 2D intensity distributions. However, we find that equalizing the intensities of cells gives additional feedback to the user and may reveal features that were not seen before normalization. In addition, it simplifies the subsequent processing. Equalization can be done in the components described above. In this case all pixels from a component are multiplied by the same number, separately for the signal stain and for the counterstain. Alternatively, normalization can be done without partitioning the image by fitting a smooth surface to the images of signal stain and counterstain. Top original cells, bottom - cells separated by watershed lines with intensities of the counterstain (blue) and signal stain (green) normalized to the maximum in each compartment. Slide: 6
7 Slide: 7 Algorithms Based on 2D Distributions of Signal Stain and Counter Stain.
8 Images and Profiles Through Model and Real Cells. To find a robust measure of nuclear translocation we have defined a model of spatial distribution of the nuclear counterstain and of the signal stain as it moves from the cytoplasm to the nucleus. The model was studied under some perturbations in order to find measures that are robust. The model of cell staining comprises a bell-shaped intensity distribution of counterstain, which is shown in blue, and a bell-shaped distribution of signal stain, which is shown in green. For the negative case the distribution of signal stain is wider and has a bellshaped crater. Profiles through the real cells show substantial similarity to the model profiles. (Profile C is plotted through two cells, profile D through three cells. All profiles are independently normalized to their intensity maxima.) A,B profiles through model distributions; C,D profiles through real cells. A,C negative, B,D positive. Blue counterstain, green signal stain. Slide: 8
9 Model and Experimental 2D Distributions of Stains. Translocation Measures Derived from 2D Distributions. To derive stable measures that characterize transitions from the negative to the positive case, we analyzed joint distributions of the stains on the model and on real cells. In the ideal case the model stain distributions are circularly symmetrical and aligned. The cross-histogram for this case is shown in panels A and B. If the model is perturbed by offsetting the centers of the two stains, by changing shape from circular to oval, or by adding noise, the distributions become fuzzy as shown in panels C and D. Typical negative and positive real cells have cross-histogram as shown in panels E and F. These distributions suggest that a translocation measure can be defined as the slope of a straight-line segment approximating the right side of the cross-histogram. This portion of the distribution corresponds to the more intense nuclear staining and is close to the center of the nucleus. The farther from the center, the more diffuse the distribution, and the less reliable the approximation becomes. Cross-histograms of counterstain (X-axis) and signal stain (Y-axis) in ideal model (A,B), perturbed model (C,D) and a real cell (E,F). Scale on both axes is Red lines show the calculated approximation. The portion of the distribution that is used for approximation with the straight line segment is found by plotting the approximated slope going from right to left and selecting the range where this approximation is the most stable. This line segment is shown in red (next slide) and its tangent multiplied by 100 is called Slope1. Top panel nuclear localization (positive), middle panel intermediate localization, bottom panel cytoplasm localization (negative) of protein. A variation on this method is to calculate two more slopes (shown by green lines). The top line is the regression line calculated on all points above the original slope segment; the bottom line is the regression line calculated on all points below the original slope segment. If all three slopes have the same sign, the result is the one with the greatest absolute value. If they have different signs, then the average of the three lines is used. This measure is called Slope3. Slide: 9
10 Translocation Measures Derived from 2D Distributions. positive intermediate negative Examples of Slope measures in cells with different protein localization. Top panel nuclear localization (positive), middle panel intermediate localization, bottom panel cytoplasm localization (negative) of protein. Slope1 - red line. Slope3 one of green lines. Slide: 10
11 Slide: 11 Algorithms Based on Measuring Signal Stain in Cellular Compartments.
12 Algorithms Based on Segmentation into Nucleus and Cytoplasm. This class of algorithms 8 is the most widely used for analysis of cytoplasm to nucleus translocation. The approach is to use the nuclear stain image to identify the nuclear mask; then to find the mask of what is some approximation of the cytoplasm, usually a ring around the nucleus as shown in the figure. After the masks representing nucleus and cytoplasm are found, the average and/or total intensity of the signal stain are calculated in each mask. The measure of translocation is the ratio or difference between these intensities. A Erosion of the nuclear mask B Gap between the eroded nuclear mask and the ring mask C Width of the ring mask Nuclear mask Eroded nuclear mask used for measurement Ring mask Slide: 12
13 Algorithms Based on Indicator Functions. Measurement Without Segmentation. Suppose the task is to measure the amount of fluorescence emitted by proteins in the nucleus of a cell. The figure on the right gives a representation of the images by their intensity profiles along a straight line through the cell Profiles through the Original image, Indicator Function image and Mask The conventional approach is to first use the blue profile to generate a mask of the nucleus (the profile through the mask is shown in black); then - to integrate intensities under the green profile within this mask. Our approach is to use the blue profile to generate an indicator function shown in red. This name was given because the function indicates the certainty we have in that a pixel belongs to the object. The indicator function may be produced by simply scaling the blue profile to the range [0-1], or [0-100%], or [0-255] (for convenience of representation as image). Integrated intensity of green in the object is produced by integrating green in the whole image, using the indicator function as weight Original Blue Profile Original Green Profile Indicator Function Mask The application of this approach does not require having two measured images; green and blue in this example. It can be applied to a single image as well. In this case the image that generates the indicator function and the image which is being integrated is the same image. This approach has several advantages: it avoids possible errors of segmentation, it is faster to perform and has better scalability with magnification. The indicator function may be produced from one or more acquired images by a complex algorithm. Slide: 13
14 Use of Indicator Functions for Analysis of Cytoplasm to Nucleus Translocation. 1. The indicator function method can be applied to the analysis of images of the cytoplasm to nucleus translocation assay. In this assay, cells are visualized in two wavelengths corresponding to nuclear stain and to protein stain. The goal of analysis is to quantify translocation of proteins between the cytoplasm and the nucleus. Images representing extreme states of this assay are shown in the next slide in Fig. A,B,C. The indicator function for the nucleus, in the simplest case, is the scaled nuclear stain image as described above and shown in Fig. D. It can also be a smoothed nuclear image or any other derivative of the nuclear image that may be beneficial. The amount of stained protein in the nucleus is the integral of image A with the weight given by image D over the area of the component containing the cell. For easier visualization, image A weighted by image D is shown as image G. Thus, the amount of stained protein in the nucleus is the sum of all pixels of G. Another useful measure is the average intensity of protein staining in the nucleus. This measure is calculated as the sum of all pixels of G divided by the sum of all pixels of D. In addition to the nucleus, the two other compartments that are useful to assess for this assay are the cytoplasm and the perinuclear region of cytoplasm, which we will call the ring. The indicator function for the cytoplasm is the complement of the indicator function for the nucleus within the compartment containing the cell (Fig. E). The indicator function for the ring is produced by dilating the indicator function for the nucleus by a given amount (width of the ring), subtracting the indicator function for the nucleus from it and normalizing to the given scale as described above. Other methods of producing the indicator function for the ring may be used too. The amount of stained protein in the cytoplasm is the sum of all pixels of H. The average intensity of protein staining in the cytoplasm is the sum of all pixels of H divided by the sum of all pixels of E. The amount of stained protein in the ring is the sum of all pixels of J. The average intensity of protein staining in the ring is the sum of all pixels of J divided by the sum of all pixels of F. Slide: 14
15 Use of Indicator Functions for Analysis of Cytoplasm to Nucleus Translocation. 2. Left panel corresponds to negative state of the assay when stained proteins are localized predominantly in cytoplasm. Right panel corresponds to positive state of the assay when stained proteins are localized predominantly in nucleus. A image of stained proteins (green), B image of nuclear stain (blue), C pseudo-color composite of A and B, D same as B normalized to [0-255] and used as indicator function of nucleus, E complement of D within the component containing the cell (shown by red contour) used as indicator function of cytoplasm, F dilation of D minus D normalized to [0-255] used as indicator function of the ring around the nucleus. G A weighted (multiplied) by D the image of protein content in nucleus, H A weighted by E the image of protein content in cytoplasm, J A weighted by F the image of protein content in a ring around the nucleus. Slide: 15
