Reproducibility of Whole-brain Structural Connectivity Networks
|
|
|
- Loren McKinney
- 5 years ago
- Views:
Transcription
1 Reproducibility of Whole-brain Structural Connectivity Networks Christopher Parker Thesis for Masters of Research in Medical and Biomedical Imaging Supervised by Prof. Sebastien Ourselin and Dr Jonathan Clayden August 2012 Words: 10,117
2 Introduction The brain is composed of billions of neurons which send and receive information between one another. Groups of functionally distinct and spatially separate neuronal populations are highly concentrated in areas of grey matter. Information transfer occurs between grey matter regions on both a local and global scale, via white matter fiber tracts, providing the physical infraastructure that supports functional interaction. Studying the whole-brain structural network using diffusion MRI tractography has recently become a popular research topic in neuroscience [1]. Here, one seeks to quantify the integrity of connections between grey matter regions via white matter pathways in vivo, on a global scale, in order to depict the topological organisation of brain structure and relate this to aspects of health and disease. The natural division of the brain into a set of connected grey matter regions forms a brain graph: allowing quantification of overall connectivity using graph theoretical analysis- a mathematical framework for analysing graphs. However, defining brain network nodes and edges is both a conceptual and practical challenge [2]. For example, how many nodes are there in the brain? and how does one construct an accurate and reproducible network of anatomical connectivity? A variety of methodological choices have been implemented to obtain the networks and it is not clear how these choices affect the anatomical accuracy of the network, the network metrics themselves, or network reproducibility. This report seeks to investigate the effect of different methodologies on the values of structural network measures and their reproducibility. Previous studies have shown that, like many other naturally occurring complex systems, such as the internet and transport network, the structural network has a small-world topology [1,3,4]. Such networks are characterised by a similar global pathlength (the average number of edges traversed to connect all nodes pairs) and a higher global clustering coefficient (the average connection density between nearest neighbours of a node) than equivalent measures found in randomly generated networks with the same degree distributions. This organisation enables efficient information transfer between any two elements of the system and simultaneously a high degree of local communication, supporting the concept of segregation (highly interconnected sub-groups of brain regions, e.g. retinotopic maps) and integration (connections between sub-groups e.g. language systems) of information transfer in the human brain. The structural network contains hubs; regions that are highly connected or highly central to network communication [5,6,7]. Occipital and parietal cortical regions are frequently highly connected and contain hub regions such as the left and right precentral cortical gyri and precuneus. In addition, networks have been split into communities of nodes (or modules), defined as sets of nodes which have relatively higher connectivity within than between communities. A comparison of diffusion-weighted imaging and resting-state fmri networks reveals a close relationship between structural and functional connections, including a significant overlap in the hub regions found in structural and functional networks [8], supporting the idea that structural networks represent the physical backbone that constrains functional interactions. Promisingly, it has been demonstrated that the structural network may reflect neurological phenotype. For example, network metrics are sensitive to the effect of age [9], gender [10] and IQ [11]. In addition, network alterations have been observed in relation to neurological diseases such as Alzheimer's disease [12,13], epilepsy [14,15] and schizophrenia [16,17,18], meaning such metrics may become useful as topological biomarkers of brain integrity. Despite a large body of literature on the topic of structural networks, there is little agreement between studies of how to define network nodes or edges. Nodes of the brain network may defined using different scales and parcellation schemes. The most commonly used technique has been to warp the subjects structural image to an anatomical atlas template, such as the AAL atlas [5,11,16,18], where the grey matter regions are labelled according to anatomical landmark criteria. Atlas warping parcellations are favoured as they are relatively quick, but because of the spatial normalisation involved, do not accurately capture the individual variation in cortical anatomy between subjects. To overcome this, some studies have used a subject-specific parcellation such as Freesurfer, where the majority of the segmentation is performed in subject native space [19,20]. The majority of whole-brain parcellations have been relatively coarse, resulting in the order of 100 nodes. With these coarse parcellations, the boundaries between nodes may intersect or combine a single tract, dividing the connectivity properties of the tract between two or more nodes. Because of the uncertainty of where to place node boundaries, a number of studies have divided parcellated regions into smaller patches (in a pseudorandom fashion) [4,21], or used a range of parcellation scales to perform network analysis [6,22,23]. Random parcellations are quick, flexible and avoid the issue of where to place node boundaries, but have no neuroscientific meaning. A recent study has reported the effect of parcellation
3 scale on network properties [24]. Network properties such as small-worldness were conserved across different parcellation scales although the values varied in a predictable but scale-dependent manner. The authors examined repeated random parcellations at each scale, and concluded that parcellation scale as opposed to parcellation scheme was the primary determinant of network topology. There is also variation in the method used to obtain network edges. In structural networks, an edge represents a neural fiber tract that connects two nodes. Measures of edge connectivity are frequently quantified based on the number of streamlines connecting regions and as such the issue of which diffusion model, tracking technique and seed location to use arises. Due to its low computational and acquisition demand, the diffusion tensor model [25], combined with deterministic tracking, is the most commonly used technique for reconstructing network edges [7,20,22,26]. The diffusion tensor model assumes there is a single fiber population in each voxel. Neural tissue however, contains a mixture of extracellular and intracellular space, isotropic and anisotropic compartments and may contain multiple fiber populations with different orientations. Streamline tracking using diffusion tensors can therefore result in erroneous tracking through regions containing crossing fibers. Other models of the diffusion signal may be fitted to the data, such as the orientation distribution function (ODF) [27], multi-compartment models [28,29] and the fiber orientation distribution [30]. Aswell as reflecting true anatomical complexity in fiber orientations, these models also incorporate measures of uncertainty into each orientation. The ODF is more commonly used for defining fiber orientations in structural network analysis [4] than multi-compartment models [31]. Multicompartment models such as the ball and two sticks model [32] may be more comprehensive than the simple diffusion tensor model, but also have some disadvantages. The number of fibers that can be represented in a voxel is commonly limited by the angular resolution of the data and they are also more computationally demanding to implement than the diffusion tensor model. A technique termed spherical deconvolution has been developed, which can resolve a higher number crossing fibers with high angular resolution of fiber orientations [30]. In contrast to other multi-fiber reconstructions, the spherical deconvolution is model-free, can represent a high number of possible fiber crossing, and does not require high angular resolution data. Deterministic or probabilistic tracking techniques may be used to track through fiber orientations. The majority of structural network reconstructions have used deterministic [10,12,20], as opposed to probabilistic streamline tractography [19,33,34]. Deterministic tractography, which simply follows the directions of maximum fiber coherence, does not track through branching fiber architecture. Probabilistic approaches on the other hand produce a dispersion of streamlines and therefore a probability of connection to other voxels from a seed voxel [28]. This enables quantification of the connection probability throughout the brain, which may be desirable for constructing structural networks when the intrinsic connection sparsity is unknown. The deterministic tracking algorithm has been shown to be inferior to probabilistic tracking approaches when considering the receiver operating characteristics of the human structural network compared a structural network of the macaque cortex derived from tract tracing techniques [31]. Also, some network metrics obtained by using probabilistic tractography [33- Vaessen et. al. 2010] were more reproducible than those using deterministic tractography [2- Bassett et. al. 2012, 35- Cheng et. al. 2012]. Different pipeline streams vary in their sensitivity to acquisition reproducibility in diffusion weighted or structural data. Intra-subject reproducibility has been investigated, using repeated diffusionweighted scanning, with respect to both diffusion signal acquisition scheme [Vaessen et. al. 2010, Bassett et. al. 2010] and parcellation scheme and scale [Bassett et. al. 2010] in a small number of subjects (n=6 and 7). [Vaessen et. al. 2010] examined the effect of varying the number of diffusion gradient directions on the network characteristics and their intra-subject reproducibility. They found that increasing the number of diffusion directions resulted in a higher number of longer streamlines, which significantly affected the global pathlength of the network. The number of gradient directions had no significant effect on the intra-subject reproducibility, although a trend of decreasing reproducibility was observed with increasing network sparsity. High coefficients of variation (CV) for the raw connection strengths were reported, indicating relatively low reproducibility across repeat scans, whereas reproducibility was high for mean network measures such as clustering and pathength. [Bassett et. al. 2010] examined the effect of varying parcellation scheme and parcellation scale on reproducibility of network properties using DTI and DSI acquisitions. While the study found that some global properties of both DTI and DSI networks were highly reproducible, more widely used metrics such as clustering coefficient, pathlength and small-worldness had poorer reproducibility. Networks obtained by DSI had a higher number of reconstructed tracts and larger mean connection strengths between nodes, agreeing with other studies showing an increased number of tracts from
4 higher angular resolution-acquired data [24, Vaessen et. al. 2010]. The parcellation and acquisition scheme giving the highest reproducibility was the AAL atlas with a DT acquisition. A plausible explanation was for this was that the longer tracts in DSI networks would be subjected to a larger cumulative noise and therefore greater variation of connection strengths across repeat scans. Another explanation was that the whole-brain templates used for parcellation would be more sensitive to poor inter-subject concordance of brain region parcellation, which would disproportionately affect the denser DSI networks. A further explanation was the fact that DSI acquisition were ~5 times longer than DTI, and used higher b-values, which decreased the signal to noise ratio of the data. A drawback of these studies is the small number of subjects used, limiting accurate measurement of connection variability. More recently, the intra- and inter-subject reproducibility of the raw connection matrix and the subsequent network metrics obtained has been studied in a larger group of subjects [Cheng et. al. 2012]. The authors used a subject-specific parcellation with deterministic tractography and used mostly measures of correlation to describe variability. Intra-subject correlation in edge weights was higher than inter-subject correlations, whereas the intra-subject correlation in global network measures was markedly lower and varied across different network metrics. The CV of edge weights between subjects was in the range of 0 160%, although this value was not reported for intra-subject edge weights. The CV for graph theoretical measures between subjects depended on which network metric was being considered, but was generally in the range of 4-10% for global metrics, and markedly higher for local (per-node) metrics. CV was not reported for intra-subject reproducibility of graph theoretical metrics. Similarly to [Bassett et. al. 2010], relatively low Intra-Class Correlation (ICC) values (below or equal to 0.5) were reported for global network metrics, indicating the between-subject variability outweighed within-subject variability. Although [Cheng et. al. 2012] used a larger number of subjects, the effects of different parcellation, diffusion modelling or fiber tracking scheme on reproducibility were not considered. Also, a simple measure of correlation was used to examine intra-subject reproducibility using only two repeat scans. Correlation may be an unsuitable measure to assess reproducibility since it assumes a statistical independence of data points. Furthermore, with the exception of [Vaessen et. al. 2010], these reproducibility studies used DT tractography to reconstruct connecting fibers. Despite using a subject-specific parcellation, [Cheng et. al. 2012] reported similarly low global network reproducibility values to that described in [Bassett et. al. 2010]. From these reproducibility studies a general trend is observable; connection strength reproducibility within and between subjects is low compared to graph theoretical measures. Global graph metrics had higher intra-subject reproducibility than local metrics, but as indicated by the moderate ICC values reported in [Bassett et. al. 2010] and [Cheng et. al. 2012], this variability is equal or only slightly smaller than betweensubject variability. Statistically, the connection strength has high CV, graph measures have a lower CV, but also an ICC around 0.5. It is clear that the values of reproducibility obtained depends on the parcellation scheme, tractography algorithm and connection weighting scheme and as admitted in [Cheng et. al. 2012], structural network reconstruction is still facing some technical challenges before being used as a reliable biomarker. A complete pipeline to maximise reproducibility by incorporating processing streams that better depict the true anatomy has not been described. Improvements in anatomical accuracy and intra-subject reproducibility of structural networks may be realised by using a more accurate model of diffusion combined with a subject-specific parcellation. In this report, the concepts of subject-specific parcellation, multi-fiber modelling, and probabilistic tracking techniques are combined and applied to the problem of network reproducibility in a large number of subjects. Two reconstruction pipelines were used to reconstruct structural networks from diffusion-weighed data acquired in triplicate. Pipeline 1 (P1) uses widely available and commonly used tools whereas pipeline 2 (P2) uses more recently developed algorithms. It is expected that with better methods for reconstructing network nodes and edges, the reproducibility of connection weights and network metrics will be higher.
