GS Analysis of Microarray Data
|
|
|
- Asher Cobb
- 6 years ago
- Views:
Transcription
1 GS Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center 22 October 2009
2 APPLIED CLUSTERING 1 Lecture 16: Applied Clustering Clustering Methods K-Means Clustering Partitioning Around Medoids Silhouette Widths Principal Components Principal Coordinates Project Normal Project Normal Clustering Abnormal Behavior Problems and Solution
3 APPLIED CLUSTERING 2 Review of hierarchical clustering Last time: we looked at hierarchical clustering of the samples. Key issues: What distance metric (euclidean, correlation, manhattan, canberra, minkowski) should we use? What linkage rule (average, complete, single, ward) should we use? Which clusters should we believe? (bootstrap resampling) How many clusters should we believe? (bootstrap) Is there any reason to believe that a hierarchical structure makes sense? Today, we ll look at some other clustering techniques.
4 APPLIED CLUSTERING 3 Simulated data To test some algorithms, we simulated data with 1000 genes and 5 different sample classes containing different numbers of samples. Here s a two-dimensional picture of the truth:
5 APPLIED CLUSTERING 4 Hierarchical clusters (correlation; average) Three of the classes (B, C, D) are mostly correct. The other two classes are less concentrated.
6 APPLIED CLUSTERING 5 K-Means Clustering Input: A data matrix, X, and the desired number of clusters, K. Output: For each sample i, a cluster assignment C(i) K. Idea: Minimize the within-cluster sum of squares K c=1 C(i)=c,C(j)=c N (x il x jl ) 2 l=1 Algorithm: 1. Make an initial guess at the centers of the clusters. 2. For each data point, find the closest cluster (Euclidean). 3. Replace each cluster center by averaging data points that are closest to it. 4. Repeat until the assignments stop changing.
7 APPLIED CLUSTERING 6 K-Means, Take 1 Perfect clustering! (Circles = starting group centers, X = final group centers)
8 APPLIED CLUSTERING 7 K-Means, Take 2 Oops: bad starting points may mean bad clusters!
9 APPLIED CLUSTERING 8 K-Means, Take 3
10 APPLIED CLUSTERING 9 Local minima may not be global K-means can be very sensitive to the choice of centers used as seeds for the algorithm. The problem is that the algorithm only converges to a local minimum for the within-cluster sum of squares, and different runs with randomly chosen centers (which is the default in the kmeans function in R) can converge to different local optima. You can see which of these three runs is better: > sum(kres1$withinss) [1] > sum(kres2$withinss) [1] > sum(kres3$withinss) [1]
11 APPLIED CLUSTERING 10 Local minima may not be global There are two ways around the fact that local minima need not be global. One is to find better starting seeds for the algorithm. For example, start with hierarchical clustering. Then cut the tree into five branches, and use the average of each branch as the starting points. Alternatively, you can run the algorithm with many random seeds, keeping track of the within-cluster sum of squares: require(classdiscovery) repeatedkmeans(t(ldata), k=5, ntimes=100)
12 APPLIED CLUSTERING 11 Multiple runs of the K-means algorithm kcent <- sample(n.samples, 5) kres <- kmeans(t(ldata), t(ldata[,kcent]) withinss <- sum(kres$withinss) for (i in 1:100) { tcent <- sample(n.samples, 5) tres <- kmeans(t(ldata), t(ldata[,tcent])) print(sum(tres$withinss)) if (sum(tres$withinss) < withinss) { kres <- tres kcent <- tcent withinss <- sum(kres$withinss) } }
13 APPLIED CLUSTERING 12 Can we use other measures of distance? The K-means clustering algorithm has another limitation. (This is not the last one we will consider). As described, it always uses Euclidean distance as the measure of dissimilarities between sample vectors. As we saw last time with hierarchical clustering, there are a large number of possible distances that we might want to use. Fortunately, a simple adjustment to the algorithm lets us work with any distance measure.
14 APPLIED CLUSTERING 13 Partitioning Around Medoids (PAM) Input: Data matrix, X, distance d, number of clusters, K. Output: For each sample i, a cluster assignment C(i) K. Idea: Minimize the within-cluster distance K c=1 C(i)=c,C(j)=c d(x i, x j ) Algorithm: 1. Make an initial guess at the centers of the clusters. 2. For each data point, find the closest cluster. 3. Replace each cluster center by the data point minimizing the total distance to other members in its cluster. 4. Repeat until the assignments stop changing.
15 APPLIED CLUSTERING 14 PAM in R To use PAM in R, you must load another package: > require(cluster) > dist.matrix <- as.dist(1-cor(ldata)/2) > pamres <- pam(dist.matrix, 5) Unlike kmeans, the implementation of pam only lets you specify the number of clusters you want, not the starting point. It also apparently always uses the same method to choose the starting point, so it does not help to run the algorithm multiple times. If their heuristic chooses a poor starting configuration, there is no way to fix it. Update: The current version of PAM does let you specify the starting points, so disregard the paragraph above.
16 APPLIED CLUSTERING 15 PAM results Not very good on our example data...
17 APPLIED CLUSTERING 16 How many clusters are there? Both kmeans and pam require you to specify the number of clusters before running the algorithm. In our example, we knew before we stated that there were five clusters. In real life, we rarely (if ever) know the number of real clusters before we start. How do we figure out the correct number of clusters? One way is to run the algorithm with different values of K, and then try to decide which methods gives the best results. The problem that remains is how we measure best.
18 APPLIED CLUSTERING 17 Silhouette Widths Kaufman and Rousseeuw (who wrote a book on clustering that describes pam along with quite a few other methods) recommend using the silhouette width as a measure of how much individual elements belong to the cluster where they are assigned. To compute the silhouette width of the i th object, define a(i) = average distance to other elements in the cluster b(i) = smallest average distance to other clusters sil(i) = (b(i) a(i))/max(a(i), b(i)). Interpretation: If sil(i) is near 1, then the object is well clustered. If sil(i) < 0, then the object is probably in the wrong cluster. If sil(i) is near 0, then it s on the border between two clusters.
19 APPLIED CLUSTERING 18 PAM : two clusters > pam2 <- pam(dmat, 2) > plot(pam2)
20 APPLIED CLUSTERING 19 PAM : three clusters > pam3 <- pam(dmat, 3) > plot(pam3)
21 APPLIED CLUSTERING 20 PAM : four clusters > pam4 <- pam(dmat, 4) > plot(pam4)
22 APPLIED CLUSTERING 21 PAM : five clusters > pam5 <- pam(dmat, 5) > plot(pam5)
23 APPLIED CLUSTERING 22 PAM : six clusters > pam6 <- pam(dmat, 6) > plot(pam6)
24 APPLIED CLUSTERING 23 PAM : seven clusters > pam7 <- pam(dmat, 7) > plot(pam7)
25 APPLIED CLUSTERING 24 > summary(silhouette(pam2))$avg.width [1] > summary(silhouette(pam3))$avg.width [1] > summary(silhouette(pam4))$avg.width [1] > summary(silhouette(pam5))$avg.width [1] > summary(silhouette(pam6))$avg.width [1] > summary(silhouette(pam7))$avg.width [1] In general, we want to choose the number of clusters that maximizes the average silhouette width. In this case, that means 4 or 5 clusters.
26 APPLIED CLUSTERING 25 Using silhouettes with K-means The silhouette function knows about pam objects, making it relatively easy to use. You can use it with other clustering routines, but you have to supply the clustering vector and the distance matrix. For example, here are the results for the best K-means clustering that we found (as measured by the within-cluster sum of squares). > euc.distance <- dist(t(ldata)) > ksil <- silhouette(kres$cluster, euc.distance) > summary(ksil)$avg.width [1] Note that the silhouette width is smaller than the one from PAM using correlation. However, the silhouette plot suggests that everything is classified correctly:
27 APPLIED CLUSTERING 26 K-means: five cluster silhouette > plot(ksil)
28 APPLIED CLUSTERING 27 Silhouettes with hclust Looking back at the hierarchical clustering, we need to cut eight branches off (including some singletons) to get down to the clusters that look real. > dmat <- as.dist((1-cor(ldata))/2) > hc <- hclust(dmat, average ) > hsil <- silhouette(cutree(hc, k=8), dmat) > summary(hsil)$avg.width [1] Why does the average silhouette width look so much better for this method?
