Parallel Characteristics of Sequence Alignment Algorithms
|
|
|
- Posy Cameron
- 5 years ago
- Views:
Transcription
1 Parallel Characteristics of Sequence Alignment Algorithms Arun I<. Iyengar Laboratory for Computer Science Massachusetts Institute of Technology Cambridge, MA Abstract Parallel algorithms for analyzing DNA and protein sequences are becoming increasingly important as sequence data continues to grow. This paper examines the parallel characteristics of four sequence alignment algorithms. The four algorithms presented are the dynamic programming algorithm developed by Needleman, Wunsch, and Sellers (the NWS algorithm), Fickett s algorithm, a parallel algorithm using some of Fickett s ideas, and an algorithm which uses some of Wilbur and Lipman s ideas for constructing alignments which are not always optimal. The NWS algorithm contains the most parallelism but also does more work than any of the other algorithms which we studied. Fickett s algorithm contains the least parallelism. However, a parallel algorithm which requires significantly fewer instructions than the NWS algorithm is obtained by modifying Fickett s algorithm. The algorithms have been implemented for a dataflow computer in the dataflow language Id. 1 Introduction This paper analyzes the parallel characteristics of four algorithms for aligning DNA and protein sequences. Biological sequence data is accumulating very rapidly. Increasingly powerful computers will be needed for analyzing DNA and proteins as databases expand. All living things transmit genetic information through DNA. Important structural and functional characteristics can be, determined from an organism s DNA. Biological sequence data provides a very powerful tool for analyzing evolutionary relationships. Many biologists want to determine the DNA sequence of the Permission to copy without fee all or part of this material is granted provided that the copies are not made or distributed for direct commercial advantage, the ACM copyright notice and the title of the publication and its date appear, and notice is given that copying is by permission of the Association for Computing Machinery. To copy otherwise, or to republish, requires a fee and/or specific permission ACM /89/001 l/o304 $1.50 entire human genome. As more information becomes available, it will become poss%le to determine important characteristics of a human being by DNA sequence analysis. The four aigorithms which we analyzed are the dynamic programming algorithm developed by Needleman, [Needleman 701, W unsch, and Sellers [Sellers 741, Fickett s algorithm [Fickett 841, a parallel algorithm similar to Fickett s algorithm, and an algorithm which is similar to Wilbur and Lipman s algorithm [Wilbur 831. A detailed presentation of these algorithms is given in [Iyengar Sequence Alignment Sequence comparison algorithms are used in several different fields including molecular biology, string editing, speech processing, and codes and error control [Kruskal 831. H owever, the algorithms presented in this paper are specifically intended for comparing biological sequences, which include DNA sequences and proteins. From a purely abstract point of view, a DNA sequence is a string defined over an alphabet consisting of four letters. A protein sequence is a string defined over an alphabet consisting of twenty-one letters. These definitions are sufficient to understand the four algorithms. The following presentation assumes no prior knowledge of biology. Of course, the reader with a strong biological background will have much more insight into the motivation behind the algorithms. An alignme?lt between two sequences defined over an alphabet C is a matrix consisting of two rows. The upper row contains the source sequence 5 1 possibly interspersed with null characters. The bottom row contains the target sequence which may also be interspersed with null characters. A null character is represented by a -. A column consisting of two null characters is not allowed. Let 2 ) y E
2 A column is a deeelion. A column is an insertion. A column 2 - ll I i-1 Y X i Y 1 is an identity if x = y; it is a substitution otherwise. A gap of length k is a series of k consecutive insertions or deletions. DNA sequences are defined over the alphabet c = {A,C,G,T}. For example, one possible alignment betweeen the sequences SI = CATGCATA and is Al : CAT-GCATA CATTGAA-A. S, = CATTGAAA A1 contains two gaps of length one, one substitution, and six identities. Two sequences are homologous if they are very similar and the degree of similarity is much higher than what would be expected by chance. 3 Sequence Alignment Algorit hms 3.1 Algorithm 1: The NWS algorithm Needleman and Wunsch [Needleman 701 were two of the first people to use computers for comparing biological sequences. Their algorithm calculates an alignment which maximizes the similarity between two sequences. The dynamic programming algorithm of Sellers calculates an alignment which minimizes the difference between the two sequences. The two approaches calculate the same alignment if parameters are selected appropriately [Smith 811. W e will henceforth refer to the NWS algorithm. We caa assign a difference score d to each alignment: i=l where s is the number of substitutions, n is the number of gaps, gsi is the size of gap i, and gp is a gap penalty function assigning positive values to all gap sizes. We will assume that gap penalties grow linearly with gap sizes. Thus, gp(gs) = gs * gap-penalty where gap-pen&y > 1. An optimal alignment with respect to a difference score is an alignment which possesses the lowest difference score. The NWS algorithm calculates an optimal alignment between Sr and S2 by memoizing optimal alignments between all prefixes of Sr and SZ. Two matrices may be allocated for storing alignments between prefixes and their difference scores. score-matriz[a, b] contains the difference score of an optimal alignment between the first a elements of Sr and the first b elements of Sa, path,matrix[a, b] contains the index of the previous cell in the alignment. The alignment is constructed by following the indices stored in path-matrix. The 0th row corresponds to alignments possessing gaps at the beginning of Sr. Thus, score-matrix[o, a] = gp(a). Similarly, the 0th column corresponds to alignments possessing gaps at the beginning of Ss. Therefore, score-matrix[b, 0] = gp(b). The remainder of the matrix is calculated inductively using the formula: score-matrix[i, j] = min(score-matrix[i - 1, j] +gap-penalty, score,matrix[i, j - l] +gap-penalty, score-matrix[i - 1, j - l] +distance(si [i], SZ [j])) where distance(x,y) = 0 if x = y = 1 if 2 # y. The indices of the previous cell are stored in path-nzatriz[i,j]. Aft er the matrices have been completely determined, an optimal alignment between the 305
3 1 2 Sl Figure 1: This figure illustrates how alignment matrices are calculated in a parallel implementation of the NWS algorithm. All cells belonging to the same diagonal wavefront may be determined concurrently once the previous wavefront has been calculated. entire sequences is constructed by following indices Fickett s algorithm calculates alignment matrices sestarting at path-.matriz[is~ (, ISzl]. The time and space quentially from lower to higher rows. If cells 0 through complexity are b)oth O(IS1( * ISzl). h- of row i- 1 possess difference scores greater than d,,,, cells 0 through k of row i do not have to be calculated. The data dependencies in the NWS algorithm de- Similarly, if cells j through IS21 of row i - 1 and cell fine diagonal wavefronts across the alignment matrices. [i, j] of score-matrix are greater than d,,,, cells j + 1 Cells lying on t:he same wavefront may be calculated through ISzJ o f row i do not have to be determined. concurrently once the previous wavefront has been determined (figure 1). An ideal parallel implementation It is frequently necessary to determine the value of runs in time 0( (S1 I + IS,l) using min( IS1 (, \&I) proces- a matrix cell, even though the value of one or two adsors [Edmiston 87]. jacent cells which define its value are unknown. For example, in figure 2, cell [i, lc + l] is defined by cells [i,b], [i-l,h], and [i-l,ic+l]. Cell [i-l,ic+l] is the 3.2 Algorithm 2: Fickett s algorithn~ only one of the three which has been calculated. Since an optimal alignment cannot pass through cells [i, k] or [i - 1, k], these cells are ignored. Thus, Fickett s algorithm requires the user to provide an upper bound cz,,,~ for the difference score of an optimal alignment. The algorithm assumes that the difference score of an optimal alignment does not exceed d maz. Under this assumption, an optimal alignment cannot pass through matrix cells which possess difference scores grea,ter than d,,,. The key to the algorithm is to keep track of a group of contiguous cells g at the beginning and end of a row which contain difference scores greater than d,,,. An optimal alignment cannot pass through any cells in g. g consists of cells containing X s in figure 2. Any alignment passing through a cell containing a circle must also pass through g. Therefore, cells containing circles do not have to be calculated. score-matrix[i, k + l] = score-matrix[i - 1, k + l]+ gap-penalty. If the score of an optimal alignment exceeds d,,,, Fickett s algorithm terminates without returning an alignment. This is useful for comparing several sequences at a time and identifying pairs which are homologous. Fickett s algorithm will spend less time comparing sequences which are not homologous than the NWS algorithm because it will terminate after a row is found in which the difference scores of all cells exceed d naz- Fickett s algorithm can be improved slightly. If score,matrix[i,j] > d,,,, 306
