On Standardizing the MR Image Intensity Scale
|
|
|
- Pamela Bailey
- 5 years ago
- Views:
Transcription
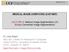
1 On Standardizing the MR Image Intensity Scale László G. Nyúl and Jayaram K. Udupa* Magnetic Resonance in Medicine 42: (1999) The lack of a standard image intensity scale in MRI causes many difficulties in image display and analysis. A two-step postprocessing method is proposed for standardizing the intensity scale in such a way that for the same MR protocol and body region, similar intensities will have similar tissue meaning. In the first step, the parameters of the standardizing transformation are learned from a set of images. In the second step, for each MR study these parameters are used to map their histogram into the standardized histogram. The method was tested quantitatively on 90 whole-brain studies of multiple sclerosis patients for several protocols and qualitatively for several other protocols and body regions. Measurements using mean squared difference showed that the standardized image intensities have statistically significantly (P F 0.01) more consistent range and meaning than the originals. Fixed gray level windows can be established for the standardized images and used for display without the need of per case adjustment. Preliminary results also indicate that the method facilitates improving the degree of automation of image segmentation. Magn Reson Med 42: , Wiley-Liss, Inc. Key words: image normalization; magnetic resonance imaging; image display; image processing Magnetic resonance imaging (MRI) has revolutionized radiological imaging of the internal structures of the human body. It has the advantage of being noninvasive, with no known health hazards. A variety of MRI protocols are available with or without the use of contrast agents. These protocols allow the setting up of different contrasts among the different tissues within the same organ system. Unfortunately, one of the major difficulties with MRI techniques has been that intensities do not have a fixed meaning, not even within the same protocol, for the same body region, for images obtained on the same scanner, for the same patient. This implies that MR images cannot be displayed at preset windows; one has to often adjust the window settings per case. The lack of a meaning for intensities also poses problems in image segmentation (1,2) and quantification (3 5), which are the main motivating factors for this work. Most visualization and analysis methods have parameters. Setting values for the parameters for these methods becomes more difficult without the same protocol-specific intensity meaning. What we need is that, for protocols that are the same or close to each other, the resulting images should be close (6). A standardizer can be incorporated at two stages of the image acquisition/processing flow. It can be built into the imaging device in order to produce images with a standard scale at the time of acquisition, or it can be used as a postprocessing step in a later phase. Attempts have been made to calibrate MR signal characteristics at the time of acquisition using phantoms (7,8). Although it is feasible to do such a calibration of all patient scans, it is somewhat cumbersome. Moreover, such a technique is not applicable to image data that have already been acquired without the required calibration phantoms. Postprocessing techniques that are applied to the image data that do not have any special acquisition requirements are, therefore, more attractive. There is a natural tendency to think that a simple scaling of the minimum to maximum intensity range of the given image to a fixed standard range may solve this problem. This usually does not help in achieving a similarity of intensities, as demonstrated in Nyúl and Udupa (9). A postprocessing technique to automatically adjust the contrast and brightness of MR images (i.e., windowing ) for image display has been presented in Wendt (10). However, although such automatic windowing may achieve display uniformity, they may not be adequate for quantitative image analysis, since the intensities still may not have tissue-specific meaning after the windowing transformation. There does not seem to have been any serious attempt to address this latter problem in the past. The method described in this article offers a simple way of transforming the images nonlinearly so that there is a significant gain in similarity of the resulting images. It is based on transforming the intensity histogram of each given volume image into a standard histogram. This is achieved in two steps a training step that is executed only once for a given protocol and body region and a transformation step that is executed for each given volume image. In the training step, certain landmarks of a standard histogram (for the body region and protocol under consideration) are estimated from a given set of volume images. In the transformation step, the actual intensity transformation from the intensity scale of the input volume image to the standard scale is computed by mapping the landmarks determined from the histogram of the given volume image to those of the standard histogram. The method is easy to implement and rapid in execution. The actual transformation itself can be stored as a lookup table in the image header. The transformed volume images permit predetermined display window settings and also facilitate quantitative image analysis. Medical Image Processing Group, Department of Radiology, University of Pennsylvania, Philadelphia, Pennsylvania. Grant sponsor: NIH; Grant number: NS 37172; Grant sponsor: Department of Army; Grant number: DAMD *Correspondence to: Jayaram K. Udupa, Medical Image Processing Group, Department of Radiology, University of Pennsylvania, 423 Guardian Drive, 4th Floor Blockley Hall, Philadelphia, PA jay@mipg.upenn.edu Received 18 May 1999; revised 20 July 1999; accepted 22 July Wiley-Liss, Inc THEORY AND ALGORITHMS The method consists of two steps. In the first ( training ) step, a set of volume images of the same body region and protocol corresponding to a population of patients is given as input. The parameters (landmarks) of a standard histogram are estimated from these image data. This step needs to be executed only once for a given protocol and
2 Standardizing the MR Image Intensity Scale 1073 body region. In the second ( transformation ) step, any given volume image acquired as per the protocol and for the body region utilized in the training step is transformed so that its histogram parameters match those of the standard histogram. In this fashion, every patient volume image histogram is deformed to match it with the standard histogram. This step is image-dependent and needs to be executed for each given volume image. This step usually results in a nonlinear intensity transformation for the given image (since the two segments are mapped independently). However, the relationship between tissue intensities is maintained and intensity comparisons can be made using the standardized images. Notation We denote the set of MRI protocols by P and the set of body regions by D. We represent a volume image by a pair V (V, g) where V is a 3-dimensional array of volume elements (voxels) covering a body region of the particular patient for whom image data V are acquired, and g is a function (called intensity function) that assigns an integer intensity value for each v V. (We will often refer to a volume image simply as an image for short. This is not to be confused with the 2D slice images. All processing operations described in this article are carried out in 3D.) We assume that g(v) 0 for all v V, and g(v) 0 if, and only if, there are no measured (and computed) data for voxel v. We denote by V PD the set of all images that can possibly be generated as per protocol P P for body region D D. The histogram of any image V is a pair H (G, h) where G is the set of all possible intensity values (gray values) in V (i.e., the range of g) and h is a function whose domain is G and whose value for each x G is the number of voxels v V for which g(v) x. Let m 1 min 5g(v)0v V and g(v) 06 and m 2 max 5g(v)0v V and g(v) 06, the minimum and maximum gray values in V, respectively. The tails of the histogram often cause problems. Usually the high intensity tail corresponds to artifacts and outlier intensities, and causes considerable inter- and intrapatient/ scanner variations. With this in mind, let pc 1 and pc 2 denote the minimum and maximum percentile values, respectively, that are used to select a range of intensity of interest (IOI). Algorithms Based on over 20 body region/protocol combinations, we have observed two types of histograms: unimodal and bimodal. We have also observed that the histograms of volume images of the same body region and protocol are always of the same type. Figure 1 illustrates them schematically. In case of bimodal histograms, we can usually use the mode (µ) that corresponds to the main foreground object in the image as a histogram landmark. With unimodal histograms, the mode usually corresponds to the background, so we need to select some other landmark. This may be, for example, the shoulder ( ) of the hump of the background intensities (identified by, for example, the point at which the histogram slope becomes 1). The locations of the histogram-specific parameters in these two cases are illustrated in Fig. 1. Since most of the protocols we studied produce bimodal histograms, we will concentrate on this FIG. 1. Location of the histogram-specific parameters (landmarks). m 1 and m 2 are the minimum and maximum intensities in the image, p 1 and p 2 are minimum and maximum percentile intensities, µ is the second mode of the histogram (in the bimodal case) and is the shoulder of the background hump (in the unimodal case). case in this article. Similar methods can be devised for the unimodal case. Our overall approach is as follows. Let the minimum and the maximum intensities on the standard scale (corresponding to the standard histogram) for the IOI be s 1 and s 2, respectively. In the training step, the landmarks p 1j, p 2j, and µ j obtained from the histogram of each image V j of a subset of images of V PD are mapped to the standard scale by mapping the intensities from [p 1j, p 2j ]to[s 1, s 2 ] linearly. The formula for mapping x [p 1j, p 2j ]tox is the following: x s 1 x p 1j p 2j p 1j (s 2 s 1 ). [1] This mapping is utilized to determine the map µ j of µ j on [s 1, s 2 ], and subsequently the rounded mean µ s of the µ j sis computed (see Algorithm 1, below). In the transformation step, for any given image V i (V i, g i ) V PD, the actual second mode µ i obtained (as described below) from the histogram of V i is matched to µ s by doing two separate
3 1074 Nyúl and Udupa linear mappings (see Algorithm 2): the first from [p 1i,µ i ]to [s 1,µ s ] and the second from [µ i, p 2i ]to[µ s, s 2 ]. Figure 2 shows the plot of the mapping function. The lower and the upper ends of the standard scale are subsequently extended to s 1i and s 2i, respectively, by mapping [m 1i, p 1i ]to [s 1i, s 1 ] and [p 2i, m 2i ]to[s 2, s 2i ], as illustrated in Fig. 2. We call this mapping from the intensities [m 1i, m 2i ]of V i to [s 1i, s 2i ] of the standard scale the standardizer of V i and denote it by Vi. The expression for Vi (from Fig. 2) is as follows: 5 aµ Vi (x) s 1 µ s s (x µ i ) p 1i µ i b, if m 1i x µ i, a µ s (x µ i ) s 2 µ s p 2i µ i b, if µ i x m 2i, [2] where a b denotes the ceiling or floor operator. (These operators convert any real number y to the closest integer Y such that Y y or Y y, respectively.) Note that s 1i V i (m 1i ), and s 2i V i (m 2i ). The image V si (V i, g si ) resulting from the standardizing mapping of V i is given by, for all v V i, g si (v) Vi (g i (v)). We point out that the free ends characterized by the values of s 1i and s 2i of the standard scale depend on the given image V i. In other words, the range [s 1i, s 2i ] may vary from image. However, [s 1, s 2 ]is independent of V i, and this is the interval within which a uniformity of intensity meaning is achieved. In order to find the second mode of the histogram, we eliminate the first mode (corresponding to the background) by thresholding. The threshold is chosen to be the overall mean intensity in the whole image. Several hundred studies have been successfully processed with this background removal process in our ongoing projects for different protocol and body region combinations. Algorithm 1. Training Input: A set of images V j (j 1,2,...,N) that is a subset of V PD, the histogram parameters pc 1, pc 2, and s 1, s 2. Output: µ s. begin 1. for j 1toNdo 2. compute the histogram H j of V j ; 3. determine intensity values p 1j and p 2j corresponding to pc 1 and pc 2 and the mode µ j on H j ; 4. map [p 1j, p 2j ]of H j onto [s 1, s 2 ]; 5. find the new mapped landmark location µ j ; 6. endfor 7. calculate the rounded mean µ s of µ j over j 1, 2,...,N; end Algorithm 2. Transformation Input: An image V i V PD, pc 1, pc 2, s 1, s 2,µ s. Output: The transformed image V si and/or a lookup table (LUT) that stores the standardizer Vi. begin 1. compute the histogram H i (G i, h i )of V i ; 2. determine intensity values p 1i and p 2i corresponding to pc 1 and pc 2 and the second mode µ i on H i ; 3. map sections of the scale of H i linearly according to Fig. 2 to the standard scale G s of the standard histogram H s (G s, h s ); 4. map the intensity value of every voxel v V i according to Vi to get V si and/or output the mapping Vi in a LUT; end Choosing the Standardization Parameters Although once the training step is completed the corresponding transformation step is fully determined, there are several possibilities to tailor the standardization to the specific needs of an application. The scale parameters may depend on the actual application. For example, s 1 should not be 0 if the values that are below pc 1 percentile need to be distinguished from nothing (i.e., value 0). Further, s 2 should be large enough not to compress the parts of the histograms (i.e., not to merge neighboring intensities after the transformation). When pc 1 0 and/or pc 2 100, the values in [m 1i, p 1i ] and [p 2i, m 2i ] must be mapped to [s 1i, s 1 ] and [s 2, s 2i ], respectively, where s 1i and s 2i are determined by applying the mapping in the two linear sections of Fig. 2. We will need the following definitions for stating certain theorems that guarantee the correct behavior of the standardizer. Let µ min min Vi V PD 5µ i 6, and µ max max V i V PD 5µ i 6. Note that µ min and µ max are dependent on s 1 and s 2. Let l, L, r, and R be indexes such that FIG. 2. The intensity mapping function for the transformation phase. µ l p 1l min V i V PD 5(µ i p 1i )6, [3]