29 APPLIED CLUSTERING 28 Hierarchical clustering: silhouette with singletons > plot(hsil)
30 APPLIED CLUSTERING 29 The silhouette width General conclusions: The average silhouette width is only a crude guide to the number of clusters present in the data. The silhouette plot seems to be more useful, but requires human intervention (in the form of cortical filtering ).
31 APPLIED CLUSTERING 30 The Gap statistic An alternative method for determining the number of clusters relies on the gap statistic. The idea is to run a clustering algorithm for different values of the number K of clusters. Let W (K) be the within-cluster error. In the case of Euclidean distance, W (K) is just the within-cluster sum of squares. For other distance measures, it is the term we minimized in the description of PAM. Because adding more clusters will always reduce this error term, W (K) is a decreasing function of K. However, it should decrease faster when K is less than the true number and slower when K is greater than the true number. The gap statistic measures the difference (on the log scale, for each K) between the observed W (K) and the expected value if the data were uniformly distributed. One then selects the K with the largest gap between the observed and expected values.
32 APPLIED CLUSTERING 31 Principal Components You may have wondered how we produced two-dimensional plots of the simulated data that involved 1000 genes and 53 samples. The short answer is: we used principal components analysis (PCA). As we have been doing throughout our discussion of clustering, we view each sample as a vector x = (x 1,..., x G ) in G-dimensional gene space. The idea behind PCA is to look for a direction (represented as a linear combination u 1 = G i=1 w ix i ) that maximizes the variability across the samples. Next, we find a second direction u 2 at right angles to the first that maximizes what remains of the variability. We keep repeating this process. The u i vectors are the principal components, and we can rewrite each sample vector as a sum of principal components instead of as a sum of separate gene expression values.
33 APPLIED CLUSTERING 32 Data reduction PCA can be used as a data reduction method. Changing from the original x-coordinates to the new u-coordinate system doesn t change the underlying structure of the sample vectors. However, it does let us focus on the directions where the data changes most rapidly. If we just use the first two or three principal components, we can produce plots that show us as much of the intrinsic variability in the data as possible. Warning: even though there is a function in R called princomp that is supposed to compute principal components, it will not work in the context of microarrays. The problem is that princomp wants to decompose the covariance matrix, which is a square matrix with size given by the number of genes. That s simply too big to manipulate.
34 APPLIED CLUSTERING 33 Sample PCA We have written a version of PCA in R using Singular Value Decomposition. The first plot of our simulated data was produced using the following commands: > spca <- SamplePCA(ldata) > plot(spca, split=group.factor) The plots that added the X s to mark the K-means centers were produced with: > plot(spca, split=factor(kres$cluster)) > x1 <- spca@scores[kcent,1] # start circles > x2 <- spca@scores[kcent,2] > points(x1, x2, col=2:6, pch=1, cex=2) > pcak <- predict(spca, t(kres$centers)) # finish X > points(pcak[,1], pcak[,2], col=2:6, pch=4)
35 APPLIED CLUSTERING 34 Principal Coordinates Using the first few principal components provides a view of the data that allows us to see as much of the variability as possible. Sometimes we have a different goal: we d like to be able to visualize the samples in a way that does as good a job as possible of preserving the distances between samples. In general, this method is called multidimensional scaling (MDS). The classical form of MDS is also known as principal coordinate analysis, and is implemented in R by the function cmdscale. If you look at the resulting graph carefully, you ll discover that it is identical to the principal components plot: classical MDS using Euclidean distance is equivalent to plotting the first few principal components.
36 APPLIED CLUSTERING 35 Euclidean classical MDS = PCA > euloc <- cmdscale(euc.distance) > plot(euloc[,1], euloc[,2], pch=16, + col=as.numeric(group.factor), + xlab= Coordinate 1, ylab= Coordinate 2 )
37 APPLIED CLUSTERING 36 Classical MDS with correlation > loc <- cmdscale(dmat) > plot(loc[,1], loc[,2], pch=16, + col=1+as.numeric(group.factor), + xlab= Coordinate 1, ylab= Coordinate 2 )
38 APPLIED CLUSTERING 37 Classical MDS with correlation Using the cloud function in the lattice package, we can also plot the first three principal coordinates: > require(lattice) > loc3d <- cmdscale(dmat, k=3) > pc1 <- loc3d[,1] > pc2 <- loc3d[,2] > pc3 <- loc3d[,3] > cloud(pc3 pc1*pc2, pch=16, cex=1.2, + col=1+as.numeric(group.factor), + screen=list(z=55, x=-70), + perspective=false)
39 APPLIED CLUSTERING 38 Three-D
40 APPLIED CLUSTERING 39 Project Normal Eighteen samples Six C57BL6 male mice Three organs: kidney, liver, testis Reference material Pool RNA from all eighteen mouse organs Replicate experiments on two-color arrays with common reference Four experiments per mouse organ Dye swaps: two red samples, two green samples
41 APPLIED CLUSTERING 40 Original analysis of Project Normal Reference: Pritchard, Hsu, Delrow, and Nelson. (2001) Project normal: defining normal variance in mouse gene expression. PNAS 98: Print-tip specific intensity dependent loess normalization Scale adjusted (using MAD) Work with log ratios (experimental/reference) Perform F-test for each gene to see if mouse-to-mouse variance exceeds the array-to-array variance.
42 APPLIED CLUSTERING 41 First steps We chose to process the data using a simple global normalization (to the 75 th percentile) instead of loess normalization, since we believed that the mixed reference RNA should have a different distribution of intensities than RNA from a single organ. We then transformed the intensities in each channel by computing their base-two logarithm. Main Question: Can we determine from the project normal data set which genes are specifically expressed in each of the three organs?
43 APPLIED CLUSTERING 42 Clustering Methods If we cluster the data, what should we expect to see? Which clustering method would be most appropriate for a first look at the data? Hierarchical clustering Partitioning around medoids K-means Multidimensional scaling Principal components analysis
44 APPLIED CLUSTERING 43 Hierarchical clustering Euclidean distance, average linkage Back to clustering methods
45 APPLIED CLUSTERING 44 Hierarchical clustering Correlation distance, average linkage Back to clustering methods
46 APPLIED CLUSTERING 45 Partitioning Around Medoids Euclidean distance, four clusters Back to clustering methods
47 APPLIED CLUSTERING 46 Partitioning Around Medoids Euclidean distance, five clusters Back to clustering methods
48 APPLIED CLUSTERING 47 Partitioning Around Medoids Correlation distance, four clusters Back to clustering methods
49 APPLIED CLUSTERING 48 Partitioning Around Medoids Correlation distance, five clusters Back to clustering methods
50 APPLIED CLUSTERING 49 Partitioning Around Medoids Correlation distance, six clusters Back to clustering methods
51 APPLIED CLUSTERING 50 K-means Number of channels in each cluster: Channel C1 C2 C3 C4 Experiment Reference Organ C1 C2 C3 C4 Kidney Liver Testis Best of 50 runs with four clusters Back to clustering methods
52 APPLIED CLUSTERING 51 K-means Number of channels in each cluster: Channel C1 C2 C3 C4 C5 Experiment Reference Best of 50 runs with five clusters Back to clustering methods Organ C1 C2 C3 C4 C5 Kidney Liver Testis
53 APPLIED CLUSTERING 52 Multidimensional Scaling Correlation distance, colored to indicate channel Back to clustering methods
54 APPLIED CLUSTERING 53 Multidimensional Scaling Correlation distance, colored to indicate organ Back to clustering methods
55 APPLIED CLUSTERING 54 Principal components analysis Euclidean distance, indicating channel and organ. Back to clustering methods Forward to second PCA
56 APPLIED CLUSTERING 55 Abnormal Behavior Regardless of which exploratory method we use to look at the data, we see that something strange is happening here. We might not have noticed this behavior if we had immediately gone to the log ratios instead of clustering the separate channels. What might explain the presence of two different kinds of reference channels? First thought: dye swaps. But this doesn t make sense, since then we would expect the experimental channels to split the same way (giving us eight clusters in total).