16 Family of Measures for Algorithms Based on Measuring Signal Stain in Cellular Compartments. A family of assay measures can be constructed as ratios of the average (A) or total (T) amounts of stained protein in the three compartments: nucleus (N), cytoplasm (C), and ring (R). For more convenient scaling we use 100*log 2 () of all ratio measures. In addition, a non-ratio measure of the amount of signal stain in the nucleus may be also useful. The measures may be produces by segmentation into masks of nucleus and cytoplasm (M) or without segmentation by the use of indicator functions (W). Each measure may be calculated for each cell and then the well measure is some statistic of the cell population, e.g. mean or median; alternatively these measures may be calculated globally on the whole image (G). Global measures are not included in this study. The family of measures is given by the formulas: N2{R C}_{A T}{W M}{ G} and N_{A T}{W M}{ G}. N2R_AW N2R_TW N2R_AM N2R_TM Nucleus-to-Ring measures depend on parameters A, B, and C. N2C_AW N2C_TW N_AW N_TW N2C_AM N2C_TM N_AM N_TM Nucleus-to-Cytoplasm measures or, more accurately, nucleus-tocompartment measures depend on parameters A and B Nucleus-only measures depend on parameter A A - Average intensity, T - Total intensity Measures based on indicator functions Measures based on masks require segmentation and depend on the threshold. Slide: 16
17 Parameters of the Indicator Function Algorithm for Analysis of Cytoplasm to Nucleus Translocation. C B A 5 3 Parameters of the indicator-function algorithm for analysis of cytoplasm to nucleus translocation: A Erosion of the nuclear indicator function B Gap between the eroded nuclear indicator function and the ring indicator function C Width of the ring indicator function 1 (solid blue profile) original nuclear indicator function 2 (dotted blue profile) eroded nuclear indicator function 3 (dotted black profile) - nuclear indicator function dilated by B-A 4 (solid black profile) - nuclear indicator function dilated by B-A+C; this is the indicator function of nucleus and ring together 5 (solid green profile) indicator function of the ring produced by subtracting profile 3 from profile 4 and normalizing to full scale. Slide: 17
18 Methodology of Algorithm Evaluation. Establish quality measure 1,2 Find a representative image set Analyze all available algorithms with as wide a range of parameters as feasible. Make conclusions about parameter dependency Choose the best algorithms/measures and a narrow range of parameters Analyze dependency of the assay measures and their quality on relevant factors. Some of the factors may be fixed in the instrumentation or in the algorithm, but some are open to the user and must be considered in the experiment design. Dependency on the number of analyzed cells. The number of cells (or fields) to analyze is the most typical decision that the user must make in a HCS experiment Dependency on the nonuniformity of illumination. The user may not have direct influence over this, but it is important because this may be a concern when comparing systems and it may be a burden to perform regular calibration. Dependency on the choice of objective. This usually applies only to wide-field systems and is often up to the user. It is important to understand the influence of the properties of microscope objectives on the assay result. Here we analyze only the influence of numerical aperture; analysis of magnification dependency has been done previously 9. Dependency on the accuracy of focusing. The inherent accuracy of focusing may not be under the user control, but sometimes the decision on how often to focus is left to the user. It is important to understand how focusing affects the measurements. By extension it also sheds light on the importance of flat field. Slide: 18
19 V-factor a Generalization of Z-factor. (1) Z = 1 SD 3( M pos pos + SD M neg neg ) -inf Hypothetical dose curve 30 (2) V = 1 6( M SD pos of _ fit M neg ) Effect inf (3) SD of = _ fit 1 n n ( fmodel fexperiment 1 2 ) Dose For a complete discussion of v-factor see ref. 2,3. Slide: 19
20 Description of Plates Used in the Algorithm Comparison. Vitra plate A B C D E F G H MCF7 A549 TNF-α concentration in log g/ml Cells were plated at about 10,000 cells per well in a 96 well microtiter plate (Packard ViewPlate) and incubated overnight. The cells were then treated with varying doses of TNFa for 30 minutes. This treatment results in the activation of NFKB and the translocation of the p65 subunit from the cytoplasm to the nucleus. The cells were subsequently fixed and immuno-stained for p65 and counterstained with the Hoechst nuclear dye. The plate was acquired at Vitra Bioscience on the CellCard reader - a microscope-based system. Image size 1360*1024 pixels, 8-bit per pixel. BioImage plate A B C D E F G H Wortmannin: 250 nm, 2 fold dilutions, 9 points LY294002: 80 µm, 2 fold dilutions, 9 points This assay quantifies the Forkhead fusion protein (FKHR-EGFP) accumulated in the nuclei of stably transfected human osteosarcoma cells, U2OS. In proliferating cells FKHR is localized in the cytoplasm. Even without stimulation, Forkhead is constantly moving into the nucleus, but is transported out by export proteins. Upon inhibition of nuclear export, FKHR accumulates in the nucleus. In this assay, export is inhibited by blocking PI3 kinase / PKB signaling by incubating cells for 1 hr with a drug as indicated in the plate map. Nuclear staining: DRAQ. The images were acquired at BioImage on the IN Cell Analyzer Image size 640*640 pixels. Slide: 20
21 Example of Dose Curve Analysis. X: Dose; CellType=A549; Erosion=1; Gap=2; 160 Y: N2C_AM; ED50=-9.91 [-10.0 : -9.8]; V=0.71; Y=[16.9 : 152.1]; 180 Y: N2C_AW; ED50=-9.78 [-9.8 : -9.7]; V=0.71; Y=[32.4 : 165.2]; -220 Y: N2C_TM; ED50=-9.76 [-9.9 : -9.7]; V=0.67; Y=[ : ]; 0 Y: N2C_TW; ED50=-9.69 [-9.8 : -9.6]; V=0.68; Y=[ : -55.6]; Y: N2C_AMG; ED50=-9.92 [-10.0 : -9.8]; V=0.70; Y=[18.9 : 153.4]; 160 Y: N2C_AWG; ED50=-9.78 [-9.8 : -9.7]; V=0.72; Y=[30.9 : 150.7]; -220 Y: N2C_TMG; ED50=-9.75 [-9.9 : -9.7]; V=0.69; Y=[ : ]; -60 Y: N2C_TWG; ED50=-9.65 [-9.8 : -9.6]; V=0.68; Y=[ : -76.6]; Slide: 21 Vitra Plate. Concentration is in log scale.
22 Evaluation of Algorithms. Parameter Dependency. Vitra Plate V-factor and ED50 for different measures and different user parameters. MCF V-factor and ED50 for different measures and different user parameters. A N2R_AM N2R_AW N2R_TM N2R_TW N2C_AM N2R_AM N2R_AW N2R_TM N2R_TW N2C_AM 0.60 N2C_AW N2C_TM 0.60 N2C_AW N2C_TM V-factor N2C_TW N_AM N_AW N_TM V-factor N2C_TW N_AM N_AW N_TM 0.45 N_TW 0.45 N_TW 0.40 Slope1 Slope Slope1 Slope SlopeG 0.35 SlopeG ED ED Concentration is in log scale Symbol code for measures: N2R (nucleus to ring) N2C (nucleus to cytoplasm) N (nucleus only) Parameters: Nuclear erosion = [0 2] A (average intensity) Red, T (total intensity) Blue Gap = [0 4] M (mask-based) wire shape, W (weight-based) solid shape Ring = [1 8] Slide: 22
23 Evaluation of Algorithms. Parameter Dependency. BioImage Plate V-factor and ED50 for different measures and different user parameters. Wortmannin V-factor and ED50 for different measures and different user parameters. LY V-factor N2R_AM N2R_AW N2R_TM N2R_TW N2C_AM N2C_AW N2C_TM N2C_TW N_AM N_AW N_TM N_TW Slope1 Slope3 SlopeG V-factor N2R_AM N2R_AW N2R_TM N2R_TW N2C_AM N2C_AW N2C_TM N2C_TW N_AM N_AW N_TM N_TW Slope1 Slope3 SlopeG ED ED50 Symbol code for measures: N2R (nucleus to ring) N2C (nucleus to cytoplasm) N (nucleus only) A (average intensity) Red, T (total intensity) Blue M (mask-based) wire shape, W (weight-based) solid shape Parameters: Nuclear erosion = [0 2] Gap = [0 4] Ring = [1 8] Log(Nuc/Cyt), where Nuc/Cyt is the InCell3000 measure Slide: 23
24 Evaluation of Algorithms. First conclusions. Measures based on assessing signal stain in the nucleus alone (N_ measures) show higher variability and greater inconsistency than other measures, both in v-factor and in ED 50. Some measures show greater parameter dependency than others. Choice of best measures is not consistent between drugs and cell types. Each measure has maximum quality at different parameter values. There is a systematic shift between ED 50 of different measures. If the goal is comparison among drugs or cell types, then a single measure should be used. For further analysis we selected all measures except N_ and narrowed the parameter range: Nuclear erosion = [0 1] Gap = [0 3] Ring = [1 8] Ero=2 and Gap=4 were excluded because these parameters almost always result in the decline of v- factor. For realistic assessment of quality of different measures we cannot assume that the parameters would always be optimal, therefore, we averaged V-factors (and ED 50 ) for each measure over the parameter range determined above (except for slope measures, which do not have parameters). Slide: 24
25 Evaluation of Algorithms. Quality Averaged over Parameter Range. Vitra Plate Average V-factor vs. Average ED50 over a range of parameters. MCF7. N2R_AM Average V-factor vs. Average ED50 over a range of parameters. A549. N2R_AM 0.78 N2R_AW N2R_TM 0.76 N2R_AW N2R_TM N2R_TW N2R_TW 0.74 N2C_AM 0.72 N2C_AM N2C_AW N2C_AW Average V-factor N2C_TM N2C_TW Slope1 Slope3 Average V-factor N2C_TM N2C_TW Slope1 Slope Slope_G Slope_G Average ED Average ED50 Concentration is in log scale Symbol code for measures: N2R (nucleus to ring) N2C (nucleus to cytoplasm) N (nucleus only) Averaged over Parameter range: Nuclear erosion = [0 1] A (average intensity) Red, T (total intensity) Blue Gap = [0 3] M (mask-based) wire shape, W (weight-based) solid shape Ring = [1 8] Slide: 25
26 Evaluation of Algorithms. Quality Averaged over Parameter Range. BioImage Plate Average V-factor vs. Average ED50 over a range of parameters. Wortmannin Average V-factor vs. Average ED50 over a range of parameters. LY Average V-factor N2R_AM N2R_AW N2R_TM N2R_TW N2C_AM N2C_AW N2C_TM N2C_TW Slope1 Slope3 Slope_G Average V-factor N2R_AM N2R_AW N2R_TM N2R_TW N2C_AM N2C_AW N2C_TM N2C_TW Slope1 Slope3 Slope_G Average ED Average ED50 Symbol code for measures: N2R (nucleus to ring) N2C (nucleus to cytoplasm) N (nucleus only) A (average intensity) Red, T (total intensity) Blue M (mask-based) wire shape, W (weight-based) solid shape Averaged over Parameter range: Nuclear erosion = [0 1] Gap = [0 3] Ring = [1 8] Slide: 26
