GeneR. JORGE ARTURO ZEPEDA MARTINEZ LOPEZ HERNANDEZ JOSE FABRICIO. October 6, 2009
|
|
|
- Cecil Todd
- 5 years ago
- Views:
Transcription
1 GeneR JORGE ARTURO ZEPEDA MARTINEZ LOPEZ HERNANDEZ JOSE FABRICIO. October 6, 2009 Abstract GeneR packages allow direct use of nucleotide sequences within R software. Functions can be used to read and write sequences from main file formats (Embl, Genbank and Fasta) in order to perform a lot of manipulations and analyses. Main functions are implemented with C extensions. For this package, the functions are in 4 categories: 1.Reading and writing sequences. 2.Handling sequences. 3.Analyzing sequences. 4.Genetic tools. INSTALLATION. > source(" > bioclite("gener") Using R version (R-devel), biocinstall version Installing Bioconductor version 2.5 packages: [1] "GeneR" Please wait... > library("gener") 1 Basic manipulation with sequences as character string Standard R commands allow to extract subpart of a character string, or append two character strings, put in upper case (functions substr, paste, toupper). GeneR also has tools for the specic use of genetic sequences: insert a sequence into a another, compute the reverse complementary, count mono, di or tri-nucleotides of sequence and the possibility to import sequence from a Embl, Fastaor GeneBank file. Examples: Insert a poly A into sequence > seq <- "gtcatgcatgctaggtgacagttaaaatgcgtctaggtgacagtctaacaa" > seq2 <- insertseq(seq, "aaaaaaaaaaaaaa", 20) > seq2 1
2 [1] "gtcatgcatgctaggtgacaaaaaaaaaaaaaaagttaaaatgcgtctaggtgacagtctaacaa" Compute the reverse complementary: > strcomp(seq) [1] "ttgttagactgtcacctagacgcattttaactgtcacctagcatgcatgac" Count mono nucleotides, di-nucleotides (wsize can be from 1 to 15 if computer has enough memory): > strcomposeq(seq2, wsize = 1) T C A G X [1,] > strcomposeq(seq2, wsize = 2) TT TC TA TG TX CT CC CA CG CX AT AC AA AG [1,] AX GT GC GA GG GX XT XC XA XG XX [1,] Get some data from the web: > seqncbi("by608190", file = "arch.fa") > strreadfasta("arch.fa", from = 10, to = 35) [1] "TGCGTTTGTTTTTTAGTGACTTCTAC" 2 Manipulation of sequences with buffers. The data is loaded into the buffer and all computation done with GeneR functions, only usefull results returns to standard R objects. Which is very useful specially if you are working with great amount of data such as an entire chromosome. Examples: Sequence read from a fasta file: > readfasta("arch.fa") > readfasta("arch.fa", seqno = 1) Show current buffer: > getseq(from = 10, to = 35) [1] "TGCGTTTGTTTTTTAGTGACTTCTAC" Sames previous functions with buffers: Extract from multiple positions. 2
3 > getseq(seqno = 0, from = c(1, 10, 360), to = c(10, 20, 0)) [1] "TGGGCTTATT" "TGCGTTTGTTT" [3] "AATAAAGCCAACTTGCAGCTGCTGTT" > assemble(seqno = 0, destseqno = 1, from = c(1, 10, 360), to = c(10, + 20, 0)) > getseq(seqno = 1) [1] "TGGGCTTATTTGCGTTTGTTTAATAAAGCCAACTTGCAGCTGCTGTT" See the length of a sequence: > sizeseq(seqno = 0) [1] 385 > size0 <- sizeseq(seqno = 0) > getseq(seqno = 0, from = size0-20, to = 0) [1] "AGCCAACTTGCAGCTGCTGTT" Append and concat: > getseq(0, from = 1, to = 35) [1] "TGGGCTTATTGCGTTTGTTTTTTAGTGACTTCTAC" > getseq(1) [1] "TGGGCTTATTTGCGTTTGTTTAATAAAGCCAACTTGCAGCTGCTGTT" > appendseq(0, 1) > getseq(seqno = 0, from = size0-20, to = size0 + 10) [1] "AGCCAACTTGCAGCTGCTGTTTGGGCTTATT" > concat(seqno1 = 0, seqno2 = 1, destseqno = 3, from1 = 2, to1 = 10, + from2 = 8, to2 = 0, strand1 = 1) [1] 3 > getseq(3) [1] "AATAAGCCCAACAGCAGCTGCAAGTTGGCTTTATTAAACAAACGCAAAT" 3 Look for motifs, composition, masked position: Look for motives > exactword("aaag", seqno = 0) 3
4 [[1]] [1] Find Orfs (returns positions of predicted ORFs start position = start codon, stop position = stop codon) > getorfs(seqno = 0) start stop [1,] [2,] [3,] [4,] [5,] [6,] [7,] [8,] > maxorf(seqno = 0) [1] 96 Mask sequences while lowering case of parts of sequence: > x = c(5, 15, 30) > y = c(10, 20, 35) > lowerseq(from = x, to = y) > getseq(from = 1, to = 35) [1] "TGGGcttattGCGTttgtttTTTAGTGACttctac" And the reverse: get masked positions from a sequence: > posmaskedseq(seqno = 0, type = "lower") from to [1,] 5 10 [2,] [3,] > posmaskedseq(seqno = 0, type = "upper") [,1] [,2] [1,] 1 4 [2,] [3,] [4,] Change from Dna to Rna alphabet: > dnatorna() 4
5 [1] "UGGGcuuauuGCGUuuguuuUUUAGUGACuucuacAUGCUGAUGUCCCAUGUAUGUAGUCUUAGACCUGUUUAAUA UCUGUAACUAUCAGCUAUAAUAUUGUGGUGACCACNUGCUAUAGGAUUUUGCCUCUGUGUUAACUACAACAUAUUG GAUUGUAUGUGUCUGUGCCUCCAUGGGGGAUUAGAGCCAGAGGGAAUGUCUUCUUUGCUCUGUUUCUUUUAUAUUU AUAACAAAGAUAUGGAUACUUUCUAGUGAAUGCAUAAGUUAGUGUGCUUUUCUUAUUUUGCUUUAAAUUUCAAGUU UUUACACUGCUGUGAUAAAAUCCUCACAGAUAGCAUCCUCUGGAUGGCACAGUACAAUAAAGCCAACUUGCAGCUG CUGUUUGGGCUUAUUUGCGUUUGUUUAAUAAAGCCAACUUGCAGCUGCUGUU" 4 Adresses manipulations: GeneR developeres have defined 3 types of addresses on a subsequence extracted from a master sequence: -Absolute addresses i.e. addresses on the master sequence, from the 5 end of the input strand refered as forward (noted A) -Real addresses, i.e. addresses on the master sequence, from the 5 end of one of strands (noted R) -Relative addresses, i.e. addresses on working subsequence, from the 5 end of one of strands (noted T) Although all functions using positions need and return absolute addresses, 6 functions allow to convert R, A, T into any other type (functions RtoA, RtoT, AtoR, AtoT, TtoR, TtoA). For example lets imagine that we work with a certain chromosome, and we study only part of chromosome between position 11 to 25: > seq <- "xxxxxxxxxxatgtgtcgttaattgxxxxxxxxxxxxxxx" > placestring(seq) > setstrand(0) > writefasta(file = "arch.fa") > readfasta(file = "arch.fa", from = 11, to = 25) [1] "ATGTGTCGTTAATTG" Orfs on Absolute and relative Adresses: > getorfs() start stop [1,]