57 APPLIED CLUSTERING 56 Data merging Data was supplied in three files, one each for kidney, liver and testis. Each row in each file contained two kinds of annotations: 1. Location (block, row, and column) 2. Genetic material (IMAGE clone, UniGene ID) For our analysis, we merged the data using the annotations of genetic material. As it turns out, the locations did not agree So, we reordered data rows and merged on location...
58 APPLIED CLUSTERING 57 PCA after merging data on location Yuck. So why are most of the testis references so weird? Back to first PCA Forward to third PCA
59 APPLIED CLUSTERING 58 Inspired guessing When the gene annotations are matched Four of the testis reference channels are close to the kidney reference Twenty of the testis reference are close to the liver reference When the location annotations are matched Kidney, liver, and 4 testis references are close The other 20 testis reference are off by themselves Conclusion: A data processing error occurred partway through the testis experiments.
60 APPLIED CLUSTERING 59 Principal components, take 3 Finally, the picture we expected to start with! Back to second PCA
61 APPLIED CLUSTERING 60 Every solution creates a new problem Solution: After reordering all liver experiments and twenty testis experiments by location Can distinguish between the three organs The reference samples all cluster together New Problem: There are now two competing ways to map from locations to genetic annotations (one from the kidney data, one from the liver data). Which one is correct?
62 APPLIED CLUSTERING 61 How big is the problem? Microarray contains 5304 spots Only 3372 (63.6%) spots have UniGene annotations that are consistent across the files So, 1932 (36.4%) spots have ambiguous UniGene annotations
63 APPLIED CLUSTERING 62 UniGene Example
64 APPLIED CLUSTERING 63 UniGene Example
65 APPLIED CLUSTERING 64 Villin Expression
66 APPLIED CLUSTERING 65 Definition of abundance If the UniGene database entry for gene expression says that the cdna sources of the clones found in a cluster included kidney, then we will say that the gene is abundant in kidney. Analogous definitions obviously apply for liver, testis, or other organs.
67 APPLIED CLUSTERING 66 Abundance by consistency Abundance All UniGene Consistent Ambiguous None Kidney Liver Testis Kidney, Liver Kidney, Testis Liver, Testis All
68 APPLIED CLUSTERING 67 Combining UniGene abundance with microarray data For each gene Let I = (K, L, T ) be the binary vector of its abundance in three organs as recorded in the UniGene database. Let Y = (k, l, t) be the measured log intensity in the three organs. Model using a 3-dimensional multivariate normal distribution Y I = N 3 (µ I, Σ I ) Average replicate experiments from same mouse with same dye to produce natural triplets of measurements.
69 APPLIED CLUSTERING 68 Use consistently annotated genes to fit the model Abundance µ K µ L µ T None Kidney Liver Testis Kidney, Liver Kidney, Testis Liver, Testis All The estimates support the idea that (UniGene) abundant genes are expressed at higher levels than (UniGene) rare genes.
70 APPLIED CLUSTERING 69 Distinguishing between competing sets of annotations Use parameters estimated from the genes with consistent annotations At the ambiguous spots, compute the log-likelihood of the observed data for each possible triple of abundance annotations Given a complete set of annotations, sum the log-likelihood values over all genes Log-likelihood that the kidney data file contains the correct annotations is equal to 52, 241 Log-likelihood that the liver data file contains the correct annotations is equal to 60, 183
71 APPLIED CLUSTERING 70 Scrambled rows Our inspired guess earlier was motivated by the idea that the rows containing the annotations had somehow been reordered. We permuted the rows 100 times to obtain empirical p-values for the observed log-likelihoods P(kidney is correct) < 0.01 P(liver is correct) = The log-likelihood of the kidney file annotations was not particularly close to the maximum of 33, 491. This suggest that we can use the array data to refine the notion of abundance on a gene-by-gene basis.
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT MD Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org 19
GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT MD Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org 19
Exploratory data analysis for microarrays
 Exploratory data analysis for microarrays Jörg Rahnenführer Computational Biology and Applied Algorithmics Max Planck Institute for Informatics D-66123 Saarbrücken Germany NGFN - Courses in Practical DNA
Exploratory data analysis for microarrays Jörg Rahnenführer Computational Biology and Applied Algorithmics Max Planck Institute for Informatics D-66123 Saarbrücken Germany NGFN - Courses in Practical DNA
Cluster Analysis for Microarray Data
 Cluster Analysis for Microarray Data Seventh International Long Oligonucleotide Microarray Workshop Tucson, Arizona January 7-12, 2007 Dan Nettleton IOWA STATE UNIVERSITY 1 Clustering Group objects that
Cluster Analysis for Microarray Data Seventh International Long Oligonucleotide Microarray Workshop Tucson, Arizona January 7-12, 2007 Dan Nettleton IOWA STATE UNIVERSITY 1 Clustering Group objects that
University of California, Berkeley
 University of California, Berkeley U.C. Berkeley Division of Biostatistics Working Paper Series Year 2002 Paper 105 A New Partitioning Around Medoids Algorithm Mark J. van der Laan Katherine S. Pollard
University of California, Berkeley U.C. Berkeley Division of Biostatistics Working Paper Series Year 2002 Paper 105 A New Partitioning Around Medoids Algorithm Mark J. van der Laan Katherine S. Pollard
MSA220 - Statistical Learning for Big Data
 MSA220 - Statistical Learning for Big Data Lecture 13 Rebecka Jörnsten Mathematical Sciences University of Gothenburg and Chalmers University of Technology Clustering Explorative analysis - finding groups
MSA220 - Statistical Learning for Big Data Lecture 13 Rebecka Jörnsten Mathematical Sciences University of Gothenburg and Chalmers University of Technology Clustering Explorative analysis - finding groups
Clustering. Chapter 10 in Introduction to statistical learning
 Clustering Chapter 10 in Introduction to statistical learning 16 14 12 10 8 6 4 2 0 2 4 6 8 10 12 14 1 Clustering ² Clustering is the art of finding groups in data (Kaufman and Rousseeuw, 1990). ² What
Clustering Chapter 10 in Introduction to statistical learning 16 14 12 10 8 6 4 2 0 2 4 6 8 10 12 14 1 Clustering ² Clustering is the art of finding groups in data (Kaufman and Rousseeuw, 1990). ² What
Chapter 6 Continued: Partitioning Methods
 Chapter 6 Continued: Partitioning Methods Partitioning methods fix the number of clusters k and seek the best possible partition for that k. The goal is to choose the partition which gives the optimal
Chapter 6 Continued: Partitioning Methods Partitioning methods fix the number of clusters k and seek the best possible partition for that k. The goal is to choose the partition which gives the optimal
Network Traffic Measurements and Analysis
 DEIB - Politecnico di Milano Fall, 2017 Introduction Often, we have only a set of features x = x 1, x 2,, x n, but no associated response y. Therefore we are not interested in prediction nor classification,
DEIB - Politecnico di Milano Fall, 2017 Introduction Often, we have only a set of features x = x 1, x 2,, x n, but no associated response y. Therefore we are not interested in prediction nor classification,
Statistics 202: Data Mining. c Jonathan Taylor. Week 8 Based in part on slides from textbook, slides of Susan Holmes. December 2, / 1
 Week 8 Based in part on slides from textbook, slides of Susan Holmes December 2, 2012 1 / 1 Part I Clustering 2 / 1 Clustering Clustering Goal: Finding groups of objects such that the objects in a group
Week 8 Based in part on slides from textbook, slides of Susan Holmes December 2, 2012 1 / 1 Part I Clustering 2 / 1 Clustering Clustering Goal: Finding groups of objects such that the objects in a group
Multivariate analyses in ecology. Cluster (part 2) Ordination (part 1 & 2)
 Multivariate analyses in ecology Cluster (part 2) Ordination (part 1 & 2) 1 Exercise 9B - solut 2 Exercise 9B - solut 3 Exercise 9B - solut 4 Exercise 9B - solut 5 Multivariate analyses in ecology Cluster
Multivariate analyses in ecology Cluster (part 2) Ordination (part 1 & 2) 1 Exercise 9B - solut 2 Exercise 9B - solut 3 Exercise 9B - solut 4 Exercise 9B - solut 5 Multivariate analyses in ecology Cluster
Clustering and Visualisation of Data
 Clustering and Visualisation of Data Hiroshi Shimodaira January-March 28 Cluster analysis aims to partition a data set into meaningful or useful groups, based on distances between data points. In some
