R/BioC Exercises & Answers: Unsupervised methods
|
|
|
- Abraham Holmes
- 5 years ago
- Views:
Transcription
1 R/BioC Exercises & Answers: Unsupervised methods Perry Moerland April 20, 2010 Z Information on how to log on to a PC in the exercise room and the UNIX server can be found here: Don t forget to enable X11 forwarding when starting up PuTTY. Z These exercises depend on Chapters 12 (Gentleman et al. 2005) and 13 (Pollard and van der Laan 2005). References to equations are with respect to these chapters. Z I would like to urge you to always closely look at the R code included, the data imported, and the values of the variables created. Also use the help function to look at all the different options functions offer. Don t forget that the goal of these exercises is to make you think! Introduction Unsupervised learning methods aim at detecting structures in data. The term unsupervised refers to the fact that these methods do not use gene or sample annotations, only the (normalized) expression intensities or ratios are used. A primary purpose of such methods is to group similar data together (clustering) and provide a visualization of the data in which structures can easily be recognized. These may be relations among genes, among samples, or even between genes and samples. The discovery of such structures can lead to the development of new hypotheses, e.g., the grouping of genes with similar expression profiles may indicate that they are co-regulated and are possibly involved in the same biological process. If one looks at arrays/samples instead of genes, the separation of expression profiles of patient tissue samples may point to a possible refinement of disease taxonomy. On the other hand, unsupervised methods are often used to confirm known differences between genes or samples. If a clustering algorithm groups samples from two different tumor types into distinct clusters, this provides evidence that the tumor types indeed show clearly detectable differences in their global expression profiles. Z If gene or sample annotations (or classes) are available the more powerful approach is to use them and apply techniques for supervised learning (tomorrow s lab). As you might have noticed, in the above the word similar was used several times. This is really a central concept in unsupervised learning be it clustering or visualization. In clustering one wants to group similar objects together, whereas in visualization one wants to find a representation of a high-dimensional data set in two or three dimensions while loosing as little 1
2 information as possible: objects that are similar in the original data space should also be similar in the low-dimensional space. See (Gentleman et al. 2005) for a more elaborate discussion. Distance measures and clustering We did a mini-experiment with four arrays (A,B,C,D) and 4 genes per array. This data set has been checked for low-quality arrays and properly normalized. The resulting log-ratios are given in file /data/home/stud00/unsup/hcexample.txt. Exercise 1: Copy this file to your home directory, start R, read in hcexample.txt and store it in a variable E for later use. Try to make a plot of the array profiles that looks more or less like the one below. You might want to look into some of the more detailed plotting commands in R: plot.new, plot.window, axis etc. > E <- read.table("hcexample.txt") > plot.new() > plot.window(xlim = c(1, 4), ylim = c(-5, 6)) > axis(1, at = 1:4, lab = c("1", "2", "3", "4")) > axis(2, pos = 0.95) > mtext("genes", side = 1, line = 3, cex = 1.5) > mtext("log2(cy5/cy3)", side = 2, line = 2, cex = 1.5) > for (i in 1:ncol(E)) { + lines(e[, i], lty = i, col = i, lwd = 3) + } > legend(3.3, 2.8, c("a", "B", "C", "D"), lty = 1:4, col = 1:4, + lwd = 3, y.intersp = 1, cex = 1.3) 2
3 log2(cy5/cy3) A B C D Genes There are many other ways to do this. One way to avoid the for statement is to replace it by a call to the function matplot, for example: > matplot(e, type = "l", add = T, col = 1:4, lty = 1:4, lwd = 3) Cluster alogrithms group similar data together. What is meant by the word similar is formally defined by the notion of a distance. In R, the biodist package gives a collection of functions for calculating distance measures. We will have a look at two of them in more detail, first load the package: > library(biodist) Now calculate the Euclidean (12.2, EUC) and the Pearson sample correlation (12.2, COR) distance between the array profiles. > d.euc <- euc(t(e)) > d.euc A B C B C D > d.cor <- cor.dist(t(e), abs = FALSE) > d.cor 3
4 A B C B 2 C 0 2 D Note that you had to transpose the data matrix E since the biodist package calculates pairwise distances for all rows of a matrix. 1 Exercise 2: Can you explain the resulting distance matrix when using the Pearson correlation distance? Profiles of A and C, respectively, B and D are identical (correlation equals one) and therefore have correlation distance equal to zero. Profiles of A and B, respectively, C and D are anticorrelated (correlation equals minus one) and their correlation distance equals two. Such a distance or dissimilarity matrix forms the basis for most clustering algorithms. In microarray literature, an agglomerative (i.e., bottum-up) hierarchical approach such as implemented in the hclust function is popular. As explained in the lecture, hierarchical clustering tries to find a tree-like representation of the data in which clusters have subclusters, which have subclusters, and so on. The number of clusters depends on in how much detail one looks at the tree. Hierarchical clustering uses two types of distances: ˆ Distance between two individual data points (Euclidean, correlation etc. ) ˆ Distance between two clusters of data points, also called linkage (single, average, etc.). Exercise 3: Draw (just with pencil on paper) the dendrograms for both the Euclidean and the correlation distance matrix generated above. Explain your results. Also write the R code using hclust for both cases when using average linkage, store the dendrograms in variables hc.euc and hc.cor respectively. Plot the resulting dendrograms. > par(mfrow = c(1, 2)) > hc.euc <- hclust(d.euc, method = "average") > plot(hc.euc,, main = "Dendrogram (Euclidean)", sub = "Average linkage", + hang = -1) > hc.cor <- hclust(d.cor, method = "average") > plot(hc.cor, main = "Dendrogram (Correlation)", sub = "Average linkage", + hang = -1) > par(mfrow = c(1, 1)) 1 Some of the help pages are wrong and claim that distances are calculated column-wise. 4
5 B C D A Dendrogram (Euclidean) Dendrogram (Correlation) A C B D Height Height d.euc Average linkage d.cor Average linkage Note that the height of the branches indeed corresponds to the distances calculated in the previous exercise. Clusters can be extracted from the resulting dendrograms by a horizontal cut: > cutree(hc.euc, 2) A B C D > cutree(hc.cor, 2) A B C D These two vectors show to which cluster each array is attributed. Clustering synthetic data One peculiarity of clustering algorithms is that they always produce clusters. This happens regardless of whether there is actually any meaningful clustering structure present in the data. Let us now simulate some unstructured data (rnorm randomly generates from a normal distribution) and see what happens. > n.genes < > n.samples <- 50 5
6 > descr <- paste("s", rep(c("0", ""), times = c(9, 41)), 1:50, + sep = "") > set.seed(13) > datamatrix <- matrix(rnorm(n.genes * n.samples), nrow = n.genes) Exercise 4: Use hclust with average, single, and complete linkage to cluster the samples and plot the resulting dendrograms. Which phenomenon do you see with single linkage? How might this make interpretation difficult? The phenomenon you observe is called chaining. This is the sequential addition of single objects to an existing cluster. Due to sequential addition of single objects, often no clear cluster structure is obtained. > deuc <- euc(t(datamatrix)) > havgeuc <- hclust(deuc, method = "average") > plot(havgeuc, labels = descr, cex = 0.6) Cluster Dendrogram Height S19 S44 S07 S23 S29 S40 S38 S46 S26 S50 S27 S48 S17 S43 S03 S49 S01 S30 S09 S13 S12 S25 S08 S31 S28 S11 S45 S21 S18 S06 S39 S32 S35 S04 S34 S02 S10 S22 S15 S41 S24 S33 S14 S36 S42 S05 S20 S16 S37 S47 deuc hclust (*, "average") > hsineuc <- hclust(deuc, method = "single") > plot(hsineuc, labels = descr, cex = 0.6) 6