6 > AtoT(getOrfs()) start stop [1,] 1 12 More Fun: [1] "ATGTGTCGTTAATTG" > exactword(word = "AA") [[1]] [1] 21 > AtoT(exactWord(word = "AA")[[1]]) start stop 1 5 Bank files: -Fasta files There are 2 functions to get sequence files from internet: sequrl (from a srs server) and seqncbi (from Ncbi). Sequences are retrieved in Fasta or Embl format for sequrl and Fasta or GenBank format for seqncib. > seqncbi("by608190", file = "bank.fa") > seqncbi("by608190", file = "BY gbk", type = "genbank") Basic tools to get or write informations from bank file > readfasta(file = "bank.fa", name = "gi gb BY BY608190", + from = 1, to = 30) [1] "TGGGCTTATTGCGTTTGTTTTTTAGTGACT" > fastadescription(file = "bank.fa", name = "gi gb BY BY608190") [1] "BY RIKEN full-length enriched, visual cortex Mus musculus cdna clone K230301C17 3', mrna sequence" And to write a to the fasta bank file: > s = "cgtagctagctagctagctagctagctagcta" > placestring(s, seqno = 0) 6
7 > writefasta(seqno = 0, file = "bank.fa", name = "MySequence", + comment = "A sequence Generated by R", append = TRUE) Show bank file > cat(paste(readlines("bank.fa"), collapse = "\n")) >gi gb BY BY BY RIKEN full-length enriched, visual cortex Mus musculus cdna clone K230301C17 3', mrna sequence TGGGCTTATTGCGTTTGTTTTTTAGTGACTTCTACATGCTGATGTCCCATGTATGTAGTCTTAGACCTGT TTAATATCTGTAACTATCAGCTATAATATTGTGGTGACCACNTGCTATAGGATTTTGCCTCTGTGTTAAC TACAACATATTGGATTGTATGTGTCTGTGCCTCCATGGGGGATTAGAGCCAGAGGGAATGTCTTCTTTGC TCTGTTTCTTTTATATTTATAACAAAGATATGGATACTTTCTAGTGAATGCATAAGTTAGTGTGCTTTTC TTATTTTGCTTTAAATTTCAAGTTTTTACACTGCTGTGATAAAATCCTCACAGATAGCATCCTCTGGATG GCACAGTACAATAAAGCCAACTTGCAGCTGCTGTT >MySequence A sequence Generated by R CGTAGCTAGCTAGCTAGCTAGCTAGCTAGCTA -Embl files Some basic tools to get sequence and write Embl files > s <- "gtcatgcatcctaggtgtcagggaaaatgcgtctacgtgacagtctaacaa" > placestring(s) Add a line with FT CDS bla bla bla > writeemblline(file = "arch.embl", code = "FT", header = "CDS", + text = "<1..12", nextfield = FALSE) Add lines with FT bla bla bla > writeemblline(file = "arch.embl", code = "FT", header = "", text = "/codonstart=2", + nextfield = FALSE) > writeemblline(file = "arch.embl", code = "FT", header = "", text = "/gene=\"toto\"", + nextfield = FALSE) Add sequence > writeemblseq(file = "arch.embl") Show file > cat(paste(readlines("arch.embl"), collapse = "\n")) 7
8 FT CDS <1..12 FT /codonstart=2 FT /gene="toto" SQ Sequence 51 BP; 15 A; 11 C; 13 G; 12 T; 0 other; gtcatgcatc ctaggtgtca gggaaaatgc gtctacgtga cagtctaaca a 51 // -GeneBank files There is a function to open any sequence from a GeneBank files > readgbk(file = "BY gbk", name = "BY608190", from = 10, + to = 30) [1] "TGCGTTTGTTTTTTAGTGACT" If you want to play with differents RandomSequence, there is a tool to do it. The sequence could be make by one, two or three letters, using probabilities or numbers of elements for each one. Examples: > randomseq(prob = c(0.2, 0.3, 0.2, 0.3), letters = c("t", "C", + "A", "G"), n = 30) [1] "GTTGTCCTCGATGGCACCGGCAGGGGAATA" > randomseq(prob = rep(0.0625, 16), letters = c("tt", "TC", "TA", + "TG", "CT", "CC", "CA", "CG", "AT", "AC", "AA", "AG", "GT", + "GC", "GA", "GG"), n = 10) [1] "ATTTTTGTCAGGAAACGGTC" > shuffleseq(count = c(7, 3, 3, 4, 0), letters = c("t", "C", "A", + "G", "N")) [1] "GATCTGTCGCGATATTT" > shuffleseq(count = c(rep(4, 4), rep(2, 4), rep(1, 4), rep(0, + 4)), letters = c("tt", "TC", "TA", "TG", "CT", "CC", "CA", + "CG", "AT", "AC", "AA", "AG", "GT", "GC", "GA", "GG")) [1] "CATGCGTATGTACTCCTCCTTGCCTTTTTTAATCATTGTATTTCAGTCCACGTAAC" Computes profile of user defined quantities around sites of interest in sequence fragments. A Profile is constituted of bins of equal size around the sites of interest named origins. It produces for each bin, and for each quantity the mean, the standard deviation and the number of valid events. Example: 8
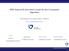
9 > s <- "" > for (i in 1:20) s <- paste(s, randomseq(n = 100), randomseq(prob = c(0.3, + 1, 1, 1, 0)/3.3, n = 100), sep = "") > placestring(s, seqno = 0) > dens <- densityprofile(ori = 200 * (1:20) - 100, from = 200 * + (0:19) + 1, to = 200 * (1:20), seqno = 0, fun = seqskew, + nbinl = 15, nbinr = 15, sizebin = 5) > plot(dens$skta, main = "TA skew") TA skew y x 9
GeneR Package. L. Cottret, A. Lucas, O. Rogier, E. Marrakchi, V. Lefort, P. Durosay, A. Viari, C. Thermes & Y. d Aubenton-Carafa October 18, 2010
 GeneR Package L. Cottret, A. Lucas, O. Rogier, E. Marrakchi, V. Lefort, P. Durosay, A. Viari, C. Thermes & Y. d Aubenton-Carafa October 18, 2010 Contents 1 Overview 1 2 Installation 2 3 Quick Start 2 4
GeneR Package L. Cottret, A. Lucas, O. Rogier, E. Marrakchi, V. Lefort, P. Durosay, A. Viari, C. Thermes & Y. d Aubenton-Carafa October 18, 2010 Contents 1 Overview 1 2 Installation 2 3 Quick Start 2 4
Biostrings. Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA June 2009
 Biostrings Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA 15-19 June 2009 Biostrings Representation DNA, RNA, amino acid, and general biological strings Manipulation
Biostrings Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA 15-19 June 2009 Biostrings Representation DNA, RNA, amino acid, and general biological strings Manipulation
Finding homologous sequences in databases
 Finding homologous sequences in databases There are multiple algorithms to search sequences databases BLAST (EMBL, NCBI, DDBJ, local) FASTA (EMBL, local) For protein only databases scan via Smith-Waterman
Finding homologous sequences in databases There are multiple algorithms to search sequences databases BLAST (EMBL, NCBI, DDBJ, local) FASTA (EMBL, local) For protein only databases scan via Smith-Waterman
Lecture 5: Markov models
 Master s course Bioinformatics Data Analysis and Tools Lecture 5: Markov models Centre for Integrative Bioinformatics Problem in biology Data and patterns are often not clear cut When we want to make a
Master s course Bioinformatics Data Analysis and Tools Lecture 5: Markov models Centre for Integrative Bioinformatics Problem in biology Data and patterns are often not clear cut When we want to make a
Using seqtools package
 Using seqtools package Wolfgang Kaisers, CBiBs HHU Dusseldorf October 30, 2017 1 seqtools package The seqtools package provides functionality for collecting and analyzing quality measures from FASTQ files.
Using seqtools package Wolfgang Kaisers, CBiBs HHU Dusseldorf October 30, 2017 1 seqtools package The seqtools package provides functionality for collecting and analyzing quality measures from FASTQ files.