27 Slide: 27 Analysis of Dependencies.
28 Dependency of V-factor on Cell Number. Vitra Plate V-factor vs. number of cell. MCF V-factor vs. number of cell. A N2R_AM 0.70 N2R_AM N2R_AW N2R_AW V-factor N2R_TM N2R_TW N2C_AM N2C_AW N2C_TM N2C_TW Slope1 V-factor N2R_TM N2R_TW N2C_AM N2C_AW N2C_TM N2C_TW Slope Slope Slope Average number of cells used in image Average number of cells used in image Symbol code for measures: N2R (nucleus to ring) N2C (nucleus to cytoplasm) A (average intensity) Red, T (total intensity) Blue M (mask-based) wire shape, W (weight-based) solid shape Parameters: Nuclear erosion = 1 Gap = 2 Ring = 4 To study the cell number dependency we selected random subsets of cells from the whole image to eliminate possible positional dependency. Slide: 28
29 Dependency of V-factor on Cell number. BioImage Plate V-factor vs. number of cells. Wortmannin V-factor vs. number of cells. LY N2R_AM 0.70 N2R_AM 0.75 N2R_AW N2R_TM 0.65 N2R_AW N2R_TM 0.70 N2R_TW 0.60 N2R_TW V-factor N2C_AM N2C_AW N2C_TM N2C_TW V-factor N2C_AM N2C_AW N2C_TM N2C_TW 0.55 Slope1 Slope Slope1 Slope Average number of cells used in image Average number of cells used in image Symbol code for measures: N2R (nucleus to ring) N2C (nucleus to cytoplasm) A (average intensity) Red, T (total intensity) Blue M (mask-based) wire shape, W (weight-based) solid shape Parameters: Nuclear erosion = 1 Gap = 2 Ring = 4 Slide: 29
30 Is Uniformity of Illumination a Factor in CNT Algorithms? C Images acquired on InCell3000 provide a convenient model to study the dependency of cellular measures on non-uniformity of illumination. Y-direction, which corresponds to the stage movement has uniform brightness, but X-direction, which corresponds to the scan of the laser beam has variable brightness. Calibration curves provided with the images are shown in Figure A. 150 B A Projection of image intensity on Y-axis, averaged over all images in the plate Actual brightness in the two channels averaged over the entire plate to eliminate random errors is shown in figures B (X-projection) and C (Y-projection). The plots show clearly uniform intensity in Y and nonuniform intensity in X. To assess if the brightness non-uniformity affects the cellular measures produced by the presented family of CNT algorithms, we plotted the measures against X-coordinates of cells and against Y-coordinates of cells. Two of the measures are shown below. The plots show that there is no dependency on either X or Y position of cells. Other measures, which are not shown, exhibit similar behavior This allows us to conclude that variation in brightness alone does not affect cellular measures if the algorithm is specifically designed to compensate for it. 50 Projection of image intensity on X-axis, averaged over all images in the plate. 500 Calibration profiles in X Slide: 30
31 Is Uniformity of Illumination a Factor in CNT Algorithms? 2. Slope1 vs. X-position of cell in image at different doses. Wortmannin. Slope1 vs. Y-position of cell in image at different doses. Wortmannin Slope Slope X_0 X_1 X_2 X_3 X_4 X_5 X_6 X_7 X_8 X_9 X-position of cell in pixels Y_0 Y_1 Y_2 Y_3 Y_4 Y_5 Y_6 Y_7 Y_8 Y_9 Y-position of cell in pixels N2R_TW vs. X-position of cell in image at different doses. Wortmannin. N2R_TW vs. Y-position of cell in image at different doses. Wortmannin N2R_TW N2R_TW X_0 X_1 X_2 X_3 X_4 X_5 X_6 X_7 X_8 X_9 X-position of cell in pixels -120 Y_0 Y_1 Y_2 Y_3 Y_4 Y_5 Y_6 Y_7 Y_8 Y_9 Y-position of cell in pixels Slide: 31
32 Dependency of CNT Measures on the Depth of Field in a Non-confocal System. 1. The Vitra plate was scanned two times: with a PlanFluor 10X 0.3NA objective and with a SFluor 10X 0.5NA objective (Nikon). The images were processed with the same parameters: Nuclear erosion=1, Gap=2, Ring=4. V-factor shows that PlanFluor always gives better results. Note that measures based on total amount of fluorescence (as opposed to average) suffer the most. V-factor on Vitra plate scanned with different objectives. MCF7. PlanFluor 10X SFluor 10X Y: Slope3; ED50=-9.78 [-9.8 : -9.7]; V=0.81; Y=[-47.6 : 67.1]; 80 Y: Slope3; ED50=-9.74 [-9.8 : -9.7]; V=0.68; Y=[-37.8 : 62.8]; V-factor N2C_AM N2C_AW N2C_TM N2C_TW N2R_AM CNT measures N2R_AW N2R_TM N2R_TW V-factor 10X PlanFluor V-factor 10X Sfluor Slope1 Slope TNFa concentration in log scale TNFa concentration in log scale Our hypothesis is that this result is due to the relative thickness of cells and the depth of field of the two objectives. The depth of field of 0.3NA PlanFluor 10X is about 8.5 µm, and the depth of field of 0.5NA SFluor 10X is about 3.6 µm. The thickness of attached cells is probably between these two values. If the cell is thicker than the depth of field of the objective it will appear fuzzy. This will make negative cells look less negative and reduce the dynamic range of the measurements. The following illustration supports this hypothesis. Analysis of the minimal and maximal values of the fitted logistic curves indeed shows reduced dynamic range with the SFluor objective. This does not explain, in full, the decrease in V-factor. The variation among replica wells also was higher with the SFluor objective. This may be attributed to focusing artifacts due to shallow depth of field. Slide: 32
33 Dependency of CNT Measures on the Depth of Field in a Non-confocal System. 2. PlanFluor10X, 0.3NA, Depth of field = 8.5 µm Distance along profile Original Normalized 2D distribution Profiles SFluor10X, 0.5NA, Depth of field = 3.6 µm Slide: 33
34 Dependency of CNT Measures on Focus Position. Images at 10X Slope1 as a function of Z position N2R as a function of Z position PlanApo10X PlanFluor10X SFluor10X -10 PlanApo10X PlanFluor10X SFluor10X Slope1 10 N2R Negative Z (step=2 um) Slope1 as a function of Z position Z (step=2 um) N2R as a function of Z position 90 PlanApo10X PlanFluor10X SFluor10X 30 Slope1 70 N2R PlanApo10X PlanFluor10X SFluor10X Positive Z (step=2 um) Z (step=2 um) CNT measures are almost identical within µm from the best focus position in 10X (less for PlanApo with N2R_AM). Objectives with lower NA are more tolerant to focusing inaccuracy. Slope1 and N2R_AM show similar dependency on Z-position on negative cells and opposite dependency on positive cells. For Slope1 PlanFluor gives the greatest range (marginally); for N2R_AM PlanApo gives the greatest range. The importance of this dependency is not only in setting focusing requirements, but also in establishing the need for flat field. Slide: 34
35 Conclusions and Recommendations. Algorithms and measures: 1. Measures based on assessing signal stain in nucleus alone show much higher variability than other measures, both in v-factor and in ED There is a systematic shift between ED50 of different measures. If the goal is comparing drugs or cell types, then a single measure should be used for the whole experiment. This points to the need for a uniformly best measure across experimental conditions. 3. Different measures have maximum quality at different parameter values. For a realistic comparison, the quality of different measures must be averaged over a reasonable parameter range. Therefore, measures which do not depend on parameters, have a big advantage. 4. Conceptually similar measures implemented by different vendors can show significant difference in quality. Choosing an instrument: 1. Illumination non-uniformity alone does not play an important role if the algorithms can compensate for it. 2. Flatness of field and accuracy of focusing are critical and cannot be easily compensated for algorithmically. Experiment design: 1. Objectives in a wide-field system should be chosen based on their depth of field and cell thickness. 2. Focusing accuracy of the system must be evaluated and focusing strategy established to provide for small error (e.g., <10µm in 10X). 3. Required number of cells can be established by analyzing a fraction of images and selecting a number above which quality does not significantly improve with cell number. Slide: 35
36 References 1. J.-H. Zhang, T.D.Y. Chung, K.R. Oldenburg A simple statistical parameter for use in evaluation and validation of high throughput screening assays, J. Biomol. Screening 4: pp , I. Ravkin "Quality Measures for Imaging-based Cellular Assays" SBS 2004 conference poster #P I. Ravkin, V. Temov, A.D. Nelson, M.A. Zarowitz, M. Hoopes, Y. Verhovsky, G. Ascue, S. Goldbard, O. Beske, B. Bhagwat, H. Marciniak "Multiplexed high-throughput image cytometry using encoded carriers", Proc. SPIE Vol. 5322, pp , 2004 (Imaging, Manipulation, and Analysis of Biomolecules, Cells, and Tissues II; Dan V. Nicolau, Joerg Enderlein, Robert C. Leif, Daniel L. Farkas; Eds.) 4. S. Beucher and F. Meyer, The Morphological Approach to Segmentation: The Watershed Transformation in: Mathematical Morphology in Image Processing, E.R. Dougherty Ed., pp , Marcel Dekker, New York, L. Vincent, P. Soille, "Watersheds in Digital Spaces: An Efficient Algorithm Based on Immersion Simulations", IEEE Transactions of Pattern Analysis and Machine Intelligence, 13, No. 6, pp , Image Processing Toolbox, The MathWorks, Inc J. Serra, Image Analysis and Mathematical Morphology, Vol. 1. Academic Press, London, US Pat. 6,671,624 Machine readable storage media for detecting distribution of macromolecules between nucleus and cytoplasm in cells. 9. Ilya Ravkin, Vladimir Temov, "Image analysis without segmentation - A new method to measure cytoplasm to nucleus translocation", poster, High Content Analysis Conference, San Francisco Slide: 36
Image Processing Fundamentals. Nicolas Vazquez Principal Software Engineer National Instruments
 Image Processing Fundamentals Nicolas Vazquez Principal Software Engineer National Instruments Agenda Objectives and Motivations Enhancing Images Checking for Presence Locating Parts Measuring Features
Image Processing Fundamentals Nicolas Vazquez Principal Software Engineer National Instruments Agenda Objectives and Motivations Enhancing Images Checking for Presence Locating Parts Measuring Features
doi: /
 Yiting Xie ; Anthony P. Reeves; Single 3D cell segmentation from optical CT microscope images. Proc. SPIE 934, Medical Imaging 214: Image Processing, 9343B (March 21, 214); doi:1.1117/12.243852. (214)
Yiting Xie ; Anthony P. Reeves; Single 3D cell segmentation from optical CT microscope images. Proc. SPIE 934, Medical Imaging 214: Image Processing, 9343B (March 21, 214); doi:1.1117/12.243852. (214)
BioImaging facility update: from multi-photon in vivo imaging to highcontent high-throughput image-based screening. Alex Laude The BioImaging Unit