7 Cluster Dendrogram S19 S44 Height S07 S23 S29 S40 S50 S28 S03 S21 S26 S27 S48 S49 S14 S09 S01 S30 S17 S43 S41 S18 S13 S08 S31 S06 S12 S11 S45 S32 S33 S25 S15 S10 S24 S39 S04 S02 S35 S22 S34 S37 S47 S36 S20 S16 S42 S05 S38 S46 deuc hclust (*, "single") > hcomeuc <- hclust(deuc, method = "complete") > plot(hcomeuc, labels = descr, cex = 0.6) Cluster Dendrogram Height S19 S44 S26 S50 S38 S46 S29 S40 S07 S23 S17 S43 S27 S48 S09 S13 S08 S31 S11 S45 S03 S49 S01 S30 S12 S25 S28 S21 S06 S39 S24 S04 S35 S14 S18 S05 S15 S02 S47 S34 S37 S32 S10 S41 S22 S36 S42 S33 S20 S16 deuc hclust (*, "complete") Some of the dendrograms generated above appear to find structure in the data. You might decide that the complete linkage dendrogram contains four clusters. Let us assume that you 7
8 stored the complete linkage dendrogram in variable hcomeuc (because of the randomness in this data set results are very likely to be different from the ones shown here). > cutree(hcomeuc, 4) [1] [39] > table(cutree(hcomeuc, 4)) Is this structure real? Since the data generated was completely random, this is not very likely. A nice way of testing the stability of a clustering result is to slightly perturb the data numerous times and see whether results are robust with respect to these small perturbations. This can be done by bootstrapping the individual genes, or rows of the data matrix (section in (Pollard and van der Laan 2005)). The idea behind the bootstrap is to create a new data matrix (of the same size as the original) by randomly selecting rows. Sampling is done with replacement, so some rows will be included multiple times and some rows will be omitted. Here is a simple example where we bootstrap row indices for a data matrix with 10 rows: > sample(1:10, replace = T) [1] We can use the ClassDiscovery package to perform bootstrap resampling for clustering. In order to test the stability of k = 4 groups for hierarchical clustering with Euclidean distance and complete linkage using 200 bootstrap samples, we do the following: > library(classdiscovery) > bc <- BootstrapClusterTest(dataMatrix, cuthclust, k = 4, method = "complete", + metric = "euclid", ntimes = 200) > image(bc, col = blueyellow(64)) 8
9 The main result of BootstrapClusterTest is that for each pair of columns (arrays in this case) in the original data, the number of times they belong to the same cluster of a bootstrap sample is calculated. This is stored in bc@result and used to create the heatmap displayed in the figure. Two samples that often cluster together are shown as yellow and two samples that cluster often apart are shown as blue. The figure clearly illustrates the absence of real structure in the data. Exercise 5: Apply the bootstrap cluster test to the Euclidean dendrogram from the toy example data stored in variable E (see previous section). How many clusters would you expect? Is there stable structure in this data set? > bc <- BootstrapClusterTest(E, cuthclust, k = 2, method = "average", + metric = "euclid", ntimes = 200) > image(bc, col = blueyellow(64), labrow = colnames(e), labcol = colnames(e)) 9
10 D C B A A B C D You would expect two clusters {A, B} and {C, D}, therefore you had to choose k = 2 in the call to BootstrapClusterTest. There is indeed stable structure, indicated by the yellow squares telling you that in all bootstrap samples {A, B}, respectively {C, D} cluster together. Occasionally, you see papers comparing two known classes of samples that perform the following analysis: ˆ Find a list of differentially expressed genes. ˆ Cluster the data using only the differentially expressed genes. ˆ Discover that you can successfully distinguish the known classes. Is this really surprising? Exercise 6: Let us try this on our simulated data stored in datamatrix. Divide the 50 samples into two classes (the first 25 and the last 25). Next, perform t-tests (using the function rowttests from the genefilter package) to see how well each gene separates the two classes, and cluster the data using the top 10 genes (hint: use the function order) with Euclidean distance and complete linkage. Use bootstrapping to test the stability of the grouping found. > classes <- as.factor(c(rep(0, 25), rep(1, 25))) > library(genefilter) > t.stats <- abs(rowttests(datamatrix, classes)$statistic) > index.top10 <- order(t.stats, decreasing = T)[1:10] > hc <- hclust(euc(t(datamatrix[index.top10, ])), method = "complete") > plot(hc, labels = descr, cex = 0.6) 10
11 Cluster Dendrogram Height S02 S22 S21 S28 S34 S41 S27 S50 S06 S09 S05 S39 S01 S38 S11 S04 S13 S08 S14 S45 S49 S31 S44 S32 S43 S26 S33 S07 S17 S18 S20 S35 S47 S40 S24 S25 S30 S46 S10 S23 S12 S15 S19 S37 S16 S42 S29 S36 S48 S03 euc(t(datamatrix[index.top10, ])) hclust (*, "complete") > bc <- BootstrapClusterTest(dataMatrix[index.top10, ], cuthclust, + k = 2, method = "complete", metric = "euclid", ntimes = 200) > cols <- rep(c("red", "green"), times = c(25, 25)) > image(bc, col = blueyellow(64), RowSideColors = cols, ColSideColors = cols)
12 Figure 1: The black line depicts one single gene expression profile and the vertical black line in the right-hand image figure illustrates how the black gene expression profile is constructed: the gene expression levels of the same spot (=gene) over multiple microarray experiments are taken. Only ten arrays are considered in this simulated dataset. Therefore, the left-hand figure has ten experiments on the x-axis. The data set consists of nine different profiles; round each profile 50 noisy profiles were generated. Therefore, each simulated array contains 9*(50+1)=459 different profiles. The log-ratio of these gene profiles is indicated on the y-axis. You can also extract the p-values from the call to rowttests and extract the 10 genes with the smallest p-values. The relatively stable clusters in this example illustrate that it is dangerous to filter on a specific contrast such as differential expression. Just due to chance, this can lead to structure even in random data. Clustering less synthetic data Figure 1 shows the gene profiles of yet another synthetic data set which is slightly closer to biological reality. The data can be found in /data/home/stud00/unsup/jq.txt. Exercise 7: When clustering genes with the Euclidean distance, how many clusters would you expect? Write the code to verify your answer. You might want to explore the heatmap function for this purpose. A heat map is a false color image with a dendrogram added to the left side and to the top (like in the previous section). Especially, explore the influence of the scale argument of heatmap. Also try to relate the clusters found to the gene expression profiles of Figure 1. > jq.data <- read.table("jq.txt") > heatmap(as.matrix(jq.data), Colv = NA, scale = "none", labrow = NA, + col = blueyellow(64)) 12
13 V1 V2 V3 V4 V5 V6 V7 V8 V9 V10 > bc <- BootstrapClusterTest(t(jq.data), cuthclust, k = 9, method = "complete", + metric = "euclid", ntimes = 200) > image(bc, col = blueyellow(64), labrow = NA, labcol = NA) 13
14 Given the way the data has been generated (see legend of Figure 1) you would expect nine clusters, corresponding to each of the nine bands. In the heatmap one can clearly recognize the characteristic up-and-down patterns and the expression levels of each of the nine bands. Exercise 8: Until now we often had to specify the number of clusters in advance. There are many methods to choose the optimal number of clusters (section in (Pollard and van der Laan 2005)). A graphical procedure could be to plot the heights of the branches of a dendrogram and then to determine where the curve becomes flatter (remember the fusion graph from the lecture). This corresponds to the point where the data is not structured anymore. Can you do this for our synthetic dataset using the function hclust? Then use the function locator to determine the optimal number of clusters in this dataset. > plot(rev(hclust(euc(as.matrix(jq.data)))$height), main = "height of the branches of the dend height of the branches of the dendrogram rev(hclust(euc(as.matrix(jq.data)))$height) Index With the command > locator(n = 1) and then clicking in the generated plot near the hinge point you can get a more or less accurate estimate of the number of clusters from the returned x-coordinate. K -means clustering Another often-used clustering algorithm is the K -means algorithm. From the lecture you might remember that the K -means algorithm tries to minimize the total within-scatter. We are going to study the behaviour of the algorithm in some more detail on a simple simulated dataset. 14