SAPLING: Suffix Array Piecewise Linear INdex for Genomics Michael Kirsche
 SAPLING: Suffix Array Piecewise Linear INdex for Genomics Michael Kirsche mkirsche@jhu.edu StringBio 2018 Outline Substring Search Problem Caching and Learned Data Structures Methods Results Ongoing work
SAPLING: Suffix Array Piecewise Linear INdex for Genomics Michael Kirsche mkirsche@jhu.edu StringBio 2018 Outline Substring Search Problem Caching and Learned Data Structures Methods Results Ongoing work
Creating a custom mappings similarity matrix
 BioNumerics Tutorial: Creating a custom mappings similarity matrix 1 Aim In BioNumerics, character values can be mapped to categorical names according to predefined criteria (see tutorial Importing non-numerical
BioNumerics Tutorial: Creating a custom mappings similarity matrix 1 Aim In BioNumerics, character values can be mapped to categorical names according to predefined criteria (see tutorial Importing non-numerical
Biostatistics and Bioinformatics Molecular Sequence Databases
 . 1 Description of Module Subject Name Paper Name Module Name/Title 13 03 Dr. Vijaya Khader Dr. MC Varadaraj 2 1. Objectives: In the present module, the students will learn about 1. Encoding linear sequences
. 1 Description of Module Subject Name Paper Name Module Name/Title 13 03 Dr. Vijaya Khader Dr. MC Varadaraj 2 1. Objectives: In the present module, the students will learn about 1. Encoding linear sequences
Wilson Leung 01/03/2018 An Introduction to NCBI BLAST. Prerequisites: Detecting and Interpreting Genetic Homology: Lecture Notes on Alignment
 An Introduction to NCBI BLAST Prerequisites: Detecting and Interpreting Genetic Homology: Lecture Notes on Alignment Resources: The BLAST web server is available at https://blast.ncbi.nlm.nih.gov/blast.cgi
An Introduction to NCBI BLAST Prerequisites: Detecting and Interpreting Genetic Homology: Lecture Notes on Alignment Resources: The BLAST web server is available at https://blast.ncbi.nlm.nih.gov/blast.cgi
Darwin: A Genomic Co-processor gives up to 15,000X speedup on long read assembly (To appear in ASPLOS 2018)
 Darwin: A Genomic Co-processor gives up to 15,000X speedup on long read assembly (To appear in ASPLOS 2018) Yatish Turakhia EE PhD candidate Stanford University Prof. Bill Dally (Electrical Engineering
Darwin: A Genomic Co-processor gives up to 15,000X speedup on long read assembly (To appear in ASPLOS 2018) Yatish Turakhia EE PhD candidate Stanford University Prof. Bill Dally (Electrical Engineering
BMI/CS 576 Fall 2015 Midterm Exam
 BMI/CS 576 Fall 2015 Midterm Exam Prof. Colin Dewey Tuesday, October 27th, 2015 11:00am-12:15pm Name: KEY Write your answers on these pages and show your work. You may use the back sides of pages as necessary.
BMI/CS 576 Fall 2015 Midterm Exam Prof. Colin Dewey Tuesday, October 27th, 2015 11:00am-12:15pm Name: KEY Write your answers on these pages and show your work. You may use the back sides of pages as necessary.
Mapping RNA sequence data (Part 1: using pathogen portal s RNAseq pipeline) Exercise 6
 Mapping RNA sequence data (Part 1: using pathogen portal s RNAseq pipeline) Exercise 6 The goal of this exercise is to retrieve an RNA-seq dataset in FASTQ format and run it through an RNA-sequence analysis
Mapping RNA sequence data (Part 1: using pathogen portal s RNAseq pipeline) Exercise 6 The goal of this exercise is to retrieve an RNA-seq dataset in FASTQ format and run it through an RNA-sequence analysis
FASTA. Besides that, FASTA package provides SSEARCH, an implementation of the optimal Smith- Waterman algorithm.
 FASTA INTRODUCTION Definition (by David J. Lipman and William R. Pearson in 1985) - Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence
FASTA INTRODUCTION Definition (by David J. Lipman and William R. Pearson in 1985) - Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence
4.1. Access the internet and log on to the UCSC Genome Bioinformatics Web Page (Figure 1-
 1. PURPOSE To provide instructions for finding rs Numbers (SNP database ID numbers) and increasing sequence length by utilizing the UCSC Genome Bioinformatics Database. 2. MATERIALS 2.1. Sequence Information
1. PURPOSE To provide instructions for finding rs Numbers (SNP database ID numbers) and increasing sequence length by utilizing the UCSC Genome Bioinformatics Database. 2. MATERIALS 2.1. Sequence Information
Topics of the talk. Biodatabases. Data types. Some sequence terminology...
 Topics of the talk Biodatabases Jarno Tuimala / Eija Korpelainen CSC What data are stored in biological databases? What constitutes a good database? Nucleic acid sequence databases Amino acid sequence
Topics of the talk Biodatabases Jarno Tuimala / Eija Korpelainen CSC What data are stored in biological databases? What constitutes a good database? Nucleic acid sequence databases Amino acid sequence
Sequence Assembly. BMI/CS 576 Mark Craven Some sequencing successes
 Sequence Assembly BMI/CS 576 www.biostat.wisc.edu/bmi576/ Mark Craven craven@biostat.wisc.edu Some sequencing successes Yersinia pestis Cannabis sativa The sequencing problem We want to determine the identity
Sequence Assembly BMI/CS 576 www.biostat.wisc.edu/bmi576/ Mark Craven craven@biostat.wisc.edu Some sequencing successes Yersinia pestis Cannabis sativa The sequencing problem We want to determine the identity
BIOINFORMATICS A PRACTICAL GUIDE TO THE ANALYSIS OF GENES AND PROTEINS
 BIOINFORMATICS A PRACTICAL GUIDE TO THE ANALYSIS OF GENES AND PROTEINS EDITED BY Genome Technology Branch National Human Genome Research Institute National Institutes of Health Bethesda, Maryland B. F.
BIOINFORMATICS A PRACTICAL GUIDE TO THE ANALYSIS OF GENES AND PROTEINS EDITED BY Genome Technology Branch National Human Genome Research Institute National Institutes of Health Bethesda, Maryland B. F.
Implementation of BDEA-A DNA cryptography for securing data in cloud
 Implementation of BDEA-A DNA cryptography for securing data in cloud Hema N 1 Sonali Ulhas Kadwadkar 2 1 Assistant Professor, Dept of Information Science Engineering, RNSIT Bangalore, Karnataka, India
Implementation of BDEA-A DNA cryptography for securing data in cloud Hema N 1 Sonali Ulhas Kadwadkar 2 1 Assistant Professor, Dept of Information Science Engineering, RNSIT Bangalore, Karnataka, India
Module 1 Artemis. Introduction. Aims IF YOU DON T UNDERSTAND, PLEASE ASK! -1-
 Module 1 Artemis Introduction Artemis is a DNA viewer and annotation tool, free to download and use, written by Kim Rutherford from the Sanger Institute (Rutherford et al., 2000). The program allows the
Module 1 Artemis Introduction Artemis is a DNA viewer and annotation tool, free to download and use, written by Kim Rutherford from the Sanger Institute (Rutherford et al., 2000). The program allows the
Algorithms for Bioinformatics
 Adapted from slides by Alexandru Tomescu, Leena Salmela and Veli Mäkinen, which are partly from http://bix.ucsd.edu/bioalgorithms/slides.php 582670 Algorithms for Bioinformatics Lecture 3: Graph Algorithms
Adapted from slides by Alexandru Tomescu, Leena Salmela and Veli Mäkinen, which are partly from http://bix.ucsd.edu/bioalgorithms/slides.php 582670 Algorithms for Bioinformatics Lecture 3: Graph Algorithms
Mapping Reads to Reference Genome
 Mapping Reads to Reference Genome DNA carries genetic information DNA is a double helix of two complementary strands formed by four nucleotides (bases): Adenine, Cytosine, Guanine and Thymine 2 of 31 Gene
Mapping Reads to Reference Genome DNA carries genetic information DNA is a double helix of two complementary strands formed by four nucleotides (bases): Adenine, Cytosine, Guanine and Thymine 2 of 31 Gene
Installation and Use. The programs here are used to index and then search a database of nucleotides.