5 Methods Subjects and Image Acquisition Twenty eight healthy young adult subjects (16 male, 12 female, mean age 28.5 standard deviation 3.9 years) participated in this study. Subjects had no prior history of neurological impairment and no brain abnormalities at the time of scanning, as determined by examination of their structural scan by an expert radiologist. Two T1-weighted images of (1mm) 3 resolution were acquired sequentially with a 3D Fast Low- Angle Shot (FLASH) sequence (176 contiguous sagittal slices, 256 x 224 mm FOV, TR=11ms, TE=4.94ms and α=15 o ) on a 1.5T Siemens Avanto MRI scanner at Great Ormand Street Hospital, London. A diffusionweighted echo planar sequence (TR = 7300ms, TE = 81ms) with 60 orthogonal diffusion directions (b = 1000s/mm -2 ) was used to acquire diffusion-weighted images of (2.5mm) 3 and three un-weighted diffusion images (b=0 images). The diffusion-weighted sequence was repeated three times for each subject to allow assessment of intra-subject reproducibility. Image Processing DICOM images were converted into NIFTI format and the brain was extracted from all images using FSL's brain extraction tool [36]. In order to increase the signal to noise ratio of the structural image, the two acquired T1-weighted images were bias field corrected, registered and averaged in Freesurfer v The diffusion-weighted volumes were corrected for eddy-current induced distortions by affine registration to the no diffusion-weighted image using the diffusion-specific FSL FDT algorithm [37]. Cortical Parcellation To define network nodes, the cortical grey matter voxels of the T1-weighted images were parcellated into regions using automated software. Pipeline 1. Freesurfer was used to parcellate the structural image into 68 cortical regions (34 for each hemisphere), as defined by the Desikan-Killiany Atlas [38]. The parcellation scheme assigns a neuroanatomical label to each location on a cortical surface model of the image, based on probabilistic information from a manually labeled training set [39]. Regions of interest (ROI) were extracted from the Freesurfer aparc+aseg volume. Pipeline 2. NiftySeg was used to parcellate the structural image into 44 brain regions (22 for each hemisphere), as defined by the Hammer's Atlas [40]. In this parcellation scheme, multi-steps first labels brain regions by propagating a set of manually labelled T1-weighted images to the subjects structural image. Multi-STEPs implements an expectation-maximization algorithm with a Markov random field regularization to produce accurate segmentations that are also computationally efficient. The LoAd algorithm was then applied to obtain the grey cortical matter of the parcellated regions [41]. For a full list of labels for both pipelines see Table A.1. Figure 1. Representative cortical parcellations shown in axial orientation. Freesurfer (left) and NiftySeg (right) were used to parcellate the grey matter of the cortex into 68 and 44 ROI, respectively. Region names and labels are listed in Table A.1.
6 Registration To segment the cortical structures of interest in native diffusion space, the structural and diffusionweighted images were co-registered. The registration field was determined as follows. An affine registration was used to register the b=0 image to the higher resolution averaged T1-weighted image. The T1-weighted image was then non-linearly registered to the b=0 image using the inverse of the transformation acquired in the previous stage as a starting transformation. The transformation field was retained and applied to each ROI to transform them into diffusion space. Label intensities were preserved in the final transformation by using nearest neighbour interpolation. Pipeline 1. FSL FLIRT was used to register b=0 image to the T1-weighted image. The registration algorithm used a normalised correlation cost function with 12 degrees of freedom and trilinear interpolation was used to estimate voxel intensities between voxels occurring due to sub-voxel transformations [37]. FSL FNIRT was then used to register from T1 to diffusion space. Pipeline 2. Niftyreg was used to perform the affine and non-linear registrations using the default settings. The non-linear registration implemented in Niftyreg is a re-factorisation of the existing non-linear registration, which uses cubic B-spline interpolation with normalised mutual information [42] as the cost function, and as such is more computationally efficient than existing non-linear registration methods [43]. Diffusion Modelling To infer the underlying orientations of fiber bundles at each voxel, a model can be fitted to or extrapolated from the diffusion signal. Pipeline 1. A ball and two sticks fibre model was fitted to the diffusion data. The Bayesian Estimation of Diffusion Parameters Obtained using Sampling Techniques (BEDPOSTX) algorithm was used to fit a 2 fibre model to each voxel. The BEDPOSTX algorithm uses Markov chain Monte Carlo sampling to estimate the fibre orientation dispersions [32]. Pipeline 2. Constrained spherical deconvolution was applied to estimate the underlying fibre orientation distributions in each voxel, using MRTrix [44]. The maximum number of allowable fiber populations was set to 8. In contrast to pipeline 1, this diffusion model makes no a priori assumptions on the number of fibre populations present in each voxel. Probabilistic Tractography In order to track the paths of fiber trajectories arising from each cortical region, 100 probabilistic streamlines were seeded for each ROI voxel. Pipeline 1. The default settings in FSL ProbTrack algorithm were used to determine the streamline trajectories [28]. The sampling interval was 0.5 mm and stopping criteria meant that tracks terminate if they curve by more than 80º. Pipeline 2. Streamlines were propagated using the default settings in MrTrix [45]. The sampling interval was 0.2 mm, maximum curvature threshold was 60º and minimum FA threshold for tracking through a voxel was 0.1. For both tracking schemes, the stopping criteria prevents streamlines from leaving the brain mask or from passing through a voxel that has already been visited. Note, using these default parameter settings, pipeline 2 initiates streamlines (from voxels with FA > 0.2) by uniform sampling from the ROI until (100 x the number of voxels in the ROI) are produced that pass the minimum acceptance criteria (a minimum streamline length of 10 mm), whereas pipeline 1 produces 100 streamlines uniformly from each voxel regardless of whether they pass the minimum acceptance criteria. Network Construction Connection Strengths To construct the network, cortical regions were represented as network nodes and the fiber connections between them as edges. The connection strength between two cortical regions was defined as the fiber density of connecting fibres (as in [46]) and was calculated as the sum of connecting streamlines divided by the mean volume of the two ROIs. Performing this calculation between N cortical regions produces an N-by-N directional association matrix of connection strengths between all regions. As diffusion MRI does not contain directional information, the value of connectivity was averaged for each region pair, resulting in an undirected graph. This network connection strength matrix was calculated for the repeat scans of all subjects.
7 Binary Networks Graph theoretical analysis was performed on binary networks derived from the connection strength matrices. Binary networks assume that connections are either absent or present. To obtain a binary network, a threshold is applied to the connection strength matrix. Thresholding allows comparability between networks and reduces the number of false positive connections. Given that graph theoretical values of binary networks are highly dependent on the number of connections in the graph, choosing a single connection strength threshold is sub-optimal for comparing networks that have differing connection strength distributions (see Fig. 2). The connection density of a graph describes the fraction of edges that exist in the network as a proportion of the total possible connections present before thresholding. Using a fixed connection density, where a given fraction of the strongest connections are retained, allows one to study the overall topology of the network without confounding effects from varying the number of edges. Once the binary network is obtained, whole-brain (global) and single region (local) network connectivity measures can be derived using graph theory. Since it is unclear which connection density to use to analyse brain networks, graph theoretical metrics were obtained for a range of densities (from 0.1 to 0.5) for each network. Figure 2. Distribution of connection strengths in the connection strength matrix of the first scan for each subject in P1 (left) and P2 (right) networks. Shown are the mean and interquartile ranges of the connection strength distributions. Outlying connection strengths are not shown. Connection strength distributions differ greatly between subjects meaning that using single value threshold to produce binary graphs will result in networks having different numbers of connections. Figure. 3. An image representation of a connection strength matrix (left) and the corresponding binary adjacency matrix (right) for subject 1, produced by thresholding the connection strength matrix so that a given number of strongest connections are retained. With the matrix origin in the bottom-left corner, entries describe the connection strength or connection presence between the seed region (matrix column) and target region (matrix row). Adjacency matrix entries of 0 and 1 are displayed as black and white, and represent the lack of or presence of a connection, respectively.
8 Graph Theoretical Anaysis Graph theoretical analysis was applied to binary networks to quantify local and global network metrics and their reproducibility. The global metrics of clustering coefficient, pathlength and smallworldness were calculated for all networks. The local metrics of clustering coefficient, pathlength and centrality were also calculated for all nodes in all networks. The clustering coefficient of a node measures the extent to which the connections from a particular node are themselves inter-connected. The local clustering coefficient was calculated as the number of connections existing between the nearest neighbours of a node divided by the total number of possible connections between neighbours given their node degrees [3]. The global clustering coefficient was calculated as the mean clustering coefficient of all nodes. The pathlength describes how close nodes are in terms of network topology. A path exists between two nodes that can be connected by some combination of edge traversal. The path length between two nodes is calculated as the minimum total number of edges traversed to travel through the network from one node to another [3]. The local pathlength of a node is the average of path lengths that connect this node to all other nodes. The global pathlength is the mean of all local pathlengths in a network. The global pathlength gives a general indication of the mean number of paths travelled to connect any two nodes in the network. Betweenness centrality (referred to hereafter as centrality), is a measure of how essential a given node is as an intermediate node in all minimum paths between other nodes in the network [47]. It is the cumulative number of shortest paths between all other nodes in the network that pass through the given node, divided by the total number of connections in the network. The small world phenomenon is a commonly reported characteristic in complex networks and signifies a network architecture which is somewhere between a random network and a regular lattice [3]. A random network will have a relatively short global pathlength and low global clustering coefficient compared to a regular lattice, which will exhibit higher mean path length and higher clustering. A small world network has either a shorter global pathlength than a random network, higher global clustering, or both. The smallworldness of a network can be calculated as the ratio of global clustering coefficient to global pathlength after both measures have been normalised (divided) by their equivalent values found in randomly generated networks. To make this calculation, networks of the same number of nodes and connection density, but a random combination of connections, were generated and their global pathlength and global clustering coefficient calculated. For each network, the random network pathlength and clustering was calculated as the mean across three trials. A small-worldness, or small-world index (SWI), greater than 1 indicates that the network has small-world properties [50]. All graph theoretical analyses used in this study were performed in-house using the R programming language (v2.14.2).
9 Network Reproducibility Reproducibility of the raw connection strengths, global network measures (global clustering coefficient, global pathlength and small-world index) and local network measures (local clustering coefficient, local pathlength and local centrality) were calculated for both pipelines. To give a general indication of local reproducibility, the reproducibility values described below are reported as an average across all nodes. Reproducibility was assessed using the Coefficient of Variation (CV) and Intra-class Correlation Coefficient (ICC). The CV measures the standard deviation of a value as a fraction of the mean, while the ICC estimates the ratio of between subject variance to total variance. Intra-subject CV was calculated as the mean intra-subject standard deviation divided by the overall subject mean. Inter-subject CV was calculated as the standard deviation of the intra-subject means, divided by the overall subject mean. ICC was calculated as the between-subject variance (variance of the intra-subject means) divided by the total variance, where the total variance is the between-subject variance plus the within-subject variance (the average intra-subject variance). An ICC above 0.5 indicates good intra-subject reproducibility, as the inter-subject variance is greater than the intra-subject variance.