15 Exercise 9: ˆ Use mvrnorm from the MASS package to create a dataset consisting of three spherical clusters of 50 datapoints each. Means of the clusters are at (0,0), (10,10), and (20,20). Covariance matrices are diagonal with ones on the diagonal. Make a scatterplot of the resulting dataset. > library(mass) > c1 <- mvrnorm(n = 50, c(0, 0), Sigma = matrix(c(1, 0, 0, 1), + 2, 2)) > c2 <- mvrnorm(n = 50, c(10, 10), Sigma = matrix(c(1, 0, 0, 1), + 2, 2)) > c3 <- mvrnorm(n = 50, c(20, 20), Sigma = matrix(c(1, 0, 0, 1), + 2, 2)) > triclust <- rbind(c1, c2, c3) > plot(triclust) triclust[,2] triclust[,1] ˆ Run the function kmeans 50 times and keep track of the total within-scatter. What do you observe? Can you relate this to the clustering solutions found and give an explanation of this behaviour? > res.kmeans <- list(na, 50) > kmeans.error <- vector("numeric", 50) > for (i in 1:50) { + res.kmeans[[i]] <- kmeans(triclust, 3) + kmeans.error[i] <- sum(res.kmeans[[i]]$withinss) 15
16 + } > kmeans.error [1] [8] [15] [22] [29] [36] [43] [50] There are a number of clustering solutions with a much higher within-scatter. These correspond to so-called local minima that K -means can get stuck in. We can plot one of these sub-optimal solutions: > indx <- which(kmeans.error > 5000) > plot(triclust) > points(res.kmeans[[indx[1]]]$centers, pch = "x", cex = 2, col = "red") triclust[,2] x x x triclust[,1] This is a solution where two cluster centers are in a single cluster, and the third is positioned between the remaining two. This is indeed a local minimum since one of the cluster centers now sharing a cluster needs to move towards the cluster without a cluster center. Such a move will result in an initial increase of the within-scatter. Since 16
17 the K -means algorithm only decreases the within-scatter, it remains stuck in this local minimum. ˆ Does hierarchical clustering also suffer from this problem? It does not suffer from this problem, since it is a deterministic algorithm, independent of random initial conditions. ˆ Compute the total within-scatter for kmeans with [2,3,5,10,20,50,100,149] clusters and then calculate the normalised within-scatter by dividing all within-scatter values by the within-scatter for two clusters. Repeat this 10 times and plot the average normalised within-scatter as a function of the number of clusters. > niter <- 10 > K <- c(2, 3, 5, 10, 20, 50, 100, 149) > ws <- vector("numeric", length(k)) > for (i in (1:length(K))) { + s <- 0 + for (j in 1:niter) { + s <- s + sum(kmeans(triclust, K[i])$withinss) + } + ws[i] <- s/niter + } > ws <- ws/ws[1] > plot(ws, xlab = "number of clusters", ylab = "normalised within-scatter", + type = "l", xaxt = "n") > axis(1, labels = K, at = 1:8) 17
18 normalised within scatter number of clusters ˆ What are the most prominent characteristics of this curve? The within-scatter decreases monotonically as the number of clusters increases. The largest decrease occurs when we go from two to three clusters, revealing that there are probably three clusters in the data. References Gentleman, R., V. Carey, W. Huber, R. Irizarry, and S. Dudoit (Eds.) (2005). Bioinformatics and Computational Biology Solutions Using R and Bioconductor. New York: Springer. Gentleman, R., B. Ding, S. Dudoit, and J. Ibrahim (2005). Chapter 12: Distance measures in DNA microarray data analysis. See Gentleman, Carey, Huber, Irizarry, and Dudoit (2005). Pollard, K. and M. van der Laan (2005). Chapter 13: Cluster analysis of genomic data. See Gentleman, Carey, Huber, Irizarry, and Dudoit (2005). This document was generated with > sessioninfo() R version ( ) i386-pc-mingw32 locale: [1] LC_COLLATE=English_United Kingdom
19 [2] LC_CTYPE=English_United Kingdom.1252 [3] LC_MONETARY=English_United Kingdom.1252 [4] LC_NUMERIC=C [5] LC_TIME=English_United Kingdom.1252 attached base packages: [1] stats graphics grdevices utils datasets methods base other attached packages: [1] MASS_7.3-4 genefilter_ ClassDiscovery_ [4] mclust_3.4.4 cluster_ PreProcess_ [7] oompabase_ biodist_ KernSmooth_ [10] Biobase_2.6.1 loaded via a namespace (and not attached): [1] annotate_ AnnotationDbi_1.8.2 DBI_0.2-5 [4] RSQLite_0.8-4 splines_ survival_ [7] xtable_
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT MD Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org 19
GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT MD Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org 19
Unsupervised learning: Clustering & Dimensionality reduction. Theo Knijnenburg Jorma de Ronde
 Unsupervised learning: Clustering & Dimensionality reduction Theo Knijnenburg Jorma de Ronde Source of slides Marcel Reinders TU Delft Lodewyk Wessels NKI Bioalgorithms.info Jeffrey D. Ullman Stanford
Unsupervised learning: Clustering & Dimensionality reduction Theo Knijnenburg Jorma de Ronde Source of slides Marcel Reinders TU Delft Lodewyk Wessels NKI Bioalgorithms.info Jeffrey D. Ullman Stanford
Exploratory data analysis for microarrays
 Exploratory data analysis for microarrays Jörg Rahnenführer Computational Biology and Applied Algorithmics Max Planck Institute for Informatics D-66123 Saarbrücken Germany NGFN - Courses in Practical DNA
Exploratory data analysis for microarrays Jörg Rahnenführer Computational Biology and Applied Algorithmics Max Planck Institute for Informatics D-66123 Saarbrücken Germany NGFN - Courses in Practical DNA
Cluster Analysis for Microarray Data
 Cluster Analysis for Microarray Data Seventh International Long Oligonucleotide Microarray Workshop Tucson, Arizona January 7-12, 2007 Dan Nettleton IOWA STATE UNIVERSITY 1 Clustering Group objects that
Cluster Analysis for Microarray Data Seventh International Long Oligonucleotide Microarray Workshop Tucson, Arizona January 7-12, 2007 Dan Nettleton IOWA STATE UNIVERSITY 1 Clustering Group objects that
Computing with large data sets
 Computing with large data sets Richard Bonneau, spring 2009 Lecture 8(week 5): clustering 1 clustering Clustering: a diverse methods for discovering groupings in unlabeled data Because these methods don
Computing with large data sets Richard Bonneau, spring 2009 Lecture 8(week 5): clustering 1 clustering Clustering: a diverse methods for discovering groupings in unlabeled data Because these methods don
How to use CNTools. Overview. Algorithms. Jianhua Zhang. April 14, 2011
 How to use CNTools Jianhua Zhang April 14, 2011 Overview Studies have shown that genomic alterations measured as DNA copy number variations invariably occur across chromosomal regions that span over several
How to use CNTools Jianhua Zhang April 14, 2011 Overview Studies have shown that genomic alterations measured as DNA copy number variations invariably occur across chromosomal regions that span over several
CQN (Conditional Quantile Normalization)
 CQN (Conditional Quantile Normalization) Kasper Daniel Hansen khansen@jhsph.edu Zhijin Wu zhijin_wu@brown.edu Modified: August 8, 2012. Compiled: April 30, 2018 Introduction This package contains the CQN
CQN (Conditional Quantile Normalization) Kasper Daniel Hansen khansen@jhsph.edu Zhijin Wu zhijin_wu@brown.edu Modified: August 8, 2012. Compiled: April 30, 2018 Introduction This package contains the CQN
9/29/13. Outline Data mining tasks. Clustering algorithms. Applications of clustering in biology
 9/9/ I9 Introduction to Bioinformatics, Clustering algorithms Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Outline Data mining tasks Predictive tasks vs descriptive tasks Example
9/9/ I9 Introduction to Bioinformatics, Clustering algorithms Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Outline Data mining tasks Predictive tasks vs descriptive tasks Example
Preliminary Figures for Renormalizing Illumina SNP Cell Line Data
 Preliminary Figures for Renormalizing Illumina SNP Cell Line Data Kevin R. Coombes 17 March 2011 Contents 1 Executive Summary 1 1.1 Introduction......................................... 1 1.1.1 Aims/Objectives..................................
Preliminary Figures for Renormalizing Illumina SNP Cell Line Data Kevin R. Coombes 17 March 2011 Contents 1 Executive Summary 1 1.1 Introduction......................................... 1 1.1.1 Aims/Objectives..................................
Distances, Clustering! Rafael Irizarry!