4 Standardizing the MR Image Intensity Scale 1075 µ L p 1L max V i V PD 5(µ i p 1i )6, [4] p 2r µ r min V i V PD 5(p 2i µ i )6, [5] p 2R µ R max V i V PD 5(p 2i µ i )6. [6] We will assume throughout that pc 1 and pc 2 are such that, for any V i V PD, p 1i µ i p 2i. We state below three theorems which are crucial to guarantee the correct behavior of the standardizer Vi for any given image V i. Their complete proofs are given in Nyúl and Udupa (9). Theorem 1 states the conditions under which it is guaranteed that no two distinct intensities in V i are merged into a single intensity in the standardized image V si. Thus, if standardizing is done respecting these conditions, then there is no loss of information and the original image can be obtained by inverting the standardizer V i. Theorem 1. For any protocol P P, any body region D D, any image V i V PD, and for any pc 1 and pc 2, such that p 1i µ i p 2i, the standardizer Vi of V i is a one-to-one mapping if µ min s 1 µ L p 1L, and s 2 µ max p 2R µ R. The following theorem gives a guidance for selecting the values of s 1 and s 2 that cause no intensity loss. Theorem 2. For any protocol P P, any body region D D, any image V i V PD, and for any pc 1 and pc 2, such that p 1i µ i p 2i, the standardizer Vi of V i is a one-to-one mapping if s 2 s 1 (µ L p 1L p 2R µ R ) max (µ L p 1L )/(µ l p 1l ), (p 2R µ R )/(p 2r µ r ). Note that Theorems 1 and 2 state conditions that require observing all images in V PD. In practice, since this is impossible to do, we estimate the right side of in the expression in Theorem 2 by examining a sufficient number of volume images and set s 2 s 1 to a number sufficiently greater than this estimated entity. Our implemented software gives a warning message should an image be encountered for which this condition is violated. Even in such cases of violation, the software can be used in such way that, when a violation is detected, s 2 s 1 is automatically updated so that this condition is indeed satisfied. In testing 100 proton density (Pd) volume images of the brain of different patients, acquired as per a fixed fast-spinecho protocol on two GE 1.5T scanners, we found that µ L p 1L 1139, p 2R µ R 960, µ l p 1l 693, and p 2r µ r 502 for pc 1 0 and pc By the condition in Theorem 2, this implies that, with s 1 1, if we set s 2 to at least 4016, we can make sure that the mapping is lossless (though probably in many cases a smaller value would be sufficient). In all our experiments (and current use), we have set s This choice has been adequate in all protocols we currently use and is highly unlikely to lead to a lossy mapping at least for any proton density brain images acquired as per the above protocol. We arrived at the choice of pc by the following experiments. We randomly selected a few Pd brain volume images acquired as per the above protocol. With pc 1 0 fixed, we computed p 2i for several values of pc 2 (in steps of 0.05, between 99.0 and 100.0) for each test image V i.we chose the value of pc 2 to be the largest value at which the variation in p 2i with respect to changes in pc 2 reached an acceptably small value. Figure 3 shows the plot of the standard deviation of p 2i for the images in the test set versus pc 2 (Fig. 3a) and the derivative of that function (Fig. 3b). The value pc corresponds to the shoulder in the graph on the right where the function starts increasing rapidly. This implies that beyond this pc 2 value, the variations in the intensities are not systematic but random. Theorem 3 states that, once the conditions of Theorem 1 are satisfied, the order of the input intensities is maintained in the output images. Theorem 3. For any protocol P P, any body region D D, any image V i V PD, any standardizer Vi of V i that satisfies the conditions in Theorem 1, and for any intensities x 1 and x 2 of V i, Vi (x 1 ) Vi (x 2 ) if, and only if, x 1 x 2. That is, the actual order of brightness of tissue regions in V i is maintained in the image V si output by Algorithm 2, although their relative contrast may change. In a Pd image FIG. 3. A plot of the standard deviation of p 2i for the images in the test set vs. pc 2 (a), and its first derivative (b). pc is approximately the largest value at which the derivative is still small (0.5).
5 1076 Nyúl and Udupa of a brain, for example, the known brightness relationship gray matter white matter CSF is maintained in V si. EVALUATION Our hypothesis is that, for any given protocol P P, and for any body region D D, the standardized images V si have more consistent tissue meaning for image intensities than the images V i before standardization. That is, after standardization voxels having the same intensity value are more likely to contain the same kind of tissue. For testing this hypothesis, for each protocol P and body region D, we need to consider the following variations in image data: (i) intrapatient (time-to-time) variation, (ii) interpatient variation, (iii) variations among different machines of the same brand, and (iv) variations among machines of different brands. Verifying this hypothesis rigorously taking all these factors into account is indeed a formidable task. Instead, we will rigorously test the hypothesis for several protocols and for only one body region, namely the brain, and for the factors indicated in only (i) and (ii) above. We will also provide qualitative evidences for the validity of the hypothesis through display examples before and after standardization for factors indicated in (i), (ii), and (iii) above for several P and D. Qualitative Comparison We conducted qualitative comparisons for the following MRI protocols and body regions: fast spin-echo (FSE) Pd, FSE T 2, spin-echo (SE) Pd, SE T 2,T 1 with Gadolinium enhancement (T 1 E), and an SPGR sequence, all for the brain; a T 1 -weighted gradient-echo sequence for the foot. The training set consisted of 10 studies (by a study we mean a volume image) of different patients in all cases. There were 30 studies each of FSE Pd, FSE T 2, and T 1 E, and 10 studies each of SE Pd, SE T 2,T 1 GRE, and SPGR were transformed using the corresponding trained parameters. Images of all protocols had bimodal histograms except SPGR, which had unimodal histograms. In the latter case, a different landmark set was used in the standardization algorithm (see Fig. 1b), but the results are not discussed here because of space limitations. The parameters for training were chosen as follows. The low and high percentiles were set to pc 1 0 and pc Experiments were also done with other high percentiles (i.e., 100, 99.5, and 99.0). For the standard scale, we chose s 1 1, s Further, in all cases we made sure that the standardization mapping was lossless. In all cases, we derive the standardizer based on the whole volume image and not on the individual slices. Histograms Histograms of 10 FSE Pd studies at different stages of the transformation are displayed in Fig. 4. The low intensity part of the histograms that corresponds to the background voxels has been removed from the display in order to show the intensity of interest on a better scale. Original histograms of the volume images are plotted in Fig. 4a. Figure 4b displays the histograms after a simple linear mapping of [p 1i, p 2i ]to[s 1, s 2 ], with pc 1 0 and pc Histograms after the final standardizing mapping are plotted in Fig. FIG. 4. Histograms at different stages of the standardization process for 10 different FSE Pd studies. Original histograms (a), histograms after intensity scaling from [p 1i, p 2i ]to[s 1, s 2 ], with s 1 1, s (b), and after final standardization (c). 4c. A visual comparison shows that the histograms are more similar in shape and location after standardization than before. This implies that the actual intensity values and their distribution in the images are more similar after standardization. Note also that the simple linear mapping illustrated in the graph in Fig. 4b that accounts for outlier
6 Standardizing the MR Image Intensity Scale 1077 FIG. 5. Original slices from three studies acquired as per the same FSE Pd protocol before standardization displayed at a fixed window that was actually set up for the first image (first row), and after standardization displayed at a fixed standard window (second row). FIG. 6. Original slices from three studies acquired as per the same T 1 protocol with Gadolinium enhancement before standardization displayed at default windows (first row), and after standardization displayed at a fixed standard window (second row). intensities is also not enough in making the histograms cluster. Display at Fixed Window Images in the first row of Fig. 5 show a slice from each of three different patient FSE Pd studies, all acquired using the same protocol. They are all displayed at a fixed gray level window that was actually set up for the first image. This window does not seem to be appropriate for the other two datasets because they have quite different intensity ranges. In the second row of Fig. 5, the same slices are displayed, after standardization (always on the whole volume image) using pc , at a fixed standard brain window that we devised after examining a few standardized images. In Table 1, we list for several different protocols such standard window settings (level and width) that we have devised in this fashion. The structures are well portrayed and the brightness and contrast are more similar than that of the originals. A similar behavior was observed on FSE T 2, SE Pd, and SE T 2 images. Figure 6 is analogous to Fig. 5. It shows images before and after standardization for a T 1 protocol with Gadolinium enhancement. All images shown are acquired using the same protocol. The first row shows the original data displayed using the default window setting (i.e., window level set to the middle of the full range, and window width set to the full range of intensities of the study). The second row shows the same slices after standardization displayed using the standard brain window settings from Table 1. Figure 7 illustrates the improvements resulting from standardization for a different body region, namely the foot. The imaging protocol in this case was a T 1 -weighted gradientecho sequence and was the same for the three studies. The following example is included to demonstrate that the standardizer still works when substantial deviations exist in the test images compared to the images used for training. Figure 8 shows a dataset of the brain of a patient with a large tumor. The first row of Fig. 8 shows a slice of a FSE Pd and a FSE T 2 dataset before standardization and the second row shows the same slices after standardization. In this case, the standardizer was arrived at from the training data utilized for the multiple sclerosis (MS) patient FSE Pd (Fig. 5) and FSE T 2 images with a similar (but not identical) protocol. In Figure 9, we illustrate that the method is tolerant to small variations in protocol settings as well as variations that may exist among machines of the same brand in different hospitals. The figure shows three different patient studies obtained from a GE 1.5T Signa scanner at the University of Colorado Health Sciences Center (data courtesy of Dr. Jack Simon) using an SE Pd protocol. Images Table 1 Standard Windows for the Brain Images for Different Protocols Arrived at From the Standardized Images Level Width FSE Pd FSE T T 1 E SE Pd SE T Similar tables can be constructed for other body regions, protocols, and for specific tissue regions. FIG. 7. Original slices of three foot studies before (first row), and after standardization (second row). The imaging protocol for all three datasets was a T 1 -weighted gradient-echo sequence with identical parameters.
