Finding sequence similarities with genes of known function is a common approach to infer a newly sequenced gene s function
|
|
|
- Theodore Nelson
- 5 years ago
- Views:
Transcription
1
2 Outline DNA Sequence Comparison: First Success Stories Change Problem Manhattan Tourist Problem Longest Paths in Graphs Sequence Alignment Edit Distance Longest Common Subsequence Problem Dot Matrices
3 Finding sequence similarities with genes of known function is a common approach to infer a newly sequenced gene s function In 1984 Russell Doolittle and colleagues found similarities between cancer-causing gene and normal growth factor (PDGF) gene
4 Cystic fibrosis (CF) is a chronic and frequently fatal genetic disease of the body's mucus glands (abnormally high level of mucus in glands). CF primarily affects the respiratory systems in children. Mucus is a slimy material that coats many epithelial surfaces and is secreted into fluids such as saliva
5 In early 1980s biologists hypothesized that CF is an autosomal recessive disorder caused by mutations in a gene that remained unknown till 1989 Heterozygous carriers are asymptomatic Must be homozygously recessive in this gene in order to be diagnosed with CF
6 ATP binding proteins are present on cell membrane and act as transport channel In 1989 biologists found similarity between the cystic fibrosis gene and ATP binding proteins A plausible function for cystic fibrosis gene, given the fact that CF involves sweet secretion with abnormally high sodium level
7 If a high % of cystic fibrosis (CF) patients have a certain mutation in the gene and the normal patients don t, then that could be an indicator of a mutation that is related to CF A certain mutation was found in 70% of CF patients, convincing evidence that it is a predominant genetic diagnostics marker for CF
8
9 CFTR (Cystic Fibrosis Transmembrane conductance Regulator) protein is acting in the cell membrane of epithelial cells that secrete mucus These cells line the airways of the nose, lungs, the stomach wall, etc.
10 The CFTR protein (1480 amino acids) regulates a chloride ion channel Adjusts the wateriness of fluids secreted by the cell Those with cystic fibrosis are missing one single amino acid in their CFTR Mucus ends up being too thick, affecting many organs
11 Gene similarities between two genes with known and unknown function alert biologists to some possibilities Computing a similarity score between two genes tells how likely it is that they have similar functions Dynamic programming is a technique for revealing similarities between genes The Change Problem is a good problem to introduce the idea of dynamic programming
12 Goal: Convert some amount of money M into given denominations, using the fewest possible number of coins Input: An amount of money M, and an array of d denominations c = (c 1, c 2,, c d ), in a decreasing order of value (c 1 > c 2 > > c d ) Output: A list of d integers i 1, i 2,, i d such that c 1 i 1 + c 2 i c d i d = M and i 1 + i i d is minimal
13 Given the denominations 1, 3, and 5, what is the minimum number of coins needed to make change for a given value? Value Min # of coins Only one coin is needed to make change for the values 1, 3, and 5
14 Given the denominations 1, 3, and 5, what is the minimum number of coins needed to make change for a given value? Value Min # of coins However, two coins are needed to make change for the values 2, 4, 6, 8, and 10.
15 Given the denominations 1, 3, and 5, what is the minimum number of coins needed to make change for a given value? Value Min # of coins Lastly, three coins are needed to make change for the values 7 and 9
16 This example is expressed by the following recurrence relation: minnumcoins(m) = min of minnumcoins(m-1) + 1 minnumcoins(m-3) + 1 minnumcoins(m-5) + 1
17 Given the denominations c: c 1, c 2,, c d, the recurrence relation is: minnumcoins(m-c 1 ) + 1 minnumcoins(m) = min of minnumcoins(m-c 2 ) + 1 minnumcoins(m-c d ) + 1
18 1. RecursiveChange(M,c,d) 2. if M = 0 3. return 0 4. bestnumcoins infinity 5. for i 1 to d 6. if M c i 7. numcoins RecursiveChange(M c i, c, d) 8. if numcoins + 1 < bestnumcoins 9. bestnumcoins numcoins return bestnumcoins
19 It recalculates the optimal coin combination for a given amount of money repeatedly i.e., M = 77, c = (1,3,7): Optimal coin combo for 70 cents is computed 9 times! (77 7), ( ), ( ),. the optimal combination for 20 cents will be recomputed billions of time this algorithm is impractical
20 Save results of each computation for 0 to M We can do a reference call to find an already computed value, instead of re-computing each time Running time M*d, where M is the value of money and d is the number of denominations
21 1. DPChange(M,c,d) 2. bestnumcoins for m 1 to M 4. bestnumcoins m infinity 5. for i 1 to d 6. if m c i 7. if bestnumcoins m ci + 1 < bestnumcoins m 8. bestnumcoins m bestnumcoins m ci return bestnumcoins M
22 c = (1,3,7) M = 9
23 Imagine seeking a path (from source to sink) to travel (only eastward and southward) with the most number of attractions (*) in the Manhattan grid Source * * * * * * * * * * * * Sink
24 Imagine seeking a path (from source to sink) to travel (only eastward and southward) with the most number of attractions (*) in the Manhattan grid Source * * * * * * * * * * * * Sink
25 Goal: Find the longest path in a weighted grid. Input: A weighted grid G with two distinct vertices, one labeled source and the other labeled sink Output: A longest path in G from source to sink
26 source j coordinate i coordinate sink
27 source promising start, but leads to bad choices! sink
28 MT(n,m) if n=0 or m=0 return MT(n,m) x MT(n-1,m)+ length of the edge from (n- 1,m) to (n,m) y MT(n,m-1)+ length of the edge from (n,m-1) to (n,m) return max{x,y}
29 MT(n,m) x MT(n-1,m)+ length of the edge from (n- 1,m) to (n,m) y MT(n,m-1)+ return min{x,y} length of the edge from (n,m-1) to (n,m) What s wrong with this approach?
30 source j 0 1 i S 0,1 = S 1,0 = 5 Calculate optimal path score for each vertex in the graph Each vertex s score is the maximum of the prior vertices score plus the weight of the respective edge in between
31 source j i S 0,2 = S 1,1 = S 2,0 = 8
32 source j i S 3,0 = S 1,2 = S 2,1 = S 3,0 = 8
33 source j i S 1,3 = S 2,2 = greedy alg. fails! 9 S 3,1 = 9
34 source j i S 2,3 = S 3,2 = 9
35 source j i Done! (showing all back-traces) S 3,3 = 16
36 Computing the score for a point (i,j) by the recurrence relation: s i, j = max s i-1, j + weight of the edge between (i-1, j) and (i, j) s i, j-1 + weight of the edge between (i, j-1) and (i, j) The running time is n x m for a n by m grid (n = # of rows, m = # of columns)
37 A 2 A 3 A 1 B What about diagonals? The score at point B is given by: s B = max of s A1 + weight of the edge (A 1, B) s A2 + weight of the edge (A 2, B) s A3 + weight of the edge (A 3, B)
38 Computing the score for point x is given by the recurrence relation: s x = max of s y + weight of vertex (y, x) where y є Predecessors(x) Predecessors (x) set of vertices that have edges leading to x The running time for a graph G(V, E) (V is the set of all vertices and E is the set of all edges) is O(E) since each edge is evaluated once
39 The only hitch is that one must decide on the order in which visit the vertices By the time the vertex x is analyzed, the values s y for all its predecessors y should be computed otherwise we are in trouble. We need to traverse the vertices in some order Try to find such order for a directed cycle???