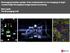 BioImaging facility update: from multi-photon in vivo imaging to highcontent high-throughput image-based screening Alex Laude The BioImaging Unit Multi-dimensional, multi-modal imaging at the sub-cellular
BioImaging facility update: from multi-photon in vivo imaging to highcontent high-throughput image-based screening Alex Laude The BioImaging Unit Multi-dimensional, multi-modal imaging at the sub-cellular
EE795: Computer Vision and Intelligent Systems
 EE795: Computer Vision and Intelligent Systems Spring 2012 TTh 17:30-18:45 WRI C225 Lecture 04 130131 http://www.ee.unlv.edu/~b1morris/ecg795/ 2 Outline Review Histogram Equalization Image Filtering Linear
EE795: Computer Vision and Intelligent Systems Spring 2012 TTh 17:30-18:45 WRI C225 Lecture 04 130131 http://www.ee.unlv.edu/~b1morris/ecg795/ 2 Outline Review Histogram Equalization Image Filtering Linear
Biomedical Image Analysis. Mathematical Morphology
 Biomedical Image Analysis Mathematical Morphology Contents: Foundation of Mathematical Morphology Structuring Elements Applications BMIA 15 V. Roth & P. Cattin 265 Foundations of Mathematical Morphology
Biomedical Image Analysis Mathematical Morphology Contents: Foundation of Mathematical Morphology Structuring Elements Applications BMIA 15 V. Roth & P. Cattin 265 Foundations of Mathematical Morphology
Colocalization Algorithm. User s Guide
 Colocalization Algorithm User s Guide Copyright 2008 Aperio Technologies, Inc. Part Number/Revision: MAN 0082, Revision A Date: March 7, 2008 This document applies to software versions Release 9.0 and
Colocalization Algorithm User s Guide Copyright 2008 Aperio Technologies, Inc. Part Number/Revision: MAN 0082, Revision A Date: March 7, 2008 This document applies to software versions Release 9.0 and
Digital Image Processing COSC 6380/4393
 Digital Image Processing COSC 6380/4393 Lecture 6 Sept 6 th, 2017 Pranav Mantini Slides from Dr. Shishir K Shah and Frank (Qingzhong) Liu Today Review Logical Operations on Binary Images Blob Coloring
Digital Image Processing COSC 6380/4393 Lecture 6 Sept 6 th, 2017 Pranav Mantini Slides from Dr. Shishir K Shah and Frank (Qingzhong) Liu Today Review Logical Operations on Binary Images Blob Coloring
09/11/2017. Morphological image processing. Morphological image processing. Morphological image processing. Morphological image processing (binary)
 Towards image analysis Goal: Describe the contents of an image, distinguishing meaningful information from irrelevant one. Perform suitable transformations of images so as to make explicit particular shape
Towards image analysis Goal: Describe the contents of an image, distinguishing meaningful information from irrelevant one. Perform suitable transformations of images so as to make explicit particular shape
Lecture 4 Image Enhancement in Spatial Domain
 Digital Image Processing Lecture 4 Image Enhancement in Spatial Domain Fall 2010 2 domains Spatial Domain : (image plane) Techniques are based on direct manipulation of pixels in an image Frequency Domain
Digital Image Processing Lecture 4 Image Enhancement in Spatial Domain Fall 2010 2 domains Spatial Domain : (image plane) Techniques are based on direct manipulation of pixels in an image Frequency Domain
Virtual Frap User Guide
 Virtual Frap User Guide http://wiki.vcell.uchc.edu/twiki/bin/view/vcell/vfrap Center for Cell Analysis and Modeling University of Connecticut Health Center 2010-1 - 1 Introduction Flourescence Photobleaching
Virtual Frap User Guide http://wiki.vcell.uchc.edu/twiki/bin/view/vcell/vfrap Center for Cell Analysis and Modeling University of Connecticut Health Center 2010-1 - 1 Introduction Flourescence Photobleaching
IN Cell Analyzer 1000 Analysis Modules Multi Target Analysis Training
 IN Cell Analyzer 1000 Analysis Modules Multi Target Analysis Training Multi Target Analysis Neurite Outgrowth Membrane Translocation Analysis Object Intensity Dual Area Object Analysis Granularity Morphology
IN Cell Analyzer 1000 Analysis Modules Multi Target Analysis Training Multi Target Analysis Neurite Outgrowth Membrane Translocation Analysis Object Intensity Dual Area Object Analysis Granularity Morphology
Background information:
 Image- based screening using subcellular localization of FOXO1A in osteosarcoma cells: A computer exercise using CellProfiler & CellProfiler Analyst software Carolina Wählby, Martin Simonsson, Megan Rokop
Image- based screening using subcellular localization of FOXO1A in osteosarcoma cells: A computer exercise using CellProfiler & CellProfiler Analyst software Carolina Wählby, Martin Simonsson, Megan Rokop
Biomedical Image Analysis. Point, Edge and Line Detection
 Biomedical Image Analysis Point, Edge and Line Detection Contents: Point and line detection Advanced edge detection: Canny Local/regional edge processing Global processing: Hough transform BMIA 15 V. Roth
Biomedical Image Analysis Point, Edge and Line Detection Contents: Point and line detection Advanced edge detection: Canny Local/regional edge processing Global processing: Hough transform BMIA 15 V. Roth
A Proposal for the Implementation of a Parallel Watershed Algorithm
 A Proposal for the Implementation of a Parallel Watershed Algorithm A. Meijster and J.B.T.M. Roerdink University of Groningen, Institute for Mathematics and Computing Science P.O. Box 800, 9700 AV Groningen,
A Proposal for the Implementation of a Parallel Watershed Algorithm A. Meijster and J.B.T.M. Roerdink University of Groningen, Institute for Mathematics and Computing Science P.O. Box 800, 9700 AV Groningen,
Image Processing: Final Exam November 10, :30 10:30
 Image Processing: Final Exam November 10, 2017-8:30 10:30 Student name: Student number: Put your name and student number on all of the papers you hand in (if you take out the staple). There are always
Image Processing: Final Exam November 10, 2017-8:30 10:30 Student name: Student number: Put your name and student number on all of the papers you hand in (if you take out the staple). There are always
T H E J O U R N A L O F C E L L B I O L O G Y
 Supplemental material Courtois et al., http://www.jcb.org/cgi/content/full/jcb.201202135/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Measurement of distribution and size of MTOCs in the
Supplemental material Courtois et al., http://www.jcb.org/cgi/content/full/jcb.201202135/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Measurement of distribution and size of MTOCs in the
SPCM Software Runs Online-FLIM at 10 Images per Second
 SPCM Software Runs Online-FLIM at 10 Images per Second Abstract: Version 9.72 SPCM software of the bh TCSPC/FLIM systems displays fluorescence lifetime images at a rate of 10 images per second. The calculation
SPCM Software Runs Online-FLIM at 10 Images per Second Abstract: Version 9.72 SPCM software of the bh TCSPC/FLIM systems displays fluorescence lifetime images at a rate of 10 images per second. The calculation
Nuclei Segmentation of Whole Slide Images in Digital Pathology
 Nuclei Segmentation of Whole Slide Images in Digital Pathology Dennis Ai Department of Electrical Engineering Stanford University Stanford, CA dennisai@stanford.edu Abstract Pathology is the study of the
Nuclei Segmentation of Whole Slide Images in Digital Pathology Dennis Ai Department of Electrical Engineering Stanford University Stanford, CA dennisai@stanford.edu Abstract Pathology is the study of the
Quantifying Three-Dimensional Deformations of Migrating Fibroblasts
 45 Chapter 4 Quantifying Three-Dimensional Deformations of Migrating Fibroblasts This chapter presents the full-field displacements and tractions of 3T3 fibroblast cells during migration on polyacrylamide
45 Chapter 4 Quantifying Three-Dimensional Deformations of Migrating Fibroblasts This chapter presents the full-field displacements and tractions of 3T3 fibroblast cells during migration on polyacrylamide
CS443: Digital Imaging and Multimedia Binary Image Analysis. Spring 2008 Ahmed Elgammal Dept. of Computer Science Rutgers University
 CS443: Digital Imaging and Multimedia Binary Image Analysis Spring 2008 Ahmed Elgammal Dept. of Computer Science Rutgers University Outlines A Simple Machine Vision System Image segmentation by thresholding
CS443: Digital Imaging and Multimedia Binary Image Analysis Spring 2008 Ahmed Elgammal Dept. of Computer Science Rutgers University Outlines A Simple Machine Vision System Image segmentation by thresholding
ECG782: Multidimensional Digital Signal Processing
 Professor Brendan Morris, SEB 3216, brendan.morris@unlv.edu ECG782: Multidimensional Digital Signal Processing Spatial Domain Filtering http://www.ee.unlv.edu/~b1morris/ecg782/ 2 Outline Background Intensity
Professor Brendan Morris, SEB 3216, brendan.morris@unlv.edu ECG782: Multidimensional Digital Signal Processing Spatial Domain Filtering http://www.ee.unlv.edu/~b1morris/ecg782/ 2 Outline Background Intensity
Image Processing. Bilkent University. CS554 Computer Vision Pinar Duygulu
 Image Processing CS 554 Computer Vision Pinar Duygulu Bilkent University Today Image Formation Point and Blob Processing Binary Image Processing Readings: Gonzalez & Woods, Ch. 3 Slides are adapted from
Image Processing CS 554 Computer Vision Pinar Duygulu Bilkent University Today Image Formation Point and Blob Processing Binary Image Processing Readings: Gonzalez & Woods, Ch. 3 Slides are adapted from
Independent Resolution Test of
 Independent Resolution Test of as conducted and published by Dr. Adam Puche, PhD University of Maryland June 2005 as presented by (formerly Thales Optem Inc.) www.qioptiqimaging.com Independent Resolution
Independent Resolution Test of as conducted and published by Dr. Adam Puche, PhD University of Maryland June 2005 as presented by (formerly Thales Optem Inc.) www.qioptiqimaging.com Independent Resolution
CITS 4402 Computer Vision
 CITS 4402 Computer Vision A/Prof Ajmal Mian Adj/A/Prof Mehdi Ravanbakhsh, CEO at Mapizy (www.mapizy.com) and InFarm (www.infarm.io) Lecture 02 Binary Image Analysis Objectives Revision of image formation
CITS 4402 Computer Vision A/Prof Ajmal Mian Adj/A/Prof Mehdi Ravanbakhsh, CEO at Mapizy (www.mapizy.com) and InFarm (www.infarm.io) Lecture 02 Binary Image Analysis Objectives Revision of image formation
Anno accademico 2006/2007. Davide Migliore
 Robotica Anno accademico 6/7 Davide Migliore migliore@elet.polimi.it Today What is a feature? Some useful information The world of features: Detectors Edges detection Corners/Points detection Descriptors?!?!?