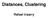 Distances, Clustering! Rafael Irizarry! Heatmaps! Distance! Clustering organizes things that are close into groups! What does it mean for two genes to be close?! What does it mean for two samples to
Distances, Clustering! Rafael Irizarry! Heatmaps! Distance! Clustering organizes things that are close into groups! What does it mean for two genes to be close?! What does it mean for two samples to
MSA220 - Statistical Learning for Big Data
 MSA220 - Statistical Learning for Big Data Lecture 13 Rebecka Jörnsten Mathematical Sciences University of Gothenburg and Chalmers University of Technology Clustering Explorative analysis - finding groups
MSA220 - Statistical Learning for Big Data Lecture 13 Rebecka Jörnsten Mathematical Sciences University of Gothenburg and Chalmers University of Technology Clustering Explorative analysis - finding groups
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 15: Microarray clustering http://compbio.pbworks.com/f/wood2.gif Some slides were adapted from Dr. Shaojie Zhang (University of Central Florida) Microarray
EECS730: Introduction to Bioinformatics Lecture 15: Microarray clustering http://compbio.pbworks.com/f/wood2.gif Some slides were adapted from Dr. Shaojie Zhang (University of Central Florida) Microarray
Analysis of two-way cell-based assays
 Analysis of two-way cell-based assays Lígia Brás, Michael Boutros and Wolfgang Huber April 16, 2015 Contents 1 Introduction 1 2 Assembling the data 2 2.1 Reading the raw intensity files..................
Analysis of two-way cell-based assays Lígia Brás, Michael Boutros and Wolfgang Huber April 16, 2015 Contents 1 Introduction 1 2 Assembling the data 2 2.1 Reading the raw intensity files..................
10701 Machine Learning. Clustering
 171 Machine Learning Clustering What is Clustering? Organizing data into clusters such that there is high intra-cluster similarity low inter-cluster similarity Informally, finding natural groupings among
171 Machine Learning Clustering What is Clustering? Organizing data into clusters such that there is high intra-cluster similarity low inter-cluster similarity Informally, finding natural groupings among
Clustering and Visualisation of Data
 Clustering and Visualisation of Data Hiroshi Shimodaira January-March 28 Cluster analysis aims to partition a data set into meaningful or useful groups, based on distances between data points. In some
Clustering and Visualisation of Data Hiroshi Shimodaira January-March 28 Cluster analysis aims to partition a data set into meaningful or useful groups, based on distances between data points. In some
Introduction for heatmap3 package
 Introduction for heatmap3 package Shilin Zhao April 6, 2015 Contents 1 Example 1 2 Highlights 4 3 Usage 5 1 Example Simulate a gene expression data set with 40 probes and 25 samples. These samples are
Introduction for heatmap3 package Shilin Zhao April 6, 2015 Contents 1 Example 1 2 Highlights 4 3 Usage 5 1 Example Simulate a gene expression data set with 40 probes and 25 samples. These samples are
Class Discovery with OOMPA
 Class Discovery with OOMPA Kevin R. Coombes October, 0 Contents Introduction Getting Started Distances and Clustering. Colored Clusters........................... Checking the Robustness of Clusters 8
Class Discovery with OOMPA Kevin R. Coombes October, 0 Contents Introduction Getting Started Distances and Clustering. Colored Clusters........................... Checking the Robustness of Clusters 8
Hierarchical Clustering
 What is clustering Partitioning of a data set into subsets. A cluster is a group of relatively homogeneous cases or observations Hierarchical Clustering Mikhail Dozmorov Fall 2016 2/61 What is clustering
What is clustering Partitioning of a data set into subsets. A cluster is a group of relatively homogeneous cases or observations Hierarchical Clustering Mikhail Dozmorov Fall 2016 2/61 What is clustering
Lecture 25: Review I
 Lecture 25: Review I Reading: Up to chapter 5 in ISLR. STATS 202: Data mining and analysis Jonathan Taylor 1 / 18 Unsupervised learning In unsupervised learning, all the variables are on equal standing,
Lecture 25: Review I Reading: Up to chapter 5 in ISLR. STATS 202: Data mining and analysis Jonathan Taylor 1 / 18 Unsupervised learning In unsupervised learning, all the variables are on equal standing,
limma: A brief introduction to R
 limma: A brief introduction to R Natalie P. Thorne September 5, 2006 R basics i R is a command line driven environment. This means you have to type in commands (line-by-line) for it to compute or calculate
limma: A brief introduction to R Natalie P. Thorne September 5, 2006 R basics i R is a command line driven environment. This means you have to type in commands (line-by-line) for it to compute or calculate
Measure of Distance. We wish to define the distance between two objects Distance metric between points:
 Measure of Distance We wish to define the distance between two objects Distance metric between points: Euclidean distance (EUC) Manhattan distance (MAN) Pearson sample correlation (COR) Angle distance
Measure of Distance We wish to define the distance between two objects Distance metric between points: Euclidean distance (EUC) Manhattan distance (MAN) Pearson sample correlation (COR) Angle distance
MATH3880 Introduction to Statistics and DNA MATH5880 Statistics and DNA Practical Session Monday, 16 November pm BRAGG Cluster
 MATH3880 Introduction to Statistics and DNA MATH5880 Statistics and DNA Practical Session Monday, 6 November 2009 3.00 pm BRAGG Cluster This document contains the tasks need to be done and completed by
MATH3880 Introduction to Statistics and DNA MATH5880 Statistics and DNA Practical Session Monday, 6 November 2009 3.00 pm BRAGG Cluster This document contains the tasks need to be done and completed by
Clustering. Chapter 10 in Introduction to statistical learning
 Clustering Chapter 10 in Introduction to statistical learning 16 14 12 10 8 6 4 2 0 2 4 6 8 10 12 14 1 Clustering ² Clustering is the art of finding groups in data (Kaufman and Rousseeuw, 1990). ² What
Clustering Chapter 10 in Introduction to statistical learning 16 14 12 10 8 6 4 2 0 2 4 6 8 10 12 14 1 Clustering ² Clustering is the art of finding groups in data (Kaufman and Rousseeuw, 1990). ² What
Differential Expression Analysis at PATRIC
 Differential Expression Analysis at PATRIC The following step- by- step workflow is intended to help users learn how to upload their differential gene expression data to their private workspace using Expression
Differential Expression Analysis at PATRIC The following step- by- step workflow is intended to help users learn how to upload their differential gene expression data to their private workspace using Expression
Gene Clustering & Classification
 BINF, Introduction to Computational Biology Gene Clustering & Classification Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Introduction to Gene Clustering
BINF, Introduction to Computational Biology Gene Clustering & Classification Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Introduction to Gene Clustering
Course on Microarray Gene Expression Analysis
 Course on Microarray Gene Expression Analysis ::: Normalization methods and data preprocessing Madrid, April 27th, 2011. Gonzalo Gómez ggomez@cnio.es Bioinformatics Unit CNIO ::: Introduction. The probe-level
Course on Microarray Gene Expression Analysis ::: Normalization methods and data preprocessing Madrid, April 27th, 2011. Gonzalo Gómez ggomez@cnio.es Bioinformatics Unit CNIO ::: Introduction. The probe-level
Multivariate Analysis
 Multivariate Analysis Cluster Analysis Prof. Dr. Anselmo E de Oliveira anselmo.quimica.ufg.br anselmo.disciplinas@gmail.com Unsupervised Learning Cluster Analysis Natural grouping Patterns in the data
Multivariate Analysis Cluster Analysis Prof. Dr. Anselmo E de Oliveira anselmo.quimica.ufg.br anselmo.disciplinas@gmail.com Unsupervised Learning Cluster Analysis Natural grouping Patterns in the data
Finding Clusters 1 / 60
 Finding Clusters Types of Clustering Approaches: Linkage Based, e.g. Hierarchical Clustering Clustering by Partitioning, e.g. k-means Density Based Clustering, e.g. DBScan Grid Based Clustering 1 / 60
Finding Clusters Types of Clustering Approaches: Linkage Based, e.g. Hierarchical Clustering Clustering by Partitioning, e.g. k-means Density Based Clustering, e.g. DBScan Grid Based Clustering 1 / 60
Examples of implementation of pre-processing method described in paper with R code snippets - Electronic Supplementary Information (ESI)
 Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2015 Examples of implementation of pre-processing method described in paper with R code snippets - Electronic
Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2015 Examples of implementation of pre-processing method described in paper with R code snippets - Electronic
Distance Measures for Gene Expression (and other high-dimensional) Data
 Distance Measures for Gene Expression (and other high-dimensional) Data Utah State University Spring 04 STAT 5570: Statistical Bioinformatics Notes. References Chapter of Bioconductor Monograph (course
Distance Measures for Gene Expression (and other high-dimensional) Data Utah State University Spring 04 STAT 5570: Statistical Bioinformatics Notes. References Chapter of Bioconductor Monograph (course
Hard clustering. Each object is assigned to one and only one cluster. Hierarchical clustering is usually hard. Soft (fuzzy) clustering