40 A numbering of vertices of the graph is called topological ordering of the DAG if every edge of the DAG connects a vertex with a smaller label to a vertex with a larger label (i.e. i < j if v i becomes before v j ) In other words, if vertices are positioned on a line in an increasing order of labels then all edges go from left to right.
41 3 different strategies: a) Column by column b) Row by row c) Along diagonals (useful for parallel computation) c) a) b)
42 Goal: Find a longest path between two vertices in a weighted DAG Input: A weighted DAG G with source and sink vertices Output: A longest path in G from source to sink
43 Suppose vertex v has indegree 3 and predecessors {u 1, u 2, u 3 } Longest path to v from source is: s v = max of In General: su 1 + u 1 su 2 + u 2 su 3 + u 3 s v = max u (s u + weight of edge from u to v)
44 Last time, we introduced Hamming distance (number of mismatches between two sequences). Edit distance: number of operations needed to transform one sequence into another Edit distance doesn t need the sequences to be of the same length (unlike, Hamming distance)
45 Hamming distance always compares i -th letter of v with i -th letter of w V = ATATATAT W = TATATATA Hamming distance: d(v, w)=8 Computing Hamming distance is a trivial task.
46 Hamming distance always compares i -th letter of v with i -th letter of w V = ATATATAT W = TATATATA Hamming distance: d(v, w)=8 Just one shift Make it all line up Computing Hamming distance is a trivial task Edit distance may compare i -th letter of v with j -th letter of w V = - ATATATAT W = TATATATA Edit distance: d(v, w)=2 Computing edit distance is a non-trivial task
47 Hamming distance always compares i -th letter of v with i -th letter of w V = ATATATAT W = TATATATA Hamming distance: d(v, w)=8 Edit distance may compare i -th letter of v with j -th letter of w V = - ATATATAT W = TATATATA Edit distance: d(v, w)=2 (one insertion and one deletion) How to find what j goes with what i???
48 TGCATAT ATCCGAT in 5 steps TGCATAT (delete last T) TGCATA (delete last A) TGCAT (insert A at front) ATGCAT (substitute C for 3 rd G) ATCCAT (insert G before last A) ATCCGAT (Done)
49 TGCATAT ATCCGAT in 5 steps TGCATAT (delete last T) TGCATA (delete last A) TGCAT (insert A at front) ATGCAT (substitute C for 3 rd G) ATCCAT (insert G before last A) ATCCGAT (Done) What is the edit distance? 5?
50 TGCATAT ATCCGAT in 4 steps TGCATAT (insert A at front) ATGCATAT (delete 6 th T) ATGCATA (substitute G for 5 th A) ATGCGTA (substitute C for 3 rd G) ATCCGAT (Done)
51 TGCATAT ATCCGAT in 4 steps TGCATAT (insert A at front) ATGCATAT (delete 6 th T) ATGCATA (substitute G for 5 th A) ATGCGTA (substitute C for 3 rd G) ATCCGAT (Done) Can it be done in 3 steps???
52 Given 2 DNA sequences v and w: v : w : AT CT GAT T GCAT A m = 7 n = 6 Alignment : 2 * k matrix ( k > m, n ) letters of v letters of w A T -- G T T A T -- A T C G T -- A -- C 4 matches 2 insertions 2 deletions
53 Given two sequences v = v 1 v 2 v m and w = w 1 w 2 w n The LCS of v and w is a sequence of positions in v: 1 < i 1 < i 2 < < i t < m and a sequence of positions in w: 1 < j 1 < j 2 < < j t < n such that i t -th letter of v equals to j t -letter of w and t is maximal
54 i coords: 0 elements of v elements of w j coords: A T -- C -- T G A T C -- T G C A T -- A -- C (0,0) (1,0) (2,1) (2,2) (3,3) (3,4) (4,5) (5,5) (6,6) (7,6) (8,7) Every common subsequence is a path in 2-D grid
55 Find the LCS of two strings Input: A weighted graph G with two distinct vertices, one labeled source one labeled sink Output: A longest path in G from source to sink Solve using an LCS edit graph with diagonals replaced with +1 edges
56 i T G C A T A C j A T C T G A T C Every path is a common subsequence. Every diagonal edge adds an extra element to common subsequence LCS Problem: Find a path with maximum number of diagonal edges
57 Let v i = prefix of v of length i: v 1 v i and w j = prefix of w of length j: w 1 w j The length of LCS(v i,w j ) is computed by: s i, j = max s i-1, j s i, j-1 s i-1, j if v i = w j
58 s i,j = MAX s i-1,j + 0 s i,j s i-1,j , if v i = w j i-1,j -1 i-1,j i,j -1 i,j
59 and represent indels in v and w with score 0. represent matches with score 1. The score of the alignment path is 5.
60 Every path in the edit graph corresponds to an alignment:
61 Old Alignment v= AT_GTTAT_ w= ATCGT_A_C New Alignment v= AT_GTTAT_ w= ATCG_TA_C
62 v= AT_GTTAT_ w= ATCGT_A_C (0,0), (1,1), (2,2), (2,3), (3,4), (4,5), (5,5), (6,6), (7,6), (7,7)
63 Initialize 1 st row and 1 st column to be all zeroes.
64 S i,j = S i-1, j-1 max S i-1, j S i, j-1 value from NW +1, if v i = w j value from North (top) value from West (left)
65 Find a match in row and column 2. i=2, j=2,5 is a match (T). j=2, i=4,5,7 is a match (T). Since v i = w j, s i,j = s i-1,j-1 +1 s 2,2 = [s 1,1 = 1] + 1 s 2,5 = [s 1,4 = 1] + 1 s 4,2 = [s 3,1 = 1] + 1 s 5,2 = [s 4,1 = 1] + 1 s 7,2 = [s 6,1 = 1] + 1
66 Continuing with the dynamic programming algorithm gives this result.
67 s i,j = s i-1, j-1 +1 if v i = w j max s i-1, j s i, j-1
68 Follow the arrows backwards from sink
69 1. LCS(v,w) 2. for i 1 to n 3. s i, for j 1 to m 5. s 0,j 0 6. for i 1 to n 7. for j 1 to m 8. s i-1,j 9. s i,j max s i,j s i-1,j-1 + 1, if v i = w j 11. if s i,j = s i-1,j b i,j if s i,j = s i,j-1 if s i,j = s i-1,j return (s n,m, b)
70 1. PrintLCS(b,v,i,j) 2. if i = 0 or j = 0 3. return 4. if b i,j = 5. PrintLCS(b,v,i-1,j-1) 6. print v i 7. else 8. if b i,j = 9. PrintLCS(b,v,i-1,j) 10. else 11. PrintLCS(b,v,i,j-1)
71 It takes O(nm) time to fill in the nxm dynamic programming matrix. Why O(nm)? The pseudocode consists of a nested for loop inside of another for loop to set up a nxm matrix.