4 k j i-l i Sl Figure 2: Cells at the beginning and end of rows are pruned by Fickett s algorithm. If the cells containing X s have been eliminated, the cells containing circles can also be eliminated. Fickett s algorithm concludes that an optimal alignment cannot pass through cell [i, j]. score-matriz[i, j] is a lower bound on the difference score for an optimal alignment passing through cell [it j]. If a greater lower bound l;,j can be calculated, more matrix cells may be ignored. a1 = J&J - i elements of & and a2 = IS21 - J- elements of 5 2 have not been added to the alignment at the time cell [i, j] is determined. We may calculate li,j by assuming that the remaining elements will constitute min(al, ~42) identities and Ial - u2 ] insertions or deletions. Thus, li,j = scorenmtriz[i, j] + [Ul - Utl * gup~penazty. In the improved version of Fickett s algorithm, we conclude that an optimal alignment cannot pass through cell [i, j] if li,j > d,,,. Our implementation uses the improved version of Fickett s algorithm. 3.3 A parallel algorithm similar to Fickett s algorithm Fickett s algorithm calculates alignment matrices sequentially. It usually requires fewer total instructions than the NWS algorithm. However, Fickett s algorithm contains far less parallelism. A parallel algorithm which only calculates part of the alignment matrices is obtained by filling in the matrices along wavefronts instead of rows (figure 1). As each wavefront is calculated, we keep track of contiguous blocks of cells at the edges of the two previous wavefronts which can be ignored because their difference scores are too high. Let wavefront Ic consist of all matrix cells whose row and column indices sum to k. If all cells along wavefronts a - 1 and k - 2 with row numbers greater than or equal to j possess difference scores which are two high, we do not have to calculate any matrix cells on wavefront L with row numbers exceeding j. Similarly, if all cells along wavefronts h - 1 with row numbers less than i and k - 2 with row numbers less than i - 1 possess difference scores which are too big, we do not have to calculate any cells on wavefront k with row numbers less than i (figure 3). 3.4 A fast algorithm which does not always find an optimal aligmnent Fast alignment algorithms exist which do not always find optimal alignments. We have implemented an algorithm which locates the matrix region containing the largest number of exact k-tuple matches between Si and 5 2 and only calculates cells in this vicinity. A k-tuple match between 5 1 and S2 is a sequence of k elements which occurs in both 5 1 and 5 2. Cells corresponding to exact k-tuple matches between S1 and Sz form diagonal line segments across the align- 307
5 k-2 k-l k L Figure 3: If the! cells on wavefronts k - 1 and k - 2 containing X s possess difference scores which are too high, the cells on wavefront k containing circles do not have to be calculated. This strategy is used by algorithm 3. -I 1 s2 Figure 4: Cells corresponding to k-tuple matches form diagonal line segments across alignment matrices. Algorithm 4 only calculates matrix cells in the vicinity of the diagonals containing the most k-tuple matches. 308
6 ment matrix (figure 4). The best alignment is likely to pass through the region of the matris containing the most k-tuple matches. Our algorithm locates a group of contiguous diagonals 9 containing the most lc-tuple matches. It then uses dynamic programming to calculate the best alignment which only passes through diagonals which are no farther than wz diagonals from the middle of g. wz is supplied by the user. The idea of locating k-tuple matches to quickly calculate alignments which are not always optimal was suggested by Wilbur and Lipman [Wilbur 831. I<-tuple matches between S1 and Sz may be located by utilizing a hash table of size ]Ck] (where ICI is the size of the alphabet over which S1 and Sz are defined). Each of the ICI different k-tuples is assigned to a different bucket of the hash table. The index identifying the position of each k-tuple belonging to Sr is stored in the appropriate hash table bucket. After this has occurred, k-tuples belonging to Sz are assigned to buckets in the hash table. If a k-tuple in Sz is assigned to a bucket containing an index into 5 1, a match has been found. 4 Results and Discussion The four algorithms were implemented in the dataflow language Id. The parallelism in Id is implicit. This means that the programmer does not have to insert any explicit parallel constructs in order to write a parallel program. Concurrency is detected automatically by the compiler. This makes Id very easy to use. Programming in Id is no more difficult than programming in a sequential language. We are currently in the process of building a dataflow computer which executes Id programs. The results presented in this paper were produced by a dataflow simulator. Our simulator is not powerful enough to align two sequences with lengths significantly longer than 100. Table 1 displays the statistics produced by the four algorithms. The same set of sequences were used to test each algorithm. In every case, algorithms 2, 3, and 4 constructed alignments using fewer instructions than algorithm 1. However, algorithm 1 invariably possessed a shorter critical path length. Furthermore, algorithm 1 is the easiest of the four to implement. Algorithm 1 is a good choice if enough processors are available because it contains the most pa.rallelism. However, it may be possible to efficiently utilize the processors on a parallel machine by constructing many alignments concurrently. In this situation, each alignment does not have to he constructed by a parallel algorithm. Any algorithm which requires fewer instructions than the NWS algorithm will produce a faster solution, even if a single iteration of the algorithm does not contain much parallelism. Algorithms 2 and 3 were tested using 3 different values for cl,,,. d,,, is an upper bound on the difference score which is supplied by the user. The instruction counts for both algorithms increase with d,,,. Maximum efficiency is obtained by making d,,, as small as possible. However, both algorithms will terminate without returning an alignment if the difference score of an optimal alignment exceeds dmar. Algorithm 3 contains significantly more parallelism than algorithm 2. This becomes more noticeable when longer sequences are compared with higher values specified for d,,,. For example, when sequences of length 15 are aligned and d,,, = G, the ratio of the critical path length of algorithm 3 to that of algorithm 2 is.678. By contrast, this ratio becomes.299 when sequences of length 100 are aligned and d,,, = 40. Algorithm 4 was tested using two different sets of values for 2~2. A trade-off exists between the instruction count and the quality of alignments constructed by the algorithm. If wz is small, a small number of matrix cells are calculated. As w2 increases, more matrix cells are calculated, and the instruction count increases. However, the probability of finding an optimal aiignment also increases. Unlike the instruction count, the critical path length does not change much with wz. The lower instruction counts algorithms 2, 3, and 4 possess relative to algorithm 1 become more apparent when longer sequences are compared. The differences are not significant for sequences of lengths 15 or less. When longer sequences are compared, algorithms 2, 3, and 4 require significantly fewer instructions than algorithm 1. This effect would become more pronounced if we were able to compare sequences significantly longer than Summary and Conclusions None of the algorithms which we studied is unequivocally better than the others. They all have strengths and weaknesses. The correct algorithm to use clearly depends upon the nature of the application. The NWS algorithm contains the most parallelism. It is also the easiest algorithm to implement. However, it requires many instructions. Fickett s algorithm does not contain much parallelism. However, it can align homologous sequences using much fewer instructions than the NWS algorithm. 309