7 1078 Nyúl and Udupa FIG. 8. Original slices, before standardization, of FSE Pd (left) and T 2 (right) datasets of a patient s brain with a large tumor displayed at default windows (first row), and after standardization, displayed at a standard window setting (second row) listed in Table 1 for the FSE Pd and T 2 case. before standardization (displayed using the default window settings) are shown in the first row. The second row shows the same slices after standardization displayed by using the standard window setting from Table 1. In this case, training was done utilizing the SE Pd image data acquired on a scanner of the same brand in our hospital. Cross and Mixed Training The example illustrated in Fig. 10 shows that the method is robust against small variations in protocol settings. The first row of Fig. 10 shows three SE Pd studies before standardization (displayed with the default window setting). The second row shows the same slices after standardization with the parameters that were derived from a training set of an identical SE Pd protocol. These images are displayed with the standard window setting for SE Pd listed in Table 1. The third row shows the same slices after standardization with the parameters that were derived from a training set of images acquired as per an FSE FIG. 9. Three SE Pd studies acquired as per the same protocol before standardization at default windows (first row), and after standardization displayed at a standard window (second row). The training data were acquired as per a similar protocol (with slightly different parameters) from a scanner of the same brand at a different hospital. FIG. 10. Original slices from three studies acquired as per the same SE Pd protocol before standardization displayed at default windows (first row), the same slices displayed at a fixed standard window after standardization with the parameters that were derived from a training set of an identical SE Pd protocol (second row), and after standardization with the parameters that were derived from a training set of images acquired as per an FSE Pd protocol displayed at the standard window for FSE Pd (third row). Pd protocol. These images are displayed with the standard window setting for FSE Pd listed in Table 1. The main difference between the two mappings is a shift of the second mode. This is silently corrected for display when the proper window setting is applied. After both transformations and by using the corresponding window settings, the images look similar and have good brightness and contrast. The training based on a set of images of a given protocol P and body region D makes the standardizer tuned tightly to the images in V PD. This is due to the fact that the variation among histograms of images in V PD is less than that of images taken from V PD and V P D where P P and/or D D. However, images of different protocols and body regions can be mixed for training and still create a standardizer that fosters consistency of tissue meaning of intensities. This is illustrated in Fig. 11, where images from four different combinations of protocol and body region were utilized in the training step and in devising the standardizer. All images (FSE Pd, T 2,T 1 E brain, and GRE foot) were collected into a pool and were not distinguished during the training. That is, a single parameter configuration (i.e., pc 1 0, pc , s 1 1, and s ) were applied to all images in the training pool. The figure quite elegantly demonstrates that the standardized images show a better consistency of brightness and contrast at fixed window displays (bottom row) than the original images displayed at a default window for each image (top row). This observation invites the question whether it is possible to devise a whole-body standardizer which will obviate the need for protocol- and body-specific standardizers. We will not pursue this investigation here.
8 Standardizing the MR Image Intensity Scale 1079 FIG. 11. Illustration of the effect of mixed training. Ten brain studies from each of FSE Pd (first column), FSE T 2 (second column), and T 1 E (third column), and 10 T 1 -weighted gradient-echo studies of the foot (fourth column) were used in training. The first row shows the display at a study-specific default window of one slice of a study from each of these protocols before standardization. The second row shows the same slices after standardization displayed at one fixed window. Quantitative Comparison In order to assess the effectiveness of the standardization method objectively, we conducted two types of quantitative tests on datasets of brain obtained from three protocols: FSE Pd, FSE T 2, and T 1 E. The first test is for assessing intrapatient variation before and after standardization. In the second test, we used an algorithm (5) to segment different tissue regions and compared intensity statistics in different segmented tissue regions for assessing interpatient variation before and after standardization. Intrapatient Variation We used the same training datasets and parameter configurations as for qualitative comparison. The test method for all three protocols was the same. Two studies (volume images) acquired at different time instances were selected randomly for each of 15 MS patients from our database. The time distance between the two scans of the same patient varied between 1 and 6 years. For each patient, we registered the first scan to the second via a rigid transformation based on intensity value correlations. Because these patients had MS, the lesions were segmented (4,5) and removed for the purpose of comparison for minimizing variations in the two images of the same patient that may have been caused by the disease. The similarity of a pair of these registered, lesion-removed images was measured by the mean squared intensity difference between the two images normalized by the range of the image intensities of the older image in the pair. This similarity measure, denoted NMSD, was computed for every pair of volume images before and after intensity standardization for each of the three protocols. Table 2 shows that the mean and the standard deviation of the NMSD over all studies after standardization are smaller than that before standardization. The values of NMSD for the 15 pairs of studies were compared using a paired t-test under the null hypothesis that there is no difference in NMSD before and after standardization. The hypothesis was rejected at a significance level of P 0.01, indicating that the change in NMSD is statistically significant. Intensities in those parts of the image that have experienced no changes due to changes in the patient over time are made considerably more repeatable after standardization than before. Further, the lower standard deviation of NMSD achieved after standardization indicates that the uniformity of intensity meaning resulting from standardization is less dependent on the patient. Interpatient Variation For this comparison, we randomly selected 12 FSE Pd and 12 FSE T 2 volume images from our database. For each patient only one time instance is considered. All images were then segmented into white mater (WM), gray matter (GM), cerebrospinal fluid (CSF), and MS lesion (LS) regions (5). For each of these regions in each image V i in each of these protocols, we calculated the normalized mean intensity (NMI) by dividing the mean intensity in the region by p 2i p 1i. This was repeated for the standardized image V si of V i wherein normalization was done by dividing by s 2 s 1. The standard deviations of the NMI values over the 12 volume images before and after standardization for both protocols and for WM, GM, and CSF tissue regions are shown in Table 3, together with the corresponding 95% confidence intervals (assuming a normal distribution for all NMI values) and percent coefficient of variation (CV) of NMI. The results indicate that for every tissue region in both protocols tested, the standard deviation of NMI values is reduced by a factor of 2 to 3 after standardization and %CV is reduced considerably. This implies that a substantially improved uniformity of tissue meaning for intensities is obtained across patient studies after standardization. CONCLUDING REMARKS We have described a method for standardizing MR image intensity scales in this article. Our goal was to devise a method that post-hoc makes an intensity transformation of images that are routinely acquired, without requiring spe- Table 2 Mean and Standard Deviation of Normalized Mean Squared Differences (NMSD) Before and After Standardization for 15 Pairs of Studies and for Three Different Protocols FSE Pd FSE T 2 T 1 E Mean SD Mean SD Mean SD Before After P-value Each pair represents the studies obtained for the same patient at two time instances. The P-values of paired t-tests are also shown.
9 1080 Nyúl and Udupa Table 3 Standard Deviation, the 95% Confidence Interval (CI) of the Normalized Mean Intensity (NMI) Values and % Coefficient of Variation (CV) of Different Tissues in FSE Pd and T 2 Images FSE Pd FSE T 2 SD CI low CI high %CV SD CI low CI high %CV WM Before After GM Before After CSF Before After cialized acquisition protocols and calibration phantoms. The standardizing method aims at achieving consistency of tissue meaning of intensities by devising a transformation that is specific to a given MRI protocol and body region. The basic idea is to deform the histogram of a given volume image so that it matches a standard histogram for that protocol-body-region group. The parameters of the standard histogram are learned in a training step. We have provided theoretical guidelines, and a practical demonstration of how to utilize them, for selecting the values of the parameters of the method and proved that lossless intensity transformation and order is guaranteed, if choices are made as per guidelines. The choice of the actual landmarks is an important factor. Other landmarks (e.g., median instead of the mode of the second hump) can be used that are more suitable, or more sophisticated curve fitting can be done to make the histograms more similar. We mention some possible changes to the basic standardization method that can be incorporated to make it better fit the type of input data and the actual application: 1) use more histogram landmarks, such as quartiles and deciles, 2) use polynomial functions to stretch the histogram segments, and 3) use splinefitting techniques instead of segment-by-segment linear stretching. We have presented two types of studies to assess the degree of uniformity of intensity meaning achieved after standardization. In qualitative studies, we have shown through image display examples for several protocols and body regions that the consistency of the brightness level and contrast of images is considerably improved after standardization. This permits standardizing and fixing windows by protocol, body region, and tissue regions. This will minimize or eliminate the human interaction required in the per-case manual window adjustments that are currently required in visualizing MR images on physician viewing stations. This may also help in filming MR studies with a uniform appearance of the images. In quantitative studies, we have assessed the scannerdependent intra- and interpatient intensity variations and demonstrated that these are substantially reduced after standardization. In a subjective examination of the image displays from over 100 studies, we have not come across any case where the method seemed to have failed. A formal observer study is currently under way to assess this component. The method is simple, fast, easy to implement, and completely automatic. In a Picture Archiving and Communication System, it can be incorporated as a DICOM Value of Interest lookup table so that images are automatically transformed or accompanied by the correct lookup table when they are downloaded to the viewing station. It can even be built into the MR scanner to automatically produce images with the standard scale. Our preliminary studies indicate that image analysis and tissue segmentation methods our main motivation to undertake this research are considerably improved in terms of their constancy of parameter settings and their degree of automation. With standardization, numerical meaning is achieved and, hence, numerical diagnosis and study of diseases may become possible. For example, for MS studies we may be able to specify an interval [t 1, t 2 ]of the standardized Pd values which correspond to normal WM tissue, another interval [t 3, t 4 ] corresponding to WM that is normal appearing but that is actually affected by the disease. Our preliminary results indicate that when the values of the imaged parameter (T 2, Pd, etc.) do not intrinsically overlap for the different tissues, then such a numerical characterization may be possible. We use the standardization method as the first step in all MR image visualization and analysis tasks in which we are currently engaged. Figures 10 and 11 indicate a possible simplification of our approach. Instead of devising the standardization transform by protocol and body region, it may be possible to devise them based only on the type of the histogram (Fig. 1). That is, the protocol- and body-region-specific training and transformation will then be obviated, hopefully achieving consistency of intensity meaning independent of the protocol and body region. If we match the shoulder of the single mode in Fig. 1b with the second mode in Fig. 1a for images with different types of histograms, it may even be possible to devise standardization schemes that are independent of the histogram type. ACKNOWLEDGMENTS The authors thank Drs. Robert Grossman, Bruce Hirsch, and David Hackney for the MRI data sets utilized in this research.