10 Results Connection Strengths Connection strengths determine which edges are included in the network and are therefore of great importance to structural network analysis. P1 and P2 networks had different raw connection densities (Fig. 4, right). P1 networks had a mean density of 0.67 ± 0.12 (± standard deviation) across all subjects, whereas P2 networks had a mean density of 0.98 ± The connection density ranged from 0.48 to 0.89 for P1 networks and 0.91 to 1.00 for P2 networks. To compare overall connectivity strengths between pipelines P1 and P2, the subject grand mean connectivity matrix was calculated by averaging all connection matrices. Fig 4 shows that the two pipelines resulted in different patterns of connectivity strengths. The average connection strength was 0.05 for P1 and 0.15 for P2. Networks in both pipelines had higher raw connection strengths between intra-hemispheric regions. The mean intra-hemispheric connection strength was 0.09 for P1 and 0.24 for P2, whereas the mean inter-hemispheric connection strength was 0.01 for P1 and 0.05 for P2. Inter-hemispheric connections between homologous left-right regions were also high for both pipelines, with a mean of 0.29 and 0.56 for P1 and P2. Figure 4. Connection strength matrices for P1 and P2. (Left) P1 grand mean connection strength matrix. (Middle) P2 grand mean connection strength matrix. (Right) Point plot of connection densities of all networks for P1 and P2. Connection density is calculated as the number of non-zero connections as a proportion of the total number of possible connections. P2 networks have higher mean connection strengths and higher raw connection densities. Intra- and inter-subject variability of connection strengths differed greatly in P1 and P2 networks (Fig. 5). The mean intra-subject CV of all connections in P1 networks was 0.66 ± 0.37, whereas the mean for P2 networks was 0.26 ± Intra-subject CV ranged from 0.05 to 1.73 in P1 networks, whereas P2 ranged from 0.04 to For both P1 and P2, the intra-subject CV was lower for intra- than inter-hemispheric connections (Fig. A.1). Inter-subject variability was higher in magnitude but showed a similar pattern to the intra-subject variability (Fig. A2). The mean inter-subject CV of all connections in P1 networks was 1.77 ± 0.82, whereas the mean for P2 networks was 0.71 ± Inter-subject CV ranged from 0.18 to 5.29 in P1 networks, whereas P2 ranged from 0.07 to For both P1 and P2, the inter-subject CV was lower for intra- compared with inter-hemispheric connections.
11 Figure 5. Reproducibility of raw connection strengths in pipelines 1 and 2. Boxplots show the distribution and range of intra- and inter-subject CV's. For both P1 (dark-red and red boxes) and P2 (dark-blue and blue boxes), inter-subject CV was generally higher than intra-subject CV. In P2 networks, both intra- and intersubject CVs of connections have smaller distributions than in P1 networks. For both pipelines, the ICC for all connections was high (all above 0.5, Fig. A.3), indicating that inter-subject variance was greater than intra-subject variance. Over all connections, the mean ICC was for both pipelines. In contrast to CV values, little difference was found in the ICC of intra- vs interhemispheric connections.
12 Graph Theory Metrics Global Metrics To describe the overall relationship between network metrics and connection density, the mean global metrics were calculated across subjects for each connection density. Fig. 6 shows that for both pipelines, global metric were dependent on the connection density. Clustering coefficient increased from 0.49 to 0.72 in P1 networks and 0.40 to 0.70 in P2 networks. Pathlength decreased from 3.0 to 1.48 in P1 and 3.44 to 1.47 in P2. SWI decreased from 3.91 to 1.44 in P1 and 3.23 to 1.39 in P2, respectively. Figure 6. Global network characteristics for P1 and P2 networks over a range of connection densities. Shown is the average metric over all subjects ± the standard error of the subject means for (Left) clustering coefficient, (middle) pathlength and (right) SWI. P1 is shown in red and P2 is shown in blue. P1 networks have higher mean clustering, lower mean pathlength and higher mean SWI over all connection densities. Intra- and inter-subject variability of global graph metrics differed in P1 and P2 networks. The intrasubject CV was low for global clustering coefficient, pathlength, centrality and SWI and decreased moderately with increasing network density in both pipelines (Fig. 7). The inter-subject CV of global metrics was higher than intra-subject CV and also decreased with increasing network density. Although the decrease was not uniform, global clustering coefficient intra-subject CV decreased from to 0.008, global pathlength decreased from to and SWI decreased from to with network densities of 0.1 to 0.5 in P1 networks. Global clustering coefficient intra-subject CV decreased from to 0.007, global pathlength decreased from to and SWI decreased from to with network densities of 0.1 to 0.5 in P2 networks. Global clustering coefficient inter-subject CV decreased from to 0.017, global pathlength decreased from to and SWI decreased from to with network densities of 0.1 to 0.5 in P1 networks. In P2 networks, inter-subject CV decreased from to for global clustering coefficient, to for global pathlength and to for SWI.
13 Figure 7. Reproducibility of global network metrics over a range of connection densities. Plotted are the intra-subject CV (triangles) and inter-subject CV (squares) for (left) clustering coefficient, (middle) pathlength and (right) SWI. P1 is plotted in blue and P2 is in red. Inter-subject CV is lower than intra-subject CV over all connection densities for all network metrics. For both pipelines, SWI has the highest variability at low connection densities whereas all metrics have lower CV at higher densities. For both pipelines, the ICC of global metrics was generally high (above 0.5) for clustering coefficient, pathlength and SWI, indicating that between-subject variation was greater than within-subject variation for these metrics. P2 networks had higher global clustering coefficient ICC than P1 networks across all network densities. For global pathlength, P1 networks had greater ICC than P2 networks. For both pipelines, the ICC of small-world index was lower at densities of 0.25 or less, whereas the ICC was high for densities of 0.3 or more (>0.75). Figure 8. Reproducibility of global network metrics over a range of connection densities. The ICC is plotted for clustering coefficient (green), pathlength (blue) and small-world index (red) for P1 (left) and P2 (right) networks. ICC is greater than 0.5 for all metrics in both pipelines (apart from SWI at densities of 0.2 or less for P1 and 0.1 for P2), indicating good intra-subject reproducibility.
14 Local Metrics Intra- and inter-subject variability of local graph metrics did not differ greatly between P1 and P2 networks. For both pipelines, local metrics had higher intra and inter-subject CVs than their equivalent global metrics (Fig. 9). As with global metrics, inter-subject was greater than intra-subject CV, suggesting good inter-scan reproducibility. For both pipelines the intra and inter-subject CV generally decreased with network density for clustering coefficient and centrality, although the relationship was less strong than with global metrics. The mean intra-subject CV of local metrics decreased from to and to for clustering coefficient and centrality respectively, for P1 networks. For P2 networks, the mean intrasubject CV decreased from to and to for clustering coefficient, and centrality. For both pipelines, the mean intra-subject CV was similar across all densities for local pathlength (~ 0.021). Figure 9. Reproducibility of local network metrics over a range of connection densities. Plotted are the mean intra-subject (triangles) and inter-subject (squares) CV over all nodes for P1 (red) and P2 (blue) ± the standard deviation of the CV across nodes. Local metrics have higher CV than global metrics, although intersubject CV is higher than intra-subject CV for (left) clustering coefficient, (middle) pathlength and (right) centrality, indicating good reproducibility. For both pipelines, the between-subject variability of local metrics was higher than within-subject variability for most nodes (Fig. 9). The proportion of total variance accounted for by between-subject variance was higher in local than global metrics. Across nodes, the mean ICC increased marginally with increasing density. In P1 networks, the local clustering coefficient, pathlength and centrality mean ICC was above 0.7 for for all network densities. For P2 networks, the local mean ICC was above 0.65 for for all connection densities.
15 Figure 10. Reproducibility of local network metrics over a range of network densities. The ICC is plotted for clustering coefficient (green), pathlength (blue) and centrality (purple). Points show the mean ICC of all nodes ± the standard deviation of the ICC across nodes. Both pipelines give high ICCs for local metrics.
16 Discussion Studying the whole-brain structural network using diffusion MRI tractography has begun to provide new insights into the anatomical organisation of nerve fibers in vivo. By quantifying local and global network architecture, these analyses have potential to detect both widely distributed and targeted alterations in nerve fiber integrity that may occur due to neurological variation or disease. However, the different parcellation and tractography schemes that have been used to define network nodes and edges have resulted in variability of network metrics between studies, meaning the effect of methodology on network architecture is a confounding factor in structural network studies. In addition, different schemes may be differently sensitive to noise and other image artefacts present in diffusion-weighted data, limiting their ability to describe true differences in network architecture between subjects. In this report we investigated the intra- and inter-subject reproducibility of network metrics obtained using two different reconstruction pipelines in a large group of subjects undergoing repeat diffusion-weighted scanning. In both of the pipelines, the network characteristics displayed similar relationships with randomly generated graphs, such as higher clustering and lower pathlength. Differences between pipelines were observed in terms of network values and their intra- and inter-subject variability in both raw connection strengths and global and local network metrics. The raw connection matrices were densely connected for both pipelines (Fig. 4). For both pipelines, intra-hemspheric connections were stronger than inter-hemispheric connections. In addition, interhemispheric connections between homologous left-right hemisphere regions were also stronger. This pattern of connectivity has been previously demonstrated in other whole-brain structural networks [5, Cheng et. al. 2012]. Due to the proximity of intra-hemispheric regions, it is not clear whether the connectivity pattern represents greater anatomical connectivity within hemispheres, or, since intra-hemispheric regions are closer, a possible distance bias in the connection strength quantification. P1 networks were sparser and had lower connection strengths than P2 networks. This could be due to the different number of nodes, and therefore the differences in region volumes between the pipelines. P2 networks had fewer nodes, meaning the region volumes were larger. Conceptually, larger regions will encompass a higher number of functionally distinct neural populations. As each of these populations connects to other regions via fiber tracts, larger regions may have a higher probability of connection to other nodes. Practically, the total number of seeded streamlines was proportional to the number of voxels in the node ROI, which may have increased the connection probability between larger regions. Another explanation for increased connectivity strength and densities in P2 networks could be the number of voxel fiber populations represented by each pipeline. P2 estimates the fiber orientations using spherical deconvolution, which is capable of representing up to eight independent fiber orientations per voxel [44]. P1 used the ball and two-sticks model which can represent a maximum of two fiber orientations per voxel [32]. This difference in orientation modelling means that P2 streamlines may have been be able to track through some regions containing multiple fiber populations where P1 streamlines otherwise terminated, resulting in a higher number of longer streamlines in P2 networks and therefore higher overall connectivity. Many previous studies have obtained more sparsely connected matrices ([5, 11] for example), which are thought to represent the true sparsity of neural fibers found in tract tracing studies [51]. Studies obtaining sparse networks (around connection density) have typically employed deterministic tractography techniques, which use a low number of seeds to quantify connectivity. Probabilistic tractography techniques however utilise a larger number of seeded streamlines to capture information regarding connection probabilities between regions, resulting in a more strongly connected network [Vaessen et. al. 2010]. Due to the difficulty in validation of fiber tracts in vivo, it is not known which technique reflects the true anatomical connectivity pattern. An advantage of using probabilistic approaches is that the more densely populated connection matrices allow for greater flexibility in the choice of connection density at which to analyse the binary network. Similarities between pipelines were found in connection strength reproducibility. For both pipelines, the intra- and inter-subject CV of connections was inversely related to the connection strength (Fig. A.1, A.2). Since connection strength was proportional to the distance between regions, this suggests that a streamline distance between regions may have affected the intra- and inter-subject connection strength variability. A distance bias may occur in streamline-based tractography because of the higher cumulative noise encountered for longer streamlines. A detailed investigation of the effect of distance on connection
17 strength and reproducibility was outside the scope of the report. However, high correlations were found between connection euclidean distance, connection strength and intra- and inter-subject reproducibility for both pipelines (Table A.2), suggesting that the effect of streamline distance should be investigated in future work. A number of previous studies have suggested the existence of a streamline distance bias in connection quantification. [Cheng et. al. 2012] showed that a connection quantification method that normalised connection strengths by the connecting streamline distance improved the correlation in connection strengths between repeat scans. An approach to correct for distance bias therefore may be to incorporate measures of distance into the connection strength measure as a penalization term, as in [Cheng et. al. 2012]. For both pipelines the reproducibility of connection strengths was higher within-subjects than between-subjects, as shown by the high ICC values and higher inter- than intra-subject CV's (Fig. 5, A.1, A.2, A.3). This finding is in agreement with previous studies by [Bassett et. al. 2010] and [Cheng et. al. 2012] where measures of correlation in connection strength between repeat scans had greater within-subject than between-subject similarity. Despite high ICC values, both pipelines had high connection strength CVs (Fig. 5). Previous studies have also reported high CVs for connection strengths. [Vaessen et. al. 2010] found that intra-subject connection strength CV was in the range of %, whereas [Cheng et. al. 2012] found inter-subject CV's up to 160%. P2 networks had lower intra- and inter-subject CV's than P1 networks, indicating superior within-subject and between-subject reproducibility for this pipeline. This may be due to the higher connection strengths of P2 networks, where the higher number of connecting streamlines increased the signal to noise ratio of the connection strengths. As previously mentioned, the number of connecting streamlines may have increased in P2 networks because of the fiber orientation modelling method, the larger node volumes, or some combination of both. [Cheng et. al. 2012] found a small negative correlation between node size and the normalised standard deviation of connection strength using an identical connection quantification scheme. This suggests that node size may have a negligible effect on connection strength variation and therefore that other factors such as tractography scheme may be the cause of the differences in reproducibility between these pipelines. Another explanation may be the registration of ROI to diffusion space. Visual inspection of registrations shows that P1 registrations were poor in frontal regions, whereas P2 registrations were of higher correspondence. Examining the reproducibility of connections from frontal regions in P1 and P2 networks would allow the influence of this mis-registration to be assessed. For both pipelines a number of similarities and trends were observable in the global network metrics obtained. All networks had a SWI greater than 1, meaning that either the clustering is higher or pathlength is shorter than that found in an equivalently connected random networks (Fig. 6). Small-worldness is a phenomenon found in almost all structural network studies and suggests the network is connected in a way that promotes local specialisation and global integration of brain regions. Global clustering coefficient, which may represent local specialisation, increased as function of connection density for both pipelines. It has been demonstrated in previous studies that the distribution of streamline lengths is skewed towards short streamlines [24, Vaessen et. al. 2010], meaning that the majority of streamlines connect regions that are nearby. The increase in clustering with density may be because higher densities include a higher number of lower strength connections, which are in turn likely to be between anatomically close regions. For both pipelines, global pathlength decreased with increasing network density. This is because the addition of more connections to the network creates new paths between nodes and therefore decreases the average pathlength in the network. [Vaessen et. al. 2010] also found that clustering coefficient increased and pathlength decreased with increasing connection density. Despite an increase in clustering coefficient and a decrease in pathlength observed with increasing density, the SWI decreases with density for both pipelines. The SWI calculation considers the characteristics of randomly generated graphs and therefore this observation must be due either a larger decrease in the pathlength or a larger increase in clustering of the equivalent random networks. In other words, the weaker connections included into the network with increasing density contain a higher number of connections (possibly false positive connections) that do not reflect the small-world characteristics of the structural network in comparison to random graphs. There were also notable differences in global metrics between pipelines. Mean clustering was higher in P1 than P2 networks for all connection densities. Also, P2 had higher pathlength than P1 over all connection densities. These findings are in contrast to [24] and [Bassett et. al. 2010] where clustering coefficient decreased and pathlength increased with nodal scale. As P2 had fewer nodes, this may be because the average distance between nodes increased as a function of node scale. P1 networks had higher SWI than