7 7 Algorithm 1 Algorithm 2 d max = 3 d ma2-4 d mcm -5 d maz -8- Algorithm 2 d mar -6 d max -8 d mar = 10 d mar = 20 Algorithm 2 d maz -9 d mox = 16 d maz = 20 d mas = 40 Algorithm 3 d max -3 d maz -4- d mar -5 d maa: =8 Algorithm 3 d mas -6 d maa -8 d ma2 = 10 d mar = 20 Algorithm 3 d mm =9 --l d mar mm = ,010, ,239 96,604 26, % 2,473 4,873 1, % 15 13, % 2, % 30 26, % 4, % 50 55, % 9, % , % 21, % 15 16, % 2, % 30 39, % 6, % 50 77, % 11, % , % 38, % 15 21, % 3, % 30 62, % 9, % , % 18, % , % 63, % 15 17, % 1, % 30 34, % 3, % 50 68, % 5, % , % 12, % 15 20, % 1, % 30 46, % 3, % 50 88, % 6, % , % 15, % 15 25, % 1, % , ,560 68, % 70.76% 52.68% 19,024 4,208 7, % 390.4% 278.1% 15 22, % 1, % 30 49, % 3, % 50 86, % 5, % , % 10, % 15 26, % 1, % 30 60, % 2, % , % 5, % , % 10, % Table 1: Simulation statistics for the four alignment algorithms. The critical path length is the longest chain of operations which must be executed sequentially in any parallel implementation. 310
8 Figure 5: The parallelism profile which results when algorithm 1 aligns two homologous sequences of length 30. The instruction count and critical path length are 96,604 and 1,513 respectively. The sequential tail beginning just before 1000 on the X-axis results from constructing the alignment after the matrices have been determined. 1fJoo ow.lllls,000 Figure G: The parallelism profile which results when algorithm 2 aligns two homologous sequences of length 30 and d,,,.,, = 16. The instruction count and critical path length are 62,321 and 9,306 respectively. 311
9 1 Figure 7: The parallelism profile which results when algorithm 3 aligns two homologous sequences of length 3C and d,,, = 16. The instruction count and critical path length are 68,360 and 4,208 respec.tively. 1 Figure 8: The parallelism profile which results when algorithm 4 aligns two homologous sequences of length 3C and 202 = 4. The instruction count and critical path length are 60,032 and 2,994 respectively. k-tuple matches art located using hash tables from 0 to about 3S0 time steps. The matrix region containing the most k-tuple matche: is located from 380 to about 1080 time steps. Matrix cells surrounding this region are calculated by dyna.mic programming from 1080 to about 2100 time steps, The alignment is constructed from the matrices from 2100 tc 2,994 time steps. 312
10 It is a good algorithm to use for searching through a database of sequences and locating pairs which are homologous. Parallelism may be obtained by aligning several pairs of sequences at the same time even though a single iteration is sequential. The third algorithm which we implemented uses some of Fickett s ideas. It contains significantly more parallelism than Fickett s algorithm. Furthermore, it requires about the same number of instructions. Both algorithms will not return an alignment if the sequences differ by too much. The fourth algorithm we implemented does not always find an optimal alignment. However, it finds a good alignment in the vast majority of cases. Unlike algorithms 2 and 3, it always returns an alignment. Furthermore, it requires fewer instructions than the NWS algorithm and contains more parallelism than Fickett s algorithm. All four algorithms were encoded in the dataflow language Id. The parallelism in Id is implicit. This greatly simplifies the programmer s job. Explicitly parallelizing the alignment algorithms is a fairly significant task. The NWS algorithm is the easiest of the four algorithms to parallelize. However, implementing the NWS algorithm efficiently on a SIMD computer such as a Connection Machine requires a lot of work [Lander 881. The other algorithms are even harder to parallelize explicitly because of their added complexity. An Id programmer does not encounter any difficulties which a sequential programmer would not also face. He is totally insulated from the characteristics of the machine executing his program. [Fickett 841 [Iyengar 881 [Kruskal 831 [Lander 881 [Needleman 701 Needleman, S. B., and C. D. Wunsch. A General Method Applicable to the Search for Similarities in the Amino Acid Sequences of Two Proteins. In Journal of Molecular Biology , pp [Nikhil 881 Fickett, J. W. Fast Optimal Alignment. In Nucleic Acids Research 18:l. 1984, pp Iyengar, A. K. Parallel DNA Sequence Analysis. Master s Thesis, Technical Report TR-428, Laboratory for Computer Science, Massachusetts Institute of Technology. Cambridge, MA 02139, Kruskal, J. B. An Overview of Sequence Comparison, Time Warps, String Edits, and Macromolecules. In Siam Review 25:2. April 1983, pp Lander, E., Mesirov, J., and Taylor, W. Protein Sequence Comparison on a Data Parallel Computer. In Proceedings of the 1988 International Conference on Parallel Processing, Volume 3. pp , Penn State Press, Pennsylvania. Nikhil, R. S. Id Version 88.1 Reference Manual. CSG Memo 284, M. I. T. Laboratory for Computer Science, August 29, Acknowledgements: The simulation tools for generating the statistics presented in this paper were developed by the Computation Structures Group at MIT. Rishiyur S. Nikhil made many useful suggestions which improved the quality of this research. The author has been supported in part by a graduate fellowship from the National Science Foundation. References [Edmiston 871 Edmiston, E., and Wagner, R. A. Parallelization of the Dynamic Programming Algorithm for Comparison of Sequences. In Proceedings of Ihe 1987 International Conference on Parallel Processing. pp , Penn State Press, Pennsylvania. [Sellers 741 [Smith 811 [Wilbur 831 Sellers, P. H. On the Theory and Computation of Evolutionary Distances. SIAM J. Appl. Math. 26:4. June 1974, pp Smith, T. F., et. al. Comparative Biosequence Metrics. Journal of Molecular Evolution , pp Wilbur, W. J., and D. J. Lipman. Rapid Similarity Searches of Nucleic Acid and Protein Data Banks, Proc. Natl. Acad. Sci. USA 80. February 1983, pp
Multiple Sequence Alignment: Multidimensional. Biological Motivation
 Multiple Sequence Alignment: Multidimensional Dynamic Programming Boston University Biological Motivation Compare a new sequence with the sequences in a protein family. Proteins can be categorized into
Multiple Sequence Alignment: Multidimensional Dynamic Programming Boston University Biological Motivation Compare a new sequence with the sequences in a protein family. Proteins can be categorized into
) I R L Press Limited, Oxford, England. The protein identification resource (PIR)
 Volume 14 Number 1 Volume 1986 Nucleic Acids Research 14 Number 1986 Nucleic Acids Research The protein identification resource (PIR) David G.George, Winona C.Barker and Lois T.Hunt National Biomedical
Volume 14 Number 1 Volume 1986 Nucleic Acids Research 14 Number 1986 Nucleic Acids Research The protein identification resource (PIR) David G.George, Winona C.Barker and Lois T.Hunt National Biomedical
Bioinformatics explained: Smith-Waterman
 Bioinformatics Explained Bioinformatics explained: Smith-Waterman May 1, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com
Bioinformatics Explained Bioinformatics explained: Smith-Waterman May 1, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com
FastA & the chaining problem
 FastA & the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 1 Sources for this lecture: Lectures by Volker Heun, Daniel Huson and Knut Reinert,
FastA & the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 1 Sources for this lecture: Lectures by Volker Heun, Daniel Huson and Knut Reinert,
FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:
 FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:56 4001 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem
FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:56 4001 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem
An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST
 An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST Alexander Chan 5075504 Biochemistry 218 Final Project An Analysis of Pairwise
An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST Alexander Chan 5075504 Biochemistry 218 Final Project An Analysis of Pairwise
Lectures by Volker Heun, Daniel Huson and Knut Reinert, in particular last years lectures
 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 4.1 Sources for this lecture Lectures by Volker Heun, Daniel Huson and Knut
4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 4.1 Sources for this lecture Lectures by Volker Heun, Daniel Huson and Knut
Computational Molecular Biology
 Computational Molecular Biology Erwin M. Bakker Lecture 3, mainly from material by R. Shamir [2] and H.J. Hoogeboom [4]. 1 Pairwise Sequence Alignment Biological Motivation Algorithmic Aspect Recursive
Computational Molecular Biology Erwin M. Bakker Lecture 3, mainly from material by R. Shamir [2] and H.J. Hoogeboom [4]. 1 Pairwise Sequence Alignment Biological Motivation Algorithmic Aspect Recursive
Computational Genomics and Molecular Biology, Fall
 Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA.
 Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA. Fasta is used to compare a protein or DNA sequence to all of the
Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA. Fasta is used to compare a protein or DNA sequence to all of the
BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio CS 466 Saurabh Sinha
 BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio. 1990. CS 466 Saurabh Sinha Motivation Sequence homology to a known protein suggest function of newly sequenced protein Bioinformatics
BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio. 1990. CS 466 Saurabh Sinha Motivation Sequence homology to a known protein suggest function of newly sequenced protein Bioinformatics
Sequence alignment theory and applications Session 3: BLAST algorithm
 Sequence alignment theory and applications Session 3: BLAST algorithm Introduction to Bioinformatics online course : IBT Sonal Henson Learning Objectives Understand the principles of the BLAST algorithm
Sequence alignment theory and applications Session 3: BLAST algorithm Introduction to Bioinformatics online course : IBT Sonal Henson Learning Objectives Understand the principles of the BLAST algorithm
CISC 889 Bioinformatics (Spring 2003) Multiple Sequence Alignment
 CISC 889 Bioinformatics (Spring 2003) Multiple Sequence Alignment Courtesy of jalview 1 Motivations Collective statistic Protein families Identification and representation of conserved sequence features
CISC 889 Bioinformatics (Spring 2003) Multiple Sequence Alignment Courtesy of jalview 1 Motivations Collective statistic Protein families Identification and representation of conserved sequence features
COS 551: Introduction to Computational Molecular Biology Lecture: Oct 17, 2000 Lecturer: Mona Singh Scribe: Jacob Brenner 1. Database Searching
 COS 551: Introduction to Computational Molecular Biology Lecture: Oct 17, 2000 Lecturer: Mona Singh Scribe: Jacob Brenner 1 Database Searching In database search, we typically have a large sequence database
COS 551: Introduction to Computational Molecular Biology Lecture: Oct 17, 2000 Lecturer: Mona Singh Scribe: Jacob Brenner 1 Database Searching In database search, we typically have a large sequence database
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014
 Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
An Efficient Algorithm to Locate All Locally Optimal Alignments Between Two Sequences Allowing for Gaps
 An Efficient Algorithm to Locate All Locally Optimal Alignments Between Two Sequences Allowing for Gaps Geoffrey J. Barton Laboratory of Molecular Biophysics University of Oxford Rex Richards Building
An Efficient Algorithm to Locate All Locally Optimal Alignments Between Two Sequences Allowing for Gaps Geoffrey J. Barton Laboratory of Molecular Biophysics University of Oxford Rex Richards Building
Research on Pairwise Sequence Alignment Needleman-Wunsch Algorithm
 5th International Conference on Mechatronics, Materials, Chemistry and Computer Engineering (ICMMCCE 2017) Research on Pairwise Sequence Alignment Needleman-Wunsch Algorithm Xiantao Jiang1, a,*,xueliang
5th International Conference on Mechatronics, Materials, Chemistry and Computer Engineering (ICMMCCE 2017) Research on Pairwise Sequence Alignment Needleman-Wunsch Algorithm Xiantao Jiang1, a,*,xueliang
Notes on Dynamic-Programming Sequence Alignment
 Notes on Dynamic-Programming Sequence Alignment Introduction. Following its introduction by Needleman and Wunsch (1970), dynamic programming has become the method of choice for rigorous alignment of DNA
Notes on Dynamic-Programming Sequence Alignment Introduction. Following its introduction by Needleman and Wunsch (1970), dynamic programming has become the method of choice for rigorous alignment of DNA
On the Efficacy of Haskell for High Performance Computational Biology
 On the Efficacy of Haskell for High Performance Computational Biology Jacqueline Addesa Academic Advisors: Jeremy Archuleta, Wu chun Feng 1. Problem and Motivation Biologists can leverage the power of
On the Efficacy of Haskell for High Performance Computational Biology Jacqueline Addesa Academic Advisors: Jeremy Archuleta, Wu chun Feng 1. Problem and Motivation Biologists can leverage the power of
FASTA. Besides that, FASTA package provides SSEARCH, an implementation of the optimal Smith- Waterman algorithm.
 FASTA INTRODUCTION Definition (by David J. Lipman and William R. Pearson in 1985) - Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence
FASTA INTRODUCTION Definition (by David J. Lipman and William R. Pearson in 1985) - Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence
PROTEIN MULTIPLE ALIGNMENT MOTIVATION: BACKGROUND: Marina Sirota
 Marina Sirota MOTIVATION: PROTEIN MULTIPLE ALIGNMENT To study evolution on the genetic level across a wide range of organisms, biologists need accurate tools for multiple sequence alignment of protein
Marina Sirota MOTIVATION: PROTEIN MULTIPLE ALIGNMENT To study evolution on the genetic level across a wide range of organisms, biologists need accurate tools for multiple sequence alignment of protein
Computational Molecular Biology
 Computational Molecular Biology Erwin M. Bakker Lecture 2 Materials used from R. Shamir [2] and H.J. Hoogeboom [4]. 1 Molecular Biology Sequences DNA A, T, C, G RNA A, U, C, G Protein A, R, D, N, C E,
Computational Molecular Biology Erwin M. Bakker Lecture 2 Materials used from R. Shamir [2] and H.J. Hoogeboom [4]. 1 Molecular Biology Sequences DNA A, T, C, G RNA A, U, C, G Protein A, R, D, N, C E,
Important Example: Gene Sequence Matching. Corrigiendum. Central Dogma of Modern Biology. Genetics. How Nucleotides code for Amino Acids
 Important Example: Gene Sequence Matching Century of Biology Two views of computer science s relationship to biology: Bioinformatics: computational methods to help discover new biology from lots of data
Important Example: Gene Sequence Matching Century of Biology Two views of computer science s relationship to biology: Bioinformatics: computational methods to help discover new biology from lots of data
Central Issues in Biological Sequence Comparison
 Central Issues in Biological Sequence Comparison Definitions: What is one trying to find or optimize? Algorithms: Can one find the proposed object optimally or in reasonable time optimize? Statistics:
Central Issues in Biological Sequence Comparison Definitions: What is one trying to find or optimize? Algorithms: Can one find the proposed object optimally or in reasonable time optimize? Statistics:
Profiles and Multiple Alignments. COMP 571 Luay Nakhleh, Rice University
 Profiles and Multiple Alignments COMP 571 Luay Nakhleh, Rice University Outline Profiles and sequence logos Profile hidden Markov models Aligning profiles Multiple sequence alignment by gradual sequence
Profiles and Multiple Alignments COMP 571 Luay Nakhleh, Rice University Outline Profiles and sequence logos Profile hidden Markov models Aligning profiles Multiple sequence alignment by gradual sequence
24 Grundlagen der Bioinformatik, SS 10, D. Huson, April 26, This lecture is based on the following papers, which are all recommended reading:
 24 Grundlagen der Bioinformatik, SS 10, D. Huson, April 26, 2010 3 BLAST and FASTA This lecture is based on the following papers, which are all recommended reading: D.J. Lipman and W.R. Pearson, Rapid
24 Grundlagen der Bioinformatik, SS 10, D. Huson, April 26, 2010 3 BLAST and FASTA This lecture is based on the following papers, which are all recommended reading: D.J. Lipman and W.R. Pearson, Rapid
As of August 15, 2008, GenBank contained bases from reported sequences. The search procedure should be
 48 Bioinformatics I, WS 09-10, S. Henz (script by D. Huson) November 26, 2009 4 BLAST and BLAT Outline of the chapter: 1. Heuristics for the pairwise local alignment of two sequences 2. BLAST: search and
48 Bioinformatics I, WS 09-10, S. Henz (script by D. Huson) November 26, 2009 4 BLAST and BLAT Outline of the chapter: 1. Heuristics for the pairwise local alignment of two sequences 2. BLAST: search and
Comparison of Sequence Similarity Measures for Distant Evolutionary Relationships
 Comparison of Sequence Similarity Measures for Distant Evolutionary Relationships Abhishek Majumdar, Peter Z. Revesz Department of Computer Science and Engineering, University of Nebraska-Lincoln, Lincoln,
Comparison of Sequence Similarity Measures for Distant Evolutionary Relationships Abhishek Majumdar, Peter Z. Revesz Department of Computer Science and Engineering, University of Nebraska-Lincoln, Lincoln,
Lecture 3: February Local Alignment: The Smith-Waterman Algorithm
 CSCI1820: Sequence Alignment Spring 2017 Lecture 3: February 7 Lecturer: Sorin Istrail Scribe: Pranavan Chanthrakumar Note: LaTeX template courtesy of UC Berkeley EECS dept. Notes are also adapted from
CSCI1820: Sequence Alignment Spring 2017 Lecture 3: February 7 Lecturer: Sorin Istrail Scribe: Pranavan Chanthrakumar Note: LaTeX template courtesy of UC Berkeley EECS dept. Notes are also adapted from
Pairwise Sequence Alignment: Dynamic Programming Algorithms. COMP Spring 2015 Luay Nakhleh, Rice University
 Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 - Spring 2015 Luay Nakhleh, Rice University DP Algorithms for Pairwise Alignment The number of all possible pairwise alignments (if
Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 - Spring 2015 Luay Nakhleh, Rice University DP Algorithms for Pairwise Alignment The number of all possible pairwise alignments (if
Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 Luay Nakhleh, Rice University
 1 Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 Luay Nakhleh, Rice University DP Algorithms for Pairwise Alignment 2 The number of all possible pairwise alignments (if gaps are allowed)
1 Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 Luay Nakhleh, Rice University DP Algorithms for Pairwise Alignment 2 The number of all possible pairwise alignments (if gaps are allowed)
Bioinformatics for Biologists
 Bioinformatics for Biologists Sequence Analysis: Part I. Pairwise alignment and database searching Fran Lewitter, Ph.D. Director Bioinformatics & Research Computing Whitehead Institute Topics to Cover
Bioinformatics for Biologists Sequence Analysis: Part I. Pairwise alignment and database searching Fran Lewitter, Ph.D. Director Bioinformatics & Research Computing Whitehead Institute Topics to Cover
Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it.