 August 28, 2018 Kevin C. O'Kane kc.okane@gmail.com https://threadsafebooks.com Installation and Use The programs here are used to index and then search a database of nucleotides. Sequence Database The
August 28, 2018 Kevin C. O'Kane kc.okane@gmail.com https://threadsafebooks.com Installation and Use The programs here are used to index and then search a database of nucleotides. Sequence Database The
Lecture 2, Introduction to Python. Python Programming Language
 BINF 3360, Introduction to Computational Biology Lecture 2, Introduction to Python Young-Rae Cho Associate Professor Department of Computer Science Baylor University Python Programming Language Script
BINF 3360, Introduction to Computational Biology Lecture 2, Introduction to Python Young-Rae Cho Associate Professor Department of Computer Science Baylor University Python Programming Language Script
Exeter Sequencing Service
 Exeter Sequencing Service A guide to your denovo RNA-seq results An overview Once your results are ready, you will receive an email with a password-protected link to them. Click the link to access your
Exeter Sequencing Service A guide to your denovo RNA-seq results An overview Once your results are ready, you will receive an email with a password-protected link to them. Click the link to access your
Graph Algorithms in Bioinformatics
 Graph Algorithms in Bioinformatics Computational Biology IST Ana Teresa Freitas 2015/2016 Sequencing Clone-by-clone shotgun sequencing Human Genome Project Whole-genome shotgun sequencing Celera Genomics
Graph Algorithms in Bioinformatics Computational Biology IST Ana Teresa Freitas 2015/2016 Sequencing Clone-by-clone shotgun sequencing Human Genome Project Whole-genome shotgun sequencing Celera Genomics
Identiyfing splice junctions from RNA-Seq data
 Identiyfing splice junctions from RNA-Seq data Joseph K. Pickrell pickrell@uchicago.edu October 4, 2010 Contents 1 Motivation 2 2 Identification of potential junction-spanning reads 2 3 Calling splice
Identiyfing splice junctions from RNA-Seq data Joseph K. Pickrell pickrell@uchicago.edu October 4, 2010 Contents 1 Motivation 2 2 Identification of potential junction-spanning reads 2 3 Calling splice
Genome 373: Genome Assembly. Doug Fowler
 Genome 373: Genome Assembly Doug Fowler What are some of the things we ve seen we can do with HTS data? We ve seen that HTS can enable a wide variety of analyses ranging from ID ing variants to genome-
Genome 373: Genome Assembly Doug Fowler What are some of the things we ve seen we can do with HTS data? We ve seen that HTS can enable a wide variety of analyses ranging from ID ing variants to genome-
Sequencing. Computational Biology IST Ana Teresa Freitas 2011/2012. (BACs) Whole-genome shotgun sequencing Celera Genomics
 Computational Biology IST Ana Teresa Freitas 2011/2012 Sequencing Clone-by-clone shotgun sequencing Human Genome Project Whole-genome shotgun sequencing Celera Genomics (BACs) 1 Must take the fragments
Computational Biology IST Ana Teresa Freitas 2011/2012 Sequencing Clone-by-clone shotgun sequencing Human Genome Project Whole-genome shotgun sequencing Celera Genomics (BACs) 1 Must take the fragments
Tutorial 1: Exploring the UCSC Genome Browser
 Last updated: May 12, 2011 Tutorial 1: Exploring the UCSC Genome Browser Open the homepage of the UCSC Genome Browser at: http://genome.ucsc.edu/ In the blue bar at the top, click on the Genomes link.
Last updated: May 12, 2011 Tutorial 1: Exploring the UCSC Genome Browser Open the homepage of the UCSC Genome Browser at: http://genome.ucsc.edu/ In the blue bar at the top, click on the Genomes link.
Package Lncident. October 8, 2016
 Type Package Package Lncident October 8, 2016 Title Long Non-Coding RNA identification and Find Open Reading Frame Version 1.3.3 Author Han Siyu Maintainer Han Siyu Description Functions
Type Package Package Lncident October 8, 2016 Title Long Non-Coding RNA identification and Find Open Reading Frame Version 1.3.3 Author Han Siyu Maintainer Han Siyu Description Functions
User Guide for DNAFORM Clone Search Engine
 User Guide for DNAFORM Clone Search Engine Document Version: 3.0 Dated from: 1 October 2010 The document is the property of K.K. DNAFORM and may not be disclosed, distributed, or replicated without the
User Guide for DNAFORM Clone Search Engine Document Version: 3.0 Dated from: 1 October 2010 The document is the property of K.K. DNAFORM and may not be disclosed, distributed, or replicated without the
# input parameters for the script my ($seq, $start, $window, $max_length) #sequence file, calculation start position, window size, max length
 #!/bin/perl use List::Util qw[min max sum]; sub TDD # hash of arrays with thermodynamic parameters for DNA/DNA duplex # hash keys are respective pairs # first array element is enthalpy (dh) # second array
#!/bin/perl use List::Util qw[min max sum]; sub TDD # hash of arrays with thermodynamic parameters for DNA/DNA duplex # hash keys are respective pairs # first array element is enthalpy (dh) # second array
Sequence Alignment. GBIO0002 Archana Bhardwaj University of Liege
 Sequence Alignment GBIO0002 Archana Bhardwaj University of Liege 1 What is Sequence Alignment? A sequence alignment is a way of arranging the sequences of DNA, RNA, or protein to identify regions of similarity.
Sequence Alignment GBIO0002 Archana Bhardwaj University of Liege 1 What is Sequence Alignment? A sequence alignment is a way of arranging the sequences of DNA, RNA, or protein to identify regions of similarity.
DNA Sequencing The Shortest Superstring & Traveling Salesman Problems Sequencing by Hybridization
 Eulerian & Hamiltonian Cycle Problems DNA Sequencing The Shortest Superstring & Traveling Salesman Problems Sequencing by Hybridization The Bridge Obsession Problem Find a tour crossing every bridge just
Eulerian & Hamiltonian Cycle Problems DNA Sequencing The Shortest Superstring & Traveling Salesman Problems Sequencing by Hybridization The Bridge Obsession Problem Find a tour crossing every bridge just
Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA.
 Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA. Fasta is used to compare a protein or DNA sequence to all of the
Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA. Fasta is used to compare a protein or DNA sequence to all of the
Introduction to Genome Browsers
 Introduction to Genome Browsers Rolando Garcia-Milian, MLS, AHIP (Rolando.milian@ufl.edu) Department of Biomedical and Health Information Services Health Sciences Center Libraries, University of Florida
Introduction to Genome Browsers Rolando Garcia-Milian, MLS, AHIP (Rolando.milian@ufl.edu) Department of Biomedical and Health Information Services Health Sciences Center Libraries, University of Florida
Filogeografía BIOL 4211, Universidad de los Andes 25 de enero a 01 de abril 2006
 Laboratory excercise written by Andrew J. Crawford with the support of CIES Fulbright Program and Fulbright Colombia. Enjoy! Filogeografía BIOL 4211, Universidad de los Andes 25 de enero
Laboratory excercise written by Andrew J. Crawford with the support of CIES Fulbright Program and Fulbright Colombia. Enjoy! Filogeografía BIOL 4211, Universidad de los Andes 25 de enero
SPAREPARTSCATALOG: CONNECTORS SPARE CONNECTORS KTM ART.-NR.: 3CM EN
 SPAREPARTSCATALOG: CONNECTORS ART.-NR.: 3CM3208201EN CONTENT SPARE CONNECTORS AA-AN SPARE CONNECTORS AO-BC SPARE CONNECTORS BD-BQ SPARE CONNECTORS BR-CD 3 4 5 6 SPARE CONNECTORS CE-CR SPARE CONNECTORS
SPAREPARTSCATALOG: CONNECTORS ART.-NR.: 3CM3208201EN CONTENT SPARE CONNECTORS AA-AN SPARE CONNECTORS AO-BC SPARE CONNECTORS BD-BQ SPARE CONNECTORS BR-CD 3 4 5 6 SPARE CONNECTORS CE-CR SPARE CONNECTORS
DNASIS MAX V2.0. Tutorial Booklet
 Sequence Analysis Software DNASIS MAX V2.0 Tutorial Booklet CONTENTS Introduction...2 1. DNASIS MAX...5 1-1: Protein Translation & Function...5 1-2: Nucleic Acid Alignments(BLAST Search)...10 1-3: Vector
Sequence Analysis Software DNASIS MAX V2.0 Tutorial Booklet CONTENTS Introduction...2 1. DNASIS MAX...5 1-1: Protein Translation & Function...5 1-2: Nucleic Acid Alignments(BLAST Search)...10 1-3: Vector
Weighted Finite-State Transducers in Computational Biology
 Weighted Finite-State Transducers in Computational Biology Mehryar Mohri Courant Institute of Mathematical Sciences mohri@cims.nyu.edu Joint work with Corinna Cortes (Google Research). 1 This Tutorial
Weighted Finite-State Transducers in Computational Biology Mehryar Mohri Courant Institute of Mathematical Sciences mohri@cims.nyu.edu Joint work with Corinna Cortes (Google Research). 1 This Tutorial
DNA Sequencing. Overview
 BINF 3350, Genomics and Bioinformatics DNA Sequencing Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Backgrounds Eulerian Cycles Problem Hamiltonian Cycles
BINF 3350, Genomics and Bioinformatics DNA Sequencing Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Backgrounds Eulerian Cycles Problem Hamiltonian Cycles
Glimmer Release Notes Version 3.01 (Beta) Arthur L. Delcher
 Glimmer Release Notes Version 3.01 (Beta) Arthur L. Delcher 10 October 2005 1 Introduction This document describes Version 3 of the Glimmer gene-finding software. This version incorporates a nearly complete
Glimmer Release Notes Version 3.01 (Beta) Arthur L. Delcher 10 October 2005 1 Introduction This document describes Version 3 of the Glimmer gene-finding software. This version incorporates a nearly complete
INTRODUCTION TO BIOINFORMATICS
 Molecular Biology-2017 1 INTRODUCTION TO BIOINFORMATICS In this section, we want to provide a simple introduction to using the web site of the National Center for Biotechnology Information NCBI) to obtain
Molecular Biology-2017 1 INTRODUCTION TO BIOINFORMATICS In this section, we want to provide a simple introduction to using the web site of the National Center for Biotechnology Information NCBI) to obtain
WSSP-10 Chapter 7 BLASTN: DNA vs DNA searches
 WSSP-10 Chapter 7 BLASTN: DNA vs DNA searches 4-3 DSAP: BLASTn Page p. 7-1 NCBI BLAST Home Page p. 7-1 NCBI BLASTN search page p. 7-2 Copy sequence from DSAP or wave form program p. 7-2 Choose a database
WSSP-10 Chapter 7 BLASTN: DNA vs DNA searches 4-3 DSAP: BLASTn Page p. 7-1 NCBI BLAST Home Page p. 7-1 NCBI BLASTN search page p. 7-2 Copy sequence from DSAP or wave form program p. 7-2 Choose a database
Lecture 5 Advanced BLAST
 Introduction to Bioinformatics for Medical Research Gideon Greenspan gdg@cs.technion.ac.il Lecture 5 Advanced BLAST BLAST Recap Sequence Alignment Complexity and indexing BLASTN and BLASTP Basic parameters
Introduction to Bioinformatics for Medical Research Gideon Greenspan gdg@cs.technion.ac.il Lecture 5 Advanced BLAST BLAST Recap Sequence Alignment Complexity and indexing BLASTN and BLASTP Basic parameters
Solutions Exercise Set 3 Author: Charmi Panchal
 Solutions Exercise Set 3 Author: Charmi Panchal Problem 1: Suppose we have following fragments: f1 = ATCCTTAACCCC f2 = TTAACTCA f3 = TTAATACTCCC f4 = ATCTTTC f5 = CACTCCCACACA f6 = CACAATCCTTAACCC f7 =
Solutions Exercise Set 3 Author: Charmi Panchal Problem 1: Suppose we have following fragments: f1 = ATCCTTAACCCC f2 = TTAACTCA f3 = TTAATACTCCC f4 = ATCTTTC f5 = CACTCCCACACA f6 = CACAATCCTTAACCC f7 =
GBS Bioinformatics Pipeline(s) Overview
 GBS Bioinformatics Pipeline(s) Overview Getting from sequence files to genotypes. Pipeline Coding: Ed Buckler Jeff Glaubitz James Harriman Presentation: Terry Casstevens With supporting information from
GBS Bioinformatics Pipeline(s) Overview Getting from sequence files to genotypes. Pipeline Coding: Ed Buckler Jeff Glaubitz James Harriman Presentation: Terry Casstevens With supporting information from
SPARE CONNECTORS KTM 2014
 SPAREPARTSCATALOG: // ENGINE ART.-NR.: 3208201EN CONTENT CONNECTORS FOR WIRING HARNESS AA-AN CONNECTORS FOR WIRING HARNESS AO-BC CONNECTORS FOR WIRING HARNESS BD-BQ CONNECTORS FOR WIRING HARNESS BR-CD
SPAREPARTSCATALOG: // ENGINE ART.-NR.: 3208201EN CONTENT CONNECTORS FOR WIRING HARNESS AA-AN CONNECTORS FOR WIRING HARNESS AO-BC CONNECTORS FOR WIRING HARNESS BD-BQ CONNECTORS FOR WIRING HARNESS BR-CD
Laboratorio di Basi di Dati per Bioinformatica
 Laboratorio di Basi di Dati per Bioinformatica Laurea in Bioinformatica Docente: Carlo Combi Email: carlo.combi@univr.it Lezione 11 Postgresql per la Bioinformatica Postbio: http://postbio.projects.postgresql.org/
Laboratorio di Basi di Dati per Bioinformatica Laurea in Bioinformatica Docente: Carlo Combi Email: carlo.combi@univr.it Lezione 11 Postgresql per la Bioinformatica Postbio: http://postbio.projects.postgresql.org/
Unix, Perl and BioPerl
 Unix, Perl and BioPerl III: Sequence Analysis with Perl - Modules and BioPerl George Bell, Ph.D. WIBR Bioinformatics and Research Computing Sequence analysis with Perl Modules and BioPerl Regular expressions
Unix, Perl and BioPerl III: Sequence Analysis with Perl - Modules and BioPerl George Bell, Ph.D. WIBR Bioinformatics and Research Computing Sequence analysis with Perl Modules and BioPerl Regular expressions
New generation of patent sequence databases Information Sources in Biotechnology Japan
 New generation of patent sequence databases Information Sources in Biotechnology Japan EBI is an Outstation of the European Molecular Biology Laboratory. Patent-related resources Patents Patent Resources
New generation of patent sequence databases Information Sources in Biotechnology Japan EBI is an Outstation of the European Molecular Biology Laboratory. Patent-related resources Patents Patent Resources
Algorithms for Bioinformatics
 Adapted from slides by Alexandru Tomescu, Leena Salmela and Veli Mäkinen, which are partly from http://bix.ucsd.edu/bioalgorithms/slides.php 58670 Algorithms for Bioinformatics Lecture 5: Graph Algorithms
Adapted from slides by Alexandru Tomescu, Leena Salmela and Veli Mäkinen, which are partly from http://bix.ucsd.edu/bioalgorithms/slides.php 58670 Algorithms for Bioinformatics Lecture 5: Graph Algorithms
TECH NOTE Improving the Sensitivity of Ultra Low Input mrna Seq
 TECH NOTE Improving the Sensitivity of Ultra Low Input mrna Seq SMART Seq v4 Ultra Low Input RNA Kit for Sequencing Powered by SMART and LNA technologies: Locked nucleic acid technology significantly improves