18 P2 networks at all densities. This finding is in agreement with [24], where it was demonstrated that increasing nodal scale results in increasing SWI. However, while the findings of [24] are due primarily to a decrease in the clustering coefficients of random graphs, our findings are likely to be primarily due to both an increase clustering coefficient and decrease in pathlength of P1 networks. The higher clustering and lower pathlength found for P1 networks demonstrates that the prediction of global metrics with node scale in [24] applies only to networks generated with similar pipelines and demonstrates that other methodological factors such as parcellation and tractography scheme also affect the global clustering and pathlength metrics independently of nodal scale. Despite large differences in the reproducibility of connection strengths between pipelines, the global network metrics were highly reproducible in both pipelines (Fig. 7, 8). These results may be explained by the relatively lower CV of higher strengths connections that are retained in the graph after thresholding, providing robustness of graph theoretical analysis to highly variable connections. For both pipelines, the intra-subject CV of of all global measures was lower than their inter-subject CV across densities, reflecting relatively good within-subject reproducibility. Both intra- and inter-subject CV were small in magnitude (generally less than 6%) and decreased with network density (Fig. 9). Global pathlength had the highest between-subject and within-subject reproducibility in both pipelines. For both pipelines, the inter-subject CV for global pathlength was less than 3% whereas intra-subject CV was less than 1%. The ICC, which measures the proportion of between-subject variability to total variability, was above 0.5 for global clustering and global pathlength across all densities for both pipelines, indicating that between-subject differences in these metrics outweighed within-subject differences. P1 global pathlengths had a higher ICC than P2, while P2 global clustering coefficient ICCs were higher than P1. The SWI ICC was approximately 0.8 for densities of 0.25 or more for both pipelines, but at densities less than 0.25 the within-subject variability was approximately equal or greater than between-subject variability for this metric. Since both global clustering and pathlength had high ICCs at this density range, the low reproducibility of SWI found at low connection densities may be due to variation in the clustering and pathlength of the random networks. Increased reproducibility of the SWI at lower connection densities may therefore be achieved by increasing the number of random networks used to calculate the random clustering and pathlength, thereby reducing their variation. Reproducibility of global network metrics was higher than some previously reported values. The intra-subject reproducibility of global metrics of both pipelines was higher than in [Bassett et. al. 2010], where lower ICC values (~0.6) were reported for global network metrics averaged over multiple nodal scales and parcellation schemes. [Vaessen et. al. 2010] reported high reproducibility, with intra-subject CVs of less than 3% and ICCs of approximately 0.7 for global clustering, pathlength and small-world index at densities of 0.2. As in the current study, the intra-subject CV of these global metrics decreased with increasing density, although the effect was also small. The reason for the difference in reported reproducibility values between studies may be due to other methodological factors, such as different atlases used for nodal parcellation (both [Vaessen et. al. 2010] and [Bassett et. al. 2010] employed different template-based atlases) or different fiber tracking techniques used ([Vaessen et. al. 2010] use probabilistic whereas [Bassett et. al. 2010] used deterministic tractography]). [Cheng et. al. 2012] obtained low ICC values for many global network metrics such as small-world index, clustering coefficient and pathlength across a range of connection densities. [Cheng et. al. 2012] employed the same parcellation scale and scheme as used in P1 but used deterministic instead of probabilistic tractography. Given the high ICC of global metrics reported for P1 and for [Vaessen et. al. 2010], where probabilstic tractography was used, it is tempting to speculate that the higher reproducibility of P1 networks compared with [Cheng et. al. 2012] may be due to the effect of the tractography scheme. However, a number of other methodological differences also exist between P1 and [Cheng et. al. 2012], such as the method used to quantify connection strength, connection strength thresholding scheme, and normalisation of graph theoretical metrics. There has been no investigations into the effect of diffusion modelling or tractography scheme on the reproducibility of structural network metrics and such work would help to explain the differences n reproducibility values reported in this and other studies. While global network metrics have some value in providing a quantitative description of the overall neural organisation of fiber tracts, local metrics can also provide valuable insight into the connection patterns of specific regions. As the definition of node boundaries varied between these pipelines, reproducibility of local metrics was compared on the basis of the mean and standard deviation of reproducibility measures
19 across all nodes. For both pipelines, the inter-subject CV was higher than intra-subject CV across all densities. The intra- and inter-subject CV was however higher for local than global metrics and decreased with increasing network density (Fig. 9). Global metrics were calculated by averaging the local metrics across all nodes and so inter-scan differences in global metrics are masked so long as differences in local metrics cancel each out overall. This averaging also increasing the signal to noise of the global metric. Similarly to global metrics, pathlength had highest mean reproducibility, of all local metrics, with inter- and intra-subject CV's of approximately 5% and 2% respectively for both pipelines. Local centrality was the most highly variable measure both within and between subjects for both pipelines, with a mean intra-subject CV of approximately 25%. The local centrality may be particularly sensitive to changes in network edges within subjects since it considers both the pathlength and nodes along the path, while the pathlength and clustering metrics consider a larger amount of more general information that are not defined by specific edges to the same extent. The ICC of all local metrics was high for both pipelines (above 0.65) across all densities. Previous studies examining intra-subject reproducibility have demonstrated higher ICC for global than local metrics. [Cheng et. al. 2012] demonstrated that the local metrics of betweenness centrality, pathlength and clustering coefficient exhibit significant inter-subject variation compared with global metrics. Similarly to these results, betweenness centrality was found to be the most highly variable metric whilst pathlength was least variable between subjects. Although [Cheng et. al. 2012] did not quantify the intra-subject reproducibility of local metrics, [Bassett et. al. 2010] reported poor reproducibility of the local clustering and pathlength metrics with mean ICC's of approximately 0.3. In [Bassett et. al. 2010], reproducibility and characteristics of local measures were dependent on the choice of atlas, although some repeatable findings such as high connectivity and reproducibility for frontal regions and poorer reproducibilty and weaker connectivity between temporal regions were apparent in all atlases used. A more in depth analysis of which regions had the highest reproducibility of network metrics would be an informative next step and it would be interesting to compare the regional reproducibility with that of [Bassett et. al. 2010]. This is however, to the best of our knowledge, the first demonstration of high intra-subject reproducibility for local network metrics. A number of general limitations exist for both pipelines that should be considered when interpreting these results. Calculation of the SWI involved normalising the global clustering and pathlength metrics by equivalent measures found in randomly generated graphs with the same number of nodes and connections. However, to ensure that the random network is a true null model, the node degree distribution of the random graphs should also be identical and this was not the case in this study. In addition, a large number of repeated generations should be made and their results averaged to get a truer value for random graph characteristics. In this study three random graphs were generated for each network at each connection density and results suggest this may have added some variability to the SWI metric. Another limitation is that a small range of graph theoretical metrics were examined and further studies would benefit from assessing the reproducibility of other network measures such as node degree, modularity, and the presence of hubs or motifs. Another general consideration is that inter-cortical connections are known to relay via sub-cortical grey matter regions such as the thalamus and hippocampus, where many streamlines may terminate. The regions were not included in the analysis and this may have resulted in lower connectivity between some cortical connections that relay via sub-cortical grey matter. Also, the pipelines used different parcellation, registration, diffusion modelling and tractography techniques, the methodological source of differences in networks and their reproducibility was difficult to identify. For example, the large differences in overall connection strengths and densities could be due to differences in the number of nodes used, node boundaries, or the way that tractography techniques define network edges. An important next step is therefore to delineate the effects of tractography and parcellation scheme on the results found here by fixing other parts of the pipeline apart from the aspect one is interested in studying. Improving the reproducibility of structural network metrics is desirable goal for studying individual network architectures. However, there are still a number of other important conceptual and practical challenges in the field of structural network analysis. One issue concerns the neurological relevance of graph theoretical metrics. The global pathlength attempts to capture the efficiency of which regions are connected to all other regions by considering the shortest path problem. However, functionally connected regions may not interact via their shortest paths and not all nodes may be functionally connected to each other, meaning consideration of all shortest paths may not reflect the true measure of local efficiency of information transfer. Other graph metrics are also available to describe network architecture, such as modularity, motif
20 composition and the presence of network hubs. An interesting challenge in the field is interpretation of existing graph theoretical measures and devising new measures of neuroscientific relevance. A further limitation of structural network analysis in general is the validation of connections determined by tractography techniques. This is particularly important as connection is quantified between many regions, meaning that discrepancies in anatomical accuracy accumulate. Some studies have attempted to validate connections found in structural networks by comparing them with equivalent connections found in tract tracing techniques of the macaque cortex [52]. However, both techniques have different limitations and the lack of correspondence between nodes used in tract tracing and structural MRI parcellation limit comparisons of validity between them. Validation of structural network connectivity is currently an ongoing challenge and potential limitation of the technique. Another topic of uncertainty in structural network analyses is the definition of network nodes. Boundaries between nodes may be defined by the underlying cytoarchitectonic composition, by anatomical landmarks or by their structural functional connectivity and it is not clear which method is most appropriate. Also, the node scale at which to perform structural network analysis is not determined. Although [24] identified differences in network metrics across parcellation scales, the study did not suggest which node scale to use and did not address the impact of node scale on network reproducibility. Since the underlying fiber tracts do no change with node scale, it would be useful to formulate a network processing stream where network metrics, such as the overall connection strength and clustering, were stable irrespective of node scale. This report has highlighted a number of common and different characteristics of whole-brain networks and their reproducibility between alternative pipelines. P2 networks had higher connection strengths, connection densities and connection reproducibility. The cause of such differences between pipelines was difficult to identify as multiple processing stages differed. However, there was some evidence of a streamline distance bias towards connection strength and their reproducibility for both pipelines. Although different pipelines had different clustering, pathlength and small-world properties, the reproducibility of both global and local metrics were high (ICC > 0.5), advocating their use as potential biomarkers for studying local and global alterations in structural connectivity. Future studies in the shortterm should focus on delineating the effects of parcellation and tractography scheme on the results, correcting for sources of connection strength biases and on assessing the reproducibility of a wider range of graph theoretical measures. A number of wider issues could also be addressed in future studies such as how to obtain network measures that represent individual connectivity reproducibly and that is not dependent on node scale. The P2 pipeline described provides reasonable reproducibility of all metrics in comparison to previously described pipelines and uses software that is faster than the more conventional techniques used in P1, which is beneficial given the obvious need for network reconstruction techniques to be assessed across variation in one or more methodological components. The high within-subject reproducibility of both pipelines suggests they would be suitable for examining differences in network architectures between individuals or groups and hence to provide insight into anatomical changes that occur due to neurological variation in health or disease.