 Sequence Alignments Overview Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it. Sequence alignment means arranging
Sequence Alignments Overview Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it. Sequence alignment means arranging
Lecture 9: Core String Edits and Alignments
 Biosequence Algorithms, Spring 2005 Lecture 9: Core String Edits and Alignments Pekka Kilpeläinen University of Kuopio Department of Computer Science BSA Lecture 9: String Edits and Alignments p.1/30 III:
Biosequence Algorithms, Spring 2005 Lecture 9: Core String Edits and Alignments Pekka Kilpeläinen University of Kuopio Department of Computer Science BSA Lecture 9: String Edits and Alignments p.1/30 III:
Lecture Overview. Sequence search & alignment. Searching sequence databases. Sequence Alignment & Search. Goals: Motivations:
 Lecture Overview Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating
Lecture Overview Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating
Sequence Alignment & Search
 Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating the first version
Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating the first version
Multiple Sequence Alignment Using Reconfigurable Computing
 Multiple Sequence Alignment Using Reconfigurable Computing Carlos R. Erig Lima, Heitor S. Lopes, Maiko R. Moroz, and Ramon M. Menezes Bioinformatics Laboratory, Federal University of Technology Paraná
Multiple Sequence Alignment Using Reconfigurable Computing Carlos R. Erig Lima, Heitor S. Lopes, Maiko R. Moroz, and Ramon M. Menezes Bioinformatics Laboratory, Federal University of Technology Paraná
Data Mining Technologies for Bioinformatics Sequences
 Data Mining Technologies for Bioinformatics Sequences Deepak Garg Computer Science and Engineering Department Thapar Institute of Engineering & Tecnology, Patiala Abstract Main tool used for sequence alignment
Data Mining Technologies for Bioinformatics Sequences Deepak Garg Computer Science and Engineering Department Thapar Institute of Engineering & Tecnology, Patiala Abstract Main tool used for sequence alignment
A CAM(Content Addressable Memory)-based architecture for molecular sequence matching
 A CAM(Content Addressable Memory)-based architecture for molecular sequence matching P.K. Lala 1 and J.P. Parkerson 2 1 Department Electrical Engineering, Texas A&M University, Texarkana, Texas, USA 2
A CAM(Content Addressable Memory)-based architecture for molecular sequence matching P.K. Lala 1 and J.P. Parkerson 2 1 Department Electrical Engineering, Texas A&M University, Texarkana, Texas, USA 2
Fast Sequence Alignment Method Using CUDA-enabled GPU
 Fast Sequence Alignment Method Using CUDA-enabled GPU Yeim-Kuan Chang Department of Computer Science and Information Engineering National Cheng Kung University Tainan, Taiwan ykchang@mail.ncku.edu.tw De-Yu
Fast Sequence Alignment Method Using CUDA-enabled GPU Yeim-Kuan Chang Department of Computer Science and Information Engineering National Cheng Kung University Tainan, Taiwan ykchang@mail.ncku.edu.tw De-Yu
Acceleration of Algorithm of Smith-Waterman Using Recursive Variable Expansion.
 www.ijarcet.org 54 Acceleration of Algorithm of Smith-Waterman Using Recursive Variable Expansion. Hassan Kehinde Bello and Kazeem Alagbe Gbolagade Abstract Biological sequence alignment is becoming popular
www.ijarcet.org 54 Acceleration of Algorithm of Smith-Waterman Using Recursive Variable Expansion. Hassan Kehinde Bello and Kazeem Alagbe Gbolagade Abstract Biological sequence alignment is becoming popular
bl,...,b, where i runs from 0 to m and j runs from 0 to n. Then with initial values
 Volume 12 Number 1 1984 Nucleic Acids Research Fast optinal alignment James W.Fickett Theoretical Division, Los Alamos National Laboratory, University of Califormia, Los Alamos, NM 87545, USA Received
Volume 12 Number 1 1984 Nucleic Acids Research Fast optinal alignment James W.Fickett Theoretical Division, Los Alamos National Laboratory, University of Califormia, Los Alamos, NM 87545, USA Received
Space Efficient Linear Time Construction of
 Space Efficient Linear Time Construction of Suffix Arrays Pang Ko and Srinivas Aluru Dept. of Electrical and Computer Engineering 1 Laurence H. Baker Center for Bioinformatics and Biological Statistics
Space Efficient Linear Time Construction of Suffix Arrays Pang Ko and Srinivas Aluru Dept. of Electrical and Computer Engineering 1 Laurence H. Baker Center for Bioinformatics and Biological Statistics
Alignment of Long Sequences
 Alignment of Long Sequences BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2009 Mark Craven craven@biostat.wisc.edu Pairwise Whole Genome Alignment: Task Definition Given a pair of genomes (or other large-scale
Alignment of Long Sequences BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2009 Mark Craven craven@biostat.wisc.edu Pairwise Whole Genome Alignment: Task Definition Given a pair of genomes (or other large-scale
Lecture 10. Sequence alignments
 Lecture 10 Sequence alignments Alignment algorithms: Overview Given a scoring system, we need to have an algorithm for finding an optimal alignment for a pair of sequences. We want to maximize the score
Lecture 10 Sequence alignments Alignment algorithms: Overview Given a scoring system, we need to have an algorithm for finding an optimal alignment for a pair of sequences. We want to maximize the score
Bioinformatics explained: BLAST. March 8, 2007
 Bioinformatics Explained Bioinformatics explained: BLAST March 8, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com Bioinformatics
Bioinformatics Explained Bioinformatics explained: BLAST March 8, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com Bioinformatics
Special course in Computer Science: Advanced Text Algorithms
 Special course in Computer Science: Advanced Text Algorithms Lecture 6: Alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Special course in Computer Science: Advanced Text Algorithms Lecture 6: Alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Dynamic Programming. Lecture Overview Introduction
 Lecture 12 Dynamic Programming 12.1 Overview Dynamic Programming is a powerful technique that allows one to solve many different types of problems in time O(n 2 ) or O(n 3 ) for which a naive approach
Lecture 12 Dynamic Programming 12.1 Overview Dynamic Programming is a powerful technique that allows one to solve many different types of problems in time O(n 2 ) or O(n 3 ) for which a naive approach
Acceleration of the Smith-Waterman algorithm for DNA sequence alignment using an FPGA platform
 Acceleration of the Smith-Waterman algorithm for DNA sequence alignment using an FPGA platform Barry Strengholt Matthijs Brobbel Delft University of Technology Faculty of Electrical Engineering, Mathematics
Acceleration of the Smith-Waterman algorithm for DNA sequence alignment using an FPGA platform Barry Strengholt Matthijs Brobbel Delft University of Technology Faculty of Electrical Engineering, Mathematics
A Special-Purpose Processor for Gene Sequence Analysis. Barry Fagin* J. GIll Watt** Thayer School of Engineering
 A Special-Purpose Processor for Gene Sequence Analysis Barry Fagin* (barry.fagin@dartmouth.edu) J. GIll Watt** Thayer School of Engineering Robert Gross (bob.gross@dartmouth.edu) Department of Biology
A Special-Purpose Processor for Gene Sequence Analysis Barry Fagin* (barry.fagin@dartmouth.edu) J. GIll Watt** Thayer School of Engineering Robert Gross (bob.gross@dartmouth.edu) Department of Biology
Biology 644: Bioinformatics
 Find the best alignment between 2 sequences with lengths n and m, respectively Best alignment is very dependent upon the substitution matrix and gap penalties The Global Alignment Problem tries to find