72 Initialization: d i,0 = i, and d j,0 = j Recurrence: d i,j = { d i 1, j + 1, d i, j-1 + 1, d i-1, j -1, if v i = w j }
Dynamic Programming: Edit Distance
 Dynamic Programming: Edit Distance Outline. DNA Sequence Comparison and CF. Change Problem. Manhattan Tourist Problem. Longest Paths in Graphs. Sequence Alignment 6. Edit Distance Section : DNA Sequence
Dynamic Programming: Edit Distance Outline. DNA Sequence Comparison and CF. Change Problem. Manhattan Tourist Problem. Longest Paths in Graphs. Sequence Alignment 6. Edit Distance Section : DNA Sequence
Dynamic Programming Comp 122, Fall 2004
 Dynamic Programming Comp 122, Fall 2004 Review: the previous lecture Principles of dynamic programming: optimization problems, optimal substructure property, overlapping subproblems, trade space for time,
Dynamic Programming Comp 122, Fall 2004 Review: the previous lecture Principles of dynamic programming: optimization problems, optimal substructure property, overlapping subproblems, trade space for time,
EECS 4425: Introductory Computational Bioinformatics Fall Suprakash Datta
 EECS 4425: Introductory Computational Bioinformatics Fall 2018 Suprakash Datta datta [at] cse.yorku.ca Office: CSEB 3043 Phone: 416-736-2100 ext 77875 Course page: http://www.cse.yorku.ca/course/4425 Many
EECS 4425: Introductory Computational Bioinformatics Fall 2018 Suprakash Datta datta [at] cse.yorku.ca Office: CSEB 3043 Phone: 416-736-2100 ext 77875 Course page: http://www.cse.yorku.ca/course/4425 Many
Dynamic Programming: Sequence alignment. CS 466 Saurabh Sinha
 Dynamic Programming: Sequence alignment CS 466 Saurabh Sinha DNA Sequence Comparison: First Success Story Finding sequence similarities with genes of known function is a common approach to infer a newly
Dynamic Programming: Sequence alignment CS 466 Saurabh Sinha DNA Sequence Comparison: First Success Story Finding sequence similarities with genes of known function is a common approach to infer a newly
Dynamic Programming Part I: Examples. Bioinfo I (Institut Pasteur de Montevideo) Dynamic Programming -class4- July 25th, / 77
 Dynamic Programming Part I: Examples Bioinfo I (Institut Pasteur de Montevideo) Dynamic Programming -class4- July 25th, 2011 1 / 77 Dynamic Programming Recall: the Change Problem Other problems: Manhattan
Dynamic Programming Part I: Examples Bioinfo I (Institut Pasteur de Montevideo) Dynamic Programming -class4- July 25th, 2011 1 / 77 Dynamic Programming Recall: the Change Problem Other problems: Manhattan
1. Basic idea: Use smaller instances of the problem to find the solution of larger instances
 Chapter 8. Dynamic Programming CmSc Intro to Algorithms. Basic idea: Use smaller instances of the problem to find the solution of larger instances Example : Fibonacci numbers F = F = F n = F n- + F n-
Chapter 8. Dynamic Programming CmSc Intro to Algorithms. Basic idea: Use smaller instances of the problem to find the solution of larger instances Example : Fibonacci numbers F = F = F n = F n- + F n-
9/29/09 Comp /Comp Fall
 9/29/9 Comp 9-9/Comp 79-9 Fall 29 1 So far we ve tried: A greedy algorithm that does not work for all inputs (it is incorrect) An exhaustive search algorithm that is correct, but can take a long time A
9/29/9 Comp 9-9/Comp 79-9 Fall 29 1 So far we ve tried: A greedy algorithm that does not work for all inputs (it is incorrect) An exhaustive search algorithm that is correct, but can take a long time A
Pairwise Sequence alignment Basic Algorithms
 Pairwise Sequence alignment Basic Algorithms Agenda - Previous Lesson: Minhala - + Biological Story on Biomolecular Sequences - + General Overview of Problems in Computational Biology - Reminder: Dynamic
Pairwise Sequence alignment Basic Algorithms Agenda - Previous Lesson: Minhala - + Biological Story on Biomolecular Sequences - + General Overview of Problems in Computational Biology - Reminder: Dynamic
6.1 The Power of DNA Sequence Comparison
 6 Dynamic Programming Algorithms We introduced dynamic programming in chapter 2 with the Rocks problem. While the Rocks problem does not appear to be related to bioinformatics, the algorithm that we described
6 Dynamic Programming Algorithms We introduced dynamic programming in chapter 2 with the Rocks problem. While the Rocks problem does not appear to be related to bioinformatics, the algorithm that we described
DynamicProgramming. September 17, 2018
 DynamicProgramming September 17, 2018 1 Lecture 11: Dynamic Programming CBIO (CSCI) 4835/6835: Introduction to Computational Biology 1.1 Overview and Objectives We ve so far discussed sequence alignment
DynamicProgramming September 17, 2018 1 Lecture 11: Dynamic Programming CBIO (CSCI) 4835/6835: Introduction to Computational Biology 1.1 Overview and Objectives We ve so far discussed sequence alignment
3/5/2018 Lecture15. Comparing Sequences. By Miguel Andrade at English Wikipedia.
 3/5/2018 Lecture15 Comparing Sequences By Miguel Andrade at English Wikipedia 1 http://localhost:8888/notebooks/comp555s18/lecture15.ipynb# 1/1 Sequence Similarity A common problem in Biology Insulin Protein
3/5/2018 Lecture15 Comparing Sequences By Miguel Andrade at English Wikipedia 1 http://localhost:8888/notebooks/comp555s18/lecture15.ipynb# 1/1 Sequence Similarity A common problem in Biology Insulin Protein
Special course in Computer Science: Advanced Text Algorithms
 Special course in Computer Science: Advanced Text Algorithms Lecture 6: Alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Special course in Computer Science: Advanced Text Algorithms Lecture 6: Alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Sequence Alignment: Mo1va1on and Algorithms. Lecture 2: August 23, 2012
 Sequence Alignment: Mo1va1on and Algorithms Lecture 2: August 23, 2012 Mo1va1on and Introduc1on Importance of Sequence Alignment For DNA, RNA and amino acid sequences, high sequence similarity usually
Sequence Alignment: Mo1va1on and Algorithms Lecture 2: August 23, 2012 Mo1va1on and Introduc1on Importance of Sequence Alignment For DNA, RNA and amino acid sequences, high sequence similarity usually
Chapter 3, Algorithms Algorithms
 CSI 2350, Discrete Structures Chapter 3, Algorithms Young-Rae Cho Associate Professor Department of Computer Science Baylor University 3.1. Algorithms Definition A finite sequence of precise instructions
CSI 2350, Discrete Structures Chapter 3, Algorithms Young-Rae Cho Associate Professor Department of Computer Science Baylor University 3.1. Algorithms Definition A finite sequence of precise instructions
DNA Alignment With Affine Gap Penalties
 DNA Alignment With Affine Gap Penalties Laurel Schuster Why Use Affine Gap Penalties? When aligning two DNA sequences, one goal may be to infer the mutations that made them different. Though it s impossible
DNA Alignment With Affine Gap Penalties Laurel Schuster Why Use Affine Gap Penalties? When aligning two DNA sequences, one goal may be to infer the mutations that made them different. Though it s impossible
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 06: Multiple Sequence Alignment https://upload.wikimedia.org/wikipedia/commons/thumb/7/79/rplp0_90_clustalw_aln.gif/575px-rplp0_90_clustalw_aln.gif Slides
EECS730: Introduction to Bioinformatics Lecture 06: Multiple Sequence Alignment https://upload.wikimedia.org/wikipedia/commons/thumb/7/79/rplp0_90_clustalw_aln.gif/575px-rplp0_90_clustalw_aln.gif Slides
Global Alignment Scoring Matrices Local Alignment Alignment with Affine Gap Penalties
 Global Alignment Scoring Matrices Local Alignment Alignment with Affine Gap Penalties From LCS to Alignment: Change the Scoring The Longest Common Subsequence (LCS) problem the simplest form of sequence
Global Alignment Scoring Matrices Local Alignment Alignment with Affine Gap Penalties From LCS to Alignment: Change the Scoring The Longest Common Subsequence (LCS) problem the simplest form of sequence
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 04: Variations of sequence alignments http://www.pitt.edu/~mcs2/teaching/biocomp/tutorials/global.html Slides adapted from Dr. Shaojie Zhang (University