21 Appendix P1 Desikan Atlas regions (Freesurfer labels) 1007 (2007) Fusiform gyrus 1008 (2008) Inferior parietal cortex 1009 (2009) Inferior temporal gyrus 1010 (2010) Isthmus cingulate cortex 1011 (2011) Lateral occipital cortex 1012 (2012) Lateral orbital frontal cortex 1013 (2013) Lingual gyrus 1014 (2014) Medial orbital frontal cortex 1015 (2015) Middle temporal gyrus 1016 (2016) Parahippocampal gyrus 1017 (2017) Paracentral lobule 1018 (2018) Pars opercularis 1019 (2019) Pars orbitalis 1020 (2020) Pars triangularis 1021 (2021) Pericalcarine cortex 1022 (2022) Postcentral gyrus 1023 (2023) Posterior-cingulate cortex 1024 (2024) Precentral gyrus 1025 (2025) Precuneus cortex 1026 (2026) Rostral anterior cingulate cortex 1027 (2027) Rostral middle frontal gyrus 1028 (2028) Superior frontal gyrus 1029 (2029) Superior parietal cortex 1030 (2030) Superior temporal gyrus 1031 (2031) Supramarginal gyrus 1032 (2032) Frontal pole 1033 (2033) Temporal pole 1034 (2034) Transverse temporal cortex 1035 (2035) Insula cortex P2 - Hammer's Atlas regions(niftyseg labels) 6 (5) Anterior temporal lobe medial part 8 (7) Anterior temporal lobe lateral part 10 (9) Parahippocampal and ambient gyri 12 (11) Superior temporal gyrus 14 (13) Middle and inferior temporal gyri 16 (15) Fusiform gyrus 20 (21) Insula 22 (23) Lateral remainder of occipital lobe 24 (25) Gyrus cinguli anterior part 26 (27) Gyrus cinguli posterior part 28 (29) Middle frontal gyrus 30 (31) Posterior temporal lobe 32 (33) Inferolateral remainder of parietal lobe 50 (51) Precentral gyrus 52 (53) Gyrus rectus 54 (55) Orbitofrontal gyri 56 (57) Inferior frontal gyrus 58 (59) Superior frontal gyrus 60 (61) Postcentral gyrus 62 (63) Superior parietal gyrus 64 (65) Lingual gyrus 66 (67) Cuneus Table A.1. Node ID numbers (left) and their corresponding names for P1 (Freesurfer, left) and P2 (Hammer's, right) grey matter parcellations. The number outside the brackets describes the left hemisphere regions while the number outiside describes the right hemisphere regions. Regions are listed in the order they appear in the connection matrices. The first half of the connection matrix indices in this report indices refer to left hemisphere regions and the second half to right hemisphere regions.
22 Figure A.1. Intra-subject reproducibility of connection strengths in pipeline 1 and 2. For each connection the intra-subject CV is calculated as the mean of the standard deviations across repeat scans divided by the overall mean for each connection. Both pipelines had high intra-subject CVs which show were inversely correlated with connection strength. P1 had higher intra-subject CVs than P2. Both Matrices are represented with the same colour bar. Figure A.2. Inter-subject reproducibility of connection strengths in pipeline 1 (left) and 2 (right). For each connection the inter-subject CV is calculated as the standard deviation of the means across repeat scans divided by the overall mean for each connection. Both pipelines had high inter-subject CVs which were inversely correlated with connection strength. P1 had higher inter-subject CVs than P2. Matrices are represented with the same colour bar.
23 Figure A.3. The ICC of all connections in pipeline 1 (left) and 2 (right). All connections in both pipelines had ICCs greater than 0.5, indicating the inter-subject variance was greater than the intra-subject variance. Matrices are using the same colour bar. P1 distance strength intra CV inter CV distance strength intra CV inter CV P2 distance strength intra CV inter CV distance strength intra CV inter CV Table A.2. Correlations between connection distance, strength, intra-subject CV and inter-subject CV for both pipelines. Connection strength was inversely correlated with distance as well as intra- and inter-subject CV, demonstrating that connections that a further apart are weaker and more variable. Connections that had high intra-subject CV tended also had high inter-subject CV. The correlations were stronger in P2 than P1 networks.
The organization of the human cerebral cortex estimated by intrinsic functional connectivity
 1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
Diffusion model fitting and tractography: A primer
 Diffusion model fitting and tractography: A primer Anastasia Yendiki HMS/MGH/MIT Athinoula A. Martinos Center for Biomedical Imaging 03/18/10 Why n how Diffusion model fitting and tractography 0/18 Why
Diffusion model fitting and tractography: A primer Anastasia Yendiki HMS/MGH/MIT Athinoula A. Martinos Center for Biomedical Imaging 03/18/10 Why n how Diffusion model fitting and tractography 0/18 Why
FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING. Francesca Pizzorni Ferrarese
 FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING Francesca Pizzorni Ferrarese Pipeline overview WM and GM Segmentation Registration Data reconstruction Tractography
FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING Francesca Pizzorni Ferrarese Pipeline overview WM and GM Segmentation Registration Data reconstruction Tractography
Lilla Zöllei A.A. Martinos Center, MGH; Boston, MA
 Lilla Zöllei lzollei@nmr.mgh.harvard.edu A.A. Martinos Center, MGH; Boston, MA Bruce Fischl Gheorghe Postelnicu Jean Augustinack Anastasia Yendiki Allison Stevens Kristen Huber Sita Kakonoori + the FreeSurfer
Lilla Zöllei lzollei@nmr.mgh.harvard.edu A.A. Martinos Center, MGH; Boston, MA Bruce Fischl Gheorghe Postelnicu Jean Augustinack Anastasia Yendiki Allison Stevens Kristen Huber Sita Kakonoori + the FreeSurfer
Fiber Selection from Diffusion Tensor Data based on Boolean Operators
 Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhof 1, G. Greiner 2, M. Buchfelder 3, C. Nimsky 4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhof 1, G. Greiner 2, M. Buchfelder 3, C. Nimsky 4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Diffusion MRI. Introduction and Modern Methods. John Plass. Department Of Psychology
 Diffusion MRI Introduction and Modern Methods John Plass Department Of Psychology Diffusion MRI Introduction and Modern Methods John Plass Department Of Psychology Overview I. Why use diffusion MRI? II.
Diffusion MRI Introduction and Modern Methods John Plass Department Of Psychology Diffusion MRI Introduction and Modern Methods John Plass Department Of Psychology Overview I. Why use diffusion MRI? II.
CHAPTER 2. Morphometry on rodent brains. A.E.H. Scheenstra J. Dijkstra L. van der Weerd
 CHAPTER 2 Morphometry on rodent brains A.E.H. Scheenstra J. Dijkstra L. van der Weerd This chapter was adapted from: Volumetry and other quantitative measurements to assess the rodent brain, In vivo NMR
CHAPTER 2 Morphometry on rodent brains A.E.H. Scheenstra J. Dijkstra L. van der Weerd This chapter was adapted from: Volumetry and other quantitative measurements to assess the rodent brain, In vivo NMR
Norbert Schuff VA Medical Center and UCSF
 Norbert Schuff Medical Center and UCSF Norbert.schuff@ucsf.edu Medical Imaging Informatics N.Schuff Course # 170.03 Slide 1/67 Objective Learn the principle segmentation techniques Understand the role
Norbert Schuff Medical Center and UCSF Norbert.schuff@ucsf.edu Medical Imaging Informatics N.Schuff Course # 170.03 Slide 1/67 Objective Learn the principle segmentation techniques Understand the role
Supplementary methods
 Supplementary methods This section provides additional technical details on the sample, the applied imaging and analysis steps and methods. Structural imaging Trained radiographers placed all participants
Supplementary methods This section provides additional technical details on the sample, the applied imaging and analysis steps and methods. Structural imaging Trained radiographers placed all participants
Evaluation of Local Filter Approaches for Diffusion Tensor based Fiber Tracking
 Evaluation of Local Filter Approaches for Diffusion Tensor based Fiber Tracking D. Merhof 1, M. Buchfelder 2, C. Nimsky 3 1 Visual Computing, University of Konstanz, Konstanz 2 Department of Neurosurgery,
Evaluation of Local Filter Approaches for Diffusion Tensor based Fiber Tracking D. Merhof 1, M. Buchfelder 2, C. Nimsky 3 1 Visual Computing, University of Konstanz, Konstanz 2 Department of Neurosurgery,
What is a network? Network Analysis
 What is a network? Network Analysis Valerie Cardenas Nicolson Associate Adjunct Professor Department of Radiology and Biomedical Imaging Complex weblike structures Cell is network of chemicals connected
What is a network? Network Analysis Valerie Cardenas Nicolson Associate Adjunct Professor Department of Radiology and Biomedical Imaging Complex weblike structures Cell is network of chemicals connected
The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy
 The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy Sokratis K. Makrogiannis, PhD From post-doctoral research at SBIA lab, Department of Radiology,
The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy Sokratis K. Makrogiannis, PhD From post-doctoral research at SBIA lab, Department of Radiology,
An Introduction To Automatic Tissue Classification Of Brain MRI. Colm Elliott Mar 2014
 An Introduction To Automatic Tissue Classification Of Brain MRI Colm Elliott Mar 2014 Tissue Classification Tissue classification is part of many processing pipelines. We often want to classify each voxel
An Introduction To Automatic Tissue Classification Of Brain MRI Colm Elliott Mar 2014 Tissue Classification Tissue classification is part of many processing pipelines. We often want to classify each voxel
Generation of Hulls Encompassing Neuronal Pathways Based on Tetrahedralization and 3D Alpha Shapes
 Generation of Hulls Encompassing Neuronal Pathways Based on Tetrahedralization and 3D Alpha Shapes Dorit Merhof 1,2, Martin Meister 1, Ezgi Bingöl 1, Peter Hastreiter 1,2, Christopher Nimsky 2,3, Günther
Generation of Hulls Encompassing Neuronal Pathways Based on Tetrahedralization and 3D Alpha Shapes Dorit Merhof 1,2, Martin Meister 1, Ezgi Bingöl 1, Peter Hastreiter 1,2, Christopher Nimsky 2,3, Günther
Normalisation of Neonatal Brain Network Measures Using Stochastic Approaches
 Normalisation of Neonatal Brain Network Measures Using Stochastic Approaches M. Schirmer 1, G. Ball 1, S.J. Counsell 1, A.D. Edwards 1, D. Rueckert 2, J.V. Hajnal 1, and P. Aljabar 1 1 Division of Imaging
Normalisation of Neonatal Brain Network Measures Using Stochastic Approaches M. Schirmer 1, G. Ball 1, S.J. Counsell 1, A.D. Edwards 1, D. Rueckert 2, J.V. Hajnal 1, and P. Aljabar 1 1 Division of Imaging
DIFFUSION TENSOR IMAGING ANALYSIS. Using Analyze
 DIFFUSION TENSOR IMAGING ANALYSIS Using Analyze 2 Table of Contents 1. Introduction page 3 2. Loading DTI Data page 4 3. Computing DTI Maps page 5 4. Defining ROIs for Fiber Tracking page 6 5. Visualizing
DIFFUSION TENSOR IMAGING ANALYSIS Using Analyze 2 Table of Contents 1. Introduction page 3 2. Loading DTI Data page 4 3. Computing DTI Maps page 5 4. Defining ROIs for Fiber Tracking page 6 5. Visualizing
NEURO M203 & BIOMED M263 WINTER 2014
 NEURO M203 & BIOMED M263 WINTER 2014 MRI Lab 2: Neuroimaging Connectivity Lab In today s lab we will work with sample diffusion imaging data and the group averaged fmri data collected during your scanning
NEURO M203 & BIOMED M263 WINTER 2014 MRI Lab 2: Neuroimaging Connectivity Lab In today s lab we will work with sample diffusion imaging data and the group averaged fmri data collected during your scanning
A Model-Independent, Multi-Image Approach to MR Inhomogeneity Correction