 An unsupervised machine learning problem Grouping a set of objects in such a way that objects in the same group (a cluster) are more similar (in some sense or another) to each other than to those in other
An unsupervised machine learning problem Grouping a set of objects in such a way that objects in the same group (a cluster) are more similar (in some sense or another) to each other than to those in other
Dimension reduction : PCA and Clustering
 Dimension reduction : PCA and Clustering By Hanne Jarmer Slides by Christopher Workman Center for Biological Sequence Analysis DTU The DNA Array Analysis Pipeline Array design Probe design Question Experimental
Dimension reduction : PCA and Clustering By Hanne Jarmer Slides by Christopher Workman Center for Biological Sequence Analysis DTU The DNA Array Analysis Pipeline Array design Probe design Question Experimental
Clustering. CE-717: Machine Learning Sharif University of Technology Spring Soleymani
 Clustering CE-717: Machine Learning Sharif University of Technology Spring 2016 Soleymani Outline Clustering Definition Clustering main approaches Partitional (flat) Hierarchical Clustering validation
Clustering CE-717: Machine Learning Sharif University of Technology Spring 2016 Soleymani Outline Clustering Definition Clustering main approaches Partitional (flat) Hierarchical Clustering validation
Analyzing Genomic Data with NOJAH
 Analyzing Genomic Data with NOJAH TAB A) GENOME WIDE ANALYSIS Step 1: Select the example dataset or upload your own. Two example datasets are available. Genome-Wide TCGA-BRCA Expression datasets and CoMMpass
Analyzing Genomic Data with NOJAH TAB A) GENOME WIDE ANALYSIS Step 1: Select the example dataset or upload your own. Two example datasets are available. Genome-Wide TCGA-BRCA Expression datasets and CoMMpass
Clustering Techniques
 Clustering Techniques Bioinformatics: Issues and Algorithms CSE 308-408 Fall 2007 Lecture 16 Lopresti Fall 2007 Lecture 16-1 - Administrative notes Your final project / paper proposal is due on Friday,
Clustering Techniques Bioinformatics: Issues and Algorithms CSE 308-408 Fall 2007 Lecture 16 Lopresti Fall 2007 Lecture 16-1 - Administrative notes Your final project / paper proposal is due on Friday,
Data Exploration with PCA and Unsupervised Learning with Clustering Paul Rodriguez, PhD PACE SDSC
 Data Exploration with PCA and Unsupervised Learning with Clustering Paul Rodriguez, PhD PACE SDSC Clustering Idea Given a set of data can we find a natural grouping? Essential R commands: D =rnorm(12,0,1)
Data Exploration with PCA and Unsupervised Learning with Clustering Paul Rodriguez, PhD PACE SDSC Clustering Idea Given a set of data can we find a natural grouping? Essential R commands: D =rnorm(12,0,1)
HIERARCHICAL clustering analysis (or HCA) is an
 A Data-Driven Approach to Estimating the Number of Clusters in Hierarchical Clustering* Antoine E. Zambelli arxiv:1608.04700v1 [q-bio.qm] 16 Aug 2016 Abstract We propose two new methods for estimating
A Data-Driven Approach to Estimating the Number of Clusters in Hierarchical Clustering* Antoine E. Zambelli arxiv:1608.04700v1 [q-bio.qm] 16 Aug 2016 Abstract We propose two new methods for estimating
How do microarrays work
 Lecture 3 (continued) Alvis Brazma European Bioinformatics Institute How do microarrays work condition mrna cdna hybridise to microarray condition Sample RNA extract labelled acid acid acid nucleic acid
Lecture 3 (continued) Alvis Brazma European Bioinformatics Institute How do microarrays work condition mrna cdna hybridise to microarray condition Sample RNA extract labelled acid acid acid nucleic acid
Clustering CS 550: Machine Learning
 Clustering CS 550: Machine Learning This slide set mainly uses the slides given in the following links: http://www-users.cs.umn.edu/~kumar/dmbook/ch8.pdf http://www-users.cs.umn.edu/~kumar/dmbook/dmslides/chap8_basic_cluster_analysis.pdf
Clustering CS 550: Machine Learning This slide set mainly uses the slides given in the following links: http://www-users.cs.umn.edu/~kumar/dmbook/ch8.pdf http://www-users.cs.umn.edu/~kumar/dmbook/dmslides/chap8_basic_cluster_analysis.pdf
Package sigqc. June 13, 2018
 Title Quality Control Metrics for Gene Signatures Version 0.1.20 Package sigqc June 13, 2018 Description Provides gene signature quality control metrics in publication ready plots. Namely, enables the
Title Quality Control Metrics for Gene Signatures Version 0.1.20 Package sigqc June 13, 2018 Description Provides gene signature quality control metrics in publication ready plots. Namely, enables the
Package SC3. November 27, 2017
 Type Package Title Single-Cell Consensus Clustering Version 1.7.1 Author Vladimir Kiselev Package SC3 November 27, 2017 Maintainer Vladimir Kiselev A tool for unsupervised
Type Package Title Single-Cell Consensus Clustering Version 1.7.1 Author Vladimir Kiselev Package SC3 November 27, 2017 Maintainer Vladimir Kiselev A tool for unsupervised
Clustering Jacques van Helden
 Statistical Analysis of Microarray Data Clustering Jacques van Helden Jacques.van.Helden@ulb.ac.be Contents Data sets Distance and similarity metrics K-means clustering Hierarchical clustering Evaluation
Statistical Analysis of Microarray Data Clustering Jacques van Helden Jacques.van.Helden@ulb.ac.be Contents Data sets Distance and similarity metrics K-means clustering Hierarchical clustering Evaluation
Cluster Analysis. Prof. Thomas B. Fomby Department of Economics Southern Methodist University Dallas, TX April 2008 April 2010
 Cluster Analysis Prof. Thomas B. Fomby Department of Economics Southern Methodist University Dallas, TX 7575 April 008 April 010 Cluster Analysis, sometimes called data segmentation or customer segmentation,
Cluster Analysis Prof. Thomas B. Fomby Department of Economics Southern Methodist University Dallas, TX 7575 April 008 April 010 Cluster Analysis, sometimes called data segmentation or customer segmentation,
Machine Learning and Data Mining. Clustering (1): Basics. Kalev Kask
 Machine Learning and Data Mining Clustering (1): Basics Kalev Kask Unsupervised learning Supervised learning Predict target value ( y ) given features ( x ) Unsupervised learning Understand patterns of
Machine Learning and Data Mining Clustering (1): Basics Kalev Kask Unsupervised learning Supervised learning Predict target value ( y ) given features ( x ) Unsupervised learning Understand patterns of
Multivariate Methods
 Multivariate Methods Cluster Analysis http://www.isrec.isb-sib.ch/~darlene/embnet/ Classification Historically, objects are classified into groups periodic table of the elements (chemistry) taxonomy (zoology,
Multivariate Methods Cluster Analysis http://www.isrec.isb-sib.ch/~darlene/embnet/ Classification Historically, objects are classified into groups periodic table of the elements (chemistry) taxonomy (zoology,
LD vignette Measures of linkage disequilibrium
 LD vignette Measures of linkage disequilibrium David Clayton June 13, 2018 Calculating linkage disequilibrium statistics We shall first load some illustrative data. > data(ld.example) The data are drawn
LD vignette Measures of linkage disequilibrium David Clayton June 13, 2018 Calculating linkage disequilibrium statistics We shall first load some illustrative data. > data(ld.example) The data are drawn
Package SC3. September 29, 2018
 Type Package Title Single-Cell Consensus Clustering Version 1.8.0 Author Vladimir Kiselev Package SC3 September 29, 2018 Maintainer Vladimir Kiselev A tool for unsupervised
Type Package Title Single-Cell Consensus Clustering Version 1.8.0 Author Vladimir Kiselev Package SC3 September 29, 2018 Maintainer Vladimir Kiselev A tool for unsupervised
Chapter 6: Cluster Analysis
 Chapter 6: Cluster Analysis The major goal of cluster analysis is to separate individual observations, or items, into groups, or clusters, on the basis of the values for the q variables measured on each
Chapter 6: Cluster Analysis The major goal of cluster analysis is to separate individual observations, or items, into groups, or clusters, on the basis of the values for the q variables measured on each
Contents. ! Data sets. ! Distance and similarity metrics. ! K-means clustering. ! Hierarchical clustering. ! Evaluation of clustering results
 Statistical Analysis of Microarray Data Contents Data sets Distance and similarity metrics K-means clustering Hierarchical clustering Evaluation of clustering results Clustering Jacques van Helden Jacques.van.Helden@ulb.ac.be
Statistical Analysis of Microarray Data Contents Data sets Distance and similarity metrics K-means clustering Hierarchical clustering Evaluation of clustering results Clustering Jacques van Helden Jacques.van.Helden@ulb.ac.be
Shrinkage of logarithmic fold changes
 Shrinkage of logarithmic fold changes Michael Love August 9, 2014 1 Comparing the posterior distribution for two genes First, we run a DE analysis on the Bottomly et al. dataset, once with shrunken LFCs
Shrinkage of logarithmic fold changes Michael Love August 9, 2014 1 Comparing the posterior distribution for two genes First, we run a DE analysis on the Bottomly et al. dataset, once with shrunken LFCs
ConsensusClusterPlus (Tutorial)
 ConsensusClusterPlus (Tutorial) Matthew D. Wilkerson April 30, 2018 1 Summary ConsensusClusterPlus is a tool for unsupervised class discovery. This document provides a tutorial of how to use ConsensusClusterPlus.