10 Standardizing the MR Image Intensity Scale 1081 REFERENCES 1. Bezdek JC, Hall LO, Clarke LP. Review of MR image segmentation techniques using pattern-recognition. Med Phys 1993;20: Clarke LP, Velthuizen RP, Phuphanich S, Schellenberg JD, Arrington JA, Silbiger M. MRI: stability of three supervised segmentation techniques. Magn Reson Imaging 1993;11: Kikinis R, Shenton ME, Gerig G, Martin J, Anderson M, Metcalf D, Guttmann CRG, McCarley RW, Lorensen W, Cline H, Jolesz FA. Routine quantitative analysis of brain and cerebrospinal-fluid spaces with MR imaging. J Magn Reson Imaging 1992;2: Samarasekera S, Udupa JK, Miki Y, Grossman RI. A new computerassisted method for the quantification of enhancing lesions in multiple sclerosis. J Comput Assist Tomogr 1997;21: Udupa JK, Wei L, Samarasekera S, Miki Y, van Buchem MA, Grossman RI. Multiple sclerosis lesion quantification using fuzzy-connectedness principles. IEEE Trans Med Imaging 1997;16: Nyúl LG, Udupa JK. On standardizing the MR image intensity scale. Radiology 1998;209P: Edelstein WA, Bottomley PA, Pfeifer LM. A signal-to-noise calibration procedure for NMR imaging systems. Med Phys 1984;11: Yamamoto T, Nambu T, Miyasaka K, Morita Y. Accurate and practical calibration of MR signal intensities by the new transmission amplitude method: application of the numerical diagnosis to MRI. Radiology 1998;209P: Nyúl LG, Udupa JK. On standardizing the MR image intensity scale. Technical Report MIPG.250, Medical Image Processing Group, Department of Radiology, University of Pennsylvania, January Wendt RE. Automatic adjustment of contrast and brightness of magnetic resonance images. J Digit Imaging 1994;7:95 97.
Whole Body MRI Intensity Standardization
 Whole Body MRI Intensity Standardization Florian Jäger 1, László Nyúl 1, Bernd Frericks 2, Frank Wacker 2 and Joachim Hornegger 1 1 Institute of Pattern Recognition, University of Erlangen, {jaeger,nyul,hornegger}@informatik.uni-erlangen.de
Whole Body MRI Intensity Standardization Florian Jäger 1, László Nyúl 1, Bernd Frericks 2, Frank Wacker 2 and Joachim Hornegger 1 1 Institute of Pattern Recognition, University of Erlangen, {jaeger,nyul,hornegger}@informatik.uni-erlangen.de
INTERPOLATION is a commonly used operation in image
 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 18, NO. 2, FEBRUARY 1999 137 A Task-Specific Evaluation of Three-Dimensional Image Interpolation Techniques George J. Grevera, Jayaram K. Udupa,* Senior Member,
IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 18, NO. 2, FEBRUARY 1999 137 A Task-Specific Evaluation of Three-Dimensional Image Interpolation Techniques George J. Grevera, Jayaram K. Udupa,* Senior Member,
Chapter 3 Set Redundancy in Magnetic Resonance Brain Images
 16 Chapter 3 Set Redundancy in Magnetic Resonance Brain Images 3.1 MRI (magnetic resonance imaging) MRI is a technique of measuring physical structure within the human anatomy. Our proposed research focuses
16 Chapter 3 Set Redundancy in Magnetic Resonance Brain Images 3.1 MRI (magnetic resonance imaging) MRI is a technique of measuring physical structure within the human anatomy. Our proposed research focuses
New Technology Allows Multiple Image Contrasts in a Single Scan
 These images were acquired with an investigational device. PD T2 T2 FLAIR T1 MAP T1 FLAIR PSIR T1 New Technology Allows Multiple Image Contrasts in a Single Scan MR exams can be time consuming. A typical
These images were acquired with an investigational device. PD T2 T2 FLAIR T1 MAP T1 FLAIR PSIR T1 New Technology Allows Multiple Image Contrasts in a Single Scan MR exams can be time consuming. A typical
Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging
 1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
Learning to Identify Fuzzy Regions in Magnetic Resonance Images
 Learning to Identify Fuzzy Regions in Magnetic Resonance Images Sarah E. Crane and Lawrence O. Hall Department of Computer Science and Engineering, ENB 118 University of South Florida 4202 E. Fowler Ave.
Learning to Identify Fuzzy Regions in Magnetic Resonance Images Sarah E. Crane and Lawrence O. Hall Department of Computer Science and Engineering, ENB 118 University of South Florida 4202 E. Fowler Ave.
MEDICAL IMAGE COMPUTING (CAP 5937) LECTURE 9: Medical Image Segmentation (III) (Fuzzy Connected Image Segmentation)
 SPRING 2017 1 MEDICAL IMAGE COMPUTING (CAP 5937) LECTURE 9: Medical Image Segmentation (III) (Fuzzy Connected Image Segmentation) Dr. Ulas Bagci HEC 221, Center for Research in Computer Vision (CRCV),
SPRING 2017 1 MEDICAL IMAGE COMPUTING (CAP 5937) LECTURE 9: Medical Image Segmentation (III) (Fuzzy Connected Image Segmentation) Dr. Ulas Bagci HEC 221, Center for Research in Computer Vision (CRCV),
MEDICAL IMAGE COMPUTING (CAP 5937) LECTURE 4: Pre-Processing Medical Images (II)
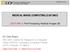 SPRING 2016 1 MEDICAL IMAGE COMPUTING (CAP 5937) LECTURE 4: Pre-Processing Medical Images (II) Dr. Ulas Bagci HEC 221, Center for Research in Computer Vision (CRCV), University of Central Florida (UCF),
SPRING 2016 1 MEDICAL IMAGE COMPUTING (CAP 5937) LECTURE 4: Pre-Processing Medical Images (II) Dr. Ulas Bagci HEC 221, Center for Research in Computer Vision (CRCV), University of Central Florida (UCF),
MR IMAGE SEGMENTATION
 MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
Classification of Abdominal Tissues by k-means Clustering for 3D Acoustic and Shear-Wave Modeling
 1 Classification of Abdominal Tissues by k-means Clustering for 3D Acoustic and Shear-Wave Modeling Kevin T. Looby klooby@stanford.edu I. ABSTRACT Clutter is an effect that degrades the quality of medical
1 Classification of Abdominal Tissues by k-means Clustering for 3D Acoustic and Shear-Wave Modeling Kevin T. Looby klooby@stanford.edu I. ABSTRACT Clutter is an effect that degrades the quality of medical
k-space Interpretation of the Rose Model: Noise Limitation on the Detectable Resolution in MRI
 k-space Interpretation of the Rose Model: Noise Limitation on the Detectable Resolution in MRI Richard Watts and Yi Wang* Magnetic Resonance in Medicine 48:550 554 (2002) Noise limitation on the detected
k-space Interpretation of the Rose Model: Noise Limitation on the Detectable Resolution in MRI Richard Watts and Yi Wang* Magnetic Resonance in Medicine 48:550 554 (2002) Noise limitation on the detected
JOURNAL OF INTERNATIONAL ACADEMIC RESEARCH FOR MULTIDISCIPLINARY Impact Factor 2.417, ISSN: , Volume 3, Issue 10, November 2015
 MEDICAL IMAGE PROCESSING ALGORITHM S.KRISHNAVENI* *Research Scholar, Dept. of Computer Science, Sengunthar Arts and Science College, Tiruchengode, Tamil Nadu, India ABSTRACT This paper makes a review on
MEDICAL IMAGE PROCESSING ALGORITHM S.KRISHNAVENI* *Research Scholar, Dept. of Computer Science, Sengunthar Arts and Science College, Tiruchengode, Tamil Nadu, India ABSTRACT This paper makes a review on
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit
 Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION
 ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION Abstract: MIP Project Report Spring 2013 Gaurav Mittal 201232644 This is a detailed report about the course project, which was to implement
ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION Abstract: MIP Project Report Spring 2013 Gaurav Mittal 201232644 This is a detailed report about the course project, which was to implement
Automatic Generation of Training Data for Brain Tissue Classification from MRI
 MICCAI-2002 1 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. Cocosco, Alex P. Zijdenbos, and Alan C. Evans McConnell Brain Imaging Centre, Montreal Neurological
MICCAI-2002 1 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. Cocosco, Alex P. Zijdenbos, and Alan C. Evans McConnell Brain Imaging Centre, Montreal Neurological
CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION
 CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION 6.1 INTRODUCTION Fuzzy logic based computational techniques are becoming increasingly important in the medical image analysis arena. The significant
CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION 6.1 INTRODUCTION Fuzzy logic based computational techniques are becoming increasingly important in the medical image analysis arena. The significant
ANALYSIS OF PULMONARY FIBROSIS IN MRI, USING AN ELASTIC REGISTRATION TECHNIQUE IN A MODEL OF FIBROSIS: Scleroderma
 ANALYSIS OF PULMONARY FIBROSIS IN MRI, USING AN ELASTIC REGISTRATION TECHNIQUE IN A MODEL OF FIBROSIS: Scleroderma ORAL DEFENSE 8 th of September 2017 Charlotte MARTIN Supervisor: Pr. MP REVEL M2 Bio Medical
ANALYSIS OF PULMONARY FIBROSIS IN MRI, USING AN ELASTIC REGISTRATION TECHNIQUE IN A MODEL OF FIBROSIS: Scleroderma ORAL DEFENSE 8 th of September 2017 Charlotte MARTIN Supervisor: Pr. MP REVEL M2 Bio Medical
CHAPTER 2. Morphometry on rodent brains. A.E.H. Scheenstra J. Dijkstra L. van der Weerd
 CHAPTER 2 Morphometry on rodent brains A.E.H. Scheenstra J. Dijkstra L. van der Weerd This chapter was adapted from: Volumetry and other quantitative measurements to assess the rodent brain, In vivo NMR
CHAPTER 2 Morphometry on rodent brains A.E.H. Scheenstra J. Dijkstra L. van der Weerd This chapter was adapted from: Volumetry and other quantitative measurements to assess the rodent brain, In vivo NMR
A Model-Independent, Multi-Image Approach to MR Inhomogeneity Correction
 Tina Memo No. 2007-003 Published in Proc. MIUA 2007 A Model-Independent, Multi-Image Approach to MR Inhomogeneity Correction P. A. Bromiley and N.A. Thacker Last updated 13 / 4 / 2007 Imaging Science and
Tina Memo No. 2007-003 Published in Proc. MIUA 2007 A Model-Independent, Multi-Image Approach to MR Inhomogeneity Correction P. A. Bromiley and N.A. Thacker Last updated 13 / 4 / 2007 Imaging Science and
Prepare a stem-and-leaf graph for the following data. In your final display, you should arrange the leaves for each stem in increasing order.