EECS730: Introduction to Bioinformatics Lecture 04: Variations of sequence alignments http://www.pitt.edu/~mcs2/teaching/biocomp/tutorials/global.html Slides adapted from Dr. Shaojie Zhang (University
Sequence Alignment: Mo1va1on and Algorithms
 Sequence Alignment: Mo1va1on and Algorithms Mo1va1on and Introduc1on Importance of Sequence Alignment For DNA, RNA and amino acid sequences, high sequence similarity usually implies significant func1onal
Sequence Alignment: Mo1va1on and Algorithms Mo1va1on and Introduc1on Importance of Sequence Alignment For DNA, RNA and amino acid sequences, high sequence similarity usually implies significant func1onal
Computational Molecular Biology
 Computational Molecular Biology Erwin M. Bakker Lecture 2 Materials used from R. Shamir [2] and H.J. Hoogeboom [4]. 1 Molecular Biology Sequences DNA A, T, C, G RNA A, U, C, G Protein A, R, D, N, C E,
Computational Molecular Biology Erwin M. Bakker Lecture 2 Materials used from R. Shamir [2] and H.J. Hoogeboom [4]. 1 Molecular Biology Sequences DNA A, T, C, G RNA A, U, C, G Protein A, R, D, N, C E,
Lecture 10: Local Alignments
 Lecture 10: Local Alignments Study Chapter 6.8-6.10 1 Outline Edit Distances Longest Common Subsequence Global Sequence Alignment Scoring Matrices Local Sequence Alignment Alignment with Affine Gap Penalties
Lecture 10: Local Alignments Study Chapter 6.8-6.10 1 Outline Edit Distances Longest Common Subsequence Global Sequence Alignment Scoring Matrices Local Sequence Alignment Alignment with Affine Gap Penalties
Presentation for use with the textbook, Algorithm Design and Applications, by M. T. Goodrich and R. Tamassia, Wiley, Dynamic Programming
 Presentation for use with the textbook, Algorithm Design and Applications, by M. T. Goodrich and R. Tamassia, Wiley, 25 Dynamic Programming Terrible Fibonacci Computation Fibonacci sequence: f = f(n) 2
Presentation for use with the textbook, Algorithm Design and Applications, by M. T. Goodrich and R. Tamassia, Wiley, 25 Dynamic Programming Terrible Fibonacci Computation Fibonacci sequence: f = f(n) 2
Dynamic Programming. Lecture Overview Introduction
 Lecture 12 Dynamic Programming 12.1 Overview Dynamic Programming is a powerful technique that allows one to solve many different types of problems in time O(n 2 ) or O(n 3 ) for which a naive approach
Lecture 12 Dynamic Programming 12.1 Overview Dynamic Programming is a powerful technique that allows one to solve many different types of problems in time O(n 2 ) or O(n 3 ) for which a naive approach
Computational Genomics and Molecular Biology, Fall
 Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
Evolutionary tree reconstruction (Chapter 10)
 Evolutionary tree reconstruction (Chapter 10) Early Evolutionary Studies Anatomical features were the dominant criteria used to derive evolutionary relationships between species since Darwin till early
Evolutionary tree reconstruction (Chapter 10) Early Evolutionary Studies Anatomical features were the dominant criteria used to derive evolutionary relationships between species since Darwin till early
Data Mining Technologies for Bioinformatics Sequences
 Data Mining Technologies for Bioinformatics Sequences Deepak Garg Computer Science and Engineering Department Thapar Institute of Engineering & Tecnology, Patiala Abstract Main tool used for sequence alignment
Data Mining Technologies for Bioinformatics Sequences Deepak Garg Computer Science and Engineering Department Thapar Institute of Engineering & Tecnology, Patiala Abstract Main tool used for sequence alignment
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014
 Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
Gene regulation. DNA is merely the blueprint Shared spatially (among all tissues) and temporally But cells manage to differentiate
 Gene regulation DNA is merely the blueprint Shared spatially (among all tissues) and temporally But cells manage to differentiate Especially but not only during developmental stage And cells respond to
Gene regulation DNA is merely the blueprint Shared spatially (among all tissues) and temporally But cells manage to differentiate Especially but not only during developmental stage And cells respond to
Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD
 Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD Assumptions: Biological sequences evolved by evolution. Micro scale changes: For short sequences (e.g. one
Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD Assumptions: Biological sequences evolved by evolution. Micro scale changes: For short sequences (e.g. one
Dynamic Programming Algorithms
 CSC 364S Notes University of Toronto, Fall 2003 Dynamic Programming Algorithms The setting is as follows. We wish to find a solution to a given problem which optimizes some quantity Q of interest; for
CSC 364S Notes University of Toronto, Fall 2003 Dynamic Programming Algorithms The setting is as follows. We wish to find a solution to a given problem which optimizes some quantity Q of interest; for
Returning to Dynamic Programming
 Returning to Dynamic Programming 1 What is an Algorithm? An algorithm is a sequence of instructions that one must perform in order to solve a well-formulated problem. 2 Correctness An algorithm is correct
Returning to Dynamic Programming 1 What is an Algorithm? An algorithm is a sequence of instructions that one must perform in order to solve a well-formulated problem. 2 Correctness An algorithm is correct
.. Fall 2011 CSC 570: Bioinformatics Alexander Dekhtyar..
 .. Fall 2011 CSC 570: Bioinformatics Alexander Dekhtyar.. PAM and BLOSUM Matrices Prepared by: Jason Banich and Chris Hoover Background As DNA sequences change and evolve, certain amino acids are more
.. Fall 2011 CSC 570: Bioinformatics Alexander Dekhtyar.. PAM and BLOSUM Matrices Prepared by: Jason Banich and Chris Hoover Background As DNA sequences change and evolve, certain amino acids are more
CMSC351 - Fall 2014, Homework #4
 CMSC351 - Fall 2014, Homework #4 Due: November 14th at the start of class PRINT Name: Grades depend on neatness and clarity. Write your answers with enough detail about your approach and concepts used,
CMSC351 - Fall 2014, Homework #4 Due: November 14th at the start of class PRINT Name: Grades depend on neatness and clarity. Write your answers with enough detail about your approach and concepts used,
CSE 417 Dynamic Programming (pt 5) Multiple Inputs
 CSE 417 Dynamic Programming (pt 5) Multiple Inputs Reminders > HW5 due Wednesday Dynamic Programming Review > Apply the steps... optimal substructure: (small) set of solutions, constructed from solutions
CSE 417 Dynamic Programming (pt 5) Multiple Inputs Reminders > HW5 due Wednesday Dynamic Programming Review > Apply the steps... optimal substructure: (small) set of solutions, constructed from solutions
BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio CS 466 Saurabh Sinha
 BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio. 1990. CS 466 Saurabh Sinha Motivation Sequence homology to a known protein suggest function of newly sequenced protein Bioinformatics
BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio. 1990. CS 466 Saurabh Sinha Motivation Sequence homology to a known protein suggest function of newly sequenced protein Bioinformatics
15-451/651: Design & Analysis of Algorithms January 26, 2015 Dynamic Programming I last changed: January 28, 2015
 15-451/651: Design & Analysis of Algorithms January 26, 2015 Dynamic Programming I last changed: January 28, 2015 Dynamic Programming is a powerful technique that allows one to solve many different types
15-451/651: Design & Analysis of Algorithms January 26, 2015 Dynamic Programming I last changed: January 28, 2015 Dynamic Programming is a powerful technique that allows one to solve many different types
5.1 The String reconstruction problem
 CS125 Lecture 5 Fall 2014 5.1 The String reconstruction problem The greedy approach doesn t always work, as we have seen. It lacks flexibility; if at some point, it makes a wrong choice, it becomes stuck.