Clustering and Visualisation of Data Hiroshi Shimodaira January-March 28 Cluster analysis aims to partition a data set into meaningful or useful groups, based on distances between data points. In some
Clustering. CS294 Practical Machine Learning Junming Yin 10/09/06
 Clustering CS294 Practical Machine Learning Junming Yin 10/09/06 Outline Introduction Unsupervised learning What is clustering? Application Dissimilarity (similarity) of objects Clustering algorithm K-means,
Clustering CS294 Practical Machine Learning Junming Yin 10/09/06 Outline Introduction Unsupervised learning What is clustering? Application Dissimilarity (similarity) of objects Clustering algorithm K-means,
Clustering K-means. Machine Learning CSEP546 Carlos Guestrin University of Washington February 18, Carlos Guestrin
 Clustering K-means Machine Learning CSEP546 Carlos Guestrin University of Washington February 18, 2014 Carlos Guestrin 2005-2014 1 Clustering images Set of Images [Goldberger et al.] Carlos Guestrin 2005-2014
Clustering K-means Machine Learning CSEP546 Carlos Guestrin University of Washington February 18, 2014 Carlos Guestrin 2005-2014 1 Clustering images Set of Images [Goldberger et al.] Carlos Guestrin 2005-2014
10701 Machine Learning. Clustering
 171 Machine Learning Clustering What is Clustering? Organizing data into clusters such that there is high intra-cluster similarity low inter-cluster similarity Informally, finding natural groupings among
171 Machine Learning Clustering What is Clustering? Organizing data into clusters such that there is high intra-cluster similarity low inter-cluster similarity Informally, finding natural groupings among
CS 1675 Introduction to Machine Learning Lecture 18. Clustering. Clustering. Groups together similar instances in the data sample
 CS 1675 Introduction to Machine Learning Lecture 18 Clustering Milos Hauskrecht milos@cs.pitt.edu 539 Sennott Square Clustering Groups together similar instances in the data sample Basic clustering problem:
CS 1675 Introduction to Machine Learning Lecture 18 Clustering Milos Hauskrecht milos@cs.pitt.edu 539 Sennott Square Clustering Groups together similar instances in the data sample Basic clustering problem:
ECS 234: Data Analysis: Clustering ECS 234
 : Data Analysis: Clustering What is Clustering? Given n objects, assign them to groups (clusters) based on their similarity Unsupervised Machine Learning Class Discovery Difficult, and maybe ill-posed
: Data Analysis: Clustering What is Clustering? Given n objects, assign them to groups (clusters) based on their similarity Unsupervised Machine Learning Class Discovery Difficult, and maybe ill-posed
Summer School in Statistics for Astronomers & Physicists June 15-17, Cluster Analysis
 Summer School in Statistics for Astronomers & Physicists June 15-17, 2005 Session on Computational Algorithms for Astrostatistics Cluster Analysis Max Buot Department of Statistics Carnegie-Mellon University
Summer School in Statistics for Astronomers & Physicists June 15-17, 2005 Session on Computational Algorithms for Astrostatistics Cluster Analysis Max Buot Department of Statistics Carnegie-Mellon University
Clustering. Lecture 6, 1/24/03 ECS289A
 Clustering Lecture 6, 1/24/03 What is Clustering? Given n objects, assign them to groups (clusters) based on their similarity Unsupervised Machine Learning Class Discovery Difficult, and maybe ill-posed
Clustering Lecture 6, 1/24/03 What is Clustering? Given n objects, assign them to groups (clusters) based on their similarity Unsupervised Machine Learning Class Discovery Difficult, and maybe ill-posed
Gene Clustering & Classification
 BINF, Introduction to Computational Biology Gene Clustering & Classification Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Introduction to Gene Clustering
BINF, Introduction to Computational Biology Gene Clustering & Classification Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Introduction to Gene Clustering
How do microarrays work
 Lecture 3 (continued) Alvis Brazma European Bioinformatics Institute How do microarrays work condition mrna cdna hybridise to microarray condition Sample RNA extract labelled acid acid acid nucleic acid
Lecture 3 (continued) Alvis Brazma European Bioinformatics Institute How do microarrays work condition mrna cdna hybridise to microarray condition Sample RNA extract labelled acid acid acid nucleic acid
A new algorithm for hybrid hierarchical clustering with visualization and the bootstrap
 A new algorithm for hybrid hierarchical clustering with visualization and the bootstrap Mark J. van der Laan and Katherine S. Pollard Abstract We propose a hybrid clustering method, Hierarchical Ordered
A new algorithm for hybrid hierarchical clustering with visualization and the bootstrap Mark J. van der Laan and Katherine S. Pollard Abstract We propose a hybrid clustering method, Hierarchical Ordered
Cluster Analysis. Mu-Chun Su. Department of Computer Science and Information Engineering National Central University 2003/3/11 1
 Cluster Analysis Mu-Chun Su Department of Computer Science and Information Engineering National Central University 2003/3/11 1 Introduction Cluster analysis is the formal study of algorithms and methods
Cluster Analysis Mu-Chun Su Department of Computer Science and Information Engineering National Central University 2003/3/11 1 Introduction Cluster analysis is the formal study of algorithms and methods
CS 2750 Machine Learning. Lecture 19. Clustering. CS 2750 Machine Learning. Clustering. Groups together similar instances in the data sample
 Lecture 9 Clustering Milos Hauskrecht milos@cs.pitt.edu 539 Sennott Square Clustering Groups together similar instances in the data sample Basic clustering problem: distribute data into k different groups
Lecture 9 Clustering Milos Hauskrecht milos@cs.pitt.edu 539 Sennott Square Clustering Groups together similar instances in the data sample Basic clustering problem: distribute data into k different groups
Clustering. Informal goal. General types of clustering. Applications: Clustering in information search and analysis. Example applications in search
 Informal goal Clustering Given set of objects and measure of similarity between them, group similar objects together What mean by similar? What is good grouping? Computation time / quality tradeoff 1 2
Informal goal Clustering Given set of objects and measure of similarity between them, group similar objects together What mean by similar? What is good grouping? Computation time / quality tradeoff 1 2
Measure of Distance. We wish to define the distance between two objects Distance metric between points:
 Measure of Distance We wish to define the distance between two objects Distance metric between points: Euclidean distance (EUC) Manhattan distance (MAN) Pearson sample correlation (COR) Angle distance
Measure of Distance We wish to define the distance between two objects Distance metric between points: Euclidean distance (EUC) Manhattan distance (MAN) Pearson sample correlation (COR) Angle distance
Clustering. CE-717: Machine Learning Sharif University of Technology Spring Soleymani
 Clustering CE-717: Machine Learning Sharif University of Technology Spring 2016 Soleymani Outline Clustering Definition Clustering main approaches Partitional (flat) Hierarchical Clustering validation
Clustering CE-717: Machine Learning Sharif University of Technology Spring 2016 Soleymani Outline Clustering Definition Clustering main approaches Partitional (flat) Hierarchical Clustering validation
K-means clustering Based in part on slides from textbook, slides of Susan Holmes. December 2, Statistics 202: Data Mining.
 K-means clustering Based in part on slides from textbook, slides of Susan Holmes December 2, 2012 1 / 1 K-means Outline K-means, K-medoids Choosing the number of clusters: Gap test, silhouette plot. Mixture
K-means clustering Based in part on slides from textbook, slides of Susan Holmes December 2, 2012 1 / 1 K-means Outline K-means, K-medoids Choosing the number of clusters: Gap test, silhouette plot. Mixture
Cluster Analysis. Angela Montanari and Laura Anderlucci
 Cluster Analysis Angela Montanari and Laura Anderlucci 1 Introduction Clustering a set of n objects into k groups is usually moved by the aim of identifying internally homogenous groups according to a
Cluster Analysis Angela Montanari and Laura Anderlucci 1 Introduction Clustering a set of n objects into k groups is usually moved by the aim of identifying internally homogenous groups according to a
Medoid Partitioning. Chapter 447. Introduction. Dissimilarities. Types of Cluster Variables. Interval Variables. Ordinal Variables.