 Chapter 2 2.1 Descriptive Statistics A stem-and-leaf graph, also called a stemplot, allows for a nice overview of quantitative data without losing information on individual observations. It can be a good
Chapter 2 2.1 Descriptive Statistics A stem-and-leaf graph, also called a stemplot, allows for a nice overview of quantitative data without losing information on individual observations. It can be a good
surface Image reconstruction: 2D Fourier Transform
 2/1/217 Chapter 2-3 K-space Intro to k-space sampling (chap 3) Frequenc encoding and Discrete sampling (chap 2) Point Spread Function K-space properties K-space sampling principles (chap 3) Basic Contrast
2/1/217 Chapter 2-3 K-space Intro to k-space sampling (chap 3) Frequenc encoding and Discrete sampling (chap 2) Point Spread Function K-space properties K-space sampling principles (chap 3) Basic Contrast
CHAPTER 3 TUMOR DETECTION BASED ON NEURO-FUZZY TECHNIQUE
 32 CHAPTER 3 TUMOR DETECTION BASED ON NEURO-FUZZY TECHNIQUE 3.1 INTRODUCTION In this chapter we present the real time implementation of an artificial neural network based on fuzzy segmentation process
32 CHAPTER 3 TUMOR DETECTION BASED ON NEURO-FUZZY TECHNIQUE 3.1 INTRODUCTION In this chapter we present the real time implementation of an artificial neural network based on fuzzy segmentation process
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit
 Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Digital Volume Correlation for Materials Characterization
 19 th World Conference on Non-Destructive Testing 2016 Digital Volume Correlation for Materials Characterization Enrico QUINTANA, Phillip REU, Edward JIMENEZ, Kyle THOMPSON, Sharlotte KRAMER Sandia National
19 th World Conference on Non-Destructive Testing 2016 Digital Volume Correlation for Materials Characterization Enrico QUINTANA, Phillip REU, Edward JIMENEZ, Kyle THOMPSON, Sharlotte KRAMER Sandia National
Technical Publications
 GE Medical Systems Technical Publications Direction 2188003-100 Revision 0 Tissue Volume Analysis DICOM for DICOM V3.0 Copyright 1997 By General Electric Co. Do not duplicate REVISION HISTORY REV DATE
GE Medical Systems Technical Publications Direction 2188003-100 Revision 0 Tissue Volume Analysis DICOM for DICOM V3.0 Copyright 1997 By General Electric Co. Do not duplicate REVISION HISTORY REV DATE
RADIOMICS: potential role in the clinics and challenges
 27 giugno 2018 Dipartimento di Fisica Università degli Studi di Milano RADIOMICS: potential role in the clinics and challenges Dr. Francesca Botta Medical Physicist Istituto Europeo di Oncologia (Milano)
27 giugno 2018 Dipartimento di Fisica Università degli Studi di Milano RADIOMICS: potential role in the clinics and challenges Dr. Francesca Botta Medical Physicist Istituto Europeo di Oncologia (Milano)
Descriptive Statistics, Standard Deviation and Standard Error
 AP Biology Calculations: Descriptive Statistics, Standard Deviation and Standard Error SBI4UP The Scientific Method & Experimental Design Scientific method is used to explore observations and answer questions.
AP Biology Calculations: Descriptive Statistics, Standard Deviation and Standard Error SBI4UP The Scientific Method & Experimental Design Scientific method is used to explore observations and answer questions.
Computer-Aided Tracking of MS lesions
 Computer-Aided Tracking of MS lesions Deborah Sturm* 1, Devorah Gurwitz Kletenik 2 Phillip Koshy 3 1 Department of Computer Science, College of Staten Island, City University of New York, 2800 Victory
Computer-Aided Tracking of MS lesions Deborah Sturm* 1, Devorah Gurwitz Kletenik 2 Phillip Koshy 3 1 Department of Computer Science, College of Staten Island, City University of New York, 2800 Victory
Technical Publications
 g GE Medical Systems Technical Publications Direction 2275362-100 Revision 0 DICOM for DICOM V3.0 Copyright 2000 By General Electric Co. Do not duplicate REVISION HISTORY REV DATE REASON FOR CHANGE 0 May
g GE Medical Systems Technical Publications Direction 2275362-100 Revision 0 DICOM for DICOM V3.0 Copyright 2000 By General Electric Co. Do not duplicate REVISION HISTORY REV DATE REASON FOR CHANGE 0 May
Learning-based Neuroimage Registration
 Learning-based Neuroimage Registration Leonid Teverovskiy and Yanxi Liu 1 October 2004 CMU-CALD-04-108, CMU-RI-TR-04-59 School of Computer Science Carnegie Mellon University Pittsburgh, PA 15213 Abstract
Learning-based Neuroimage Registration Leonid Teverovskiy and Yanxi Liu 1 October 2004 CMU-CALD-04-108, CMU-RI-TR-04-59 School of Computer Science Carnegie Mellon University Pittsburgh, PA 15213 Abstract
Histograms. h(r k ) = n k. p(r k )= n k /NM. Histogram: number of times intensity level rk appears in the image
 Histograms h(r k ) = n k Histogram: number of times intensity level rk appears in the image p(r k )= n k /NM normalized histogram also a probability of occurence 1 Histogram of Image Intensities Create
Histograms h(r k ) = n k Histogram: number of times intensity level rk appears in the image p(r k )= n k /NM normalized histogram also a probability of occurence 1 Histogram of Image Intensities Create
arxiv: v1 [cs.cv] 11 Apr 2018
![arxiv: v1 [cs.cv] 11 Apr 2018 arxiv: v1 [cs.cv] 11 Apr 2018](/thumbs/83/88598484.jpg) Unsupervised Segmentation of 3D Medical Images Based on Clustering and Deep Representation Learning Takayasu Moriya a, Holger R. Roth a, Shota Nakamura b, Hirohisa Oda c, Kai Nagara c, Masahiro Oda a,
Unsupervised Segmentation of 3D Medical Images Based on Clustering and Deep Representation Learning Takayasu Moriya a, Holger R. Roth a, Shota Nakamura b, Hirohisa Oda c, Kai Nagara c, Masahiro Oda a,
Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00
 Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00 Marcel Prastawa 1 and Guido Gerig 1 Abstract July 17, 2008 1 Scientific Computing and Imaging
Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00 Marcel Prastawa 1 and Guido Gerig 1 Abstract July 17, 2008 1 Scientific Computing and Imaging
Chapter 1. Looking at Data-Distribution
 Chapter 1. Looking at Data-Distribution Statistics is the scientific discipline that provides methods to draw right conclusions: 1)Collecting the data 2)Describing the data 3)Drawing the conclusions Raw
Chapter 1. Looking at Data-Distribution Statistics is the scientific discipline that provides methods to draw right conclusions: 1)Collecting the data 2)Describing the data 3)Drawing the conclusions Raw
Supplementary methods
 Supplementary methods This section provides additional technical details on the sample, the applied imaging and analysis steps and methods. Structural imaging Trained radiographers placed all participants
Supplementary methods This section provides additional technical details on the sample, the applied imaging and analysis steps and methods. Structural imaging Trained radiographers placed all participants
Acquisition Description Exploration Examination Understanding what data is collected. Characterizing properties of data.
 Summary Statistics Acquisition Description Exploration Examination what data is collected Characterizing properties of data. Exploring the data distribution(s). Identifying data quality problems. Selecting
Summary Statistics Acquisition Description Exploration Examination what data is collected Characterizing properties of data. Exploring the data distribution(s). Identifying data quality problems. Selecting
Segmenting Glioma in Multi-Modal Images using a Generative Model for Brain Lesion Segmentation
 Segmenting Glioma in Multi-Modal Images using a Generative Model for Brain Lesion Segmentation Bjoern H. Menze 1,2, Koen Van Leemput 3, Danial Lashkari 4 Marc-André Weber 5, Nicholas Ayache 2, and Polina
Segmenting Glioma in Multi-Modal Images using a Generative Model for Brain Lesion Segmentation Bjoern H. Menze 1,2, Koen Van Leemput 3, Danial Lashkari 4 Marc-André Weber 5, Nicholas Ayache 2, and Polina
Segmenting Lesions in Multiple Sclerosis Patients James Chen, Jason Su
 Segmenting Lesions in Multiple Sclerosis Patients James Chen, Jason Su Radiologists and researchers spend countless hours tediously segmenting white matter lesions to diagnose and study brain diseases.