Robotica Anno accademico 6/7 Davide Migliore migliore@elet.polimi.it Today What is a feature? Some useful information The world of features: Detectors Edges detection Corners/Points detection Descriptors?!?!?
New Edge-Enhanced Error Diffusion Algorithm Based on the Error Sum Criterion
 New Edge-Enhanced Error Diffusion Algorithm Based on the Error Sum Criterion Jae Ho Kim* Tae Il Chung Hyung Soon Kim* Kyung Sik Son* Pusan National University Image and Communication Laboratory San 3,
New Edge-Enhanced Error Diffusion Algorithm Based on the Error Sum Criterion Jae Ho Kim* Tae Il Chung Hyung Soon Kim* Kyung Sik Son* Pusan National University Image and Communication Laboratory San 3,
45 µm polystyrene bead embedded in scattering tissue phantom. (a,b) raw images under oblique
 Phase gradient microscopy in thick tissue with oblique back-illumination Tim N Ford, Kengyeh K Chu & Jerome Mertz Supplementary Figure 1: Comparison of added versus subtracted raw OBM images 45 µm polystyrene
Phase gradient microscopy in thick tissue with oblique back-illumination Tim N Ford, Kengyeh K Chu & Jerome Mertz Supplementary Figure 1: Comparison of added versus subtracted raw OBM images 45 µm polystyrene
CHAPTER 6 DETECTION OF MASS USING NOVEL SEGMENTATION, GLCM AND NEURAL NETWORKS
 130 CHAPTER 6 DETECTION OF MASS USING NOVEL SEGMENTATION, GLCM AND NEURAL NETWORKS A mass is defined as a space-occupying lesion seen in more than one projection and it is described by its shapes and margin
130 CHAPTER 6 DETECTION OF MASS USING NOVEL SEGMENTATION, GLCM AND NEURAL NETWORKS A mass is defined as a space-occupying lesion seen in more than one projection and it is described by its shapes and margin
Network Snakes for the Segmentation of Adjacent Cells in Confocal Images
 Network Snakes for the Segmentation of Adjacent Cells in Confocal Images Matthias Butenuth 1 and Fritz Jetzek 2 1 Institut für Photogrammetrie und GeoInformation, Leibniz Universität Hannover, 30167 Hannover
Network Snakes for the Segmentation of Adjacent Cells in Confocal Images Matthias Butenuth 1 and Fritz Jetzek 2 1 Institut für Photogrammetrie und GeoInformation, Leibniz Universität Hannover, 30167 Hannover
Digital Volume Correlation for Materials Characterization
 19 th World Conference on Non-Destructive Testing 2016 Digital Volume Correlation for Materials Characterization Enrico QUINTANA, Phillip REU, Edward JIMENEZ, Kyle THOMPSON, Sharlotte KRAMER Sandia National
19 th World Conference on Non-Destructive Testing 2016 Digital Volume Correlation for Materials Characterization Enrico QUINTANA, Phillip REU, Edward JIMENEZ, Kyle THOMPSON, Sharlotte KRAMER Sandia National
ND Processing Tools in NIS-Elements
 ND Processing Tools in NIS-Elements Overview This technical note describes basic uses of the ND processing tools available in NIS-Elements. These tools are specifically designed for arithmetic functions
ND Processing Tools in NIS-Elements Overview This technical note describes basic uses of the ND processing tools available in NIS-Elements. These tools are specifically designed for arithmetic functions
Types of Edges. Why Edge Detection? Types of Edges. Edge Detection. Gradient. Edge Detection
 Why Edge Detection? How can an algorithm extract relevant information from an image that is enables the algorithm to recognize objects? The most important information for the interpretation of an image
Why Edge Detection? How can an algorithm extract relevant information from an image that is enables the algorithm to recognize objects? The most important information for the interpretation of an image
Research Article Image Segmentation Using Gray-Scale Morphology and Marker-Controlled Watershed Transformation
 Discrete Dynamics in Nature and Society Volume 2008, Article ID 384346, 8 pages doi:10.1155/2008/384346 Research Article Image Segmentation Using Gray-Scale Morphology and Marker-Controlled Watershed Transformation
Discrete Dynamics in Nature and Society Volume 2008, Article ID 384346, 8 pages doi:10.1155/2008/384346 Research Article Image Segmentation Using Gray-Scale Morphology and Marker-Controlled Watershed Transformation
Supplementary Figures
 Supplementary Figures re co rd ed ch annels darkf ield MPM2 PI cell cycle phases G1/S/G2 prophase metaphase anaphase telophase bri ghtfield 55 pixel Supplementary Figure 1 Images of Jurkat cells captured
Supplementary Figures re co rd ed ch annels darkf ield MPM2 PI cell cycle phases G1/S/G2 prophase metaphase anaphase telophase bri ghtfield 55 pixel Supplementary Figure 1 Images of Jurkat cells captured
Effect of OTF attenuation on single-slice SR-SIM reconstruction
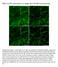 Effect of OTF attenuation on single-slice SR-SIM reconstruction Supplementary Figure 1: Actin filaments in U2OS cells, labelled with Phalloidin-Atto488, measured on a DeltaVision OMX, excited at 488 nm
Effect of OTF attenuation on single-slice SR-SIM reconstruction Supplementary Figure 1: Actin filaments in U2OS cells, labelled with Phalloidin-Atto488, measured on a DeltaVision OMX, excited at 488 nm
MACHINE VISION AS A METHOD FOR CHARACTERIZING SOLAR TRACKER PERFORMANCE
 MACHINE VISION AS A METHOD FOR CHARACTERIZING SOLAR TRACKER PERFORMANCE M. Davis, J. Lawler, J. Coyle, A. Reich, T. Williams GreenMountain Engineering, LLC ABSTRACT This paper describes an approach to
MACHINE VISION AS A METHOD FOR CHARACTERIZING SOLAR TRACKER PERFORMANCE M. Davis, J. Lawler, J. Coyle, A. Reich, T. Williams GreenMountain Engineering, LLC ABSTRACT This paper describes an approach to
Contour LS-K Optical Surface Profiler
 Contour LS-K Optical Surface Profiler LightSpeed Focus Variation Provides High-Speed Metrology without Compromise Innovation with Integrity Optical & Stylus Metrology Deeper Understanding More Quickly
Contour LS-K Optical Surface Profiler LightSpeed Focus Variation Provides High-Speed Metrology without Compromise Innovation with Integrity Optical & Stylus Metrology Deeper Understanding More Quickly
SENSITIVITY DIFFERENTLY TO SEE THINGS THE SPEED AND. Opera Phenix High Content Screening System
 THE SPEED AND SENSITIVITY TO SEE THINGS DIFFERENTLY Opera Phenix High Content Screening System For research use only. Not for use in diagnostic procedures. HIGH CONTENT SCREENING WITHOUT THE COMPROMISE
THE SPEED AND SENSITIVITY TO SEE THINGS DIFFERENTLY Opera Phenix High Content Screening System For research use only. Not for use in diagnostic procedures. HIGH CONTENT SCREENING WITHOUT THE COMPROMISE
Binary Shape Characterization using Morphological Boundary Class Distribution Functions
 Binary Shape Characterization using Morphological Boundary Class Distribution Functions Marcin Iwanowski Institute of Control and Industrial Electronics, Warsaw University of Technology, ul.koszykowa 75,
Binary Shape Characterization using Morphological Boundary Class Distribution Functions Marcin Iwanowski Institute of Control and Industrial Electronics, Warsaw University of Technology, ul.koszykowa 75,
Simplified model of ray propagation and outcoupling in TFs.