TECH NOTE Improving the Sensitivity of Ultra Low Input mrna Seq SMART Seq v4 Ultra Low Input RNA Kit for Sequencing Powered by SMART and LNA technologies: Locked nucleic acid technology significantly improves
Welcome to. Python 2. Session #5. Michael Purcaro, Chris MacKay, Nick Hathaway, and the GSBS Bootstrappers February 2014
 Welcome to Python 2 Session #5 Michael Purcaro, Chris MacKay, Nick Hathaway, and the GSBS Bootstrappers February 2014 michael.purcaro@umassmed.edu 1 Building Blocks: modules To more easily reuse code,
Welcome to Python 2 Session #5 Michael Purcaro, Chris MacKay, Nick Hathaway, and the GSBS Bootstrappers February 2014 michael.purcaro@umassmed.edu 1 Building Blocks: modules To more easily reuse code,
Sequence analysis with Perl Modules and BioPerl. Unix, Perl and BioPerl. Regular expressions. Objectives. Some uses of regular expressions
 Unix, Perl and BioPerl III: Sequence Analysis with Perl - Modules and BioPerl George Bell, Ph.D. WIBR Bioinformatics and Research Computing Sequence analysis with Perl Modules and BioPerl Regular expressions
Unix, Perl and BioPerl III: Sequence Analysis with Perl - Modules and BioPerl George Bell, Ph.D. WIBR Bioinformatics and Research Computing Sequence analysis with Perl Modules and BioPerl Regular expressions
Package gquad. June 7, 2017
 Type Package Package gquad June 7, 2017 Title Prediction of G Quadruplees and Other Non-B DNA Motifs Version 2.1-1 Date 2017-06-06 Author Maintainer Genomic biology is not limited to
Type Package Package gquad June 7, 2017 Title Prediction of G Quadruplees and Other Non-B DNA Motifs Version 2.1-1 Date 2017-06-06 Author Maintainer Genomic biology is not limited to
GRASPm: an efficient algorithm for exact pattern-matching in genomic sequences
 Int. J. Bioinformatics Research and Applications, Vol. GRASPm: an efficient algorithm for exact pattern-matching in genomic sequences Sérgio Deusdado* Centre for Mountain Research (CIMO), Polytechnic Institute
Int. J. Bioinformatics Research and Applications, Vol. GRASPm: an efficient algorithm for exact pattern-matching in genomic sequences Sérgio Deusdado* Centre for Mountain Research (CIMO), Polytechnic Institute
CISC 636 Computational Biology & Bioinformatics (Fall 2016)
 CISC 636 Computational Biology & Bioinformatics (Fall 2016) Sequence pairwise alignment Score statistics: E-value and p-value Heuristic algorithms: BLAST and FASTA Database search: gene finding and annotations
CISC 636 Computational Biology & Bioinformatics (Fall 2016) Sequence pairwise alignment Score statistics: E-value and p-value Heuristic algorithms: BLAST and FASTA Database search: gene finding and annotations
v0.3.0 May 18, 2016 SNPsplit operates in two stages:
 May 18, 2016 v0.3.0 SNPsplit is an allele-specific alignment sorter which is designed to read alignment files in SAM/ BAM format and determine the allelic origin of reads that cover known SNP positions.
May 18, 2016 v0.3.0 SNPsplit is an allele-specific alignment sorter which is designed to read alignment files in SAM/ BAM format and determine the allelic origin of reads that cover known SNP positions.
Sequence Assembly Required!
 Sequence Assembly Required! 1 October 3, ISMB 20172007 1 Sequence Assembly Genome Sequenced Fragments (reads) Assembled Contigs Finished Genome 2 Greedy solution is bounded 3 Typical assembly strategy
Sequence Assembly Required! 1 October 3, ISMB 20172007 1 Sequence Assembly Genome Sequenced Fragments (reads) Assembled Contigs Finished Genome 2 Greedy solution is bounded 3 Typical assembly strategy
Biostrings Lab (BioC2008)
 Biostrings Lab (BioC2008) H. Pagès Gentleman Lab Fred Hutchinson Cancer Research Center Seattle, WA July 31, 2008 1 Lab overview 1.1 Goals Learn the basics of the Biostrings package and how to use it together
Biostrings Lab (BioC2008) H. Pagès Gentleman Lab Fred Hutchinson Cancer Research Center Seattle, WA July 31, 2008 1 Lab overview 1.1 Goals Learn the basics of the Biostrings package and how to use it together
Advanced UCSC Browser Functions
 Advanced UCSC Browser Functions Dr. Thomas Randall tarandal@email.unc.edu bioinformatics.unc.edu UCSC Browser: genome.ucsc.edu Overview Custom Tracks adding your own datasets Utilities custom tools for
Advanced UCSC Browser Functions Dr. Thomas Randall tarandal@email.unc.edu bioinformatics.unc.edu UCSC Browser: genome.ucsc.edu Overview Custom Tracks adding your own datasets Utilities custom tools for
mgu74a.db November 2, 2013 Map Manufacturer identifiers to Accession Numbers
 mgu74a.db November 2, 2013 mgu74aaccnum Map Manufacturer identifiers to Accession Numbers mgu74aaccnum is an R object that contains mappings between a manufacturer s identifiers and manufacturers accessions.
mgu74a.db November 2, 2013 mgu74aaccnum Map Manufacturer identifiers to Accession Numbers mgu74aaccnum is an R object that contains mappings between a manufacturer s identifiers and manufacturers accessions.
EBI services. Jennifer McDowall EMBL-EBI
 EBI services Jennifer McDowall EMBL-EBI The SLING project is funded by the European Commission within Research Infrastructures of the FP7 Capacities Specific Programme, grant agreement number 226073 (Integrating
EBI services Jennifer McDowall EMBL-EBI The SLING project is funded by the European Commission within Research Infrastructures of the FP7 Capacities Specific Programme, grant agreement number 226073 (Integrating
MacVector for Mac OS X
 MacVector 10.6 for Mac OS X System Requirements MacVector 10.6 runs on any PowerPC or Intel Macintosh running Mac OS X 10.4 or higher. It is a Universal Binary, meaning that it runs natively on both PowerPC
MacVector 10.6 for Mac OS X System Requirements MacVector 10.6 runs on any PowerPC or Intel Macintosh running Mac OS X 10.4 or higher. It is a Universal Binary, meaning that it runs natively on both PowerPC
Release Notes. Version Gene Codes Corporation
 Version 4.10.1 Release Notes 2010 Gene Codes Corporation Gene Codes Corporation 775 Technology Drive, Ann Arbor, MI 48108 USA 1.800.497.4939 (USA) +1.734.769.7249 (elsewhere) +1.734.769.7074 (fax) www.genecodes.com
Version 4.10.1 Release Notes 2010 Gene Codes Corporation Gene Codes Corporation 775 Technology Drive, Ann Arbor, MI 48108 USA 1.800.497.4939 (USA) +1.734.769.7249 (elsewhere) +1.734.769.7074 (fax) www.genecodes.com
Chaotic Image Encryption Technique using S-box based on DNA Approach
 Chaotic Image Encryption Technique using S-box based on DNA Approach Anchal Jain Pooja Agarwal Rashi Jain Vyomesh Singh ABSTRACT In recent years many DNA approach based encryption algorithms have been
Chaotic Image Encryption Technique using S-box based on DNA Approach Anchal Jain Pooja Agarwal Rashi Jain Vyomesh Singh ABSTRACT In recent years many DNA approach based encryption algorithms have been
Sh-miR designer. a tool for construction of RNA interference reagents: sh-mirs
 Sh-miR designer a tool for construction of RNA interference reagents: sh-mirs AMU-Poznan Team Instructors: Marcin Borkowski Michał Ren Student Members: Martyna Urbanek Mateusz Flieger Sylwester Brzeczkowski
Sh-miR designer a tool for construction of RNA interference reagents: sh-mirs AMU-Poznan Team Instructors: Marcin Borkowski Michał Ren Student Members: Martyna Urbanek Mateusz Flieger Sylwester Brzeczkowski
from scratch A primer for scientists working with Next-Generation- Sequencing data CHAPTER 8 biopython
 from scratch A primer for scientists working with Next-Generation- Sequencing data CHAPTER 8 biopython Chapter 8: Biopython Biopython is a collection of modules that implement common bioinformatical tasks
from scratch A primer for scientists working with Next-Generation- Sequencing data CHAPTER 8 biopython Chapter 8: Biopython Biopython is a collection of modules that implement common bioinformatical tasks
m6aviewer Version Documentation
 m6aviewer Version 1.6.0 Documentation Contents 1. About 2. Requirements 3. Launching m6aviewer 4. Running Time Estimates 5. Basic Peak Calling 6. Running Modes 7. Multiple Samples/Sample Replicates 8.
m6aviewer Version 1.6.0 Documentation Contents 1. About 2. Requirements 3. Launching m6aviewer 4. Running Time Estimates 5. Basic Peak Calling 6. Running Modes 7. Multiple Samples/Sample Replicates 8.