Find the best alignment between 2 sequences with lengths n and m, respectively Best alignment is very dependent upon the substitution matrix and gap penalties The Global Alignment Problem tries to find
Special course in Computer Science: Advanced Text Algorithms
 Special course in Computer Science: Advanced Text Algorithms Lecture 8: Multiple alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Special course in Computer Science: Advanced Text Algorithms Lecture 8: Multiple alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Pairwise Sequence Alignment. Zhongming Zhao, PhD
 Pairwise Sequence Alignment Zhongming Zhao, PhD Email: zhongming.zhao@vanderbilt.edu http://bioinfo.mc.vanderbilt.edu/ Sequence Similarity match mismatch A T T A C G C G T A C C A T A T T A T G C G A T
Pairwise Sequence Alignment Zhongming Zhao, PhD Email: zhongming.zhao@vanderbilt.edu http://bioinfo.mc.vanderbilt.edu/ Sequence Similarity match mismatch A T T A C G C G T A C C A T A T T A T G C G A T
Highly Scalable and Accurate Seeds for Subsequence Alignment
 Highly Scalable and Accurate Seeds for Subsequence Alignment Abhijit Pol Tamer Kahveci Department of Computer and Information Science and Engineering, University of Florida, Gainesville, FL, USA, 32611
Highly Scalable and Accurate Seeds for Subsequence Alignment Abhijit Pol Tamer Kahveci Department of Computer and Information Science and Engineering, University of Florida, Gainesville, FL, USA, 32611
DNA Alignment With Affine Gap Penalties
 DNA Alignment With Affine Gap Penalties Laurel Schuster Why Use Affine Gap Penalties? When aligning two DNA sequences, one goal may be to infer the mutations that made them different. Though it s impossible
DNA Alignment With Affine Gap Penalties Laurel Schuster Why Use Affine Gap Penalties? When aligning two DNA sequences, one goal may be to infer the mutations that made them different. Though it s impossible
Distributed Protein Sequence Alignment
 Distributed Protein Sequence Alignment ABSTRACT J. Michael Meehan meehan@wwu.edu James Hearne hearne@wwu.edu Given the explosive growth of biological sequence databases and the computational complexity
Distributed Protein Sequence Alignment ABSTRACT J. Michael Meehan meehan@wwu.edu James Hearne hearne@wwu.edu Given the explosive growth of biological sequence databases and the computational complexity
From Smith-Waterman to BLAST
 From Smith-Waterman to BLAST Jeremy Buhler July 23, 2015 Smith-Waterman is the fundamental tool that we use to decide how similar two sequences are. Isn t that all that BLAST does? In principle, it is
From Smith-Waterman to BLAST Jeremy Buhler July 23, 2015 Smith-Waterman is the fundamental tool that we use to decide how similar two sequences are. Isn t that all that BLAST does? In principle, it is
Computational Biology Lecture 4: Overlap detection, Local Alignment, Space Efficient Needleman-Wunsch Saad Mneimneh
 Computational Biology Lecture 4: Overlap detection, Local Alignment, Space Efficient Needleman-Wunsch Saad Mneimneh Overlap detection: Semi-Global Alignment An overlap of two sequences is considered an
Computational Biology Lecture 4: Overlap detection, Local Alignment, Space Efficient Needleman-Wunsch Saad Mneimneh Overlap detection: Semi-Global Alignment An overlap of two sequences is considered an
Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD
 Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD Assumptions: Biological sequences evolved by evolution. Micro scale changes: For short sequences (e.g. one
Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD Assumptions: Biological sequences evolved by evolution. Micro scale changes: For short sequences (e.g. one
Principles of Bioinformatics. BIO540/STA569/CSI660 Fall 2010
 Principles of Bioinformatics BIO540/STA569/CSI660 Fall 2010 Lecture 11 Multiple Sequence Alignment I Administrivia Administrivia The midterm examination will be Monday, October 18 th, in class. Closed
Principles of Bioinformatics BIO540/STA569/CSI660 Fall 2010 Lecture 11 Multiple Sequence Alignment I Administrivia Administrivia The midterm examination will be Monday, October 18 th, in class. Closed
Global Alignment Scoring Matrices Local Alignment Alignment with Affine Gap Penalties
 Global Alignment Scoring Matrices Local Alignment Alignment with Affine Gap Penalties From LCS to Alignment: Change the Scoring The Longest Common Subsequence (LCS) problem the simplest form of sequence
Global Alignment Scoring Matrices Local Alignment Alignment with Affine Gap Penalties From LCS to Alignment: Change the Scoring The Longest Common Subsequence (LCS) problem the simplest form of sequence
Comparison MICHAEL S. WATERMAN*
 ADVANCBS IN APPLIED MATHEMATICS 6,129-134 (1985) : Dynamic Programming Algorithms for Picture Comparison MICHAEL S. WATERMAN* Departments of Mathematics and Biological Sciences, Univeniry of Southern California.
ADVANCBS IN APPLIED MATHEMATICS 6,129-134 (1985) : Dynamic Programming Algorithms for Picture Comparison MICHAEL S. WATERMAN* Departments of Mathematics and Biological Sciences, Univeniry of Southern California.
.. Fall 2011 CSC 570: Bioinformatics Alexander Dekhtyar..
 .. Fall 2011 CSC 570: Bioinformatics Alexander Dekhtyar.. PAM and BLOSUM Matrices Prepared by: Jason Banich and Chris Hoover Background As DNA sequences change and evolve, certain amino acids are more
.. Fall 2011 CSC 570: Bioinformatics Alexander Dekhtyar.. PAM and BLOSUM Matrices Prepared by: Jason Banich and Chris Hoover Background As DNA sequences change and evolve, certain amino acids are more
BLAST, Profile, and PSI-BLAST
 BLAST, Profile, and PSI-BLAST Jianlin Cheng, PhD School of Electrical Engineering and Computer Science University of Central Florida 26 Free for academic use Copyright @ Jianlin Cheng & original sources
BLAST, Profile, and PSI-BLAST Jianlin Cheng, PhD School of Electrical Engineering and Computer Science University of Central Florida 26 Free for academic use Copyright @ Jianlin Cheng & original sources
Exhaustive Generation and Visual Browsing for Radiation Patterns of Linear Array Antennas
 MITSUBISHI ELECTRIC RESEARCH LABORATORIES http://www.merl.com Exhaustive Generation and Visual Browsing for Radiation Patterns of Linear Array Antennas Darren Leigh, Tom Lanning, Neal Lesh, Kathy Ryall
MITSUBISHI ELECTRIC RESEARCH LABORATORIES http://www.merl.com Exhaustive Generation and Visual Browsing for Radiation Patterns of Linear Array Antennas Darren Leigh, Tom Lanning, Neal Lesh, Kathy Ryall
Cost Partitioning Techniques for Multiple Sequence Alignment. Mirko Riesterer,
 Cost Partitioning Techniques for Multiple Sequence Alignment Mirko Riesterer, 10.09.18 Agenda. 1 Introduction 2 Formal Definition 3 Solving MSA 4 Combining Multiple Pattern Databases 5 Cost Partitioning
Cost Partitioning Techniques for Multiple Sequence Alignment Mirko Riesterer, 10.09.18 Agenda. 1 Introduction 2 Formal Definition 3 Solving MSA 4 Combining Multiple Pattern Databases 5 Cost Partitioning
A FAST LONGEST COMMON SUBSEQUENCE ALGORITHM FOR BIOSEQUENCES ALIGNMENT
 A FAST LONGEST COMMON SUBSEQUENCE ALGORITHM FOR BIOSEQUENCES ALIGNMENT Wei Liu 1,*, Lin Chen 2, 3 1 Institute of Information Science and Technology, Nanjing University of Aeronautics and Astronautics,
A FAST LONGEST COMMON SUBSEQUENCE ALGORITHM FOR BIOSEQUENCES ALIGNMENT Wei Liu 1,*, Lin Chen 2, 3 1 Institute of Information Science and Technology, Nanjing University of Aeronautics and Astronautics,
Software Implementation of Smith-Waterman Algorithm in FPGA
 Software Implementation of Smith-Waterman lgorithm in FP NUR FRH IN SLIMN, NUR DLILH HMD SBRI, SYED BDUL MULIB L JUNID, ZULKIFLI BD MJID, BDUL KRIMI HLIM Faculty of Electrical Engineering Universiti eknologi