 Supplementary Figure 1 Simplified model of ray propagation and outcoupling in TFs. After total reflection at the core boundary with an angle α, a ray entering the taper (blue line) hits the taper sidewalls
Supplementary Figure 1 Simplified model of ray propagation and outcoupling in TFs. After total reflection at the core boundary with an angle α, a ray entering the taper (blue line) hits the taper sidewalls
Study on the Signboard Region Detection in Natural Image
 , pp.179-184 http://dx.doi.org/10.14257/astl.2016.140.34 Study on the Signboard Region Detection in Natural Image Daeyeong Lim 1, Youngbaik Kim 2, Incheol Park 1, Jihoon seung 1, Kilto Chong 1,* 1 1567
, pp.179-184 http://dx.doi.org/10.14257/astl.2016.140.34 Study on the Signboard Region Detection in Natural Image Daeyeong Lim 1, Youngbaik Kim 2, Incheol Park 1, Jihoon seung 1, Kilto Chong 1,* 1 1567
EECS490: Digital Image Processing. Lecture #22
 Lecture #22 Gold Standard project images Otsu thresholding Local thresholding Region segmentation Watershed segmentation Frequency-domain techniques Project Images 1 Project Images 2 Project Images 3 Project
Lecture #22 Gold Standard project images Otsu thresholding Local thresholding Region segmentation Watershed segmentation Frequency-domain techniques Project Images 1 Project Images 2 Project Images 3 Project
ECG782: Multidimensional Digital Signal Processing
 Professor Brendan Morris, SEB 3216, brendan.morris@unlv.edu ECG782: Multidimensional Digital Signal Processing Spring 2014 TTh 14:30-15:45 CBC C313 Lecture 03 Image Processing Basics 13/01/28 http://www.ee.unlv.edu/~b1morris/ecg782/
Professor Brendan Morris, SEB 3216, brendan.morris@unlv.edu ECG782: Multidimensional Digital Signal Processing Spring 2014 TTh 14:30-15:45 CBC C313 Lecture 03 Image Processing Basics 13/01/28 http://www.ee.unlv.edu/~b1morris/ecg782/
Interactive Math Glossary Terms and Definitions
 Terms and Definitions Absolute Value the magnitude of a number, or the distance from 0 on a real number line Addend any number or quantity being added addend + addend = sum Additive Property of Area the
Terms and Definitions Absolute Value the magnitude of a number, or the distance from 0 on a real number line Addend any number or quantity being added addend + addend = sum Additive Property of Area the
Cellular Imaging Solutions Imaging with a vision
 Cellular Imaging Solutions Imaging with a vision www.moleculardevices.com High content screening (HCS) utilizes automated, high-resolution microscopy systems to assay and visualize phenotypic responses
Cellular Imaging Solutions Imaging with a vision www.moleculardevices.com High content screening (HCS) utilizes automated, high-resolution microscopy systems to assay and visualize phenotypic responses
An Intuitive Explanation of Fourier Theory
 An Intuitive Explanation of Fourier Theory Steven Lehar slehar@cns.bu.edu Fourier theory is pretty complicated mathematically. But there are some beautifully simple holistic concepts behind Fourier theory
An Intuitive Explanation of Fourier Theory Steven Lehar slehar@cns.bu.edu Fourier theory is pretty complicated mathematically. But there are some beautifully simple holistic concepts behind Fourier theory
Coupling of surface roughness to the performance of computer-generated holograms
 Coupling of surface roughness to the performance of computer-generated holograms Ping Zhou* and Jim Burge College of Optical Sciences, University of Arizona, Tucson, Arizona 85721, USA *Corresponding author:
Coupling of surface roughness to the performance of computer-generated holograms Ping Zhou* and Jim Burge College of Optical Sciences, University of Arizona, Tucson, Arizona 85721, USA *Corresponding author:
Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging
 1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
Impact of 3D Laser Data Resolution and Accuracy on Pipeline Dents Strain Analysis
 More Info at Open Access Database www.ndt.net/?id=15137 Impact of 3D Laser Data Resolution and Accuracy on Pipeline Dents Strain Analysis Jean-Simon Fraser, Pierre-Hugues Allard Creaform, 5825 rue St-Georges,
More Info at Open Access Database www.ndt.net/?id=15137 Impact of 3D Laser Data Resolution and Accuracy on Pipeline Dents Strain Analysis Jean-Simon Fraser, Pierre-Hugues Allard Creaform, 5825 rue St-Georges,
SECTION 5 IMAGE PROCESSING 2
 SECTION 5 IMAGE PROCESSING 2 5.1 Resampling 3 5.1.1 Image Interpolation Comparison 3 5.2 Convolution 3 5.3 Smoothing Filters 3 5.3.1 Mean Filter 3 5.3.2 Median Filter 4 5.3.3 Pseudomedian Filter 6 5.3.4
SECTION 5 IMAGE PROCESSING 2 5.1 Resampling 3 5.1.1 Image Interpolation Comparison 3 5.2 Convolution 3 5.3 Smoothing Filters 3 5.3.1 Mean Filter 3 5.3.2 Median Filter 4 5.3.3 Pseudomedian Filter 6 5.3.4
Colocalization in Confocal Microscopy. Quantitative Colocalization Analysis
 Quantitative Colocalization Analysis 1 Quantitative Colocalization Analysis Colocalization of fluorescently labeled molecules (immunostainings, fluorescent proteins, etc.) is often used as the first indicator
Quantitative Colocalization Analysis 1 Quantitative Colocalization Analysis Colocalization of fluorescently labeled molecules (immunostainings, fluorescent proteins, etc.) is often used as the first indicator
2, the coefficient of variation R 2, and properties of the photon counts traces
 Supplementary Figure 1 Quality control of FCS traces. (a) Typical trace that passes the quality control (QC) according to the parameters shown in f. The QC is based on thresholds applied to fitting parameters
Supplementary Figure 1 Quality control of FCS traces. (a) Typical trace that passes the quality control (QC) according to the parameters shown in f. The QC is based on thresholds applied to fitting parameters
WCIF COLOCALIS ATION PLUGINS
 WCIF COLOCALIS ATION PLUGINS Colocalisation Test Background When a coefficient is calculated for two images, it is often unclear quite what this means, in particular for intermediate values. This raises
WCIF COLOCALIS ATION PLUGINS Colocalisation Test Background When a coefficient is calculated for two images, it is often unclear quite what this means, in particular for intermediate values. This raises
GENERAL AUTOMATED FLAW DETECTION SCHEME FOR NDE X-RAY IMAGES
 GENERAL AUTOMATED FLAW DETECTION SCHEME FOR NDE X-RAY IMAGES Karl W. Ulmer and John P. Basart Center for Nondestructive Evaluation Department of Electrical and Computer Engineering Iowa State University
GENERAL AUTOMATED FLAW DETECTION SCHEME FOR NDE X-RAY IMAGES Karl W. Ulmer and John P. Basart Center for Nondestructive Evaluation Department of Electrical and Computer Engineering Iowa State University
EyeTech. Particle Size Particle Shape Particle concentration Analyzer ANKERSMID
 EyeTech Particle Size Particle Shape Particle concentration Analyzer A new technology for measuring particle size in combination with particle shape and concentration. COMBINED LASERTECHNOLOGY & DIA Content
EyeTech Particle Size Particle Shape Particle concentration Analyzer A new technology for measuring particle size in combination with particle shape and concentration. COMBINED LASERTECHNOLOGY & DIA Content
Descriptive Statistics, Standard Deviation and Standard Error
 AP Biology Calculations: Descriptive Statistics, Standard Deviation and Standard Error SBI4UP The Scientific Method & Experimental Design Scientific method is used to explore observations and answer questions.
AP Biology Calculations: Descriptive Statistics, Standard Deviation and Standard Error SBI4UP The Scientific Method & Experimental Design Scientific method is used to explore observations and answer questions.
Vocabulary. 5-number summary Rule. Area principle. Bar chart. Boxplot. Categorical data condition. Categorical variable.
 5-number summary 68-95-99.7 Rule Area principle Bar chart Bimodal Boxplot Case Categorical data Categorical variable Center Changing center and spread Conditional distribution Context Contingency table
5-number summary 68-95-99.7 Rule Area principle Bar chart Bimodal Boxplot Case Categorical data Categorical variable Center Changing center and spread Conditional distribution Context Contingency table
Image Enhancement Using Fuzzy Morphology
 Image Enhancement Using Fuzzy Morphology Dillip Ranjan Nayak, Assistant Professor, Department of CSE, GCEK Bhwanipatna, Odissa, India Ashutosh Bhoi, Lecturer, Department of CSE, GCEK Bhawanipatna, Odissa,
Image Enhancement Using Fuzzy Morphology Dillip Ranjan Nayak, Assistant Professor, Department of CSE, GCEK Bhwanipatna, Odissa, India Ashutosh Bhoi, Lecturer, Department of CSE, GCEK Bhawanipatna, Odissa,
Sample study by 3D optical profiler Contour Elite K for KTH university.