Segmenting Lesions in Multiple Sclerosis Patients James Chen, Jason Su Radiologists and researchers spend countless hours tediously segmenting white matter lesions to diagnose and study brain diseases.
Abstract. 1. Introduction
 A New Automated Method for Three- Dimensional Registration of Medical Images* P. Kotsas, M. Strintzis, D.W. Piraino Department of Electrical and Computer Engineering, Aristotelian University, 54006 Thessaloniki,
A New Automated Method for Three- Dimensional Registration of Medical Images* P. Kotsas, M. Strintzis, D.W. Piraino Department of Electrical and Computer Engineering, Aristotelian University, 54006 Thessaloniki,
Learner Expectations UNIT 1: GRAPICAL AND NUMERIC REPRESENTATIONS OF DATA. Sept. Fathom Lab: Distributions and Best Methods of Display
 CURRICULUM MAP TEMPLATE Priority Standards = Approximately 70% Supporting Standards = Approximately 20% Additional Standards = Approximately 10% HONORS PROBABILITY AND STATISTICS Essential Questions &
CURRICULUM MAP TEMPLATE Priority Standards = Approximately 70% Supporting Standards = Approximately 20% Additional Standards = Approximately 10% HONORS PROBABILITY AND STATISTICS Essential Questions &
Compressed Sensing Reconstructions for Dynamic Contrast Enhanced MRI
 1 Compressed Sensing Reconstructions for Dynamic Contrast Enhanced MRI Kevin T. Looby klooby@stanford.edu ABSTRACT The temporal resolution necessary for dynamic contrast enhanced (DCE) magnetic resonance
1 Compressed Sensing Reconstructions for Dynamic Contrast Enhanced MRI Kevin T. Looby klooby@stanford.edu ABSTRACT The temporal resolution necessary for dynamic contrast enhanced (DCE) magnetic resonance
Slide 1. Technical Aspects of Quality Control in Magnetic Resonance Imaging. Slide 2. Annual Compliance Testing. of MRI Systems.
 Slide 1 Technical Aspects of Quality Control in Magnetic Resonance Imaging Slide 2 Compliance Testing of MRI Systems, Ph.D. Department of Radiology Henry Ford Hospital, Detroit, MI Slide 3 Compliance Testing
Slide 1 Technical Aspects of Quality Control in Magnetic Resonance Imaging Slide 2 Compliance Testing of MRI Systems, Ph.D. Department of Radiology Henry Ford Hospital, Detroit, MI Slide 3 Compliance Testing
The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy
 The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy Sokratis K. Makrogiannis, PhD From post-doctoral research at SBIA lab, Department of Radiology,
The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy Sokratis K. Makrogiannis, PhD From post-doctoral research at SBIA lab, Department of Radiology,
Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE)
 Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE) Mahnaz Maddah, Kelly H. Zou, William M. Wells, Ron Kikinis, and Simon K. Warfield
Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE) Mahnaz Maddah, Kelly H. Zou, William M. Wells, Ron Kikinis, and Simon K. Warfield
Statistical Analysis of Metabolomics Data. Xiuxia Du Department of Bioinformatics & Genomics University of North Carolina at Charlotte
 Statistical Analysis of Metabolomics Data Xiuxia Du Department of Bioinformatics & Genomics University of North Carolina at Charlotte Outline Introduction Data pre-treatment 1. Normalization 2. Centering,
Statistical Analysis of Metabolomics Data Xiuxia Du Department of Bioinformatics & Genomics University of North Carolina at Charlotte Outline Introduction Data pre-treatment 1. Normalization 2. Centering,
NIH Public Access Author Manuscript Proc IEEE Int Symp Biomed Imaging. Author manuscript; available in PMC 2014 November 15.
 NIH Public Access Author Manuscript Published in final edited form as: Proc IEEE Int Symp Biomed Imaging. 2013 April ; 2013: 748 751. doi:10.1109/isbi.2013.6556583. BRAIN TUMOR SEGMENTATION WITH SYMMETRIC
NIH Public Access Author Manuscript Published in final edited form as: Proc IEEE Int Symp Biomed Imaging. 2013 April ; 2013: 748 751. doi:10.1109/isbi.2013.6556583. BRAIN TUMOR SEGMENTATION WITH SYMMETRIC
An ITK Filter for Bayesian Segmentation: itkbayesianclassifierimagefilter
 An ITK Filter for Bayesian Segmentation: itkbayesianclassifierimagefilter John Melonakos 1, Karthik Krishnan 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332, USA {jmelonak,
An ITK Filter for Bayesian Segmentation: itkbayesianclassifierimagefilter John Melonakos 1, Karthik Krishnan 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332, USA {jmelonak,
NIH Public Access Author Manuscript Proc SPIE. Author manuscript; available in PMC 2013 December 30.
 NIH Public Access Author Manuscript Published in final edited form as: Proc SPIE. 2013 March 12; 8669:. doi:10.1117/12.2006682. Longitudinal Intensity Normalization of Magnetic Resonance Images using Patches
NIH Public Access Author Manuscript Published in final edited form as: Proc SPIE. 2013 March 12; 8669:. doi:10.1117/12.2006682. Longitudinal Intensity Normalization of Magnetic Resonance Images using Patches
OPTIMIZING A VIDEO PREPROCESSOR FOR OCR. MR IBM Systems Dev Rochester, elopment Division Minnesota
 OPTIMIZING A VIDEO PREPROCESSOR FOR OCR MR IBM Systems Dev Rochester, elopment Division Minnesota Summary This paper describes how optimal video preprocessor performance can be achieved using a software
OPTIMIZING A VIDEO PREPROCESSOR FOR OCR MR IBM Systems Dev Rochester, elopment Division Minnesota Summary This paper describes how optimal video preprocessor performance can be achieved using a software
Fast Fuzzy Clustering of Infrared Images. 2. brfcm
 Fast Fuzzy Clustering of Infrared Images Steven Eschrich, Jingwei Ke, Lawrence O. Hall and Dmitry B. Goldgof Department of Computer Science and Engineering, ENB 118 University of South Florida 4202 E.
Fast Fuzzy Clustering of Infrared Images Steven Eschrich, Jingwei Ke, Lawrence O. Hall and Dmitry B. Goldgof Department of Computer Science and Engineering, ENB 118 University of South Florida 4202 E.
ECSE 626 Project Report Multimodality Image Registration by Maximization of Mutual Information
 ECSE 626 Project Report Multimodality Image Registration by Maximization of Mutual Information Emmanuel Piuze McGill University Montreal, Qc, Canada. epiuze@cim.mcgill.ca Abstract In 1997, Maes et al.
ECSE 626 Project Report Multimodality Image Registration by Maximization of Mutual Information Emmanuel Piuze McGill University Montreal, Qc, Canada. epiuze@cim.mcgill.ca Abstract In 1997, Maes et al.
3. Data Analysis and Statistics
 3. Data Analysis and Statistics 3.1 Visual Analysis of Data 3.2.1 Basic Statistics Examples 3.2.2 Basic Statistical Theory 3.3 Normal Distributions 3.4 Bivariate Data 3.1 Visual Analysis of Data Visual
3. Data Analysis and Statistics 3.1 Visual Analysis of Data 3.2.1 Basic Statistics Examples 3.2.2 Basic Statistical Theory 3.3 Normal Distributions 3.4 Bivariate Data 3.1 Visual Analysis of Data Visual
GENERAL AUTOMATED FLAW DETECTION SCHEME FOR NDE X-RAY IMAGES
 GENERAL AUTOMATED FLAW DETECTION SCHEME FOR NDE X-RAY IMAGES Karl W. Ulmer and John P. Basart Center for Nondestructive Evaluation Department of Electrical and Computer Engineering Iowa State University
GENERAL AUTOMATED FLAW DETECTION SCHEME FOR NDE X-RAY IMAGES Karl W. Ulmer and John P. Basart Center for Nondestructive Evaluation Department of Electrical and Computer Engineering Iowa State University
Image Enhancement 3-1
 Image Enhancement The goal of image enhancement is to improve the usefulness of an image for a given task, such as providing a more subjectively pleasing image for human viewing. In image enhancement,
Image Enhancement The goal of image enhancement is to improve the usefulness of an image for a given task, such as providing a more subjectively pleasing image for human viewing. In image enhancement,
Norbert Schuff VA Medical Center and UCSF
 Norbert Schuff Medical Center and UCSF Norbert.schuff@ucsf.edu Medical Imaging Informatics N.Schuff Course # 170.03 Slide 1/67 Objective Learn the principle segmentation techniques Understand the role
Norbert Schuff Medical Center and UCSF Norbert.schuff@ucsf.edu Medical Imaging Informatics N.Schuff Course # 170.03 Slide 1/67 Objective Learn the principle segmentation techniques Understand the role
3 Graphical Displays of Data
 3 Graphical Displays of Data Reading: SW Chapter 2, Sections 1-6 Summarizing and Displaying Qualitative Data The data below are from a study of thyroid cancer, using NMTR data. The investigators looked
3 Graphical Displays of Data Reading: SW Chapter 2, Sections 1-6 Summarizing and Displaying Qualitative Data The data below are from a study of thyroid cancer, using NMTR data. The investigators looked
COMPREHENSIVE QUALITY CONTROL OF NMR TOMOGRAPHY USING 3D PRINTED PHANTOM
 COMPREHENSIVE QUALITY CONTROL OF NMR TOMOGRAPHY USING 3D PRINTED PHANTOM Mažena MACIUSOVIČ *, Marius BURKANAS *, Jonas VENIUS *, ** * Medical Physics Department, National Cancer Institute, Vilnius, Lithuania
COMPREHENSIVE QUALITY CONTROL OF NMR TOMOGRAPHY USING 3D PRINTED PHANTOM Mažena MACIUSOVIČ *, Marius BURKANAS *, Jonas VENIUS *, ** * Medical Physics Department, National Cancer Institute, Vilnius, Lithuania
Fiber Selection from Diffusion Tensor Data based on Boolean Operators
 Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhof 1, G. Greiner 2, M. Buchfelder 3, C. Nimsky 4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhof 1, G. Greiner 2, M. Buchfelder 3, C. Nimsky 4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Optimizing Flip Angle Selection in Breast MRI for Accurate Extraction and Visualization of T1 Tissue Relaxation Time
 Optimizing Flip Angle Selection in Breast MRI for Accurate Extraction and Visualization of T1 Tissue Relaxation Time GEORGIOS KETSETZIS AND MICHAEL BRADY Medical Vision Laboratory Department of Engineering
Optimizing Flip Angle Selection in Breast MRI for Accurate Extraction and Visualization of T1 Tissue Relaxation Time GEORGIOS KETSETZIS AND MICHAEL BRADY Medical Vision Laboratory Department of Engineering
Vocabulary. 5-number summary Rule. Area principle. Bar chart. Boxplot. Categorical data condition. Categorical variable.