CS125 Lecture 5 Fall 2014 5.1 The String reconstruction problem The greedy approach doesn t always work, as we have seen. It lacks flexibility; if at some point, it makes a wrong choice, it becomes stuck.
CSED233: Data Structures (2017F) Lecture12: Strings and Dynamic Programming
 (2017F) Lecture12: Strings and Dynamic Programming Daijin Kim CSE, POSTECH dkim@postech.ac.kr Strings A string is a sequence of characters Examples of strings: Python program HTML document DNA sequence
(2017F) Lecture12: Strings and Dynamic Programming Daijin Kim CSE, POSTECH dkim@postech.ac.kr Strings A string is a sequence of characters Examples of strings: Python program HTML document DNA sequence
Divide and Conquer. Bioinformatics: Issues and Algorithms. CSE Fall 2007 Lecture 12
 Divide and Conquer Bioinformatics: Issues and Algorithms CSE 308-408 Fall 007 Lecture 1 Lopresti Fall 007 Lecture 1-1 - Outline MergeSort Finding mid-point in alignment matrix in linear space Linear space
Divide and Conquer Bioinformatics: Issues and Algorithms CSE 308-408 Fall 007 Lecture 1 Lopresti Fall 007 Lecture 1-1 - Outline MergeSort Finding mid-point in alignment matrix in linear space Linear space
Lecture 12: Divide and Conquer Algorithms
 Lecture 12: Divide and Conquer Algorithms Study Chapter 7.1 7.4 1 Divide and Conquer Algorithms Divide problem into sub-problems Conquer by solving sub-problems recursively. If the sub-problems are small
Lecture 12: Divide and Conquer Algorithms Study Chapter 7.1 7.4 1 Divide and Conquer Algorithms Divide problem into sub-problems Conquer by solving sub-problems recursively. If the sub-problems are small
Divide & Conquer Algorithms
 Divide & Conquer Algorithms Outline 1. MergeSort 2. Finding the middle vertex 3. Linear space sequence alignment 4. Block alignment 5. Four-Russians speedup 6. LCS in sub-quadratic time Section 1: MergeSort
Divide & Conquer Algorithms Outline 1. MergeSort 2. Finding the middle vertex 3. Linear space sequence alignment 4. Block alignment 5. Four-Russians speedup 6. LCS in sub-quadratic time Section 1: MergeSort
CMPS 102 Solutions to Homework 7
 CMPS 102 Solutions to Homework 7 Kuzmin, Cormen, Brown, lbrown@soe.ucsc.edu November 17, 2005 Problem 1. 15.4-1 p.355 LCS Determine an LCS of x = (1, 0, 0, 1, 0, 1, 0, 1) and y = (0, 1, 0, 1, 1, 0, 1,
CMPS 102 Solutions to Homework 7 Kuzmin, Cormen, Brown, lbrown@soe.ucsc.edu November 17, 2005 Problem 1. 15.4-1 p.355 LCS Determine an LCS of x = (1, 0, 0, 1, 0, 1, 0, 1) and y = (0, 1, 0, 1, 1, 0, 1,
Lecture 9: Core String Edits and Alignments
 Biosequence Algorithms, Spring 2005 Lecture 9: Core String Edits and Alignments Pekka Kilpeläinen University of Kuopio Department of Computer Science BSA Lecture 9: String Edits and Alignments p.1/30 III:
Biosequence Algorithms, Spring 2005 Lecture 9: Core String Edits and Alignments Pekka Kilpeläinen University of Kuopio Department of Computer Science BSA Lecture 9: String Edits and Alignments p.1/30 III:
Parsimony-Based Approaches to Inferring Phylogenetic Trees
 Parsimony-Based Approaches to Inferring Phylogenetic Trees BMI/CS 576 www.biostat.wisc.edu/bmi576.html Mark Craven craven@biostat.wisc.edu Fall 0 Phylogenetic tree approaches! three general types! distance:
Parsimony-Based Approaches to Inferring Phylogenetic Trees BMI/CS 576 www.biostat.wisc.edu/bmi576.html Mark Craven craven@biostat.wisc.edu Fall 0 Phylogenetic tree approaches! three general types! distance:
Divide & Conquer Algorithms
 Divide & Conquer Algorithms Outline MergeSort Finding the middle point in the alignment matrix in linear space Linear space sequence alignment Block Alignment Four-Russians speedup Constructing LCS in
Divide & Conquer Algorithms Outline MergeSort Finding the middle point in the alignment matrix in linear space Linear space sequence alignment Block Alignment Four-Russians speedup Constructing LCS in
Robert Collins CSE586 CSE 586, Spring 2009 Advanced Computer Vision
 CSE 586, Spring 2009 Advanced Computer Vision Dynamic Programming Decision Sequences Let D be a decision sequence {d1,d2,,dn}. (e.g. in a shortest path problem, d1 is the start node, d2 is the second node
CSE 586, Spring 2009 Advanced Computer Vision Dynamic Programming Decision Sequences Let D be a decision sequence {d1,d2,,dn}. (e.g. in a shortest path problem, d1 is the start node, d2 is the second node
Dynamic Programming. An Enumeration Approach. Matrix Chain-Products. Matrix Chain-Products (not in book)
 Matrix Chain-Products (not in book) is a general algorithm design paradigm. Rather than give the general structure, let us first give a motivating example: Matrix Chain-Products Review: Matrix Multiplication.
Matrix Chain-Products (not in book) is a general algorithm design paradigm. Rather than give the general structure, let us first give a motivating example: Matrix Chain-Products Review: Matrix Multiplication.