 Sample study by 3D optical profiler Contour Elite K for KTH university Samuel.lesko@bruker.com Objectives Objectives Main goals for the visit consist of evaluating 3D optical profiler: Confirm capability
Sample study by 3D optical profiler Contour Elite K for KTH university Samuel.lesko@bruker.com Objectives Objectives Main goals for the visit consist of evaluating 3D optical profiler: Confirm capability
CHAPTER 3 IMAGE ENHANCEMENT IN THE SPATIAL DOMAIN
 CHAPTER 3 IMAGE ENHANCEMENT IN THE SPATIAL DOMAIN CHAPTER 3: IMAGE ENHANCEMENT IN THE SPATIAL DOMAIN Principal objective: to process an image so that the result is more suitable than the original image
CHAPTER 3 IMAGE ENHANCEMENT IN THE SPATIAL DOMAIN CHAPTER 3: IMAGE ENHANCEMENT IN THE SPATIAL DOMAIN Principal objective: to process an image so that the result is more suitable than the original image
ECEN 447 Digital Image Processing
 ECEN 447 Digital Image Processing Lecture 7: Mathematical Morphology Ulisses Braga-Neto ECE Department Texas A&M University Basics of Mathematical Morphology Mathematical Morphology (MM) is a discipline
ECEN 447 Digital Image Processing Lecture 7: Mathematical Morphology Ulisses Braga-Neto ECE Department Texas A&M University Basics of Mathematical Morphology Mathematical Morphology (MM) is a discipline
Algorithm User Guide:
 Algorithm User Guide: Membrane Quantification Use the Aperio algorithms to adjust (tune) the parameters until the quantitative results are sufficiently accurate for the purpose for which you intend to
Algorithm User Guide: Membrane Quantification Use the Aperio algorithms to adjust (tune) the parameters until the quantitative results are sufficiently accurate for the purpose for which you intend to
Topic 4 Image Segmentation
 Topic 4 Image Segmentation What is Segmentation? Why? Segmentation important contributing factor to the success of an automated image analysis process What is Image Analysis: Processing images to derive
Topic 4 Image Segmentation What is Segmentation? Why? Segmentation important contributing factor to the success of an automated image analysis process What is Image Analysis: Processing images to derive
CHAPTER 3 RETINAL OPTIC DISC SEGMENTATION
 60 CHAPTER 3 RETINAL OPTIC DISC SEGMENTATION 3.1 IMPORTANCE OF OPTIC DISC Ocular fundus images provide information about ophthalmic, retinal and even systemic diseases such as hypertension, diabetes, macular
60 CHAPTER 3 RETINAL OPTIC DISC SEGMENTATION 3.1 IMPORTANCE OF OPTIC DISC Ocular fundus images provide information about ophthalmic, retinal and even systemic diseases such as hypertension, diabetes, macular
Digital image processing
 Digital image processing Image enhancement algorithms: grey scale transformations Any digital image can be represented mathematically in matrix form. The number of lines in the matrix is the number of
Digital image processing Image enhancement algorithms: grey scale transformations Any digital image can be represented mathematically in matrix form. The number of lines in the matrix is the number of
Critique: Efficient Iris Recognition by Characterizing Key Local Variations
 Critique: Efficient Iris Recognition by Characterizing Key Local Variations Authors: L. Ma, T. Tan, Y. Wang, D. Zhang Published: IEEE Transactions on Image Processing, Vol. 13, No. 6 Critique By: Christopher
Critique: Efficient Iris Recognition by Characterizing Key Local Variations Authors: L. Ma, T. Tan, Y. Wang, D. Zhang Published: IEEE Transactions on Image Processing, Vol. 13, No. 6 Critique By: Christopher
Lecture 4. Digital Image Enhancement. 1. Principle of image enhancement 2. Spatial domain transformation. Histogram processing
 Lecture 4 Digital Image Enhancement 1. Principle of image enhancement 2. Spatial domain transformation Basic intensity it tranfomation ti Histogram processing Principle Objective of Enhancement Image enhancement
Lecture 4 Digital Image Enhancement 1. Principle of image enhancement 2. Spatial domain transformation Basic intensity it tranfomation ti Histogram processing Principle Objective of Enhancement Image enhancement
Bioimage Informatics
 Bioimage Informatics Lecture 14, Spring 2012 Bioimage Data Analysis (IV) Image Segmentation (part 3) Lecture 14 March 07, 2012 1 Outline Review: intensity thresholding based image segmentation Morphological
Bioimage Informatics Lecture 14, Spring 2012 Bioimage Data Analysis (IV) Image Segmentation (part 3) Lecture 14 March 07, 2012 1 Outline Review: intensity thresholding based image segmentation Morphological
A Tutorial Guide to Tribology Plug-in
 Supplementary Material A Tutorial Guide to Tribology Plug-in Tribology An ImageJ Plugin for surface topography analysis of laser textured surfaces. General Description This plugin presents an easy-to-use
Supplementary Material A Tutorial Guide to Tribology Plug-in Tribology An ImageJ Plugin for surface topography analysis of laser textured surfaces. General Description This plugin presents an easy-to-use
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature10934 Supplementary Methods Mathematical implementation of the EST method. The EST method begins with padding each projection with zeros (that is, embedding
SUPPLEMENTARY INFORMATION doi:10.1038/nature10934 Supplementary Methods Mathematical implementation of the EST method. The EST method begins with padding each projection with zeros (that is, embedding
Edges and Binary Images
 CS 699: Intro to Computer Vision Edges and Binary Images Prof. Adriana Kovashka University of Pittsburgh September 5, 205 Plan for today Edge detection Binary image analysis Homework Due on 9/22, :59pm
CS 699: Intro to Computer Vision Edges and Binary Images Prof. Adriana Kovashka University of Pittsburgh September 5, 205 Plan for today Edge detection Binary image analysis Homework Due on 9/22, :59pm
Introduction. Loading Images
 Introduction CellProfiler is a free Open Source software for automated image analysis. Versions for Mac, Windows and Linux are available and can be downloaded at: http://www.cellprofiler.org/. CellProfiler
Introduction CellProfiler is a free Open Source software for automated image analysis. Versions for Mac, Windows and Linux are available and can be downloaded at: http://www.cellprofiler.org/. CellProfiler
AnalySIS Tutorial part 2
 AnalySIS Tutorial part 2 Sveinung Lillehaug Neural Systems and Graphics Computing Laboratory Department of Anatomy University of Oslo N-0317 Oslo Norway www.nesys.uio.no Using AnalySIS to automatically
AnalySIS Tutorial part 2 Sveinung Lillehaug Neural Systems and Graphics Computing Laboratory Department of Anatomy University of Oslo N-0317 Oslo Norway www.nesys.uio.no Using AnalySIS to automatically
Algorithm User Guide:
 Algorithm User Guide: Microvessel Analysis Use the Aperio algorithms to adjust (tune) the parameters until the quantitative results are sufficiently accurate for the purpose for which you intend to use
Algorithm User Guide: Microvessel Analysis Use the Aperio algorithms to adjust (tune) the parameters until the quantitative results are sufficiently accurate for the purpose for which you intend to use
Range Imaging Through Triangulation. Range Imaging Through Triangulation. Range Imaging Through Triangulation. Range Imaging Through Triangulation
 Obviously, this is a very slow process and not suitable for dynamic scenes. To speed things up, we can use a laser that projects a vertical line of light onto the scene. This laser rotates around its vertical
Obviously, this is a very slow process and not suitable for dynamic scenes. To speed things up, we can use a laser that projects a vertical line of light onto the scene. This laser rotates around its vertical
Counting Particles or Cells Using IMAQ Vision
 Application Note 107 Counting Particles or Cells Using IMAQ Vision John Hanks Introduction To count objects, you use a common image processing technique called particle analysis, often referred to as blob
Application Note 107 Counting Particles or Cells Using IMAQ Vision John Hanks Introduction To count objects, you use a common image processing technique called particle analysis, often referred to as blob
LEGENDplex Data Analysis Software Version 8 User Guide
 LEGENDplex Data Analysis Software Version 8 User Guide Introduction Welcome to the user s guide for Version 8 of the LEGENDplex data analysis software for Windows based computers 1. This tutorial will
LEGENDplex Data Analysis Software Version 8 User Guide Introduction Welcome to the user s guide for Version 8 of the LEGENDplex data analysis software for Windows based computers 1. This tutorial will
Detection of Sub-resolution Dots in Microscopy Images
 Detection of Sub-resolution Dots in Microscopy Images Karel Štěpka, 2012 Centre for Biomedical Image Analysis, FI MU supervisor: prof. RNDr. Michal Kozubek, Ph.D. Outline Introduction Existing approaches
Detection of Sub-resolution Dots in Microscopy Images Karel Štěpka, 2012 Centre for Biomedical Image Analysis, FI MU supervisor: prof. RNDr. Michal Kozubek, Ph.D. Outline Introduction Existing approaches
Morphological Image Processing
 Morphological Image Processing Binary image processing In binary images, we conventionally take background as black (0) and foreground objects as white (1 or 255) Morphology Figure 4.1 objects on a conveyor
Morphological Image Processing Binary image processing In binary images, we conventionally take background as black (0) and foreground objects as white (1 or 255) Morphology Figure 4.1 objects on a conveyor
Mu lt i s p e c t r a l
 Viewing Angle Analyser Revolutionary system for full spectral and polarization measurement in the entire viewing angle EZContrastMS80 & EZContrastMS88 ADVANCED LIGHT ANALYSIS by Field iris Fourier plane