 Chapter 447 Introduction The objective of cluster analysis is to partition a set of objects into two or more clusters such that objects within a cluster are similar and objects in different clusters are
Chapter 447 Introduction The objective of cluster analysis is to partition a set of objects into two or more clusters such that objects within a cluster are similar and objects in different clusters are
Clustering and Dimensionality Reduction
 Clustering and Dimensionality Reduction Some material on these is slides borrowed from Andrew Moore's excellent machine learning tutorials located at: Data Mining Automatically extracting meaning from
Clustering and Dimensionality Reduction Some material on these is slides borrowed from Andrew Moore's excellent machine learning tutorials located at: Data Mining Automatically extracting meaning from
Linear and Non-linear Dimentionality Reduction Applied to Gene Expression Data of Cancer Tissue Samples
 Linear and Non-linear Dimentionality Reduction Applied to Gene Expression Data of Cancer Tissue Samples Franck Olivier Ndjakou Njeunje Applied Mathematics, Statistics, and Scientific Computation University
Linear and Non-linear Dimentionality Reduction Applied to Gene Expression Data of Cancer Tissue Samples Franck Olivier Ndjakou Njeunje Applied Mathematics, Statistics, and Scientific Computation University
Dimension reduction : PCA and Clustering
 Dimension reduction : PCA and Clustering By Hanne Jarmer Slides by Christopher Workman Center for Biological Sequence Analysis DTU The DNA Array Analysis Pipeline Array design Probe design Question Experimental
Dimension reduction : PCA and Clustering By Hanne Jarmer Slides by Christopher Workman Center for Biological Sequence Analysis DTU The DNA Array Analysis Pipeline Array design Probe design Question Experimental
Gene expression & Clustering (Chapter 10)
 Gene expression & Clustering (Chapter 10) Determining gene function Sequence comparison tells us if a gene is similar to another gene, e.g., in a new species Dynamic programming Approximate pattern matching
Gene expression & Clustering (Chapter 10) Determining gene function Sequence comparison tells us if a gene is similar to another gene, e.g., in a new species Dynamic programming Approximate pattern matching
STATS306B STATS306B. Clustering. Jonathan Taylor Department of Statistics Stanford University. June 3, 2010
 STATS306B Jonathan Taylor Department of Statistics Stanford University June 3, 2010 Spring 2010 Outline K-means, K-medoids, EM algorithm choosing number of clusters: Gap test hierarchical clustering spectral
STATS306B Jonathan Taylor Department of Statistics Stanford University June 3, 2010 Spring 2010 Outline K-means, K-medoids, EM algorithm choosing number of clusters: Gap test hierarchical clustering spectral
Clustering Algorithms. Margareta Ackerman
 Clustering Algorithms Margareta Ackerman A sea of algorithms As we discussed last class, there are MANY clustering algorithms, and new ones are proposed all the time. They are very different from each
Clustering Algorithms Margareta Ackerman A sea of algorithms As we discussed last class, there are MANY clustering algorithms, and new ones are proposed all the time. They are very different from each
Cluster Analysis. Summer School on Geocomputation. 27 June July 2011 Vysoké Pole
 Cluster Analysis Summer School on Geocomputation 27 June 2011 2 July 2011 Vysoké Pole Lecture delivered by: doc. Mgr. Radoslav Harman, PhD. Faculty of Mathematics, Physics and Informatics Comenius University,
Cluster Analysis Summer School on Geocomputation 27 June 2011 2 July 2011 Vysoké Pole Lecture delivered by: doc. Mgr. Radoslav Harman, PhD. Faculty of Mathematics, Physics and Informatics Comenius University,
Supervised vs. Unsupervised Learning
 Clustering Supervised vs. Unsupervised Learning So far we have assumed that the training samples used to design the classifier were labeled by their class membership (supervised learning) We assume now
Clustering Supervised vs. Unsupervised Learning So far we have assumed that the training samples used to design the classifier were labeled by their class membership (supervised learning) We assume now
Unsupervised Learning : Clustering
 Unsupervised Learning : Clustering Things to be Addressed Traditional Learning Models. Cluster Analysis K-means Clustering Algorithm Drawbacks of traditional clustering algorithms. Clustering as a complex
Unsupervised Learning : Clustering Things to be Addressed Traditional Learning Models. Cluster Analysis K-means Clustering Algorithm Drawbacks of traditional clustering algorithms. Clustering as a complex
1 Case study of SVM (Rob)
 DRAFT a final version will be posted shortly COS 424: Interacting with Data Lecturer: Rob Schapire and David Blei Lecture # 8 Scribe: Indraneel Mukherjee March 1, 2007 In the previous lecture we saw how
DRAFT a final version will be posted shortly COS 424: Interacting with Data Lecturer: Rob Schapire and David Blei Lecture # 8 Scribe: Indraneel Mukherjee March 1, 2007 In the previous lecture we saw how
Clustering. K-means clustering
 Clustering K-means clustering Clustering Motivation: Identify clusters of data points in a multidimensional space, i.e. partition the data set {x 1,...,x N } into K clusters. Intuition: A cluster is a
Clustering K-means clustering Clustering Motivation: Identify clusters of data points in a multidimensional space, i.e. partition the data set {x 1,...,x N } into K clusters. Intuition: A cluster is a
Week 7 Picturing Network. Vahe and Bethany
 Week 7 Picturing Network Vahe and Bethany Freeman (2005) - Graphic Techniques for Exploring Social Network Data The two main goals of analyzing social network data are identification of cohesive groups
Week 7 Picturing Network Vahe and Bethany Freeman (2005) - Graphic Techniques for Exploring Social Network Data The two main goals of analyzing social network data are identification of cohesive groups
Unsupervised Learning and Clustering
 Unsupervised Learning and Clustering Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2009 CS 551, Spring 2009 c 2009, Selim Aksoy (Bilkent University)
Unsupervised Learning and Clustering Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2009 CS 551, Spring 2009 c 2009, Selim Aksoy (Bilkent University)
9/29/13. Outline Data mining tasks. Clustering algorithms. Applications of clustering in biology
 9/9/ I9 Introduction to Bioinformatics, Clustering algorithms Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Outline Data mining tasks Predictive tasks vs descriptive tasks Example
9/9/ I9 Introduction to Bioinformatics, Clustering algorithms Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Outline Data mining tasks Predictive tasks vs descriptive tasks Example
Cluster Analysis and Visualization. Workshop on Statistics and Machine Learning 2004/2/6
 Cluster Analysis and Visualization Workshop on Statistics and Machine Learning 2004/2/6 Outlines Introduction Stages in Clustering Clustering Analysis and Visualization One/two-dimensional Data Histogram,
Cluster Analysis and Visualization Workshop on Statistics and Machine Learning 2004/2/6 Outlines Introduction Stages in Clustering Clustering Analysis and Visualization One/two-dimensional Data Histogram,
Clustering CS 550: Machine Learning
 Clustering CS 550: Machine Learning This slide set mainly uses the slides given in the following links: http://www-users.cs.umn.edu/~kumar/dmbook/ch8.pdf http://www-users.cs.umn.edu/~kumar/dmbook/dmslides/chap8_basic_cluster_analysis.pdf
Clustering CS 550: Machine Learning This slide set mainly uses the slides given in the following links: http://www-users.cs.umn.edu/~kumar/dmbook/ch8.pdf http://www-users.cs.umn.edu/~kumar/dmbook/dmslides/chap8_basic_cluster_analysis.pdf
Understanding Clustering Supervising the unsupervised
 Understanding Clustering Supervising the unsupervised Janu Verma IBM T.J. Watson Research Center, New York http://jverma.github.io/ jverma@us.ibm.com @januverma Clustering Grouping together similar data
Understanding Clustering Supervising the unsupervised Janu Verma IBM T.J. Watson Research Center, New York http://jverma.github.io/ jverma@us.ibm.com @januverma Clustering Grouping together similar data
CLUSTERING IN BIOINFORMATICS
 CLUSTERING IN BIOINFORMATICS CSE/BIMM/BENG 8 MAY 4, 0 OVERVIEW Define the clustering problem Motivation: gene expression and microarrays Types of clustering Clustering algorithms Other applications of
CLUSTERING IN BIOINFORMATICS CSE/BIMM/BENG 8 MAY 4, 0 OVERVIEW Define the clustering problem Motivation: gene expression and microarrays Types of clustering Clustering algorithms Other applications of
Lesson 3. Prof. Enza Messina
 Lesson 3 Prof. Enza Messina Clustering techniques are generally classified into these classes: PARTITIONING ALGORITHMS Directly divides data points into some prespecified number of clusters without a hierarchical
Lesson 3 Prof. Enza Messina Clustering techniques are generally classified into these classes: PARTITIONING ALGORITHMS Directly divides data points into some prespecified number of clusters without a hierarchical
Unsupervised Learning
 Unsupervised Learning Learning without Class Labels (or correct outputs) Density Estimation Learn P(X) given training data for X Clustering Partition data into clusters Dimensionality Reduction Discover
Unsupervised Learning Learning without Class Labels (or correct outputs) Density Estimation Learn P(X) given training data for X Clustering Partition data into clusters Dimensionality Reduction Discover
Computational Statistics The basics of maximum likelihood estimation, Bayesian estimation, object recognitions
 Computational Statistics The basics of maximum likelihood estimation, Bayesian estimation, object recognitions Thomas Giraud Simon Chabot October 12, 2013 Contents 1 Discriminant analysis 3 1.1 Main idea................................