Dynamic Programming. CIS 110, Fall University of Pennsylvania
 Dynamic Programming CIS 110, Fall 2012 University of Pennsylvania Dynamic Programming Dynamic programming records saves computation for reuse later. Programming: in the optimization sense ( Linear Programming
Dynamic Programming CIS 110, Fall 2012 University of Pennsylvania Dynamic Programming Dynamic programming records saves computation for reuse later. Programming: in the optimization sense ( Linear Programming
Shortest Path Algorithm
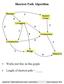 Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
Algorithms IV. Dynamic Programming. Guoqiang Li. School of Software, Shanghai Jiao Tong University
 Algorithms IV Dynamic Programming Guoqiang Li School of Software, Shanghai Jiao Tong University Dynamic Programming Shortest Paths in Dags, Revisited Shortest Paths in Dags, Revisited The special distinguishing
Algorithms IV Dynamic Programming Guoqiang Li School of Software, Shanghai Jiao Tong University Dynamic Programming Shortest Paths in Dags, Revisited Shortest Paths in Dags, Revisited The special distinguishing
Gene expression & Clustering (Chapter 10)
 Gene expression & Clustering (Chapter 10) Determining gene function Sequence comparison tells us if a gene is similar to another gene, e.g., in a new species Dynamic programming Approximate pattern matching
Gene expression & Clustering (Chapter 10) Determining gene function Sequence comparison tells us if a gene is similar to another gene, e.g., in a new species Dynamic programming Approximate pattern matching
Subset sum problem and dynamic programming
 Lecture Notes: Dynamic programming We will discuss the subset sum problem (introduced last time), and introduce the main idea of dynamic programming. We illustrate it further using a variant of the so-called
Lecture Notes: Dynamic programming We will discuss the subset sum problem (introduced last time), and introduce the main idea of dynamic programming. We illustrate it further using a variant of the so-called
Computational Molecular Biology
 Computational Molecular Biology Erwin M. Bakker Lecture 3, mainly from material by R. Shamir [2] and H.J. Hoogeboom [4]. 1 Pairwise Sequence Alignment Biological Motivation Algorithmic Aspect Recursive
Computational Molecular Biology Erwin M. Bakker Lecture 3, mainly from material by R. Shamir [2] and H.J. Hoogeboom [4]. 1 Pairwise Sequence Alignment Biological Motivation Algorithmic Aspect Recursive
Dynamic Programming II
 June 9, 214 DP: Longest common subsequence biologists often need to find out how similar are 2 DNA sequences DNA sequences are strings of bases: A, C, T and G how to define similarity? DP: Longest common
June 9, 214 DP: Longest common subsequence biologists often need to find out how similar are 2 DNA sequences DNA sequences are strings of bases: A, C, T and G how to define similarity? DP: Longest common
Memoization/Dynamic Programming. The String reconstruction problem. CS124 Lecture 11 Spring 2018
 CS124 Lecture 11 Spring 2018 Memoization/Dynamic Programming Today s lecture discusses memoization, which is a method for speeding up algorithms based on recursion, by using additional memory to remember
CS124 Lecture 11 Spring 2018 Memoization/Dynamic Programming Today s lecture discusses memoization, which is a method for speeding up algorithms based on recursion, by using additional memory to remember
BIOE 198MI Biomedical Data Analysis. Spring Semester Dynamic programming: finding the shortest path
 BIOE 98MI Biomedical Data Analysis. Spring Semester 09. Dynamic programming: finding the shortest path Page Problem Statement: we re going to learn how to convert real life problem into a graphical diagram
BIOE 98MI Biomedical Data Analysis. Spring Semester 09. Dynamic programming: finding the shortest path Page Problem Statement: we re going to learn how to convert real life problem into a graphical diagram
Dynamic Programming part 2
 Dynamic Programming part 2 Week 7 Objectives More dynamic programming examples - Matrix Multiplication Parenthesis - Longest Common Subsequence Subproblem Optimal structure Defining the dynamic recurrence
Dynamic Programming part 2 Week 7 Objectives More dynamic programming examples - Matrix Multiplication Parenthesis - Longest Common Subsequence Subproblem Optimal structure Defining the dynamic recurrence
Source. Sink. Chapter 10: Iterative Programming Maximum Flow Problem. CmSc250 Intro to Algorithms
 Chapter 10: Iterative Programming Maximum Flow Problem CmSc20 Intro to Algorithms A flow network is a model of a system where some material is produced at its source, travels through the system, and is
Chapter 10: Iterative Programming Maximum Flow Problem CmSc20 Intro to Algorithms A flow network is a model of a system where some material is produced at its source, travels through the system, and is
MULTIPLE SEQUENCE ALIGNMENT
 MULTIPLE SEQUENCE ALIGNMENT Multiple Alignment versus Pairwise Alignment Up until now we have only tried to align two sequences. What about more than two? A faint similarity between two sequences becomes
MULTIPLE SEQUENCE ALIGNMENT Multiple Alignment versus Pairwise Alignment Up until now we have only tried to align two sequences. What about more than two? A faint similarity between two sequences becomes
So far... Finished looking at lower bounds and linear sorts.
 So far... Finished looking at lower bounds and linear sorts. Next: Memoization -- Optimization problems - Dynamic programming A scheduling problem Matrix multiplication optimization Longest Common Subsequence
So far... Finished looking at lower bounds and linear sorts. Next: Memoization -- Optimization problems - Dynamic programming A scheduling problem Matrix multiplication optimization Longest Common Subsequence
Lecture 11: Dynamic Programming Steven Skiena. skiena
 Lecture 11: Dynamic Programming Steven Skiena Department of Computer Science State University of New York Stony Brook, NY 11794 4400 http://www.cs.sunysb.edu/ skiena Dynamic Programming Dynamic programming
Lecture 11: Dynamic Programming Steven Skiena Department of Computer Science State University of New York Stony Brook, NY 11794 4400 http://www.cs.sunysb.edu/ skiena Dynamic Programming Dynamic programming
Longest Common Subsequence, Knapsack, Independent Set Scribe: Wilbur Yang (2016), Mary Wootters (2017) Date: November 6, 2017
 CS161 Lecture 13 Longest Common Subsequence, Knapsack, Independent Set Scribe: Wilbur Yang (2016), Mary Wootters (2017) Date: November 6, 2017 1 Overview Last lecture, we talked about dynamic programming
CS161 Lecture 13 Longest Common Subsequence, Knapsack, Independent Set Scribe: Wilbur Yang (2016), Mary Wootters (2017) Date: November 6, 2017 1 Overview Last lecture, we talked about dynamic programming
Sequence Alignment. Ulf Leser
 Sequence Alignment Ulf Leser his Lecture Approximate String Matching Edit distance and alignment Computing global alignments Local alignment Ulf Leser: Bioinformatics, Summer Semester 2016 2 ene Function
Sequence Alignment Ulf Leser his Lecture Approximate String Matching Edit distance and alignment Computing global alignments Local alignment Ulf Leser: Bioinformatics, Summer Semester 2016 2 ene Function
Chapter 6. Dynamic Programming
 Chapter 6 Dynamic Programming CS 573: Algorithms, Fall 203 September 2, 203 6. Maximum Weighted Independent Set in Trees 6..0. Maximum Weight Independent Set Problem Input Graph G = (V, E) and weights
Chapter 6 Dynamic Programming CS 573: Algorithms, Fall 203 September 2, 203 6. Maximum Weighted Independent Set in Trees 6..0. Maximum Weight Independent Set Problem Input Graph G = (V, E) and weights
Lecture 3, Review of Algorithms. What is Algorithm?