ConsensusClusterPlus (Tutorial) Matthew D. Wilkerson April 30, 2018 1 Summary ConsensusClusterPlus is a tool for unsupervised class discovery. This document provides a tutorial of how to use ConsensusClusterPlus.
Gene expression & Clustering (Chapter 10)
 Gene expression & Clustering (Chapter 10) Determining gene function Sequence comparison tells us if a gene is similar to another gene, e.g., in a new species Dynamic programming Approximate pattern matching
Gene expression & Clustering (Chapter 10) Determining gene function Sequence comparison tells us if a gene is similar to another gene, e.g., in a new species Dynamic programming Approximate pattern matching
Package pvclust. October 23, 2015
 Version 2.0-0 Date 2015-10-23 Package pvclust October 23, 2015 Title Hierarchical Clustering with P-Values via Multiscale Bootstrap Resampling Author Ryota Suzuki , Hidetoshi Shimodaira
Version 2.0-0 Date 2015-10-23 Package pvclust October 23, 2015 Title Hierarchical Clustering with P-Values via Multiscale Bootstrap Resampling Author Ryota Suzuki , Hidetoshi Shimodaira
11/2/2017 MIST.6060 Business Intelligence and Data Mining 1. Clustering. Two widely used distance metrics to measure the distance between two records
 11/2/2017 MIST.6060 Business Intelligence and Data Mining 1 An Example Clustering X 2 X 1 Objective of Clustering The objective of clustering is to group the data into clusters such that the records within
11/2/2017 MIST.6060 Business Intelligence and Data Mining 1 An Example Clustering X 2 X 1 Objective of Clustering The objective of clustering is to group the data into clusters such that the records within
Network Traffic Measurements and Analysis
 DEIB - Politecnico di Milano Fall, 2017 Introduction Often, we have only a set of features x = x 1, x 2,, x n, but no associated response y. Therefore we are not interested in prediction nor classification,
DEIB - Politecnico di Milano Fall, 2017 Introduction Often, we have only a set of features x = x 1, x 2,, x n, but no associated response y. Therefore we are not interested in prediction nor classification,
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org
Clustering. Lecture 6, 1/24/03 ECS289A
 Clustering Lecture 6, 1/24/03 What is Clustering? Given n objects, assign them to groups (clusters) based on their similarity Unsupervised Machine Learning Class Discovery Difficult, and maybe ill-posed
Clustering Lecture 6, 1/24/03 What is Clustering? Given n objects, assign them to groups (clusters) based on their similarity Unsupervised Machine Learning Class Discovery Difficult, and maybe ill-posed
University of California, Berkeley
 University of California, Berkeley U.C. Berkeley Division of Biostatistics Working Paper Series Year 2002 Paper 105 A New Partitioning Around Medoids Algorithm Mark J. van der Laan Katherine S. Pollard
University of California, Berkeley U.C. Berkeley Division of Biostatistics Working Paper Series Year 2002 Paper 105 A New Partitioning Around Medoids Algorithm Mark J. van der Laan Katherine S. Pollard
Step-by-Step Guide to Relatedness and Association Mapping Contents
 Step-by-Step Guide to Relatedness and Association Mapping Contents OBJECTIVES... 2 INTRODUCTION... 2 RELATEDNESS MEASURES... 2 POPULATION STRUCTURE... 6 Q-K ASSOCIATION ANALYSIS... 10 K MATRIX COMPRESSION...
Step-by-Step Guide to Relatedness and Association Mapping Contents OBJECTIVES... 2 INTRODUCTION... 2 RELATEDNESS MEASURES... 2 POPULATION STRUCTURE... 6 Q-K ASSOCIATION ANALYSIS... 10 K MATRIX COMPRESSION...
Cluster analysis. Agnieszka Nowak - Brzezinska
 Cluster analysis Agnieszka Nowak - Brzezinska Outline of lecture What is cluster analysis? Clustering algorithms Measures of Cluster Validity What is Cluster Analysis? Finding groups of objects such that
Cluster analysis Agnieszka Nowak - Brzezinska Outline of lecture What is cluster analysis? Clustering algorithms Measures of Cluster Validity What is Cluster Analysis? Finding groups of objects such that
BBS654 Data Mining. Pinar Duygulu. Slides are adapted from Nazli Ikizler
 BBS654 Data Mining Pinar Duygulu Slides are adapted from Nazli Ikizler 1 Classification Classification systems: Supervised learning Make a rational prediction given evidence There are several methods for
BBS654 Data Mining Pinar Duygulu Slides are adapted from Nazli Ikizler 1 Classification Classification systems: Supervised learning Make a rational prediction given evidence There are several methods for
Unsupervised Learning and Clustering
 Unsupervised Learning and Clustering Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2009 CS 551, Spring 2009 c 2009, Selim Aksoy (Bilkent University)
Unsupervised Learning and Clustering Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2009 CS 551, Spring 2009 c 2009, Selim Aksoy (Bilkent University)
10601 Machine Learning. Hierarchical clustering. Reading: Bishop: 9-9.2
 161 Machine Learning Hierarchical clustering Reading: Bishop: 9-9.2 Second half: Overview Clustering - Hierarchical, semi-supervised learning Graphical models - Bayesian networks, HMMs, Reasoning under
161 Machine Learning Hierarchical clustering Reading: Bishop: 9-9.2 Second half: Overview Clustering - Hierarchical, semi-supervised learning Graphical models - Bayesian networks, HMMs, Reasoning under
3. Cluster analysis Overview
 Université Laval Multivariate analysis - February 2006 1 3.1. Overview 3. Cluster analysis Clustering requires the recognition of discontinuous subsets in an environment that is sometimes discrete (as
Université Laval Multivariate analysis - February 2006 1 3.1. Overview 3. Cluster analysis Clustering requires the recognition of discontinuous subsets in an environment that is sometimes discrete (as
SVM Classification in -Arrays
 SVM Classification in -Arrays SVM classification and validation of cancer tissue samples using microarray expression data Furey et al, 2000 Special Topics in Bioinformatics, SS10 A. Regl, 7055213 What
SVM Classification in -Arrays SVM classification and validation of cancer tissue samples using microarray expression data Furey et al, 2000 Special Topics in Bioinformatics, SS10 A. Regl, 7055213 What
Statistics 202: Data Mining. c Jonathan Taylor. Week 8 Based in part on slides from textbook, slides of Susan Holmes. December 2, / 1
 Week 8 Based in part on slides from textbook, slides of Susan Holmes December 2, 2012 1 / 1 Part I Clustering 2 / 1 Clustering Clustering Goal: Finding groups of objects such that the objects in a group
Week 8 Based in part on slides from textbook, slides of Susan Holmes December 2, 2012 1 / 1 Part I Clustering 2 / 1 Clustering Clustering Goal: Finding groups of objects such that the objects in a group
Multivariate analyses in ecology. Cluster (part 2) Ordination (part 1 & 2)
 Multivariate analyses in ecology Cluster (part 2) Ordination (part 1 & 2) 1 Exercise 9B - solut 2 Exercise 9B - solut 3 Exercise 9B - solut 4 Exercise 9B - solut 5 Multivariate analyses in ecology Cluster