Viewing Angle Analyser Revolutionary system for full spectral and polarization measurement in the entire viewing angle EZContrastMS80 & EZContrastMS88 ADVANCED LIGHT ANALYSIS by Field iris Fourier plane
A. K. Srivastava, K.C. Sati, Satyander Kumar alaser Science and Technology Center, Metcalfe House, Civil Lines, Delhi , INDIA
 International Journal of Scientific & Engineering Research Volume 8, Issue 7, July-2017 1752 Optical method for measurement of radius of curvature of large diameter mirrors A. K. Srivastava, K.C. Sati,
International Journal of Scientific & Engineering Research Volume 8, Issue 7, July-2017 1752 Optical method for measurement of radius of curvature of large diameter mirrors A. K. Srivastava, K.C. Sati,
Histograms. h(r k ) = n k. p(r k )= n k /NM. Histogram: number of times intensity level rk appears in the image
 Histograms h(r k ) = n k Histogram: number of times intensity level rk appears in the image p(r k )= n k /NM normalized histogram also a probability of occurence 1 Histogram of Image Intensities Create
Histograms h(r k ) = n k Histogram: number of times intensity level rk appears in the image p(r k )= n k /NM normalized histogram also a probability of occurence 1 Histogram of Image Intensities Create
Original grey level r Fig.1
 Point Processing: In point processing, we work with single pixels i.e. T is 1 x 1 operator. It means that the new value f(x, y) depends on the operator T and the present f(x, y). Some of the common examples
Point Processing: In point processing, we work with single pixels i.e. T is 1 x 1 operator. It means that the new value f(x, y) depends on the operator T and the present f(x, y). Some of the common examples
EECS490: Digital Image Processing. Lecture #19
 Lecture #19 Shading and texture analysis using morphology Gray scale reconstruction Basic image segmentation: edges v. regions Point and line locators, edge types and noise Edge operators: LoG, DoG, Canny
Lecture #19 Shading and texture analysis using morphology Gray scale reconstruction Basic image segmentation: edges v. regions Point and line locators, edge types and noise Edge operators: LoG, DoG, Canny
REMOTE SENSING OF SURFACE STRUCTURES
 REMOTE SENSING OF SURFACE STRUCTURES A.W. Koch, P. Evanschitzky and M. Jakobi Technische Universität München Institute for Measurement Systems and Sensor Technology D-8090 München, Germany Abstract: The
REMOTE SENSING OF SURFACE STRUCTURES A.W. Koch, P. Evanschitzky and M. Jakobi Technische Universität München Institute for Measurement Systems and Sensor Technology D-8090 München, Germany Abstract: The
DESIGNER S NOTEBOOK Proximity Detection and Link Budget By Tom Dunn July 2011
 INTELLIGENT OPTO SENSOR Number 38 DESIGNER S NOTEBOOK Proximity Detection and Link Budget By Tom Dunn July 2011 Overview TAOS proximity sensors operate by flashing an infrared (IR) light towards a surface
INTELLIGENT OPTO SENSOR Number 38 DESIGNER S NOTEBOOK Proximity Detection and Link Budget By Tom Dunn July 2011 Overview TAOS proximity sensors operate by flashing an infrared (IR) light towards a surface
Morphological Image Processing
 Morphological Image Processing Binary dilation and erosion" Set-theoretic interpretation" Opening, closing, morphological edge detectors" Hit-miss filter" Morphological filters for gray-level images" Cascading
Morphological Image Processing Binary dilation and erosion" Set-theoretic interpretation" Opening, closing, morphological edge detectors" Hit-miss filter" Morphological filters for gray-level images" Cascading
Math 7 Glossary Terms
 Math 7 Glossary Terms Absolute Value Absolute value is the distance, or number of units, a number is from zero. Distance is always a positive value; therefore, absolute value is always a positive value.
Math 7 Glossary Terms Absolute Value Absolute value is the distance, or number of units, a number is from zero. Distance is always a positive value; therefore, absolute value is always a positive value.
Optimization of optical systems for LED spot lights concerning the color uniformity
 Optimization of optical systems for LED spot lights concerning the color uniformity Anne Teupner* a, Krister Bergenek b, Ralph Wirth b, Juan C. Miñano a, Pablo Benítez a a Technical University of Madrid,
Optimization of optical systems for LED spot lights concerning the color uniformity Anne Teupner* a, Krister Bergenek b, Ralph Wirth b, Juan C. Miñano a, Pablo Benítez a a Technical University of Madrid,
LOCALIZATION OF FACIAL REGIONS AND FEATURES IN COLOR IMAGES. Karin Sobottka Ioannis Pitas
 LOCALIZATION OF FACIAL REGIONS AND FEATURES IN COLOR IMAGES Karin Sobottka Ioannis Pitas Department of Informatics, University of Thessaloniki 540 06, Greece e-mail:fsobottka, pitasg@zeus.csd.auth.gr Index
LOCALIZATION OF FACIAL REGIONS AND FEATURES IN COLOR IMAGES Karin Sobottka Ioannis Pitas Department of Informatics, University of Thessaloniki 540 06, Greece e-mail:fsobottka, pitasg@zeus.csd.auth.gr Index
Raster Image Correlation Spectroscopy and Number and Brightness Analysis
 Raster Image Correlation Spectroscopy and Number and Brightness Analysis 15 Principles of Fluorescence Course Paolo Annibale, PhD On behalf of Dr. Enrico Gratton lfd Raster Image Correlation Spectroscopy
Raster Image Correlation Spectroscopy and Number and Brightness Analysis 15 Principles of Fluorescence Course Paolo Annibale, PhD On behalf of Dr. Enrico Gratton lfd Raster Image Correlation Spectroscopy
Morphology-form and structure. Who am I? structuring element (SE) Today s lecture. Morphological Transformation. Mathematical Morphology
 Mathematical Morphology Morphology-form and structure Sonka 13.1-13.6 Ida-Maria Sintorn Ida.sintorn@cb.uu.se mathematical framework used for: pre-processing - noise filtering, shape simplification,...
Mathematical Morphology Morphology-form and structure Sonka 13.1-13.6 Ida-Maria Sintorn Ida.sintorn@cb.uu.se mathematical framework used for: pre-processing - noise filtering, shape simplification,...
QUALITY ASSURANCE OF MULTIFIBER CONNECTORS
 QUALITY ASSURANCE OF MULTIFIBER CONNECTORS Eric A. Norland/ Daniel Beerbohm Norland Products Inc. Cranbury, NJ 08512 ABSTRACT Industry requirements for compact, high fiber count connectors in both telecom
QUALITY ASSURANCE OF MULTIFIBER CONNECTORS Eric A. Norland/ Daniel Beerbohm Norland Products Inc. Cranbury, NJ 08512 ABSTRACT Industry requirements for compact, high fiber count connectors in both telecom
COLOCALISATION. Alexia Ferrand. Imaging Core Facility Biozentrum Basel
 COLOCALISATION Alexia Ferrand Imaging Core Facility Biozentrum Basel OUTLINE Introduction How to best prepare your samples for colocalisation How to acquire the images for colocalisation How to analyse
COLOCALISATION Alexia Ferrand Imaging Core Facility Biozentrum Basel OUTLINE Introduction How to best prepare your samples for colocalisation How to acquire the images for colocalisation How to analyse
Fast and accurate automated cell boundary determination for fluorescence microscopy
 Fast and accurate automated cell boundary determination for fluorescence microscopy Stephen Hugo Arce, Pei-Hsun Wu &, and Yiider Tseng Department of Chemical Engineering, University of Florida and National
Fast and accurate automated cell boundary determination for fluorescence microscopy Stephen Hugo Arce, Pei-Hsun Wu &, and Yiider Tseng Department of Chemical Engineering, University of Florida and National
Image Restoration and Reconstruction
 Image Restoration and Reconstruction Image restoration Objective process to improve an image, as opposed to the subjective process of image enhancement Enhancement uses heuristics to improve the image
Image Restoration and Reconstruction Image restoration Objective process to improve an image, as opposed to the subjective process of image enhancement Enhancement uses heuristics to improve the image
Single-particle electron microscopy (cryo-electron microscopy) CS/CME/BioE/Biophys/BMI 279 Nov. 16 and 28, 2017 Ron Dror
 Single-particle electron microscopy (cryo-electron microscopy) CS/CME/BioE/Biophys/BMI 279 Nov. 16 and 28, 2017 Ron Dror 1 Last month s Nobel Prize in Chemistry Awarded to Jacques Dubochet, Joachim Frank
Single-particle electron microscopy (cryo-electron microscopy) CS/CME/BioE/Biophys/BMI 279 Nov. 16 and 28, 2017 Ron Dror 1 Last month s Nobel Prize in Chemistry Awarded to Jacques Dubochet, Joachim Frank
Edge and local feature detection - 2. Importance of edge detection in computer vision
 Edge and local feature detection Gradient based edge detection Edge detection by function fitting Second derivative edge detectors Edge linking and the construction of the chain graph Edge and local feature
Edge and local feature detection Gradient based edge detection Edge detection by function fitting Second derivative edge detectors Edge linking and the construction of the chain graph Edge and local feature
Segmentation
 Lecture 6: Segmentation 24--4 Robin Strand Centre for Image Analysis Dept. of IT Uppsala University Today What is image segmentation? A smörgåsbord of methods for image segmentation: Thresholding Edge-based
Lecture 6: Segmentation 24--4 Robin Strand Centre for Image Analysis Dept. of IT Uppsala University Today What is image segmentation? A smörgåsbord of methods for image segmentation: Thresholding Edge-based
University of Minnesota Nano Fabrication Center Standard Operating Procedure
 Equipment Name: University of Minnesota Nano Fabrication Center Coral Name: hs-scope Revision Number: 1.5 Model: HS200A Revisionist: M. Fisher Location: Bay 1 Date: 9/12/2013 1 Description The Hyphenated
Equipment Name: University of Minnesota Nano Fabrication Center Coral Name: hs-scope Revision Number: 1.5 Model: HS200A Revisionist: M. Fisher Location: Bay 1 Date: 9/12/2013 1 Description The Hyphenated