 5-number summary 68-95-99.7 Rule Area principle Bar chart Bimodal Boxplot Case Categorical data Categorical variable Center Changing center and spread Conditional distribution Context Contingency table
5-number summary 68-95-99.7 Rule Area principle Bar chart Bimodal Boxplot Case Categorical data Categorical variable Center Changing center and spread Conditional distribution Context Contingency table
A MORPHOLOGY-BASED FILTER STRUCTURE FOR EDGE-ENHANCING SMOOTHING
 Proceedings of the 1994 IEEE International Conference on Image Processing (ICIP-94), pp. 530-534. (Austin, Texas, 13-16 November 1994.) A MORPHOLOGY-BASED FILTER STRUCTURE FOR EDGE-ENHANCING SMOOTHING
Proceedings of the 1994 IEEE International Conference on Image Processing (ICIP-94), pp. 530-534. (Austin, Texas, 13-16 November 1994.) A MORPHOLOGY-BASED FILTER STRUCTURE FOR EDGE-ENHANCING SMOOTHING
Automatic Quantification of DTI Parameters along Fiber Bundles
 Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein 1, Simon Hermann 1, Olaf Konrad 1, Horst K. Hahn 1, and Heinz-Otto Peitgen 1 1 MeVis Research, 28359 Bremen Email: klein@mevis.de
Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein 1, Simon Hermann 1, Olaf Konrad 1, Horst K. Hahn 1, and Heinz-Otto Peitgen 1 1 MeVis Research, 28359 Bremen Email: klein@mevis.de
2.1 Signal Production. RF_Coil. Scanner. Phantom. Image. Image Production
 An Extensible MRI Simulator for Post-Processing Evaluation Remi K.-S. Kwan?, Alan C. Evans, and G. Bruce Pike McConnell Brain Imaging Centre, Montreal Neurological Institute, McGill University, Montreal,
An Extensible MRI Simulator for Post-Processing Evaluation Remi K.-S. Kwan?, Alan C. Evans, and G. Bruce Pike McConnell Brain Imaging Centre, Montreal Neurological Institute, McGill University, Montreal,
mritc: A Package for MRI Tissue Classification
 mritc: A Package for MRI Tissue Classification Dai Feng 1 Luke Tierney 2 1 Merck Research Labratories 2 University of Iowa July 2010 Feng & Tierney (Merck & U of Iowa) MRI Tissue Classification July 2010
mritc: A Package for MRI Tissue Classification Dai Feng 1 Luke Tierney 2 1 Merck Research Labratories 2 University of Iowa July 2010 Feng & Tierney (Merck & U of Iowa) MRI Tissue Classification July 2010
A Virtual MR Scanner for Education
 A Virtual MR Scanner for Education Hackländer T, Schalla C, Trümper A, Mertens H, Hiltner J, Cramer BM Hospitals of the University Witten/Herdecke, Department of Radiology Wuppertal, Germany Purpose A
A Virtual MR Scanner for Education Hackländer T, Schalla C, Trümper A, Mertens H, Hiltner J, Cramer BM Hospitals of the University Witten/Herdecke, Department of Radiology Wuppertal, Germany Purpose A
Unsupervised Learning : Clustering
 Unsupervised Learning : Clustering Things to be Addressed Traditional Learning Models. Cluster Analysis K-means Clustering Algorithm Drawbacks of traditional clustering algorithms. Clustering as a complex
Unsupervised Learning : Clustering Things to be Addressed Traditional Learning Models. Cluster Analysis K-means Clustering Algorithm Drawbacks of traditional clustering algorithms. Clustering as a complex
Chapter 2 Describing, Exploring, and Comparing Data
 Slide 1 Chapter 2 Describing, Exploring, and Comparing Data Slide 2 2-1 Overview 2-2 Frequency Distributions 2-3 Visualizing Data 2-4 Measures of Center 2-5 Measures of Variation 2-6 Measures of Relative
Slide 1 Chapter 2 Describing, Exploring, and Comparing Data Slide 2 2-1 Overview 2-2 Frequency Distributions 2-3 Visualizing Data 2-4 Measures of Center 2-5 Measures of Variation 2-6 Measures of Relative
Generalized Scale-Based Image Filtering
 Generalized Scale-Based Image Filtering Andre Souza a, Jayaram K. Udupa a, and Anant Madabhushi a a Medical Image Processing Group, Department of Radiology University of Pennsylvania, Philadelphia, PA
Generalized Scale-Based Image Filtering Andre Souza a, Jayaram K. Udupa a, and Anant Madabhushi a a Medical Image Processing Group, Department of Radiology University of Pennsylvania, Philadelphia, PA
Week 2: Frequency distributions
 Types of data Health Sciences M.Sc. Programme Applied Biostatistics Week 2: distributions Data can be summarised to help to reveal information they contain. We do this by calculating numbers from the data
Types of data Health Sciences M.Sc. Programme Applied Biostatistics Week 2: distributions Data can be summarised to help to reveal information they contain. We do this by calculating numbers from the data
Atlas Based Segmentation of the prostate in MR images
 Atlas Based Segmentation of the prostate in MR images Albert Gubern-Merida and Robert Marti Universitat de Girona, Computer Vision and Robotics Group, Girona, Spain {agubern,marly}@eia.udg.edu Abstract.
Atlas Based Segmentation of the prostate in MR images Albert Gubern-Merida and Robert Marti Universitat de Girona, Computer Vision and Robotics Group, Girona, Spain {agubern,marly}@eia.udg.edu Abstract.
Brain Portion Peeling from T2 Axial MRI Head Scans using Clustering and Morphological Operation
 159 Brain Portion Peeling from T2 Axial MRI Head Scans using Clustering and Morphological Operation K. Somasundaram Image Processing Lab Dept. of Computer Science and Applications Gandhigram Rural Institute
159 Brain Portion Peeling from T2 Axial MRI Head Scans using Clustering and Morphological Operation K. Somasundaram Image Processing Lab Dept. of Computer Science and Applications Gandhigram Rural Institute
Simon K. Warfield, Michael Kaus, Ferenc A. Jolesz and Ron Kikinis
 Medical Image Analysis (2000) volume 4, number 1, pp 43 55 c Elsevier Science BV Adaptive, Template Moderated, Spatially Varying Statistical Classification Simon K. Warfield, Michael Kaus, Ferenc A. Jolesz
Medical Image Analysis (2000) volume 4, number 1, pp 43 55 c Elsevier Science BV Adaptive, Template Moderated, Spatially Varying Statistical Classification Simon K. Warfield, Michael Kaus, Ferenc A. Jolesz
Comparison Study of Clinical 3D MRI Brain Segmentation Evaluation
 Comparison Study of Clinical 3D MRI Brain Segmentation Evaluation Ting Song 1, Elsa D. Angelini 2, Brett D. Mensh 3, Andrew Laine 1 1 Heffner Biomedical Imaging Laboratory Department of Biomedical Engineering,
Comparison Study of Clinical 3D MRI Brain Segmentation Evaluation Ting Song 1, Elsa D. Angelini 2, Brett D. Mensh 3, Andrew Laine 1 1 Heffner Biomedical Imaging Laboratory Department of Biomedical Engineering,
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge
 Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
Effect of age and dementia on topology of brain functional networks. Paul McCarthy, Luba Benuskova, Liz Franz University of Otago, New Zealand
 Effect of age and dementia on topology of brain functional networks Paul McCarthy, Luba Benuskova, Liz Franz University of Otago, New Zealand 1 Structural changes in aging brain Age-related changes in
Effect of age and dementia on topology of brain functional networks Paul McCarthy, Luba Benuskova, Liz Franz University of Otago, New Zealand 1 Structural changes in aging brain Age-related changes in
Automatic Generation of Training Data for Brain Tissue Classification from MRI
 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. COCOSCO, Alex P. ZIJDENBOS, and Alan C. EVANS http://www.bic.mni.mcgill.ca/users/crisco/ McConnell Brain Imaging
Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. COCOSCO, Alex P. ZIJDENBOS, and Alan C. EVANS http://www.bic.mni.mcgill.ca/users/crisco/ McConnell Brain Imaging
Lecture 4. Digital Image Enhancement. 1. Principle of image enhancement 2. Spatial domain transformation. Histogram processing
 Lecture 4 Digital Image Enhancement 1. Principle of image enhancement 2. Spatial domain transformation Basic intensity it tranfomation ti Histogram processing Principle Objective of Enhancement Image enhancement
Lecture 4 Digital Image Enhancement 1. Principle of image enhancement 2. Spatial domain transformation Basic intensity it tranfomation ti Histogram processing Principle Objective of Enhancement Image enhancement
Improved Spatial Localization in 3D MRSI with a Sequence Combining PSF-Choice, EPSI and a Resolution Enhancement Algorithm
 Improved Spatial Localization in 3D MRSI with a Sequence Combining PSF-Choice, EPSI and a Resolution Enhancement Algorithm L.P. Panych 1,3, B. Madore 1,3, W.S. Hoge 1,3, R.V. Mulkern 2,3 1 Brigham and
Improved Spatial Localization in 3D MRSI with a Sequence Combining PSF-Choice, EPSI and a Resolution Enhancement Algorithm L.P. Panych 1,3, B. Madore 1,3, W.S. Hoge 1,3, R.V. Mulkern 2,3 1 Brigham and
Statistical Analysis of MRI Data
 Statistical Analysis of MRI Data Shelby Cummings August 1, 2012 Abstract Every day, numerous people around the country go under medical testing with the use of MRI technology. Developed in the late twentieth
Statistical Analysis of MRI Data Shelby Cummings August 1, 2012 Abstract Every day, numerous people around the country go under medical testing with the use of MRI technology. Developed in the late twentieth
Using the Kolmogorov-Smirnov Test for Image Segmentation
 Using the Kolmogorov-Smirnov Test for Image Segmentation Yong Jae Lee CS395T Computational Statistics Final Project Report May 6th, 2009 I. INTRODUCTION Image segmentation is a fundamental task in computer