Multivariate analyses in ecology Cluster (part 2) Ordination (part 1 & 2) 1 Exercise 9B - solut 2 Exercise 9B - solut 3 Exercise 9B - solut 4 Exercise 9B - solut 5 Multivariate analyses in ecology Cluster
Package Binarize. February 6, 2017
 Type Package Title Binarization of One-Dimensional Data Version 1.2 Date 2017-01-29 Package Binarize February 6, 2017 Author Stefan Mundus, Christoph Müssel, Florian Schmid, Ludwig Lausser, Tamara J. Blätte,
Type Package Title Binarization of One-Dimensional Data Version 1.2 Date 2017-01-29 Package Binarize February 6, 2017 Author Stefan Mundus, Christoph Müssel, Florian Schmid, Ludwig Lausser, Tamara J. Blätte,
SimBindProfiles: Similar Binding Profiles, identifies common and unique regions in array genome tiling array data
 SimBindProfiles: Similar Binding Profiles, identifies common and unique regions in array genome tiling array data Bettina Fischer October 30, 2018 Contents 1 Introduction 1 2 Reading data and normalisation
SimBindProfiles: Similar Binding Profiles, identifies common and unique regions in array genome tiling array data Bettina Fischer October 30, 2018 Contents 1 Introduction 1 2 Reading data and normalisation
Exploring cdna Data. Achim Tresch, Andreas Buness, Wolfgang Huber, Tim Beißbarth
 Exploring cdna Data Achim Tresch, Andreas Buness, Wolfgang Huber, Tim Beißbarth Practical DNA Microarray Analysis http://compdiag.molgen.mpg.de/ngfn/pma0nov.shtml The following exercise will guide you
Exploring cdna Data Achim Tresch, Andreas Buness, Wolfgang Huber, Tim Beißbarth Practical DNA Microarray Analysis http://compdiag.molgen.mpg.de/ngfn/pma0nov.shtml The following exercise will guide you
3. Cluster analysis Overview
 Université Laval Analyse multivariable - mars-avril 2008 1 3.1. Overview 3. Cluster analysis Clustering requires the recognition of discontinuous subsets in an environment that is sometimes discrete (as
Université Laval Analyse multivariable - mars-avril 2008 1 3.1. Overview 3. Cluster analysis Clustering requires the recognition of discontinuous subsets in an environment that is sometimes discrete (as
CLUSTERING IN BIOINFORMATICS
 CLUSTERING IN BIOINFORMATICS CSE/BIMM/BENG 8 MAY 4, 0 OVERVIEW Define the clustering problem Motivation: gene expression and microarrays Types of clustering Clustering algorithms Other applications of
CLUSTERING IN BIOINFORMATICS CSE/BIMM/BENG 8 MAY 4, 0 OVERVIEW Define the clustering problem Motivation: gene expression and microarrays Types of clustering Clustering algorithms Other applications of
Cluster Analysis. Ying Shen, SSE, Tongji University
 Cluster Analysis Ying Shen, SSE, Tongji University Cluster analysis Cluster analysis groups data objects based only on the attributes in the data. The main objective is that The objects within a group
Cluster Analysis Ying Shen, SSE, Tongji University Cluster analysis Cluster analysis groups data objects based only on the attributes in the data. The main objective is that The objects within a group
Clustering. CS294 Practical Machine Learning Junming Yin 10/09/06
 Clustering CS294 Practical Machine Learning Junming Yin 10/09/06 Outline Introduction Unsupervised learning What is clustering? Application Dissimilarity (similarity) of objects Clustering algorithm K-means,
Clustering CS294 Practical Machine Learning Junming Yin 10/09/06 Outline Introduction Unsupervised learning What is clustering? Application Dissimilarity (similarity) of objects Clustering algorithm K-means,
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
Clustering. Dick de Ridder 6/10/2018
 Clustering Dick de Ridder 6/10/2018 In these exercises, you will continue to work with the Arabidopsis ST vs. HT RNAseq dataset. First, you will select a subset of the data and inspect it; then cluster
Clustering Dick de Ridder 6/10/2018 In these exercises, you will continue to work with the Arabidopsis ST vs. HT RNAseq dataset. First, you will select a subset of the data and inspect it; then cluster
Exploring cdna Data. Achim Tresch, Andreas Buness, Tim Beißbarth, Wolfgang Huber
 Exploring cdna Data Achim Tresch, Andreas Buness, Tim Beißbarth, Wolfgang Huber Practical DNA Microarray Analysis http://compdiag.molgen.mpg.de/ngfn/pma0nov.shtml The following exercise will guide you
Exploring cdna Data Achim Tresch, Andreas Buness, Tim Beißbarth, Wolfgang Huber Practical DNA Microarray Analysis http://compdiag.molgen.mpg.de/ngfn/pma0nov.shtml The following exercise will guide you
5/15/16. Computational Methods for Data Analysis. Massimo Poesio UNSUPERVISED LEARNING. Clustering. Unsupervised learning introduction
 Computational Methods for Data Analysis Massimo Poesio UNSUPERVISED LEARNING Clustering Unsupervised learning introduction 1 Supervised learning Training set: Unsupervised learning Training set: 2 Clustering
Computational Methods for Data Analysis Massimo Poesio UNSUPERVISED LEARNING Clustering Unsupervised learning introduction 1 Supervised learning Training set: Unsupervised learning Training set: 2 Clustering
Unsupervised learning, Clustering CS434
 Unsupervised learning, Clustering CS434 Unsupervised learning and pattern discovery So far, our data has been in this form: We will be looking at unlabeled data: x 11,x 21, x 31,, x 1 m x 12,x 22, x 32,,
Unsupervised learning, Clustering CS434 Unsupervised learning and pattern discovery So far, our data has been in this form: We will be looking at unlabeled data: x 11,x 21, x 31,, x 1 m x 12,x 22, x 32,,
Contents. Introduction 2. PerturbationClustering 2. SubtypingOmicsData 5. References 8
 PINSPlus: Clustering Algorithm for Data Integration and Disease Subtyping Hung Nguyen, Sangam Shrestha, and Tin Nguyen Department of Computer Science and Engineering University of Nevada, Reno, NV 89557
PINSPlus: Clustering Algorithm for Data Integration and Disease Subtyping Hung Nguyen, Sangam Shrestha, and Tin Nguyen Department of Computer Science and Engineering University of Nevada, Reno, NV 89557
Tutorial script for whole-cell MALDI-TOF analysis
 Tutorial script for whole-cell MALDI-TOF analysis Julien Textoris June 19, 2013 Contents 1 Required libraries 2 2 Data loading 2 3 Spectrum visualization and pre-processing 4 4 Analysis and comparison
Tutorial script for whole-cell MALDI-TOF analysis Julien Textoris June 19, 2013 Contents 1 Required libraries 2 2 Data loading 2 3 Spectrum visualization and pre-processing 4 4 Analysis and comparison
Plotting Segment Calls From SNP Assay
 Plotting Segment Calls From SNP Assay Kevin R. Coombes 17 March 2011 Contents 1 Executive Summary 1 1.1 Introduction......................................... 1 1.1.1 Aims/Objectives..................................
Plotting Segment Calls From SNP Assay Kevin R. Coombes 17 March 2011 Contents 1 Executive Summary 1 1.1 Introduction......................................... 1 1.1.1 Aims/Objectives..................................