Computational Statistics The basics of maximum likelihood estimation, Bayesian estimation, object recognitions Thomas Giraud Simon Chabot October 12, 2013 Contents 1 Discriminant analysis 3 1.1 Main idea................................
CSE 5243 INTRO. TO DATA MINING
 CSE 5243 INTRO. TO DATA MINING Cluster Analysis: Basic Concepts and Methods Huan Sun, CSE@The Ohio State University 09/25/2017 Slides adapted from UIUC CS412, Fall 2017, by Prof. Jiawei Han 2 Chapter 10.
CSE 5243 INTRO. TO DATA MINING Cluster Analysis: Basic Concepts and Methods Huan Sun, CSE@The Ohio State University 09/25/2017 Slides adapted from UIUC CS412, Fall 2017, by Prof. Jiawei Han 2 Chapter 10.
Supervised vs unsupervised clustering
 Classification Supervised vs unsupervised clustering Cluster analysis: Classes are not known a- priori. Classification: Classes are defined a-priori Sometimes called supervised clustering Extract useful
Classification Supervised vs unsupervised clustering Cluster analysis: Classes are not known a- priori. Classification: Classes are defined a-priori Sometimes called supervised clustering Extract useful
Multivariate Methods
 Multivariate Methods Cluster Analysis http://www.isrec.isb-sib.ch/~darlene/embnet/ Classification Historically, objects are classified into groups periodic table of the elements (chemistry) taxonomy (zoology,
Multivariate Methods Cluster Analysis http://www.isrec.isb-sib.ch/~darlene/embnet/ Classification Historically, objects are classified into groups periodic table of the elements (chemistry) taxonomy (zoology,
Supplementary text S6 Comparison studies on simulated data
 Supplementary text S Comparison studies on simulated data Peter Langfelder, Rui Luo, Michael C. Oldham, and Steve Horvath Corresponding author: shorvath@mednet.ucla.edu Overview In this document we illustrate
Supplementary text S Comparison studies on simulated data Peter Langfelder, Rui Luo, Michael C. Oldham, and Steve Horvath Corresponding author: shorvath@mednet.ucla.edu Overview In this document we illustrate
Cluster Analysis. Ying Shen, SSE, Tongji University
 Cluster Analysis Ying Shen, SSE, Tongji University Cluster analysis Cluster analysis groups data objects based only on the attributes in the data. The main objective is that The objects within a group
Cluster Analysis Ying Shen, SSE, Tongji University Cluster analysis Cluster analysis groups data objects based only on the attributes in the data. The main objective is that The objects within a group
HIERARCHICAL clustering analysis (or HCA) is an
 A Data-Driven Approach to Estimating the Number of Clusters in Hierarchical Clustering* Antoine E. Zambelli arxiv:1608.04700v1 [q-bio.qm] 16 Aug 2016 Abstract We propose two new methods for estimating
A Data-Driven Approach to Estimating the Number of Clusters in Hierarchical Clustering* Antoine E. Zambelli arxiv:1608.04700v1 [q-bio.qm] 16 Aug 2016 Abstract We propose two new methods for estimating
Micro-array Image Analysis using Clustering Methods
 Micro-array Image Analysis using Clustering Methods Mrs Rekha A Kulkarni PICT PUNE kulkarni_rekha@hotmail.com Abstract Micro-array imaging is an emerging technology and several experimental procedures
Micro-array Image Analysis using Clustering Methods Mrs Rekha A Kulkarni PICT PUNE kulkarni_rekha@hotmail.com Abstract Micro-array imaging is an emerging technology and several experimental procedures
Olmo S. Zavala Romero. Clustering Hierarchical Distance Group Dist. K-means. Center of Atmospheric Sciences, UNAM.
 Center of Atmospheric Sciences, UNAM November 16, 2016 Cluster Analisis Cluster analysis or clustering is the task of grouping a set of objects in such a way that objects in the same group (called a cluster)
Center of Atmospheric Sciences, UNAM November 16, 2016 Cluster Analisis Cluster analysis or clustering is the task of grouping a set of objects in such a way that objects in the same group (called a cluster)
AN IMPROVED HYBRIDIZED K- MEANS CLUSTERING ALGORITHM (IHKMCA) FOR HIGHDIMENSIONAL DATASET & IT S PERFORMANCE ANALYSIS
 AN IMPROVED HYBRIDIZED K- MEANS CLUSTERING ALGORITHM (IHKMCA) FOR HIGHDIMENSIONAL DATASET & IT S PERFORMANCE ANALYSIS H.S Behera Department of Computer Science and Engineering, Veer Surendra Sai University
AN IMPROVED HYBRIDIZED K- MEANS CLUSTERING ALGORITHM (IHKMCA) FOR HIGHDIMENSIONAL DATASET & IT S PERFORMANCE ANALYSIS H.S Behera Department of Computer Science and Engineering, Veer Surendra Sai University
Mixture Models and the EM Algorithm
 Mixture Models and the EM Algorithm Padhraic Smyth, Department of Computer Science University of California, Irvine c 2017 1 Finite Mixture Models Say we have a data set D = {x 1,..., x N } where x i is
Mixture Models and the EM Algorithm Padhraic Smyth, Department of Computer Science University of California, Irvine c 2017 1 Finite Mixture Models Say we have a data set D = {x 1,..., x N } where x i is
/ Computational Genomics. Normalization
 10-810 /02-710 Computational Genomics Normalization Genes and Gene Expression Technology Display of Expression Information Yeast cell cycle expression Experiments (over time) baseline expression program
10-810 /02-710 Computational Genomics Normalization Genes and Gene Expression Technology Display of Expression Information Yeast cell cycle expression Experiments (over time) baseline expression program
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 15: Microarray clustering http://compbio.pbworks.com/f/wood2.gif Some slides were adapted from Dr. Shaojie Zhang (University of Central Florida) Microarray
EECS730: Introduction to Bioinformatics Lecture 15: Microarray clustering http://compbio.pbworks.com/f/wood2.gif Some slides were adapted from Dr. Shaojie Zhang (University of Central Florida) Microarray
BCLUST -- A program to assess reliability of gene clusters from expression data by using consensus tree and bootstrap resampling method
 BCLUST -- A program to assess reliability of gene clusters from expression data by using consensus tree and bootstrap resampling method Introduction This program is developed in the lab of Hongyu Zhao
BCLUST -- A program to assess reliability of gene clusters from expression data by using consensus tree and bootstrap resampling method Introduction This program is developed in the lab of Hongyu Zhao
Machine Learning and Data Mining. Clustering (1): Basics. Kalev Kask
 Machine Learning and Data Mining Clustering (1): Basics Kalev Kask Unsupervised learning Supervised learning Predict target value ( y ) given features ( x ) Unsupervised learning Understand patterns of
Machine Learning and Data Mining Clustering (1): Basics Kalev Kask Unsupervised learning Supervised learning Predict target value ( y ) given features ( x ) Unsupervised learning Understand patterns of
Data Science and Statistics in Research: unlocking the power of your data Session 3.4: Clustering
 Data Science and Statistics in Research: unlocking the power of your data Session 3.4: Clustering 1/ 1 OUTLINE 2/ 1 Overview 3/ 1 CLUSTERING Clustering is a statistical technique which creates groupings
Data Science and Statistics in Research: unlocking the power of your data Session 3.4: Clustering 1/ 1 OUTLINE 2/ 1 Overview 3/ 1 CLUSTERING Clustering is a statistical technique which creates groupings
Multivariate Analysis
 Multivariate Analysis Cluster Analysis Prof. Dr. Anselmo E de Oliveira anselmo.quimica.ufg.br anselmo.disciplinas@gmail.com Unsupervised Learning Cluster Analysis Natural grouping Patterns in the data
Multivariate Analysis Cluster Analysis Prof. Dr. Anselmo E de Oliveira anselmo.quimica.ufg.br anselmo.disciplinas@gmail.com Unsupervised Learning Cluster Analysis Natural grouping Patterns in the data
MATH5745 Multivariate Methods Lecture 13
 MATH5745 Multivariate Methods Lecture 13 April 24, 2018 MATH5745 Multivariate Methods Lecture 13 April 24, 2018 1 / 33 Cluster analysis. Example: Fisher iris data Fisher (1936) 1 iris data consists of
MATH5745 Multivariate Methods Lecture 13 April 24, 2018 MATH5745 Multivariate Methods Lecture 13 April 24, 2018 1 / 33 Cluster analysis. Example: Fisher iris data Fisher (1936) 1 iris data consists of
High throughput Data Analysis 2. Cluster Analysis
 High throughput Data Analysis 2 Cluster Analysis Overview Why clustering? Hierarchical clustering K means clustering Issues with above two Other methods Quality of clustering results Introduction WHY DO
High throughput Data Analysis 2 Cluster Analysis Overview Why clustering? Hierarchical clustering K means clustering Issues with above two Other methods Quality of clustering results Introduction WHY DO
Lecture 4 Hierarchical clustering
 CSE : Unsupervised learning Spring 00 Lecture Hierarchical clustering. Multiple levels of granularity So far we ve talked about the k-center, k-means, and k-medoid problems, all of which involve pre-specifying
CSE : Unsupervised learning Spring 00 Lecture Hierarchical clustering. Multiple levels of granularity So far we ve talked about the k-center, k-means, and k-medoid problems, all of which involve pre-specifying
Distances, Clustering! Rafael Irizarry!