Practical Course in Genome Bioinformatics
 Practical Course in Genome Bioinformatics 20/01/2017 Exercises - Day 1 http://ekhidna.biocenter.helsinki.fi/downloads/teaching/spring2017/ Answer questions Q1-Q3 below and include requested Figures 1-5
Practical Course in Genome Bioinformatics 20/01/2017 Exercises - Day 1 http://ekhidna.biocenter.helsinki.fi/downloads/teaching/spring2017/ Answer questions Q1-Q3 below and include requested Figures 1-5
(DNA#): Molecular Biology Computation Language Proposal
 (DNA#): Molecular Biology Computation Language Proposal Aalhad Patankar, Min Fan, Nan Yu, Oriana Fuentes, Stan Peceny {ap3536, mf3084, ny2263, oif2102, skp2140} @columbia.edu Motivation Inspired by the
(DNA#): Molecular Biology Computation Language Proposal Aalhad Patankar, Min Fan, Nan Yu, Oriana Fuentes, Stan Peceny {ap3536, mf3084, ny2263, oif2102, skp2140} @columbia.edu Motivation Inspired by the
[Parallel] processing pipelines in Python with generators/coroutines and multiprocessing
![[Parallel] processing pipelines in Python with generators/coroutines and multiprocessing [Parallel] processing pipelines in Python with generators/coroutines and multiprocessing](/thumbs/76/73273036.jpg) [Parallel] processing pipelines in Python with generators/coroutines and multiprocessing Example: Fastq processing utility Summaries: Qualities, base composition, redundancy,... Filters: read quality,
[Parallel] processing pipelines in Python with generators/coroutines and multiprocessing Example: Fastq processing utility Summaries: Qualities, base composition, redundancy,... Filters: read quality,
the remaiis c p i j fi r a lent'tli of time, can havetspeclii and case any affi :er shall refuse" or neglect <to «J3,b.01Jllis p. m.
 C «? G «C YY «C C C C C C G z «q Y C G «C Q «q C& qc C q q 6 «z C 6 6 «6 6 «} q % z Y 6 Y G Y & 6 Q q C«G C & C % G G 6 C 6 C 6 C q } 6 C 6 6 6 Q6 «G G q & q 6 C «C 6 G Y z666 «Y 66 ««6 «6? 6 «q q q «6?
C «? G «C YY «C C C C C C G z «q Y C G «C Q «q C& qc C q q 6 «z C 6 6 «6 6 «} q % z Y 6 Y G Y & 6 Q q C«G C & C % G G 6 C 6 C 6 C q } 6 C 6 6 6 Q6 «G G q & q 6 C «C 6 G Y z666 «Y 66 ««6 «6? 6 «q q q «6?
CLC Server. End User USER MANUAL
 CLC Server End User USER MANUAL Manual for CLC Server 10.0.1 Windows, macos and Linux March 8, 2018 This software is for research purposes only. QIAGEN Aarhus Silkeborgvej 2 Prismet DK-8000 Aarhus C Denmark
CLC Server End User USER MANUAL Manual for CLC Server 10.0.1 Windows, macos and Linux March 8, 2018 This software is for research purposes only. QIAGEN Aarhus Silkeborgvej 2 Prismet DK-8000 Aarhus C Denmark
Welcome to GenomeView 101!
 Welcome to GenomeView 101! 1. Start your computer 2. Download and extract the example data http://www.broadinstitute.org/~tabeel/broade.zip Suggestion: - Linux, Mac: make new folder in your home directory
Welcome to GenomeView 101! 1. Start your computer 2. Download and extract the example data http://www.broadinstitute.org/~tabeel/broade.zip Suggestion: - Linux, Mac: make new folder in your home directory
Wilson Leung 05/27/2008 A Simple Introduction to NCBI BLAST
 A Simple Introduction to NCBI BLAST Prerequisites: Detecting and Interpreting Genetic Homology: Lecture Notes on Alignment Resources: The BLAST web server is available at http://www.ncbi.nih.gov/blast/
A Simple Introduction to NCBI BLAST Prerequisites: Detecting and Interpreting Genetic Homology: Lecture Notes on Alignment Resources: The BLAST web server is available at http://www.ncbi.nih.gov/blast/
DNA Fragment Assembly
 Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri DNA Fragment Assembly Overlap
Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri DNA Fragment Assembly Overlap
How to submit nucleotide sequence data to the EMBL Data Library: Information for Authors
 727 How to submit nucleotide sequence data to the EMBL Data Library: Information for Authors l\i»jhe EMBL Data Library, Postfach 10.2209, D-6900 Heidelberg, Federal Republic of Germany ii I i ii January
727 How to submit nucleotide sequence data to the EMBL Data Library: Information for Authors l\i»jhe EMBL Data Library, Postfach 10.2209, D-6900 Heidelberg, Federal Republic of Germany ii I i ii January
The Kodon quickguide
 The Kodon quickguide Version 3.5 Copyright 2002-2007, Applied Maths NV. All rights reserved. Kodon is a registered trademark of Applied Maths NV. All other product names or trademarks are the property
The Kodon quickguide Version 3.5 Copyright 2002-2007, Applied Maths NV. All rights reserved. Kodon is a registered trademark of Applied Maths NV. All other product names or trademarks are the property
RESEARCH TOPIC IN BIOINFORMANTIC
 RESEARCH TOPIC IN BIOINFORMANTIC GENOME ASSEMBLY Instructor: Dr. Yufeng Wu Noted by: February 25, 2012 Genome Assembly is a kind of string sequencing problems. As we all know, the human genome is very
RESEARCH TOPIC IN BIOINFORMANTIC GENOME ASSEMBLY Instructor: Dr. Yufeng Wu Noted by: February 25, 2012 Genome Assembly is a kind of string sequencing problems. As we all know, the human genome is very
EBI patent related services
 EBI patent related services 4 th Annual Forum for SMEs October 18-19 th 2010 Jennifer McDowall Senior Scientist, EMBL-EBI EBI is an Outstation of the European Molecular Biology Laboratory. Overview Patent
EBI patent related services 4 th Annual Forum for SMEs October 18-19 th 2010 Jennifer McDowall Senior Scientist, EMBL-EBI EBI is an Outstation of the European Molecular Biology Laboratory. Overview Patent
I519 Introduction to Bioinformatics. Indexing techniques. Yuzhen Ye School of Informatics & Computing, IUB
 I519 Introduction to Bioinformatics Indexing techniques Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents We have seen indexing technique used in BLAST Applications that rely
I519 Introduction to Bioinformatics Indexing techniques Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents We have seen indexing technique used in BLAST Applications that rely
Package RWebLogo. August 29, 2016
 Type Package Title plotting custom sequence logos Version 1.0.3 Date 2014-04-14 Author Omar Wagih Maintainer Omar Wagih Package RWebLogo August 29, 2016 Description RWebLogo is a wrapper
Type Package Title plotting custom sequence logos Version 1.0.3 Date 2014-04-14 Author Omar Wagih Maintainer Omar Wagih Package RWebLogo August 29, 2016 Description RWebLogo is a wrapper
Managing big biological sequence data with Biostrings and DECIPHER. Erik Wright University of Wisconsin-Madison
 Managing big biological sequence data with Biostrings and DECIPHER Erik Wright University of Wisconsin-Madison What you should learn How to use the Biostrings and DECIPHER packages Creating a database
Managing big biological sequence data with Biostrings and DECIPHER Erik Wright University of Wisconsin-Madison What you should learn How to use the Biostrings and DECIPHER packages Creating a database
Scientific Programming Practical 10
 Scientific Programming Practical 10 Introduction Luca Bianco - Academic Year 2017-18 luca.bianco@fmach.it Biopython FROM Biopython s website: The Biopython Project is an international association of developers
Scientific Programming Practical 10 Introduction Luca Bianco - Academic Year 2017-18 luca.bianco@fmach.it Biopython FROM Biopython s website: The Biopython Project is an international association of developers
Read Mapping. de Novo Assembly. Genomics: Lecture #2 WS 2014/2015
 Mapping de Novo Assembly Institut für Medizinische Genetik und Humangenetik Charité Universitätsmedizin Berlin Genomics: Lecture #2 WS 2014/2015 Today Genome assembly: the basics Hamiltonian and Eulerian
Mapping de Novo Assembly Institut für Medizinische Genetik und Humangenetik Charité Universitätsmedizin Berlin Genomics: Lecture #2 WS 2014/2015 Today Genome assembly: the basics Hamiltonian and Eulerian
Pre-processing and quality control of sequence data. Barbera van Schaik KEBB - Bioinformatics Laboratory
 Pre-processing and quality control of sequence data Barbera van Schaik KEBB - Bioinformatics Laboratory b.d.vanschaik@amc.uva.nl Topic: quality control and prepare data for the interesting stuf Keep Throw
Pre-processing and quality control of sequence data Barbera van Schaik KEBB - Bioinformatics Laboratory b.d.vanschaik@amc.uva.nl Topic: quality control and prepare data for the interesting stuf Keep Throw
SAM / BAM Tutorial. EMBL Heidelberg. Course Materials. Tobias Rausch September 2012
 SAM / BAM Tutorial EMBL Heidelberg Course Materials Tobias Rausch September 2012 Contents 1 SAM / BAM 3 1.1 Introduction................................... 3 1.2 Tasks.......................................