 BINF 336, Introduction to Computational Biology Lecture 3, Review of Algorithms Young-Rae Cho Associate Professor Department of Computer Science Baylor University What is Algorithm? Definition A process
BINF 336, Introduction to Computational Biology Lecture 3, Review of Algorithms Young-Rae Cho Associate Professor Department of Computer Science Baylor University What is Algorithm? Definition A process
Machine Learning. Computational biology: Sequence alignment and profile HMMs
 10-601 Machine Learning Computational biology: Sequence alignment and profile HMMs Central dogma DNA CCTGAGCCAACTATTGATGAA transcription mrna CCUGAGCCAACUAUUGAUGAA translation Protein PEPTIDE 2 Growth
10-601 Machine Learning Computational biology: Sequence alignment and profile HMMs Central dogma DNA CCTGAGCCAACTATTGATGAA transcription mrna CCUGAGCCAACUAUUGAUGAA translation Protein PEPTIDE 2 Growth
Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it.
 Sequence Alignments Overview Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it. Sequence alignment means arranging
Sequence Alignments Overview Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it. Sequence alignment means arranging
Divide and Conquer Algorithms. Problem Set #3 is graded Problem Set #4 due on Thursday
 Divide and Conquer Algorithms Problem Set #3 is graded Problem Set #4 due on Thursday 1 The Essence of Divide and Conquer Divide problem into sub-problems Conquer by solving sub-problems recursively. If
Divide and Conquer Algorithms Problem Set #3 is graded Problem Set #4 due on Thursday 1 The Essence of Divide and Conquer Divide problem into sub-problems Conquer by solving sub-problems recursively. If
Special course in Computer Science: Advanced Text Algorithms
 Special course in Computer Science: Advanced Text Algorithms Lecture 8: Multiple alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Special course in Computer Science: Advanced Text Algorithms Lecture 8: Multiple alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Efficient Sequential Algorithms, Comp309. Motivation. Longest Common Subsequence. Part 3. String Algorithms
 Efficient Sequential Algorithms, Comp39 Part 3. String Algorithms University of Liverpool References: T. H. Cormen, C. E. Leiserson, R. L. Rivest Introduction to Algorithms, Second Edition. MIT Press (21).
Efficient Sequential Algorithms, Comp39 Part 3. String Algorithms University of Liverpool References: T. H. Cormen, C. E. Leiserson, R. L. Rivest Introduction to Algorithms, Second Edition. MIT Press (21).
FastA & the chaining problem
 FastA & the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 1 Sources for this lecture: Lectures by Volker Heun, Daniel Huson and Knut Reinert,
FastA & the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 1 Sources for this lecture: Lectures by Volker Heun, Daniel Huson and Knut Reinert,
Computer Science 385 Design and Analysis of Algorithms Siena College Spring Topic Notes: Dynamic Programming
 Computer Science 385 Design and Analysis of Algorithms Siena College Spring 29 Topic Notes: Dynamic Programming We next consider dynamic programming, a technique for designing algorithms to solve problems
Computer Science 385 Design and Analysis of Algorithms Siena College Spring 29 Topic Notes: Dynamic Programming We next consider dynamic programming, a technique for designing algorithms to solve problems
FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:
 FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:56 4001 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem
FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:56 4001 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem
Longest Common Subsequence. Definitions
 Longest Common Subsequence LCS is an interesting variation on the classical string matching problem: the task is that of finding the common portion of two strings (more precise definition in a couple of
Longest Common Subsequence LCS is an interesting variation on the classical string matching problem: the task is that of finding the common portion of two strings (more precise definition in a couple of
Clustering Techniques
 Clustering Techniques Bioinformatics: Issues and Algorithms CSE 308-408 Fall 2007 Lecture 16 Lopresti Fall 2007 Lecture 16-1 - Administrative notes Your final project / paper proposal is due on Friday,
Clustering Techniques Bioinformatics: Issues and Algorithms CSE 308-408 Fall 2007 Lecture 16 Lopresti Fall 2007 Lecture 16-1 - Administrative notes Your final project / paper proposal is due on Friday,
Biology 644: Bioinformatics
 Find the best alignment between 2 sequences with lengths n and m, respectively Best alignment is very dependent upon the substitution matrix and gap penalties The Global Alignment Problem tries to find
Find the best alignment between 2 sequences with lengths n and m, respectively Best alignment is very dependent upon the substitution matrix and gap penalties The Global Alignment Problem tries to find
Lectures by Volker Heun, Daniel Huson and Knut Reinert, in particular last years lectures
 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 4.1 Sources for this lecture Lectures by Volker Heun, Daniel Huson and Knut
4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 4.1 Sources for this lecture Lectures by Volker Heun, Daniel Huson and Knut
11/22/2016. Chapter 9 Graph Algorithms. Introduction. Definitions. Definitions. Definitions. Definitions
 Introduction Chapter 9 Graph Algorithms graph theory useful in practice represent many real-life problems can be slow if not careful with data structures 2 Definitions an undirected graph G = (V, E) is
Introduction Chapter 9 Graph Algorithms graph theory useful in practice represent many real-life problems can be slow if not careful with data structures 2 Definitions an undirected graph G = (V, E) is
Introduction to Programming in C Department of Computer Science and Engineering. Lecture No. #16 Loops: Matrix Using Nested for Loop
 Introduction to Programming in C Department of Computer Science and Engineering Lecture No. #16 Loops: Matrix Using Nested for Loop In this section, we will use the, for loop to code of the matrix problem.
Introduction to Programming in C Department of Computer Science and Engineering Lecture No. #16 Loops: Matrix Using Nested for Loop In this section, we will use the, for loop to code of the matrix problem.