 Distances, Clustering! Rafael Irizarry! Heatmaps! Distance! Clustering organizes things that are close into groups! What does it mean for two genes to be close?! What does it mean for two samples to
Distances, Clustering! Rafael Irizarry! Heatmaps! Distance! Clustering organizes things that are close into groups! What does it mean for two genes to be close?! What does it mean for two samples to
TELCOM2125: Network Science and Analysis
 School of Information Sciences University of Pittsburgh TELCOM2125: Network Science and Analysis Konstantinos Pelechrinis Spring 2015 2 Part 4: Dividing Networks into Clusters The problem l Graph partitioning
School of Information Sciences University of Pittsburgh TELCOM2125: Network Science and Analysis Konstantinos Pelechrinis Spring 2015 2 Part 4: Dividing Networks into Clusters The problem l Graph partitioning
INF 4300 Classification III Anne Solberg The agenda today:
 INF 4300 Classification III Anne Solberg 28.10.15 The agenda today: More on estimating classifier accuracy Curse of dimensionality and simple feature selection knn-classification K-means clustering 28.10.15
INF 4300 Classification III Anne Solberg 28.10.15 The agenda today: More on estimating classifier accuracy Curse of dimensionality and simple feature selection knn-classification K-means clustering 28.10.15
Dimension Reduction CS534
 Dimension Reduction CS534 Why dimension reduction? High dimensionality large number of features E.g., documents represented by thousands of words, millions of bigrams Images represented by thousands of
Dimension Reduction CS534 Why dimension reduction? High dimensionality large number of features E.g., documents represented by thousands of words, millions of bigrams Images represented by thousands of
INF4820. Clustering. Erik Velldal. Nov. 17, University of Oslo. Erik Velldal INF / 22
 INF4820 Clustering Erik Velldal University of Oslo Nov. 17, 2009 Erik Velldal INF4820 1 / 22 Topics for Today More on unsupervised machine learning for data-driven categorization: clustering. The task
INF4820 Clustering Erik Velldal University of Oslo Nov. 17, 2009 Erik Velldal INF4820 1 / 22 Topics for Today More on unsupervised machine learning for data-driven categorization: clustering. The task
Clustering Gene Expression Data: Acknowledgement: Elizabeth Garrett-Mayer; Shirley Liu; Robert Tibshirani; Guenther Walther; Trevor Hastie
 Clustering Gene Expression Data: Acknowledgement: Elizabeth Garrett-Mayer; Shirley Liu; Robert Tibshirani; Guenther Walther; Trevor Hastie Data from Garber et al. PNAS (98), 2001. Clustering Clustering
Clustering Gene Expression Data: Acknowledgement: Elizabeth Garrett-Mayer; Shirley Liu; Robert Tibshirani; Guenther Walther; Trevor Hastie Data from Garber et al. PNAS (98), 2001. Clustering Clustering
Hard clustering. Each object is assigned to one and only one cluster. Hierarchical clustering is usually hard. Soft (fuzzy) clustering
 An unsupervised machine learning problem Grouping a set of objects in such a way that objects in the same group (a cluster) are more similar (in some sense or another) to each other than to those in other
An unsupervised machine learning problem Grouping a set of objects in such a way that objects in the same group (a cluster) are more similar (in some sense or another) to each other than to those in other
Chapter 6: Cluster Analysis
 Chapter 6: Cluster Analysis The major goal of cluster analysis is to separate individual observations, or items, into groups, or clusters, on the basis of the values for the q variables measured on each
Chapter 6: Cluster Analysis The major goal of cluster analysis is to separate individual observations, or items, into groups, or clusters, on the basis of the values for the q variables measured on each
BBS654 Data Mining. Pinar Duygulu. Slides are adapted from Nazli Ikizler
 BBS654 Data Mining Pinar Duygulu Slides are adapted from Nazli Ikizler 1 Classification Classification systems: Supervised learning Make a rational prediction given evidence There are several methods for
BBS654 Data Mining Pinar Duygulu Slides are adapted from Nazli Ikizler 1 Classification Classification systems: Supervised learning Make a rational prediction given evidence There are several methods for
CSE 5243 INTRO. TO DATA MINING
 CSE 5243 INTRO. TO DATA MINING Cluster Analysis: Basic Concepts and Methods Huan Sun, CSE@The Ohio State University Slides adapted from UIUC CS412, Fall 2017, by Prof. Jiawei Han 2 Chapter 10. Cluster
CSE 5243 INTRO. TO DATA MINING Cluster Analysis: Basic Concepts and Methods Huan Sun, CSE@The Ohio State University Slides adapted from UIUC CS412, Fall 2017, by Prof. Jiawei Han 2 Chapter 10. Cluster
Exploratory data analysis for microarrays
 Exploratory data analysis for microarrays Adrian Alexa Computational Biology and Applied Algorithmics Max Planck Institute for Informatics D-66123 Saarbrücken slides by Jörg Rahnenführer NGFN - Courses
Exploratory data analysis for microarrays Adrian Alexa Computational Biology and Applied Algorithmics Max Planck Institute for Informatics D-66123 Saarbrücken slides by Jörg Rahnenführer NGFN - Courses
Clustering Color/Intensity. Group together pixels of similar color/intensity.