 Tina Memo No. 2007-003 Published in Proc. MIUA 2007 A Model-Independent, Multi-Image Approach to MR Inhomogeneity Correction P. A. Bromiley and N.A. Thacker Last updated 13 / 4 / 2007 Imaging Science and
Tina Memo No. 2007-003 Published in Proc. MIUA 2007 A Model-Independent, Multi-Image Approach to MR Inhomogeneity Correction P. A. Bromiley and N.A. Thacker Last updated 13 / 4 / 2007 Imaging Science and
Head motion in diffusion MRI
 Head motion in diffusion MRI Anastasia Yendiki HMS/MGH/MIT Athinoula A. Martinos Center for Biomedical Imaging 11/06/13 Head motion in diffusion MRI 0/33 Diffusion contrast Basic principle of diffusion
Head motion in diffusion MRI Anastasia Yendiki HMS/MGH/MIT Athinoula A. Martinos Center for Biomedical Imaging 11/06/13 Head motion in diffusion MRI 0/33 Diffusion contrast Basic principle of diffusion
Attention modulates spatial priority maps in human occipital, parietal, and frontal cortex
 Attention modulates spatial priority maps in human occipital, parietal, and frontal cortex Thomas C. Sprague 1 and John T. Serences 1,2 1 Neuroscience Graduate Program, University of California San Diego
Attention modulates spatial priority maps in human occipital, parietal, and frontal cortex Thomas C. Sprague 1 and John T. Serences 1,2 1 Neuroscience Graduate Program, University of California San Diego
HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Statistical Analysis of Neuroimaging Data. Phebe Kemmer BIOS 516 Sept 24, 2015
 Statistical Analysis of Neuroimaging Data Phebe Kemmer BIOS 516 Sept 24, 2015 Review from last time Structural Imaging modalities MRI, CAT, DTI (diffusion tensor imaging) Functional Imaging modalities
Statistical Analysis of Neuroimaging Data Phebe Kemmer BIOS 516 Sept 24, 2015 Review from last time Structural Imaging modalities MRI, CAT, DTI (diffusion tensor imaging) Functional Imaging modalities
Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases
 Quantities Measured by MR - Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases Static parameters (influenced by molecular environment): T, T* (transverse relaxation)
Quantities Measured by MR - Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases Static parameters (influenced by molecular environment): T, T* (transverse relaxation)
Effect of age and dementia on topology of brain functional networks. Paul McCarthy, Luba Benuskova, Liz Franz University of Otago, New Zealand
 Effect of age and dementia on topology of brain functional networks Paul McCarthy, Luba Benuskova, Liz Franz University of Otago, New Zealand 1 Structural changes in aging brain Age-related changes in
Effect of age and dementia on topology of brain functional networks Paul McCarthy, Luba Benuskova, Liz Franz University of Otago, New Zealand 1 Structural changes in aging brain Age-related changes in
TRACULA: Troubleshooting, visualization, and group analysis
 TRACULA: Troubleshooting, visualization, and group analysis Anastasia Yendiki HMS/MGH/MIT Athinoula A. Martinos Center for Biomedical Imaging 18/11/13 TRACULA: troubleshooting, visualization, group analysis
TRACULA: Troubleshooting, visualization, and group analysis Anastasia Yendiki HMS/MGH/MIT Athinoula A. Martinos Center for Biomedical Imaging 18/11/13 TRACULA: troubleshooting, visualization, group analysis
ACCEPTED MANUSCRIPT. Inter-site and inter-scanner diffusion MRI data harmonization
 Inter-site and inter-scanner diffusion MRI data harmonization H. Mirzaalian 1, L. Ning 1, P. Savadjiev 1, O. Pasternak 1, S. Bouix 1, O. Michailovich 2, M. Kubicki 1, C-F Westin 1, M.E.Shenton 1, Y. Rathi
Inter-site and inter-scanner diffusion MRI data harmonization H. Mirzaalian 1, L. Ning 1, P. Savadjiev 1, O. Pasternak 1, S. Bouix 1, O. Michailovich 2, M. Kubicki 1, C-F Westin 1, M.E.Shenton 1, Y. Rathi
Brain Extraction, Registration & EPI Distortion Correction
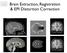 Brain Extraction, Registration & EPI Distortion Correction What use is Registration? Some common uses of registration: Combining across individuals in group studies: including fmri & diffusion Quantifying
Brain Extraction, Registration & EPI Distortion Correction What use is Registration? Some common uses of registration: Combining across individuals in group studies: including fmri & diffusion Quantifying
Computational Neuroanatomy
 Computational Neuroanatomy John Ashburner john@fil.ion.ucl.ac.uk Smoothing Motion Correction Between Modality Co-registration Spatial Normalisation Segmentation Morphometry Overview fmri time-series kernel
Computational Neuroanatomy John Ashburner john@fil.ion.ucl.ac.uk Smoothing Motion Correction Between Modality Co-registration Spatial Normalisation Segmentation Morphometry Overview fmri time-series kernel
Fmri Spatial Processing
 Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge
 Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
TractoR and Other Software
 TractoR and Other Software Jon Clayden DIBS Teaching Seminar, 11 Dec 2015 Photo by José Martín Ramírez Carrasco https://www.behance.net/martini_rc TractoR A set of R packages Additional
TractoR and Other Software Jon Clayden DIBS Teaching Seminar, 11 Dec 2015 Photo by José Martín Ramírez Carrasco https://www.behance.net/martini_rc TractoR A set of R packages Additional
MR IMAGE SEGMENTATION
 MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
Resting state network estimation in individual subjects
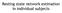 Resting state network estimation in individual subjects Data 3T NIL(21,17,10), Havard-MGH(692) Young adult fmri BOLD Method Machine learning algorithm MLP DR LDA Network image Correlation Spatial Temporal
Resting state network estimation in individual subjects Data 3T NIL(21,17,10), Havard-MGH(692) Young adult fmri BOLD Method Machine learning algorithm MLP DR LDA Network image Correlation Spatial Temporal
Introduction to fmri. Pre-processing
 Introduction to fmri Pre-processing Tibor Auer Department of Psychology Research Fellow in MRI Data Types Anatomical data: T 1 -weighted, 3D, 1/subject or session - (ME)MPRAGE/FLASH sequence, undistorted
Introduction to fmri Pre-processing Tibor Auer Department of Psychology Research Fellow in MRI Data Types Anatomical data: T 1 -weighted, 3D, 1/subject or session - (ME)MPRAGE/FLASH sequence, undistorted
This exercise uses one anatomical data set (ANAT1) and two functional data sets (FUNC1 and FUNC2).
 Exploring Brain Anatomy This week s exercises will let you explore the anatomical organization of the brain to learn some of its basic properties, as well as the location of different structures. The human
Exploring Brain Anatomy This week s exercises will let you explore the anatomical organization of the brain to learn some of its basic properties, as well as the location of different structures. The human
Basic principles of MR image analysis. Basic principles of MR image analysis. Basic principles of MR image analysis
 Basic principles of MR image analysis Basic principles of MR image analysis Julien Milles Leiden University Medical Center Terminology of fmri Brain extraction Registration Linear registration Non-linear
Basic principles of MR image analysis Basic principles of MR image analysis Julien Milles Leiden University Medical Center Terminology of fmri Brain extraction Registration Linear registration Non-linear
Supplementary Figure 1. Decoding results broken down for different ROIs
 Supplementary Figure 1 Decoding results broken down for different ROIs Decoding results for areas V1, V2, V3, and V1 V3 combined. (a) Decoded and presented orientations are strongly correlated in areas
Supplementary Figure 1 Decoding results broken down for different ROIs Decoding results for areas V1, V2, V3, and V1 V3 combined. (a) Decoded and presented orientations are strongly correlated in areas
MITK-DI. A new Diffusion Imaging Component for MITK. Klaus Fritzsche, Hans-Peter Meinzer
 MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
Anatomic parcellation based on DTI data with FSL taking the example of SMA/preSMA
 Groupe de Travail IRMf/MEG 02/2014 Anatomic parcellation based on DTI data with FSL taking the example of SMA/preSMA Magdalena Wutte, Lucile Brun & Boris Burle SMA/preSMA Parcellation - what for? - connectivity
Groupe de Travail IRMf/MEG 02/2014 Anatomic parcellation based on DTI data with FSL taking the example of SMA/preSMA Magdalena Wutte, Lucile Brun & Boris Burle SMA/preSMA Parcellation - what for? - connectivity
Where are we now? Structural MRI processing and analysis
 Where are we now? Structural MRI processing and analysis Pierre-Louis Bazin bazin@cbs.mpg.de Leipzig, Germany Structural MRI processing: why bother? Just use the standards? SPM FreeSurfer FSL However:
Where are we now? Structural MRI processing and analysis Pierre-Louis Bazin bazin@cbs.mpg.de Leipzig, Germany Structural MRI processing: why bother? Just use the standards? SPM FreeSurfer FSL However:
Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques
 Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques Sea Chen Department of Biomedical Engineering Advisors: Dr. Charles A. Bouman and Dr. Mark J. Lowe S. Chen Final Exam October
Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques Sea Chen Department of Biomedical Engineering Advisors: Dr. Charles A. Bouman and Dr. Mark J. Lowe S. Chen Final Exam October
Methods for data preprocessing
 Methods for data preprocessing John Ashburner Wellcome Trust Centre for Neuroimaging, 12 Queen Square, London, UK. Overview Voxel-Based Morphometry Morphometry in general Volumetrics VBM preprocessing
Methods for data preprocessing John Ashburner Wellcome Trust Centre for Neuroimaging, 12 Queen Square, London, UK. Overview Voxel-Based Morphometry Morphometry in general Volumetrics VBM preprocessing
SISCOM (Subtraction Ictal SPECT CO-registered to MRI)
 SISCOM (Subtraction Ictal SPECT CO-registered to MRI) Introduction A method for advanced imaging of epilepsy patients has been developed with Analyze at the Mayo Foundation which uses a combination of
SISCOM (Subtraction Ictal SPECT CO-registered to MRI) Introduction A method for advanced imaging of epilepsy patients has been developed with Analyze at the Mayo Foundation which uses a combination of
Network connectivity via inference over curvature-regularizing line graphs
 Network connectivity via inference over curvature-regularizing line graphs Asian Conference on Computer Vision Maxwell D. Collins 1,2, Vikas Singh 2,1, Andrew L. Alexander 3 1 Department of Computer Sciences
Network connectivity via inference over curvature-regularizing line graphs Asian Conference on Computer Vision Maxwell D. Collins 1,2, Vikas Singh 2,1, Andrew L. Alexander 3 1 Department of Computer Sciences
Diffusion MRI Acquisition. Karla Miller FMRIB Centre, University of Oxford
 Diffusion MRI Acquisition Karla Miller FMRIB Centre, University of Oxford karla@fmrib.ox.ac.uk Diffusion Imaging How is diffusion weighting achieved? How is the image acquired? What are the limitations,
Diffusion MRI Acquisition Karla Miller FMRIB Centre, University of Oxford karla@fmrib.ox.ac.uk Diffusion Imaging How is diffusion weighting achieved? How is the image acquired? What are the limitations,
Fiber Selection from Diffusion Tensor Data based on Boolean Operators
 Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhofl, G. Greiner 2, M. Buchfelder 3, C. Nimsky4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhofl, G. Greiner 2, M. Buchfelder 3, C. Nimsky4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Chapter 3 Set Redundancy in Magnetic Resonance Brain Images