SAM / BAM Tutorial EMBL Heidelberg Course Materials Tobias Rausch September 2012 Contents 1 SAM / BAM 3 1.1 Introduction................................... 3 1.2 Tasks.......................................
Using many concepts related to bioinformatics, an application was created to
 Patrick Graves Bioinformatics Thursday, April 26, 2007 1 - ABSTRACT Using many concepts related to bioinformatics, an application was created to visually display EST s. Each EST was displayed in the correct
Patrick Graves Bioinformatics Thursday, April 26, 2007 1 - ABSTRACT Using many concepts related to bioinformatics, an application was created to visually display EST s. Each EST was displayed in the correct
When we search a nucleic acid databases, there is no need for you to carry out your own six frame translation. Mascot always performs a 6 frame
 1 When we search a nucleic acid databases, there is no need for you to carry out your own six frame translation. Mascot always performs a 6 frame translation on the fly. That is, 3 reading frames from
1 When we search a nucleic acid databases, there is no need for you to carry out your own six frame translation. Mascot always performs a 6 frame translation on the fly. That is, 3 reading frames from
10/15/2009 Comp 590/Comp Fall
 Lecture 13: Graph Algorithms Study Chapter 8.1 8.8 10/15/2009 Comp 590/Comp 790-90 Fall 2009 1 The Bridge Obsession Problem Find a tour crossing every bridge just once Leonhard Euler, 1735 Bridges of Königsberg
Lecture 13: Graph Algorithms Study Chapter 8.1 8.8 10/15/2009 Comp 590/Comp 790-90 Fall 2009 1 The Bridge Obsession Problem Find a tour crossing every bridge just once Leonhard Euler, 1735 Bridges of Königsberg
2) NCBI BLAST tutorial This is a users guide written by the education department at NCBI.
 Web resources -- Tour. page 1 of 8 This is a guided tour. Any homework is separate. In fact, this exercise is used for multiple classes and is publicly available to everyone. The entire tour will take
Web resources -- Tour. page 1 of 8 This is a guided tour. Any homework is separate. In fact, this exercise is used for multiple classes and is publicly available to everyone. The entire tour will take
DNA Inspired Bi-directional Lempel-Ziv-like Compression Algorithms
 DNA Inspired Bi-directional Lempel-Ziv-like Compression Algorithms Transmission Systems Group (TrSys) http://trsys.faculty.jacobs-university.de Attiya Mahmood a.mahmood@jacobs-university.de Nazia Islam
DNA Inspired Bi-directional Lempel-Ziv-like Compression Algorithms Transmission Systems Group (TrSys) http://trsys.faculty.jacobs-university.de Attiya Mahmood a.mahmood@jacobs-university.de Nazia Islam
Package LncFinder. February 6, 2017
 Type Package Package LncFinder February 6, 2017 Title Long Non-Coding RNA Identification Based on Features of Sequence, EIIP and Secondary Structure Version 1.0.0 Author Han Siyu [aut, cre], Li Ying [aut],
Type Package Package LncFinder February 6, 2017 Title Long Non-Coding RNA Identification Based on Features of Sequence, EIIP and Secondary Structure Version 1.0.0 Author Han Siyu [aut, cre], Li Ying [aut],
10/8/13 Comp 555 Fall
 10/8/13 Comp 555 Fall 2013 1 Find a tour crossing every bridge just once Leonhard Euler, 1735 Bridges of Königsberg 10/8/13 Comp 555 Fall 2013 2 Find a cycle that visits every edge exactly once Linear
10/8/13 Comp 555 Fall 2013 1 Find a tour crossing every bridge just once Leonhard Euler, 1735 Bridges of Königsberg 10/8/13 Comp 555 Fall 2013 2 Find a cycle that visits every edge exactly once Linear
Genome Browsers - The UCSC Genome Browser
 Genome Browsers - The UCSC Genome Browser Background The UCSC Genome Browser is a well-curated site that provides users with a view of gene or sequence information in genomic context for a specific species,
Genome Browsers - The UCSC Genome Browser Background The UCSC Genome Browser is a well-curated site that provides users with a view of gene or sequence information in genomic context for a specific species,
Long Read RNA-seq Mapper
 UNIVERSITY OF ZAGREB FACULTY OF ELECTRICAL ENGENEERING AND COMPUTING MASTER THESIS no. 1005 Long Read RNA-seq Mapper Josip Marić Zagreb, February 2015. Table of Contents 1. Introduction... 1 2. RNA Sequencing...
UNIVERSITY OF ZAGREB FACULTY OF ELECTRICAL ENGENEERING AND COMPUTING MASTER THESIS no. 1005 Long Read RNA-seq Mapper Josip Marić Zagreb, February 2015. Table of Contents 1. Introduction... 1 2. RNA Sequencing...
Copyright 2014 Regents of the University of Minnesota
 Quality Control of Illumina Data using Galaxy August 18, 2014 Contents 1 Introduction 2 1.1 What is Galaxy?..................................... 2 1.2 Galaxy at MSI......................................
Quality Control of Illumina Data using Galaxy August 18, 2014 Contents 1 Introduction 2 1.1 What is Galaxy?..................................... 2 1.2 Galaxy at MSI......................................
Research Article An Improved Scoring Matrix for Multiple Sequence Alignment
 Hindawi Publishing Corporation Mathematical Problems in Engineering Volume 2012, Article ID 490649, 9 pages doi:10.1155/2012/490649 Research Article An Improved Scoring Matrix for Multiple Sequence Alignment
Hindawi Publishing Corporation Mathematical Problems in Engineering Volume 2012, Article ID 490649, 9 pages doi:10.1155/2012/490649 Research Article An Improved Scoring Matrix for Multiple Sequence Alignment