Chapter 9 Graph Algorithms
 Chapter 9 Graph Algorithms 2 Introduction graph theory useful in practice represent many real-life problems can be slow if not careful with data structures 3 Definitions an undirected graph G = (V, E)
Chapter 9 Graph Algorithms 2 Introduction graph theory useful in practice represent many real-life problems can be slow if not careful with data structures 3 Definitions an undirected graph G = (V, E)
Subsequence Definition. CS 461, Lecture 8. Today s Outline. Example. Assume given sequence X = x 1, x 2,..., x m. Jared Saia University of New Mexico
 Subsequence Definition CS 461, Lecture 8 Jared Saia University of New Mexico Assume given sequence X = x 1, x 2,..., x m Let Z = z 1, z 2,..., z l Then Z is a subsequence of X if there exists a strictly
Subsequence Definition CS 461, Lecture 8 Jared Saia University of New Mexico Assume given sequence X = x 1, x 2,..., x m Let Z = z 1, z 2,..., z l Then Z is a subsequence of X if there exists a strictly
CSE 101- Winter 18 Discussion Section Week 6
 CSE 101- Winter 18 Discussion Section Week 6 Administrative Introducing 1:1 Sessions: https://docs.google.com/spreadsheets/d/1kgxt_rzbzlibbdijiczs_ o1wxdwa9hhvxccprn8_bwk/edit?usp=sharing Please see the
CSE 101- Winter 18 Discussion Section Week 6 Administrative Introducing 1:1 Sessions: https://docs.google.com/spreadsheets/d/1kgxt_rzbzlibbdijiczs_ o1wxdwa9hhvxccprn8_bwk/edit?usp=sharing Please see the
Acceleration of the Smith-Waterman algorithm for DNA sequence alignment using an FPGA platform
 Acceleration of the Smith-Waterman algorithm for DNA sequence alignment using an FPGA platform Barry Strengholt Matthijs Brobbel Delft University of Technology Faculty of Electrical Engineering, Mathematics
Acceleration of the Smith-Waterman algorithm for DNA sequence alignment using an FPGA platform Barry Strengholt Matthijs Brobbel Delft University of Technology Faculty of Electrical Engineering, Mathematics
OPEN MP-BASED PARALLEL AND SCALABLE GENETIC SEQUENCE ALIGNMENT
 OPEN MP-BASED PARALLEL AND SCALABLE GENETIC SEQUENCE ALIGNMENT Asif Ali Khan*, Laiq Hassan*, Salim Ullah* ABSTRACT: In bioinformatics, sequence alignment is a common and insistent task. Biologists align
OPEN MP-BASED PARALLEL AND SCALABLE GENETIC SEQUENCE ALIGNMENT Asif Ali Khan*, Laiq Hassan*, Salim Ullah* ABSTRACT: In bioinformatics, sequence alignment is a common and insistent task. Biologists align
Sequence Alignment 1
 Sequence Alignment 1 Nucleotide and Base Pairs Purine: A and G Pyrimidine: T and C 2 DNA 3 For this course DNA is double-helical, with two complementary strands. Complementary bases: Adenine (A) - Thymine
Sequence Alignment 1 Nucleotide and Base Pairs Purine: A and G Pyrimidine: T and C 2 DNA 3 For this course DNA is double-helical, with two complementary strands. Complementary bases: Adenine (A) - Thymine
1 Dynamic Programming
 CS161 Lecture 13 Dynamic Programming and Greedy Algorithms Scribe by: Eric Huang Date: May 13, 2015 1 Dynamic Programming The idea of dynamic programming is to have a table of solutions of subproblems
CS161 Lecture 13 Dynamic Programming and Greedy Algorithms Scribe by: Eric Huang Date: May 13, 2015 1 Dynamic Programming The idea of dynamic programming is to have a table of solutions of subproblems
New Implementation for the Multi-sequence All-Against-All Substring Matching Problem
 New Implementation for the Multi-sequence All-Against-All Substring Matching Problem Oana Sandu Supervised by Ulrike Stege In collaboration with Chris Upton, Alex Thomo, and Marina Barsky University of
New Implementation for the Multi-sequence All-Against-All Substring Matching Problem Oana Sandu Supervised by Ulrike Stege In collaboration with Chris Upton, Alex Thomo, and Marina Barsky University of
String Patterns and Algorithms on Strings
 String Patterns and Algorithms on Strings Lecture delivered by: Venkatanatha Sarma Y Assistant Professor MSRSAS-Bangalore 11 Objectives To introduce the pattern matching problem and the important of algorithms
String Patterns and Algorithms on Strings Lecture delivered by: Venkatanatha Sarma Y Assistant Professor MSRSAS-Bangalore 11 Objectives To introduce the pattern matching problem and the important of algorithms
Lecture Overview. Sequence search & alignment. Searching sequence databases. Sequence Alignment & Search. Goals: Motivations:
 Lecture Overview Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating
Lecture Overview Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating
CMPSC 250 Analysis of Algorithms Spring 2018 Dr. Aravind Mohan Shortest Paths April 16, 2018
 1 CMPSC 250 Analysis of Algorithms Spring 2018 Dr. Aravind Mohan Shortest Paths April 16, 2018 Shortest Paths The discussion in these notes captures the essence of Dijkstra s algorithm discussed in textbook
1 CMPSC 250 Analysis of Algorithms Spring 2018 Dr. Aravind Mohan Shortest Paths April 16, 2018 Shortest Paths The discussion in these notes captures the essence of Dijkstra s algorithm discussed in textbook
Dynamic Programming in 3-D Progressive Alignment Profile Progressive Alignment (ClustalW) Scoring Multiple Alignments Entropy Sum of Pairs Alignment
 Dynamic Programming in 3-D Progressive Alignment Profile Progressive Alignment (ClustalW) Scoring Multiple Alignments Entropy Sum of Pairs Alignment Partial Order Alignment (POA) A-Bruijin (ABA) Approach
Dynamic Programming in 3-D Progressive Alignment Profile Progressive Alignment (ClustalW) Scoring Multiple Alignments Entropy Sum of Pairs Alignment Partial Order Alignment (POA) A-Bruijin (ABA) Approach
Sequence alignment algorithms
 Sequence alignment algorithms Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 23 rd 27 After this lecture, you can decide when to use local and global sequence alignments
Sequence alignment algorithms Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 23 rd 27 After this lecture, you can decide when to use local and global sequence alignments
17 dicembre Luca Bortolussi SUFFIX TREES. From exact to approximate string matching.
 17 dicembre 2003 Luca Bortolussi SUFFIX TREES From exact to approximate string matching. An introduction to string matching String matching is an important branch of algorithmica, and it has applications
17 dicembre 2003 Luca Bortolussi SUFFIX TREES From exact to approximate string matching. An introduction to string matching String matching is an important branch of algorithmica, and it has applications
Sequence Alignment & Search
 Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating the first version
Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating the first version
Reminder: sequence alignment in sub-quadratic time
 Reminder: sequence alignment in sub-quadratic time Last week: Sequence alignment in sub-quadratic time for unrestricted Scoring Schemes. ) utilize LZ78 parsing 2) utilize Total Monotonicity property of
Reminder: sequence alignment in sub-quadratic time Last week: Sequence alignment in sub-quadratic time for unrestricted Scoring Schemes. ) utilize LZ78 parsing 2) utilize Total Monotonicity property of
Project and Production Management Prof. Arun Kanda Department of Mechanical Engineering Indian Institute of Technology, Delhi
 Project and Production Management Prof. Arun Kanda Department of Mechanical Engineering Indian Institute of Technology, Delhi Lecture - 8 Consistency and Redundancy in Project networks In today s lecture
Project and Production Management Prof. Arun Kanda Department of Mechanical Engineering Indian Institute of Technology, Delhi Lecture - 8 Consistency and Redundancy in Project networks In today s lecture
Sequence Alignment (chapter 6) p The biological problem p Global alignment p Local alignment p Multiple alignment
 Sequence lignment (chapter 6) p The biological problem p lobal alignment p Local alignment p Multiple alignment Local alignment: rationale p Otherwise dissimilar proteins may have local regions of similarity
Sequence lignment (chapter 6) p The biological problem p lobal alignment p Local alignment p Multiple alignment Local alignment: rationale p Otherwise dissimilar proteins may have local regions of similarity
Bioinformatics for Biologists
 Bioinformatics for Biologists Sequence Analysis: Part I. Pairwise alignment and database searching Fran Lewitter, Ph.D. Director Bioinformatics & Research Computing Whitehead Institute Topics to Cover
Bioinformatics for Biologists Sequence Analysis: Part I. Pairwise alignment and database searching Fran Lewitter, Ph.D. Director Bioinformatics & Research Computing Whitehead Institute Topics to Cover
Biology 644: Bioinformatics
 A statistical Markov model in which the system being modeled is assumed to be a Markov process with unobserved (hidden) states in the training data. First used in speech and handwriting recognition In
A statistical Markov model in which the system being modeled is assumed to be a Markov process with unobserved (hidden) states in the training data. First used in speech and handwriting recognition In
CS 231: Algorithmic Problem Solving
 CS 231: Algorithmic Problem Solving Naomi Nishimura Module 5 Date of this version: June 14, 2018 WARNING: Drafts of slides are made available prior to lecture for your convenience. After lecture, slides
CS 231: Algorithmic Problem Solving Naomi Nishimura Module 5 Date of this version: June 14, 2018 WARNING: Drafts of slides are made available prior to lecture for your convenience. After lecture, slides