 16 Chapter 3 Set Redundancy in Magnetic Resonance Brain Images 3.1 MRI (magnetic resonance imaging) MRI is a technique of measuring physical structure within the human anatomy. Our proposed research focuses
16 Chapter 3 Set Redundancy in Magnetic Resonance Brain Images 3.1 MRI (magnetic resonance imaging) MRI is a technique of measuring physical structure within the human anatomy. Our proposed research focuses
RIGID IMAGE REGISTRATION
 RIGID IMAGE REGISTRATION Duygu Tosun-Turgut, Ph.D. Center for Imaging of Neurodegenerative Diseases Department of Radiology and Biomedical Imaging duygu.tosun@ucsf.edu What is registration? Image registration
RIGID IMAGE REGISTRATION Duygu Tosun-Turgut, Ph.D. Center for Imaging of Neurodegenerative Diseases Department of Radiology and Biomedical Imaging duygu.tosun@ucsf.edu What is registration? Image registration
Quality Checking an fmri Group Result (art_groupcheck)
 Quality Checking an fmri Group Result (art_groupcheck) Paul Mazaika, Feb. 24, 2009 A statistical parameter map of fmri group analyses relies on the assumptions of the General Linear Model (GLM). The assumptions
Quality Checking an fmri Group Result (art_groupcheck) Paul Mazaika, Feb. 24, 2009 A statistical parameter map of fmri group analyses relies on the assumptions of the General Linear Model (GLM). The assumptions
syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets
 syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets Julien Gervais; Lisa Chuah Siemens Healthcare, Magnetic
syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets Julien Gervais; Lisa Chuah Siemens Healthcare, Magnetic
Learning-based Neuroimage Registration
 Learning-based Neuroimage Registration Leonid Teverovskiy and Yanxi Liu 1 October 2004 CMU-CALD-04-108, CMU-RI-TR-04-59 School of Computer Science Carnegie Mellon University Pittsburgh, PA 15213 Abstract
Learning-based Neuroimage Registration Leonid Teverovskiy and Yanxi Liu 1 October 2004 CMU-CALD-04-108, CMU-RI-TR-04-59 School of Computer Science Carnegie Mellon University Pittsburgh, PA 15213 Abstract
Functional MRI data preprocessing. Cyril Pernet, PhD
 Functional MRI data preprocessing Cyril Pernet, PhD Data have been acquired, what s s next? time No matter the design, multiple volumes (made from multiple slices) have been acquired in time. Before getting
Functional MRI data preprocessing Cyril Pernet, PhD Data have been acquired, what s s next? time No matter the design, multiple volumes (made from multiple slices) have been acquired in time. Before getting
ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION
 ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION Abstract: MIP Project Report Spring 2013 Gaurav Mittal 201232644 This is a detailed report about the course project, which was to implement
ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION Abstract: MIP Project Report Spring 2013 Gaurav Mittal 201232644 This is a detailed report about the course project, which was to implement
I.e. Sex differences in child appetitive traits and Eating in the Absence of Hunger:
 Supplementary Materials I. Evidence of sex differences on eating behavior in children I.e. Sex differences in child appetitive traits and Eating in the Absence of Hunger: Table 2. Parent Report for Child
Supplementary Materials I. Evidence of sex differences on eating behavior in children I.e. Sex differences in child appetitive traits and Eating in the Absence of Hunger: Table 2. Parent Report for Child
Blood Particle Trajectories in Phase-Contrast-MRI as Minimal Paths Computed with Anisotropic Fast Marching
 Blood Particle Trajectories in Phase-Contrast-MRI as Minimal Paths Computed with Anisotropic Fast Marching Michael Schwenke 1, Anja Hennemuth 1, Bernd Fischer 2, Ola Friman 1 1 Fraunhofer MEVIS, Institute
Blood Particle Trajectories in Phase-Contrast-MRI as Minimal Paths Computed with Anisotropic Fast Marching Michael Schwenke 1, Anja Hennemuth 1, Bernd Fischer 2, Ola Friman 1 1 Fraunhofer MEVIS, Institute
Test-Retest Reliability of Graph Theory Measures of Structural Brain Connectivity
 Test-Retest Reliability of Graph Theory Measures of Structural Brain Connectivity Emily L. Dennis 1, Neda Jahanshad 1, Arthur W. Toga 2, Katie L. McMahon 3, Greig I. de Zubicaray 5, Nicholas G. Martin
Test-Retest Reliability of Graph Theory Measures of Structural Brain Connectivity Emily L. Dennis 1, Neda Jahanshad 1, Arthur W. Toga 2, Katie L. McMahon 3, Greig I. de Zubicaray 5, Nicholas G. Martin
Whole Body MRI Intensity Standardization
 Whole Body MRI Intensity Standardization Florian Jäger 1, László Nyúl 1, Bernd Frericks 2, Frank Wacker 2 and Joachim Hornegger 1 1 Institute of Pattern Recognition, University of Erlangen, {jaeger,nyul,hornegger}@informatik.uni-erlangen.de
Whole Body MRI Intensity Standardization Florian Jäger 1, László Nyúl 1, Bernd Frericks 2, Frank Wacker 2 and Joachim Hornegger 1 1 Institute of Pattern Recognition, University of Erlangen, {jaeger,nyul,hornegger}@informatik.uni-erlangen.de
3. Cluster analysis Overview
 Université Laval Multivariate analysis - February 2006 1 3.1. Overview 3. Cluster analysis Clustering requires the recognition of discontinuous subsets in an environment that is sometimes discrete (as
Université Laval Multivariate analysis - February 2006 1 3.1. Overview 3. Cluster analysis Clustering requires the recognition of discontinuous subsets in an environment that is sometimes discrete (as
Title: Wired for function: Anatomical connectivity patterns predict face-selectivity in the
 Title: Wired for function: Anatomical connectivity patterns predict face-selectivity in the fusiform gyrus Authors: Zeynep M. Saygin *, David E. Osher *, Kami Koldewyn, Gretchen Reynolds, John D.E. Gabrieli,
Title: Wired for function: Anatomical connectivity patterns predict face-selectivity in the fusiform gyrus Authors: Zeynep M. Saygin *, David E. Osher *, Kami Koldewyn, Gretchen Reynolds, John D.E. Gabrieli,
Atelier 2 : Calcul Haute Performance et Sciences du Vivant Forum er juillet, Paris, France
 From Diffusion MR Image Analysis to Whole Brain Connectivity Simulation Jean-Philippe Thiran EPFL Lausanne, Switzerland EPFL - Lausanne HPC in life sciences at EPFL The Blue Brain project: create a biologically
From Diffusion MR Image Analysis to Whole Brain Connectivity Simulation Jean-Philippe Thiran EPFL Lausanne, Switzerland EPFL - Lausanne HPC in life sciences at EPFL The Blue Brain project: create a biologically
Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry
 Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry Nivedita Agarwal, MD Nivedita.agarwal@apss.tn.it Nivedita.agarwal@unitn.it Volume and surface morphometry Brain volume White matter
Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry Nivedita Agarwal, MD Nivedita.agarwal@apss.tn.it Nivedita.agarwal@unitn.it Volume and surface morphometry Brain volume White matter
Detecting Changes In Non-Isotropic Images
 Detecting Changes In Non-Isotropic Images K.J. Worsley 1, M. Andermann 1, T. Koulis 1, D. MacDonald, 2 and A.C. Evans 2 August 4, 1999 1 Department of Mathematics and Statistics, 2 Montreal Neurological
Detecting Changes In Non-Isotropic Images K.J. Worsley 1, M. Andermann 1, T. Koulis 1, D. MacDonald, 2 and A.C. Evans 2 August 4, 1999 1 Department of Mathematics and Statistics, 2 Montreal Neurological
Structural MRI analysis
 Structural MRI analysis volumetry and voxel-based morphometry cortical thickness measurements structural covariance network mapping Boris Bernhardt, PhD Department of Social Neuroscience, MPI-CBS bernhardt@cbs.mpg.de
Structural MRI analysis volumetry and voxel-based morphometry cortical thickness measurements structural covariance network mapping Boris Bernhardt, PhD Department of Social Neuroscience, MPI-CBS bernhardt@cbs.mpg.de
Saturn User Manual. Rubén Cárdenes. 29th January 2010 Image Processing Laboratory, University of Valladolid. Abstract
 Saturn User Manual Rubén Cárdenes 29th January 2010 Image Processing Laboratory, University of Valladolid Abstract Saturn is a software package for DTI processing and visualization, provided with a graphic
Saturn User Manual Rubén Cárdenes 29th January 2010 Image Processing Laboratory, University of Valladolid Abstract Saturn is a software package for DTI processing and visualization, provided with a graphic
Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging
 1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
CS/NEUR125 Brains, Minds, and Machines. Due: Wednesday, April 5
 CS/NEUR125 Brains, Minds, and Machines Lab 8: Using fmri to Discover Language Areas in the Brain Due: Wednesday, April 5 In this lab, you will analyze fmri data from an experiment that was designed to
CS/NEUR125 Brains, Minds, and Machines Lab 8: Using fmri to Discover Language Areas in the Brain Due: Wednesday, April 5 In this lab, you will analyze fmri data from an experiment that was designed to
Diffusion Imaging Visualization
 Diffusion Imaging Visualization Thomas Schultz URL: http://cg.cs.uni-bonn.de/schultz/ E-Mail: schultz@cs.uni-bonn.de 1 Outline Introduction to Diffusion Imaging Basic techniques Glyph-based Visualization
Diffusion Imaging Visualization Thomas Schultz URL: http://cg.cs.uni-bonn.de/schultz/ E-Mail: schultz@cs.uni-bonn.de 1 Outline Introduction to Diffusion Imaging Basic techniques Glyph-based Visualization
INDEPENDENT COMPONENT ANALYSIS APPLIED TO fmri DATA: A GENERATIVE MODEL FOR VALIDATING RESULTS
 INDEPENDENT COMPONENT ANALYSIS APPLIED TO fmri DATA: A GENERATIVE MODEL FOR VALIDATING RESULTS V. Calhoun 1,2, T. Adali, 2 and G. Pearlson 1 1 Johns Hopkins University Division of Psychiatric Neuro-Imaging,
INDEPENDENT COMPONENT ANALYSIS APPLIED TO fmri DATA: A GENERATIVE MODEL FOR VALIDATING RESULTS V. Calhoun 1,2, T. Adali, 2 and G. Pearlson 1 1 Johns Hopkins University Division of Psychiatric Neuro-Imaging,
BDP: BrainSuite Diffusion Pipeline. Chitresh Bhushan
 BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
Multimodal Imaging Brain Connectivity Analysis (MIBCA)
 Multimodal Imaging Brain Connectivity Analysis (MIBCA) Andre Santos Ribeiro, Luis Miguel Lacerda, Hugo Ferreira April 23, 2015 Abstract In recent years, connectivity studies using neuroimaging data have
Multimodal Imaging Brain Connectivity Analysis (MIBCA) Andre Santos Ribeiro, Luis Miguel Lacerda, Hugo Ferreira April 23, 2015 Abstract In recent years, connectivity studies using neuroimaging data have
Kernel Regression Estimation of Fiber Orientation Mixtures in Di usion MRI
 Ryan P. Cabeen, Mark E. Bastin, David H. Laidlaw, Kernel regression estimation of fiber orientation mixtures in diffusion MRI, NeuroImage, Volume 127, 15 February 2016, Pages 158-172, ISSN 1053-8119, http://dx.doi.org/10.1016/j.neuroimage.2015.11.061
Ryan P. Cabeen, Mark E. Bastin, David H. Laidlaw, Kernel regression estimation of fiber orientation mixtures in diffusion MRI, NeuroImage, Volume 127, 15 February 2016, Pages 158-172, ISSN 1053-8119, http://dx.doi.org/10.1016/j.neuroimage.2015.11.061
Diffusion Tensor Imaging and Reading Development
 Diffusion Tensor Imaging and Reading Development Bob Dougherty Stanford Institute for Reading and Learning Reading and Anatomy Every brain is different... Not all brains optimized for highly proficient
Diffusion Tensor Imaging and Reading Development Bob Dougherty Stanford Institute for Reading and Learning Reading and Anatomy Every brain is different... Not all brains optimized for highly proficient
BDP: BrainSuite Diffusion Pipeline. Chitresh Bhushan
 BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
MITK-DI. A new Diffusion Imaging Component for MITK. Klaus Fritzsche, Hans-Peter Meinzer
 MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
Reconstruction of Fiber Trajectories via Population-Based Estimation of Local Orientations
 IDEA Reconstruction of Fiber Trajectories via Population-Based Estimation of Local Orientations Pew-Thian Yap, John H. Gilmore, Weili Lin, Dinggang Shen Email: ptyap@med.unc.edu 2011-09-21 Poster: P2-46-
IDEA Reconstruction of Fiber Trajectories via Population-Based Estimation of Local Orientations Pew-Thian Yap, John H. Gilmore, Weili Lin, Dinggang Shen Email: ptyap@med.unc.edu 2011-09-21 Poster: P2-46-
Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow
 Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow Abstract. Finding meaningful 1-1 correspondences between hippocampal (HP) surfaces is an important but difficult
Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow Abstract. Finding meaningful 1-1 correspondences between hippocampal (HP) surfaces is an important but difficult
CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS
 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS CHAPTER 4 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS 4.1 Introduction Optical character recognition is one of
CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS CHAPTER 4 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS 4.1 Introduction Optical character recognition is one of
A Generic Lie Group Model for Computer Vision
 A Generic Lie Group Model for Computer Vision Within this research track we follow a generic Lie group approach to computer vision based on recent physiological research on how the primary visual cortex
A Generic Lie Group Model for Computer Vision Within this research track we follow a generic Lie group approach to computer vision based on recent physiological research on how the primary visual cortex
Appendix E1. Supplementary Methods. MR Image Acquisition. MR Image Analysis
 RSNA, 2015 10.1148/radiol.2015150532 Appendix E1 Supplementary Methods MR Image Acquisition By using a 1.5-T system (Avanto, Siemens Medical, Erlangen, Germany) under a program of regular maintenance (no
RSNA, 2015 10.1148/radiol.2015150532 Appendix E1 Supplementary Methods MR Image Acquisition By using a 1.5-T system (Avanto, Siemens Medical, Erlangen, Germany) under a program of regular maintenance (no
Lucy Phantom MR Grid Evaluation
 Lucy Phantom MR Grid Evaluation Anil Sethi, PhD Loyola University Medical Center, Maywood, IL 60153 November 2015 I. Introduction: The MR distortion grid, used as an insert with Lucy 3D QA phantom, is
Lucy Phantom MR Grid Evaluation Anil Sethi, PhD Loyola University Medical Center, Maywood, IL 60153 November 2015 I. Introduction: The MR distortion grid, used as an insert with Lucy 3D QA phantom, is
Lecture on Modeling Tools for Clustering & Regression
 Lecture on Modeling Tools for Clustering & Regression CS 590.21 Analysis and Modeling of Brain Networks Department of Computer Science University of Crete Data Clustering Overview Organizing data into
Lecture on Modeling Tools for Clustering & Regression CS 590.21 Analysis and Modeling of Brain Networks Department of Computer Science University of Crete Data Clustering Overview Organizing data into
Samuel Coolidge, Dan Simon, Dennis Shasha, Technical Report NYU/CIMS/TR
 Detecting Missing and Spurious Edges in Large, Dense Networks Using Parallel Computing Samuel Coolidge, sam.r.coolidge@gmail.com Dan Simon, des480@nyu.edu Dennis Shasha, shasha@cims.nyu.edu Technical Report
Detecting Missing and Spurious Edges in Large, Dense Networks Using Parallel Computing Samuel Coolidge, sam.r.coolidge@gmail.com Dan Simon, des480@nyu.edu Dennis Shasha, shasha@cims.nyu.edu Technical Report
Surface-based Analysis: Inter-subject Registration and Smoothing
 Surface-based Analysis: Inter-subject Registration and Smoothing Outline Exploratory Spatial Analysis Coordinate Systems 3D (Volumetric) 2D (Surface-based) Inter-subject registration Volume-based Surface-based
Surface-based Analysis: Inter-subject Registration and Smoothing Outline Exploratory Spatial Analysis Coordinate Systems 3D (Volumetric) 2D (Surface-based) Inter-subject registration Volume-based Surface-based
University of Florida CISE department Gator Engineering. Clustering Part 5
 Clustering Part 5 Dr. Sanjay Ranka Professor Computer and Information Science and Engineering University of Florida, Gainesville SNN Approach to Clustering Ordinary distance measures have problems Euclidean
Clustering Part 5 Dr. Sanjay Ranka Professor Computer and Information Science and Engineering University of Florida, Gainesville SNN Approach to Clustering Ordinary distance measures have problems Euclidean
UNC 4D Infant Cortical Surface Atlases, from Neonate to 6 Years of Age
 UNC 4D Infant Cortical Surface Atlases, from Neonate to 6 Years of Age Version 1.0 UNC 4D infant cortical surface atlases from neonate to 6 years of age contain 11 time points, including 1, 3, 6, 9, 12,
UNC 4D Infant Cortical Surface Atlases, from Neonate to 6 Years of Age Version 1.0 UNC 4D infant cortical surface atlases from neonate to 6 years of age contain 11 time points, including 1, 3, 6, 9, 12,
Automatic Quantification of DTI Parameters along Fiber Bundles
 Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein 1, Simon Hermann 1, Olaf Konrad 1, Horst K. Hahn 1, and Heinz-Otto Peitgen 1 1 MeVis Research, 28359 Bremen Email: klein@mevis.de
Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein 1, Simon Hermann 1, Olaf Konrad 1, Horst K. Hahn 1, and Heinz-Otto Peitgen 1 1 MeVis Research, 28359 Bremen Email: klein@mevis.de
Basic fmri Design and Analysis. Preprocessing
 Basic fmri Design and Analysis Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial filtering
Basic fmri Design and Analysis Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial filtering
2 Michael E. Leventon and Sarah F. F. Gibson a b c d Fig. 1. (a, b) Two MR scans of a person's knee. Both images have high resolution in-plane, but ha
 Model Generation from Multiple Volumes using Constrained Elastic SurfaceNets Michael E. Leventon and Sarah F. F. Gibson 1 MIT Artificial Intelligence Laboratory, Cambridge, MA 02139, USA leventon@ai.mit.edu
Model Generation from Multiple Volumes using Constrained Elastic SurfaceNets Michael E. Leventon and Sarah F. F. Gibson 1 MIT Artificial Intelligence Laboratory, Cambridge, MA 02139, USA leventon@ai.mit.edu
FMA901F: Machine Learning Lecture 3: Linear Models for Regression. Cristian Sminchisescu
 FMA901F: Machine Learning Lecture 3: Linear Models for Regression Cristian Sminchisescu Machine Learning: Frequentist vs. Bayesian In the frequentist setting, we seek a fixed parameter (vector), with value(s)
FMA901F: Machine Learning Lecture 3: Linear Models for Regression Cristian Sminchisescu Machine Learning: Frequentist vs. Bayesian In the frequentist setting, we seek a fixed parameter (vector), with value(s)
Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question.
 (1) conn - Functional connectivity toolbox v1.0 Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question. 2) in "Basic" enter "1" subject, "6" seconds
(1) conn - Functional connectivity toolbox v1.0 Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question. 2) in "Basic" enter "1" subject, "6" seconds
Analysis of fmri data within Brainvisa Example with the Saccades database
 Analysis of fmri data within Brainvisa Example with the Saccades database 18/11/2009 Note : All the sentences in italic correspond to informations relative to the specific dataset under study TP participants
Analysis of fmri data within Brainvisa Example with the Saccades database 18/11/2009 Note : All the sentences in italic correspond to informations relative to the specific dataset under study TP participants
n o r d i c B r a i n E x Tutorial DTI Module
 m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DTI Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DTI Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
White Pixel Artifact. Caused by a noise spike during acquisition Spike in K-space <--> sinusoid in image space
 White Pixel Artifact Caused by a noise spike during acquisition Spike in K-space sinusoid in image space Susceptibility Artifacts Off-resonance artifacts caused by adjacent regions with different
White Pixel Artifact Caused by a noise spike during acquisition Spike in K-space sinusoid in image space Susceptibility Artifacts Off-resonance artifacts caused by adjacent regions with different
The Diverse Club: The Integrative Core of Complex Networks M.A. Bertolero, B.T.T. Yeo, & M. D Esposito
 The Diverse Club: The Integrative Core of Complex Networks M.A. Bertolero, B.T.T. Yeo, & M. D Esposito A complex system can be represented and analyzed as a network, where nodes represent the units of
The Diverse Club: The Integrative Core of Complex Networks M.A. Bertolero, B.T.T. Yeo, & M. D Esposito A complex system can be represented and analyzed as a network, where nodes represent the units of
Ultrasonic Multi-Skip Tomography for Pipe Inspection
 18 th World Conference on Non destructive Testing, 16-2 April 212, Durban, South Africa Ultrasonic Multi-Skip Tomography for Pipe Inspection Arno VOLKER 1, Rik VOS 1 Alan HUNTER 1 1 TNO, Stieltjesweg 1,
18 th World Conference on Non destructive Testing, 16-2 April 212, Durban, South Africa Ultrasonic Multi-Skip Tomography for Pipe Inspection Arno VOLKER 1, Rik VOS 1 Alan HUNTER 1 1 TNO, Stieltjesweg 1,
FSL Workshop Session 3 David Smith & John Clithero
 FSL Workshop 12.09.08 Session 3 David Smith & John Clithero What is MELODIC? Probabilistic ICA Improves upon standard ICA Allows for inference Avoids over-fitting Three stage process ( ppca ) 1.) Dimension
FSL Workshop 12.09.08 Session 3 David Smith & John Clithero What is MELODIC? Probabilistic ICA Improves upon standard ICA Allows for inference Avoids over-fitting Three stage process ( ppca ) 1.) Dimension
Automatic Quantification of DTI Parameters along Fiber Bundles
 Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein, Simon Hermann, Olaf Konrad, Horst K. Hahn, Heinz-Otto Peitgen MeVis Research, 28359 Bremen Email: klein@mevis.de Abstract. We introduce
Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein, Simon Hermann, Olaf Konrad, Horst K. Hahn, Heinz-Otto Peitgen MeVis Research, 28359 Bremen Email: klein@mevis.de Abstract. We introduce
Clustering CS 550: Machine Learning
 Clustering CS 550: Machine Learning This slide set mainly uses the slides given in the following links: http://www-users.cs.umn.edu/~kumar/dmbook/ch8.pdf http://www-users.cs.umn.edu/~kumar/dmbook/dmslides/chap8_basic_cluster_analysis.pdf
Clustering CS 550: Machine Learning This slide set mainly uses the slides given in the following links: http://www-users.cs.umn.edu/~kumar/dmbook/ch8.pdf http://www-users.cs.umn.edu/~kumar/dmbook/dmslides/chap8_basic_cluster_analysis.pdf
1 Introduction Motivation and Aims Functional Imaging Computational Neuroanatomy... 12
 Contents 1 Introduction 10 1.1 Motivation and Aims....... 10 1.1.1 Functional Imaging.... 10 1.1.2 Computational Neuroanatomy... 12 1.2 Overview of Chapters... 14 2 Rigid Body Registration 18 2.1 Introduction.....
Contents 1 Introduction 10 1.1 Motivation and Aims....... 10 1.1.1 Functional Imaging.... 10 1.1.2 Computational Neuroanatomy... 12 1.2 Overview of Chapters... 14 2 Rigid Body Registration 18 2.1 Introduction.....