Software Implementation of Smith-Waterman lgorithm in FP NUR FRH IN SLIMN, NUR DLILH HMD SBRI, SYED BDUL MULIB L JUNID, ZULKIFLI BD MJID, BDUL KRIMI HLIM Faculty of Electrical Engineering Universiti eknologi
A Study On Pair-Wise Local Alignment Of Protein Sequence For Identifying The Structural Similarity
 A Study On Pair-Wise Local Alignment Of Protein Sequence For Identifying The Structural Similarity G. Pratyusha, Department of Computer Science & Engineering, V.R.Siddhartha Engineering College(Autonomous)
A Study On Pair-Wise Local Alignment Of Protein Sequence For Identifying The Structural Similarity G. Pratyusha, Department of Computer Science & Engineering, V.R.Siddhartha Engineering College(Autonomous)
Research Article International Journals of Advanced Research in Computer Science and Software Engineering ISSN: X (Volume-7, Issue-6)
 International Journals of Advanced Research in Computer Science and Software Engineering ISSN: 77-18X (Volume-7, Issue-6) Research Article June 017 DDGARM: Dotlet Driven Global Alignment with Reduced Matrix
International Journals of Advanced Research in Computer Science and Software Engineering ISSN: 77-18X (Volume-7, Issue-6) Research Article June 017 DDGARM: Dotlet Driven Global Alignment with Reduced Matrix
The Effect of Inverse Document Frequency Weights on Indexed Sequence Retrieval. Kevin C. O'Kane. Department of Computer Science
 The Effect of Inverse Document Frequency Weights on Indexed Sequence Retrieval Kevin C. O'Kane Department of Computer Science The University of Northern Iowa Cedar Falls, Iowa okane@cs.uni.edu http://www.cs.uni.edu/~okane
The Effect of Inverse Document Frequency Weights on Indexed Sequence Retrieval Kevin C. O'Kane Department of Computer Science The University of Northern Iowa Cedar Falls, Iowa okane@cs.uni.edu http://www.cs.uni.edu/~okane
Shortest Path Algorithm
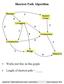 Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
Sequence alignment algorithms
 Sequence alignment algorithms Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 23 rd 27 After this lecture, you can decide when to use local and global sequence alignments
Sequence alignment algorithms Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 23 rd 27 After this lecture, you can decide when to use local and global sequence alignments
Similarity Matrix Based Session Clustering by Sequence Alignment Using Dynamic Programming
 Similarity Matrix Based Session Clustering by Sequence Alignment Using Dynamic Programming Dr.K.Duraiswamy Dean, Academic K.S.Rangasamy College of Technology Tiruchengode, India V. Valli Mayil (Corresponding
Similarity Matrix Based Session Clustering by Sequence Alignment Using Dynamic Programming Dr.K.Duraiswamy Dean, Academic K.S.Rangasamy College of Technology Tiruchengode, India V. Valli Mayil (Corresponding
Metric Indexing of Protein Databases and Promising Approaches
 WDS'07 Proceedings of Contributed Papers, Part I, 91 97, 2007. ISBN 978-80-7378-023-4 MATFYZPRESS Metric Indexing of Protein Databases and Promising Approaches D. Hoksza Charles University, Faculty of
WDS'07 Proceedings of Contributed Papers, Part I, 91 97, 2007. ISBN 978-80-7378-023-4 MATFYZPRESS Metric Indexing of Protein Databases and Promising Approaches D. Hoksza Charles University, Faculty of
Jyoti Lakhani 1, Ajay Khunteta 2, Dharmesh Harwani *3 1 Poornima University, Jaipur & Maharaja Ganga Singh University, Bikaner, Rajasthan, India
 International Journal of Scientific Research in Computer Science, Engineering and Information Technology 2017 IJSRCSEIT Volume 2 Issue 6 ISSN : 2456-3307 Improvisation of Global Pairwise Sequence Alignment
International Journal of Scientific Research in Computer Science, Engineering and Information Technology 2017 IJSRCSEIT Volume 2 Issue 6 ISSN : 2456-3307 Improvisation of Global Pairwise Sequence Alignment
OPEN MP-BASED PARALLEL AND SCALABLE GENETIC SEQUENCE ALIGNMENT
 OPEN MP-BASED PARALLEL AND SCALABLE GENETIC SEQUENCE ALIGNMENT Asif Ali Khan*, Laiq Hassan*, Salim Ullah* ABSTRACT: In bioinformatics, sequence alignment is a common and insistent task. Biologists align
OPEN MP-BASED PARALLEL AND SCALABLE GENETIC SEQUENCE ALIGNMENT Asif Ali Khan*, Laiq Hassan*, Salim Ullah* ABSTRACT: In bioinformatics, sequence alignment is a common and insistent task. Biologists align
HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT
 HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT - Swarbhanu Chatterjee. Hidden Markov models are a sophisticated and flexible statistical tool for the study of protein models. Using HMMs to analyze proteins
HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT - Swarbhanu Chatterjee. Hidden Markov models are a sophisticated and flexible statistical tool for the study of protein models. Using HMMs to analyze proteins
BLAST - Basic Local Alignment Search Tool
 Lecture for ic Bioinformatics (DD2450) April 11, 2013 Searching 1. Input: Query Sequence 2. Database of sequences 3. Subject Sequence(s) 4. Output: High Segment Pairs (HSPs) Sequence Similarity Measures:
Lecture for ic Bioinformatics (DD2450) April 11, 2013 Searching 1. Input: Query Sequence 2. Database of sequences 3. Subject Sequence(s) 4. Output: High Segment Pairs (HSPs) Sequence Similarity Measures:
Accurate Long-Read Alignment using Similarity Based Multiple Pattern Alignment and Prefix Tree Indexing
 Proposal for diploma thesis Accurate Long-Read Alignment using Similarity Based Multiple Pattern Alignment and Prefix Tree Indexing Astrid Rheinländer 01-09-2010 Supervisor: Prof. Dr. Ulf Leser Motivation
Proposal for diploma thesis Accurate Long-Read Alignment using Similarity Based Multiple Pattern Alignment and Prefix Tree Indexing Astrid Rheinländer 01-09-2010 Supervisor: Prof. Dr. Ulf Leser Motivation
Divide and Conquer Algorithms
 CSE341T 09/13/2017 Lecture 5 Divide and Conquer Algorithms We have already seen a couple of divide and conquer algorithms in this lecture. The reduce algorithm and the algorithm to copy elements of the
CSE341T 09/13/2017 Lecture 5 Divide and Conquer Algorithms We have already seen a couple of divide and conquer algorithms in this lecture. The reduce algorithm and the algorithm to copy elements of the
Reconstructing long sequences from overlapping sequence fragment. Searching databases for related sequences and subsequences
 SEQUENCE ALIGNMENT ALGORITHMS 1 Why compare sequences? Reconstructing long sequences from overlapping sequence fragment Searching databases for related sequences and subsequences Storing, retrieving and
SEQUENCE ALIGNMENT ALGORITHMS 1 Why compare sequences? Reconstructing long sequences from overlapping sequence fragment Searching databases for related sequences and subsequences Storing, retrieving and
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 04: Variations of sequence alignments http://www.pitt.edu/~mcs2/teaching/biocomp/tutorials/global.html Slides adapted from Dr. Shaojie Zhang (University
EECS730: Introduction to Bioinformatics Lecture 04: Variations of sequence alignments http://www.pitt.edu/~mcs2/teaching/biocomp/tutorials/global.html Slides adapted from Dr. Shaojie Zhang (University
An Approach to Identify the Number of Clusters
 An Approach to Identify the Number of Clusters Katelyn Gao Heather Hardeman Edward Lim Cristian Potter Carl Meyer Ralph Abbey July 11, 212 Abstract In this technological age, vast amounts of data are generated.
An Approach to Identify the Number of Clusters Katelyn Gao Heather Hardeman Edward Lim Cristian Potter Carl Meyer Ralph Abbey July 11, 212 Abstract In this technological age, vast amounts of data are generated.