 Clustering Color/Intensity Group together pixels of similar color/intensity. Agglomerative Clustering Cluster = connected pixels with similar color. Optimal decomposition may be hard. For example, find
Clustering Color/Intensity Group together pixels of similar color/intensity. Agglomerative Clustering Cluster = connected pixels with similar color. Optimal decomposition may be hard. For example, find
What to come. There will be a few more topics we will cover on supervised learning
 Summary so far Supervised learning learn to predict Continuous target regression; Categorical target classification Linear Regression Classification Discriminative models Perceptron (linear) Logistic regression
Summary so far Supervised learning learn to predict Continuous target regression; Categorical target classification Linear Regression Classification Discriminative models Perceptron (linear) Logistic regression
Note Set 4: Finite Mixture Models and the EM Algorithm
 Note Set 4: Finite Mixture Models and the EM Algorithm Padhraic Smyth, Department of Computer Science University of California, Irvine Finite Mixture Models A finite mixture model with K components, for
Note Set 4: Finite Mixture Models and the EM Algorithm Padhraic Smyth, Department of Computer Science University of California, Irvine Finite Mixture Models A finite mixture model with K components, for
Bumptrees for Efficient Function, Constraint, and Classification Learning
 umptrees for Efficient Function, Constraint, and Classification Learning Stephen M. Omohundro International Computer Science Institute 1947 Center Street, Suite 600 erkeley, California 94704 Abstract A
umptrees for Efficient Function, Constraint, and Classification Learning Stephen M. Omohundro International Computer Science Institute 1947 Center Street, Suite 600 erkeley, California 94704 Abstract A
Clustering (Basic concepts and Algorithms) Entscheidungsunterstützungssysteme
 Clustering (Basic concepts and Algorithms) Entscheidungsunterstützungssysteme Why do we need to find similarity? Similarity underlies many data science methods and solutions to business problems. Some
Clustering (Basic concepts and Algorithms) Entscheidungsunterstützungssysteme Why do we need to find similarity? Similarity underlies many data science methods and solutions to business problems. Some
Chemometrics. Description of Pirouette Algorithms. Technical Note. Abstract
 19-1214 Chemometrics Technical Note Description of Pirouette Algorithms Abstract This discussion introduces the three analysis realms available in Pirouette and briefly describes each of the algorithms
19-1214 Chemometrics Technical Note Description of Pirouette Algorithms Abstract This discussion introduces the three analysis realms available in Pirouette and briefly describes each of the algorithms
Data Exploration with PCA and Unsupervised Learning with Clustering Paul Rodriguez, PhD PACE SDSC
 Data Exploration with PCA and Unsupervised Learning with Clustering Paul Rodriguez, PhD PACE SDSC Clustering Idea Given a set of data can we find a natural grouping? Essential R commands: D =rnorm(12,0,1)
Data Exploration with PCA and Unsupervised Learning with Clustering Paul Rodriguez, PhD PACE SDSC Clustering Idea Given a set of data can we find a natural grouping? Essential R commands: D =rnorm(12,0,1)
Unsupervised Learning and Clustering
 Unsupervised Learning and Clustering Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2008 CS 551, Spring 2008 c 2008, Selim Aksoy (Bilkent University)
Unsupervised Learning and Clustering Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2008 CS 551, Spring 2008 c 2008, Selim Aksoy (Bilkent University)
Unsupervised Learning
 Unsupervised Learning Pierre Gaillard ENS Paris September 28, 2018 1 Supervised vs unsupervised learning Two main categories of machine learning algorithms: - Supervised learning: predict output Y from
Unsupervised Learning Pierre Gaillard ENS Paris September 28, 2018 1 Supervised vs unsupervised learning Two main categories of machine learning algorithms: - Supervised learning: predict output Y from
( ) =cov X Y = W PRINCIPAL COMPONENT ANALYSIS. Eigenvectors of the covariance matrix are the principal components
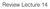 Review Lecture 14 ! PRINCIPAL COMPONENT ANALYSIS Eigenvectors of the covariance matrix are the principal components 1. =cov X Top K principal components are the eigenvectors with K largest eigenvalues
Review Lecture 14 ! PRINCIPAL COMPONENT ANALYSIS Eigenvectors of the covariance matrix are the principal components 1. =cov X Top K principal components are the eigenvectors with K largest eigenvalues
Introduction to Machine Learning CMU-10701
 Introduction to Machine Learning CMU-10701 Clustering and EM Barnabás Póczos & Aarti Singh Contents Clustering K-means Mixture of Gaussians Expectation Maximization Variational Methods 2 Clustering 3 K-
Introduction to Machine Learning CMU-10701 Clustering and EM Barnabás Póczos & Aarti Singh Contents Clustering K-means Mixture of Gaussians Expectation Maximization Variational Methods 2 Clustering 3 K-
Clustering. Supervised vs. Unsupervised Learning
 Clustering Supervised vs. Unsupervised Learning So far we have assumed that the training samples used to design the classifier were labeled by their class membership (supervised learning) We assume now
Clustering Supervised vs. Unsupervised Learning So far we have assumed that the training samples used to design the classifier were labeled by their class membership (supervised learning) We assume now
Today s lecture. Clustering and unsupervised learning. Hierarchical clustering. K-means, K-medoids, VQ
 Clustering CS498 Today s lecture Clustering and unsupervised learning Hierarchical clustering K-means, K-medoids, VQ Unsupervised learning Supervised learning Use labeled data to do something smart What
Clustering CS498 Today s lecture Clustering and unsupervised learning Hierarchical clustering K-means, K-medoids, VQ Unsupervised learning Supervised learning Use labeled data to do something smart What
Lecture 10: Semantic Segmentation and Clustering
 Lecture 10: Semantic Segmentation and Clustering Vineet Kosaraju, Davy Ragland, Adrien Truong, Effie Nehoran, Maneekwan Toyungyernsub Department of Computer Science Stanford University Stanford, CA 94305
Lecture 10: Semantic Segmentation and Clustering Vineet Kosaraju, Davy Ragland, Adrien Truong, Effie Nehoran, Maneekwan Toyungyernsub Department of Computer Science Stanford University Stanford, CA 94305
DATA MINING LECTURE 7. Hierarchical Clustering, DBSCAN The EM Algorithm
 DATA MINING LECTURE 7 Hierarchical Clustering, DBSCAN The EM Algorithm CLUSTERING What is a Clustering? In general a grouping of objects such that the objects in a group (cluster) are similar (or related)
DATA MINING LECTURE 7 Hierarchical Clustering, DBSCAN The EM Algorithm CLUSTERING What is a Clustering? In general a grouping of objects such that the objects in a group (cluster) are similar (or related)
3 Simultaneous Clustering Parameter
 1 Motivation Gene expression studies are swiftly becoming a very significant and prevalent tool in biomedical research. The microarray and gene-chip technologies allow researchers to monitor the expression
1 Motivation Gene expression studies are swiftly becoming a very significant and prevalent tool in biomedical research. The microarray and gene-chip technologies allow researchers to monitor the expression
Scaling Techniques in Political Science
 Scaling Techniques in Political Science Eric Guntermann March 14th, 2014 Eric Guntermann Scaling Techniques in Political Science March 14th, 2014 1 / 19 What you need R RStudio R code file Datasets You
Scaling Techniques in Political Science Eric Guntermann March 14th, 2014 Eric Guntermann Scaling Techniques in Political Science March 14th, 2014 1 / 19 What you need R RStudio R code file Datasets You
10601 Machine Learning. Hierarchical clustering. Reading: Bishop: 9-9.2
 161 Machine Learning Hierarchical clustering Reading: Bishop: 9-9.2 Second half: Overview Clustering - Hierarchical, semi-supervised learning Graphical models - Bayesian networks, HMMs, Reasoning under
161 Machine Learning Hierarchical clustering Reading: Bishop: 9-9.2 Second half: Overview Clustering - Hierarchical, semi-supervised learning Graphical models - Bayesian networks, HMMs, Reasoning under
Evaluation and comparison of gene clustering methods in microarray analysis
 Evaluation and comparison of gene clustering methods in microarray analysis Anbupalam Thalamuthu 1 Indranil Mukhopadhyay 1 Xiaojing Zheng 1 George C. Tseng 1,2 1 Department of Human Genetics 2 Department
Evaluation and comparison of gene clustering methods in microarray analysis Anbupalam Thalamuthu 1 Indranil Mukhopadhyay 1 Xiaojing Zheng 1 George C. Tseng 1,2 1 Department of Human Genetics 2 Department
Clustering and Dissimilarity Measures. Clustering. Dissimilarity Measures. Cluster Analysis. Perceptually-Inspired Measures
 Clustering and Dissimilarity Measures Clustering APR Course, Delft, The Netherlands Marco Loog May 19, 2008 1 What salient structures exist in the data? How many clusters? May 19, 2008 2 Cluster Analysis
Clustering and Dissimilarity Measures Clustering APR Course, Delft, The Netherlands Marco Loog May 19, 2008 1 What salient structures exist in the data? How many clusters? May 19, 2008 2 Cluster Analysis