Introduction to Data Mining
 Introduction to Data Mining Lecture #14: Clustering Seoul National University 1 In This Lecture Learn the motivation, applications, and goal of clustering Understand the basic methods of clustering (bottom-up
Introduction to Data Mining Lecture #14: Clustering Seoul National University 1 In This Lecture Learn the motivation, applications, and goal of clustering Understand the basic methods of clustering (bottom-up
Correlation. January 12, 2019
 Correlation January 12, 2019 Contents Correlations The Scattterplot The Pearson correlation The computational raw-score formula Survey data Fun facts about r Sensitivity to outliers Spearman rank-order
Correlation January 12, 2019 Contents Correlations The Scattterplot The Pearson correlation The computational raw-score formula Survey data Fun facts about r Sensitivity to outliers Spearman rank-order
Introduction: microarray quality assessment with arrayqualitymetrics
 Introduction: microarray quality assessment with arrayqualitymetrics Audrey Kauffmann, Wolfgang Huber April 4, 2013 Contents 1 Basic use 3 1.1 Affymetrix data - before preprocessing.......................
Introduction: microarray quality assessment with arrayqualitymetrics Audrey Kauffmann, Wolfgang Huber April 4, 2013 Contents 1 Basic use 3 1.1 Affymetrix data - before preprocessing.......................
What to come. There will be a few more topics we will cover on supervised learning
 Summary so far Supervised learning learn to predict Continuous target regression; Categorical target classification Linear Regression Classification Discriminative models Perceptron (linear) Logistic regression
Summary so far Supervised learning learn to predict Continuous target regression; Categorical target classification Linear Regression Classification Discriminative models Perceptron (linear) Logistic regression
Machine Learning (BSMC-GA 4439) Wenke Liu
 Machine Learning (BSMC-GA 4439) Wenke Liu 01-25-2018 Outline Background Defining proximity Clustering methods Determining number of clusters Other approaches Cluster analysis as unsupervised Learning Unsupervised
Machine Learning (BSMC-GA 4439) Wenke Liu 01-25-2018 Outline Background Defining proximity Clustering methods Determining number of clusters Other approaches Cluster analysis as unsupervised Learning Unsupervised
UNSUPERVISED LEARNING IN R. Introduction to hierarchical clustering
 UNSUPERVISED LEARNING IN R Introduction to hierarchical clustering Hierarchical clustering Number of clusters is not known ahead of time Two kinds: bottom-up and top-down, this course bottom-up Hierarchical
UNSUPERVISED LEARNING IN R Introduction to hierarchical clustering Hierarchical clustering Number of clusters is not known ahead of time Two kinds: bottom-up and top-down, this course bottom-up Hierarchical
Exploring cdna Data. Achim Tresch, Andreas Buness, Tim Beißbarth, Wolfgang Huber
 Exploring cdna Data Achim Tresch, Andreas Buness, Tim Beißbarth, Wolfgang Huber Practical DNA Microarray Analysis, Heidelberg, March 2005 http://compdiag.molgen.mpg.de/ngfn/pma2005mar.shtml The following
Exploring cdna Data Achim Tresch, Andreas Buness, Tim Beißbarth, Wolfgang Huber Practical DNA Microarray Analysis, Heidelberg, March 2005 http://compdiag.molgen.mpg.de/ngfn/pma2005mar.shtml The following
Cluster Analysis. Mu-Chun Su. Department of Computer Science and Information Engineering National Central University 2003/3/11 1
 Cluster Analysis Mu-Chun Su Department of Computer Science and Information Engineering National Central University 2003/3/11 1 Introduction Cluster analysis is the formal study of algorithms and methods
Cluster Analysis Mu-Chun Su Department of Computer Science and Information Engineering National Central University 2003/3/11 1 Introduction Cluster analysis is the formal study of algorithms and methods
Clustering algorithms 6CCS3WSN-7CCSMWAL
 Clustering algorithms 6CCS3WSN-7CCSMWAL Contents Introduction: Types of clustering Hierarchical clustering Spatial clustering (k means etc) Community detection (next week) What are we trying to cluster
Clustering algorithms 6CCS3WSN-7CCSMWAL Contents Introduction: Types of clustering Hierarchical clustering Spatial clustering (k means etc) Community detection (next week) What are we trying to cluster
Application of Hierarchical Clustering to Find Expression Modules in Cancer
 Application of Hierarchical Clustering to Find Expression Modules in Cancer T. M. Murali August 18, 2008 Innovative Application of Hierarchical Clustering A module map showing conditional activity of expression
Application of Hierarchical Clustering to Find Expression Modules in Cancer T. M. Murali August 18, 2008 Innovative Application of Hierarchical Clustering A module map showing conditional activity of expression
Machine Learning (BSMC-GA 4439) Wenke Liu
 Machine Learning (BSMC-GA 4439) Wenke Liu 01-31-017 Outline Background Defining proximity Clustering methods Determining number of clusters Comparing two solutions Cluster analysis as unsupervised Learning
Machine Learning (BSMC-GA 4439) Wenke Liu 01-31-017 Outline Background Defining proximity Clustering methods Determining number of clusters Comparing two solutions Cluster analysis as unsupervised Learning
Stats 170A: Project in Data Science Exploratory Data Analysis: Clustering Algorithms
 Stats 170A: Project in Data Science Exploratory Data Analysis: Clustering Algorithms Padhraic Smyth Department of Computer Science Bren School of Information and Computer Sciences University of California,
Stats 170A: Project in Data Science Exploratory Data Analysis: Clustering Algorithms Padhraic Smyth Department of Computer Science Bren School of Information and Computer Sciences University of California,
(1) where, l. denotes the number of nodes to which both i and j are connected, and k is. the number of connections of a node, with.
 A simulated gene co-expression network to illustrate the use of the topological overlap matrix for module detection Steve Horvath, Mike Oldham Correspondence to shorvath@mednet.ucla.edu Abstract Here we
A simulated gene co-expression network to illustrate the use of the topological overlap matrix for module detection Steve Horvath, Mike Oldham Correspondence to shorvath@mednet.ucla.edu Abstract Here we
Points Lines Connected points X-Y Scatter. X-Y Matrix Star Plot Histogram Box Plot. Bar Group Bar Stacked H-Bar Grouped H-Bar Stacked
 Plotting Menu: QCExpert Plotting Module graphs offers various tools for visualization of uni- and multivariate data. Settings and options in different types of graphs allow for modifications and customizations
Plotting Menu: QCExpert Plotting Module graphs offers various tools for visualization of uni- and multivariate data. Settings and options in different types of graphs allow for modifications and customizations
Clustering. Supervised vs. Unsupervised Learning
 Clustering Supervised vs. Unsupervised Learning So far we have assumed that the training samples used to design the classifier were labeled by their class membership (supervised learning) We assume now
Clustering Supervised vs. Unsupervised Learning So far we have assumed that the training samples used to design the classifier were labeled by their class membership (supervised learning) We assume now
Supplementary text S6 Comparison studies on simulated data
 Supplementary text S Comparison studies on simulated data Peter Langfelder, Rui Luo, Michael C. Oldham, and Steve Horvath Corresponding author: shorvath@mednet.ucla.edu Overview In this document we illustrate
Supplementary text S Comparison studies on simulated data Peter Langfelder, Rui Luo, Michael C. Oldham, and Steve Horvath Corresponding author: shorvath@mednet.ucla.edu Overview In this document we illustrate
Cluster Analysis. Angela Montanari and Laura Anderlucci
 Cluster Analysis Angela Montanari and Laura Anderlucci 1 Introduction Clustering a set of n objects into k groups is usually moved by the aim of identifying internally homogenous groups according to a
Cluster Analysis Angela Montanari and Laura Anderlucci 1 Introduction Clustering a set of n objects into k groups is usually moved by the aim of identifying internally homogenous groups according to a
Hierarchical clustering
 Hierarchical clustering Rebecca C. Steorts, Duke University STA 325, Chapter 10 ISL 1 / 63 Agenda K-means versus Hierarchical clustering Agglomerative vs divisive clustering Dendogram (tree) Hierarchical
Hierarchical clustering Rebecca C. Steorts, Duke University STA 325, Chapter 10 ISL 1 / 63 Agenda K-means versus Hierarchical clustering Agglomerative vs divisive clustering Dendogram (tree) Hierarchical