CS313 Exercise 4 Cover Page Fall 2017
 CS313 Exercise 4 Cover Page Fall 2017 Due by the start of class on Thursday, October 12, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
CS313 Exercise 4 Cover Page Fall 2017 Due by the start of class on Thursday, October 12, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
17 dicembre Luca Bortolussi SUFFIX TREES. From exact to approximate string matching.
 17 dicembre 2003 Luca Bortolussi SUFFIX TREES From exact to approximate string matching. An introduction to string matching String matching is an important branch of algorithmica, and it has applications
17 dicembre 2003 Luca Bortolussi SUFFIX TREES From exact to approximate string matching. An introduction to string matching String matching is an important branch of algorithmica, and it has applications
USING AN EXTENDED SUFFIX TREE TO SPEED-UP SEQUENCE ALIGNMENT
 IADIS International Conference Applied Computing 2006 USING AN EXTENDED SUFFIX TREE TO SPEED-UP SEQUENCE ALIGNMENT Divya R. Singh Software Engineer Microsoft Corporation, Redmond, WA 98052, USA Abdullah
IADIS International Conference Applied Computing 2006 USING AN EXTENDED SUFFIX TREE TO SPEED-UP SEQUENCE ALIGNMENT Divya R. Singh Software Engineer Microsoft Corporation, Redmond, WA 98052, USA Abdullah
Long Read RNA-seq Mapper
 UNIVERSITY OF ZAGREB FACULTY OF ELECTRICAL ENGENEERING AND COMPUTING MASTER THESIS no. 1005 Long Read RNA-seq Mapper Josip Marić Zagreb, February 2015. Table of Contents 1. Introduction... 1 2. RNA Sequencing...
UNIVERSITY OF ZAGREB FACULTY OF ELECTRICAL ENGENEERING AND COMPUTING MASTER THESIS no. 1005 Long Read RNA-seq Mapper Josip Marić Zagreb, February 2015. Table of Contents 1. Introduction... 1 2. RNA Sequencing...
Biological Sequence Matching Using Fuzzy Logic
 International Journal of Scientific & Engineering Research Volume 2, Issue 7, July-2011 1 Biological Sequence Matching Using Fuzzy Logic Nivit Gill, Shailendra Singh Abstract: Sequence alignment is the
International Journal of Scientific & Engineering Research Volume 2, Issue 7, July-2011 1 Biological Sequence Matching Using Fuzzy Logic Nivit Gill, Shailendra Singh Abstract: Sequence alignment is the
Basic Local Alignment Search Tool (BLAST)
 BLAST 26.04.2018 Basic Local Alignment Search Tool (BLAST) BLAST (Altshul-1990) is an heuristic Pairwise Alignment composed by six-steps that search for local similarities. The most used access point to
BLAST 26.04.2018 Basic Local Alignment Search Tool (BLAST) BLAST (Altshul-1990) is an heuristic Pairwise Alignment composed by six-steps that search for local similarities. The most used access point to
Hashing. Hashing Procedures
 Hashing Hashing Procedures Let us denote the set of all possible key values (i.e., the universe of keys) used in a dictionary application by U. Suppose an application requires a dictionary in which elements
Hashing Hashing Procedures Let us denote the set of all possible key values (i.e., the universe of keys) used in a dictionary application by U. Suppose an application requires a dictionary in which elements
Multiple Sequence Alignment. Mark Whitsitt - NCSA
 Multiple Sequence Alignment Mark Whitsitt - NCSA What is a Multiple Sequence Alignment (MA)? GMHGTVYANYAVDSSDLLLAFGVRFDDRVTGKLEAFASRAKIVHIDIDSAEIGKNKQPHV GMHGTVYANYAVEHSDLLLAFGVRFDDRVTGKLEAFASRAKIVHIDIDSAEIGKNKTPHV
Multiple Sequence Alignment Mark Whitsitt - NCSA What is a Multiple Sequence Alignment (MA)? GMHGTVYANYAVDSSDLLLAFGVRFDDRVTGKLEAFASRAKIVHIDIDSAEIGKNKQPHV GMHGTVYANYAVEHSDLLLAFGVRFDDRVTGKLEAFASRAKIVHIDIDSAEIGKNKTPHV
Programming assignment for the course Sequence Analysis (2006)
 Programming assignment for the course Sequence Analysis (2006) Original text by John W. Romein, adapted by Bart van Houte (bart@cs.vu.nl) Introduction Please note: This assignment is only obligatory for
Programming assignment for the course Sequence Analysis (2006) Original text by John W. Romein, adapted by Bart van Houte (bart@cs.vu.nl) Introduction Please note: This assignment is only obligatory for
A Fast Algorithm for Optimal Alignment between Similar Ordered Trees
 Fundamenta Informaticae 56 (2003) 105 120 105 IOS Press A Fast Algorithm for Optimal Alignment between Similar Ordered Trees Jesper Jansson Department of Computer Science Lund University, Box 118 SE-221
Fundamenta Informaticae 56 (2003) 105 120 105 IOS Press A Fast Algorithm for Optimal Alignment between Similar Ordered Trees Jesper Jansson Department of Computer Science Lund University, Box 118 SE-221
Combinatorial Problems on Strings with Applications to Protein Folding
 Combinatorial Problems on Strings with Applications to Protein Folding Alantha Newman 1 and Matthias Ruhl 2 1 MIT Laboratory for Computer Science Cambridge, MA 02139 alantha@theory.lcs.mit.edu 2 IBM Almaden
Combinatorial Problems on Strings with Applications to Protein Folding Alantha Newman 1 and Matthias Ruhl 2 1 MIT Laboratory for Computer Science Cambridge, MA 02139 alantha@theory.lcs.mit.edu 2 IBM Almaden
SimSearch: A new variant of dynamic programming based on distance series for optimal and near-optimal similarity discovery in biological sequences
 SimSearch: A new variant of dynamic programming based on distance series for optimal and near-optimal similarity discovery in biological sequences Sérgio A. D. Deusdado 1 and Paulo M. M. Carvalho 2 1 ESA,
SimSearch: A new variant of dynamic programming based on distance series for optimal and near-optimal similarity discovery in biological sequences Sérgio A. D. Deusdado 1 and Paulo M. M. Carvalho 2 1 ESA,
3.4 Multiple sequence alignment
 3.4 Multiple sequence alignment Why produce a multiple sequence alignment? Using more than two sequences results in a more convincing alignment by revealing conserved regions in ALL of the sequences Aligned
3.4 Multiple sequence alignment Why produce a multiple sequence alignment? Using more than two sequences results in a more convincing alignment by revealing conserved regions in ALL of the sequences Aligned
Sequence Alignment (chapter 6) p The biological problem p Global alignment p Local alignment p Multiple alignment
 Sequence lignment (chapter 6) p The biological problem p lobal alignment p Local alignment p Multiple alignment Local alignment: rationale p Otherwise dissimilar proteins may have local regions of similarity
Sequence lignment (chapter 6) p The biological problem p lobal alignment p Local alignment p Multiple alignment Local alignment: rationale p Otherwise dissimilar proteins may have local regions of similarity
QueryLines: Approximate Query for Visual Browsing
 MITSUBISHI ELECTRIC RESEARCH LABORATORIES http://www.merl.com QueryLines: Approximate Query for Visual Browsing Kathy Ryall, Neal Lesh, Tom Lanning, Darren Leigh, Hiroaki Miyashita and Shigeru Makino TR2005-015
MITSUBISHI ELECTRIC RESEARCH LABORATORIES http://www.merl.com QueryLines: Approximate Query for Visual Browsing Kathy Ryall, Neal Lesh, Tom Lanning, Darren Leigh, Hiroaki Miyashita and Shigeru Makino TR2005-015
Analysis and performance of a UPC implementation of a parallel longest common subsequence algorithm
 Michigan Technological University Digital Commons @ Michigan Tech Dissertations, Master's Theses and Master's Reports - Open Dissertations, Master's Theses and Master's Reports 2009 Analysis and performance
Michigan Technological University Digital Commons @ Michigan Tech Dissertations, Master's Theses and Master's Reports - Open Dissertations, Master's Theses and Master's Reports 2009 Analysis and performance