Using the Kolmogorov-Smirnov Test for Image Segmentation Yong Jae Lee CS395T Computational Statistics Final Project Report May 6th, 2009 I. INTRODUCTION Image segmentation is a fundamental task in computer
coding of various parts showing different features, the possibility of rotation or of hiding covering parts of the object's surface to gain an insight
 Three-Dimensional Object Reconstruction from Layered Spatial Data Michael Dangl and Robert Sablatnig Vienna University of Technology, Institute of Computer Aided Automation, Pattern Recognition and Image
Three-Dimensional Object Reconstruction from Layered Spatial Data Michael Dangl and Robert Sablatnig Vienna University of Technology, Institute of Computer Aided Automation, Pattern Recognition and Image
CHAPTER 6. 6 Huffman Coding Based Image Compression Using Complex Wavelet Transform. 6.3 Wavelet Transform based compression technique 106
 CHAPTER 6 6 Huffman Coding Based Image Compression Using Complex Wavelet Transform Page No 6.1 Introduction 103 6.2 Compression Techniques 104 103 6.2.1 Lossless compression 105 6.2.2 Lossy compression
CHAPTER 6 6 Huffman Coding Based Image Compression Using Complex Wavelet Transform Page No 6.1 Introduction 103 6.2 Compression Techniques 104 103 6.2.1 Lossless compression 105 6.2.2 Lossy compression
Averages and Variation
 Averages and Variation 3 Copyright Cengage Learning. All rights reserved. 3.1-1 Section 3.1 Measures of Central Tendency: Mode, Median, and Mean Copyright Cengage Learning. All rights reserved. 3.1-2 Focus
Averages and Variation 3 Copyright Cengage Learning. All rights reserved. 3.1-1 Section 3.1 Measures of Central Tendency: Mode, Median, and Mean Copyright Cengage Learning. All rights reserved. 3.1-2 Focus
Chapter 6: DESCRIPTIVE STATISTICS
 Chapter 6: DESCRIPTIVE STATISTICS Random Sampling Numerical Summaries Stem-n-Leaf plots Histograms, and Box plots Time Sequence Plots Normal Probability Plots Sections 6-1 to 6-5, and 6-7 Random Sampling
Chapter 6: DESCRIPTIVE STATISTICS Random Sampling Numerical Summaries Stem-n-Leaf plots Histograms, and Box plots Time Sequence Plots Normal Probability Plots Sections 6-1 to 6-5, and 6-7 Random Sampling
Diffusion Mapping with FireVoxel Quick Start Guide
 Diffusion Mapping with FireVoxel Quick Start Guide Medical image analysis tool developed by Artem Mikheev and Henry Rusinek Radiology Department, NYU School of Medicine Original version prepared by Jinyu
Diffusion Mapping with FireVoxel Quick Start Guide Medical image analysis tool developed by Artem Mikheev and Henry Rusinek Radiology Department, NYU School of Medicine Original version prepared by Jinyu
Adaptive Fuzzy Connectedness-Based Medical Image Segmentation
 Adaptive Fuzzy Connectedness-Based Medical Image Segmentation Amol Pednekar Ioannis A. Kakadiaris Uday Kurkure Visual Computing Lab, Dept. of Computer Science, Univ. of Houston, Houston, TX, USA apedneka@bayou.uh.edu
Adaptive Fuzzy Connectedness-Based Medical Image Segmentation Amol Pednekar Ioannis A. Kakadiaris Uday Kurkure Visual Computing Lab, Dept. of Computer Science, Univ. of Houston, Houston, TX, USA apedneka@bayou.uh.edu
3 Graphical Displays of Data
 3 Graphical Displays of Data Reading: SW Chapter 2, Sections 1-6 Summarizing and Displaying Qualitative Data The data below are from a study of thyroid cancer, using NMTR data. The investigators looked
3 Graphical Displays of Data Reading: SW Chapter 2, Sections 1-6 Summarizing and Displaying Qualitative Data The data below are from a study of thyroid cancer, using NMTR data. The investigators looked
SPM8 for Basic and Clinical Investigators. Preprocessing. fmri Preprocessing
 SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
The Bootstrap and Jackknife
 The Bootstrap and Jackknife Summer 2017 Summer Institutes 249 Bootstrap & Jackknife Motivation In scientific research Interest often focuses upon the estimation of some unknown parameter, θ. The parameter
The Bootstrap and Jackknife Summer 2017 Summer Institutes 249 Bootstrap & Jackknife Motivation In scientific research Interest often focuses upon the estimation of some unknown parameter, θ. The parameter
Automatic Quantification of DTI Parameters along Fiber Bundles
 Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein, Simon Hermann, Olaf Konrad, Horst K. Hahn, Heinz-Otto Peitgen MeVis Research, 28359 Bremen Email: klein@mevis.de Abstract. We introduce
Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein, Simon Hermann, Olaf Konrad, Horst K. Hahn, Heinz-Otto Peitgen MeVis Research, 28359 Bremen Email: klein@mevis.de Abstract. We introduce
Nonrigid Motion Compensation of Free Breathing Acquired Myocardial Perfusion Data
 Nonrigid Motion Compensation of Free Breathing Acquired Myocardial Perfusion Data Gert Wollny 1, Peter Kellman 2, Andrés Santos 1,3, María-Jesus Ledesma 1,3 1 Biomedical Imaging Technologies, Department
Nonrigid Motion Compensation of Free Breathing Acquired Myocardial Perfusion Data Gert Wollny 1, Peter Kellman 2, Andrés Santos 1,3, María-Jesus Ledesma 1,3 1 Biomedical Imaging Technologies, Department
A Transmission Line Matrix Model for Shielding Effects in Stents
 A Transmission Line Matrix Model for hielding Effects in tents Razvan Ciocan (1), Nathan Ida (2) (1) Clemson University Physics and Astronomy Department Clemson University, C 29634-0978 ciocan@clemon.edu
A Transmission Line Matrix Model for hielding Effects in tents Razvan Ciocan (1), Nathan Ida (2) (1) Clemson University Physics and Astronomy Department Clemson University, C 29634-0978 ciocan@clemon.edu
Application of Clustering Techniques to Energy Data to Enhance Analysts Productivity
 Application of Clustering Techniques to Energy Data to Enhance Analysts Productivity Wendy Foslien, Honeywell Labs Valerie Guralnik, Honeywell Labs Steve Harp, Honeywell Labs William Koran, Honeywell Atrium
Application of Clustering Techniques to Energy Data to Enhance Analysts Productivity Wendy Foslien, Honeywell Labs Valerie Guralnik, Honeywell Labs Steve Harp, Honeywell Labs William Koran, Honeywell Atrium
ECLT 5810 Data Preprocessing. Prof. Wai Lam
 ECLT 5810 Data Preprocessing Prof. Wai Lam Why Data Preprocessing? Data in the real world is imperfect incomplete: lacking attribute values, lacking certain attributes of interest, or containing only aggregate
ECLT 5810 Data Preprocessing Prof. Wai Lam Why Data Preprocessing? Data in the real world is imperfect incomplete: lacking attribute values, lacking certain attributes of interest, or containing only aggregate
EPI Data Are Acquired Serially. EPI Data Are Acquired Serially 10/23/2011. Functional Connectivity Preprocessing. fmri Preprocessing
 Functional Connectivity Preprocessing Geometric distortion Head motion Geometric distortion Head motion EPI Data Are Acquired Serially EPI Data Are Acquired Serially descending 1 EPI Data Are Acquired
Functional Connectivity Preprocessing Geometric distortion Head motion Geometric distortion Head motion EPI Data Are Acquired Serially EPI Data Are Acquired Serially descending 1 EPI Data Are Acquired
Functional MRI in Clinical Research and Practice Preprocessing
 Functional MRI in Clinical Research and Practice Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization
Functional MRI in Clinical Research and Practice Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization
Processing math: 100% Intensity Normalization
 Intensity Normalization Overall Pipeline 2/21 Intensity normalization Conventional MRI intensites (T1-w, T2-w, PD, FLAIR) are acquired in arbitrary units Images are not comparable across scanners, subjects,
Intensity Normalization Overall Pipeline 2/21 Intensity normalization Conventional MRI intensites (T1-w, T2-w, PD, FLAIR) are acquired in arbitrary units Images are not comparable across scanners, subjects,
Elastically Deforming a Three-Dimensional Atlas to Match Anatomical Brain Images
 University of Pennsylvania ScholarlyCommons Technical Reports (CIS) Department of Computer & Information Science May 1993 Elastically Deforming a Three-Dimensional Atlas to Match Anatomical Brain Images
University of Pennsylvania ScholarlyCommons Technical Reports (CIS) Department of Computer & Information Science May 1993 Elastically Deforming a Three-Dimensional Atlas to Match Anatomical Brain Images
A ranklet-based CAD for digital mammography
 A ranklet-based CAD for digital mammography Enrico Angelini 1, Renato Campanini 1, Emiro Iampieri 1, Nico Lanconelli 1, Matteo Masotti 1, Todor Petkov 1, and Matteo Roffilli 2 1 Physics Department, University
A ranklet-based CAD for digital mammography Enrico Angelini 1, Renato Campanini 1, Emiro Iampieri 1, Nico Lanconelli 1, Matteo Masotti 1, Todor Petkov 1, and Matteo Roffilli 2 1 Physics Department, University
MRI Segmentation MIDAS, 2007, 2010
 MRI Segmentation MIDAS, 2007, 2010 Lawrence O. Hall, Dmitry Goldgof, Yuhua Gu, Prodip Hore Dept. of Computer Science & Engineering University of South Florida CONTENTS: 1. Introduction... 1 2. Installing
MRI Segmentation MIDAS, 2007, 2010 Lawrence O. Hall, Dmitry Goldgof, Yuhua Gu, Prodip Hore Dept. of Computer Science & Engineering University of South Florida CONTENTS: 1. Introduction... 1 2. Installing
