EECS 4425: Introductory Computational Bioinformatics Fall Suprakash Datta
|
|
|
- Belinda Austin
- 5 years ago
- Views:
Transcription
1 EECS 4425: Introductory Computational Bioinformatics Fall 2018 Suprakash Datta datta [at] cse.yorku.ca Office: CSEB 3043 Phone: ext Course page: Many of the slides are taken from 9/18/2018 EECS 4425, Fall
2 Next: sequence alignment Why align? Picture from 9/18/2018 EECS 4425, Fall
3 Local vs Global Alignment Picture from Wikipedia (By Yz cs Own work, CC BY-SA 4.0, 9/18/2018 EECS 4425, Fall
4 Dynamic programming (DP) Typically used for optimization problems Often results in efficient algorithms Not applicable to all problems Caveats: Need not yield poly-time algorithms No unique formulations for most problems May not rule out greedy algorithms 9/18/2018 EECS 4425, Fall
5 Example Counting the number of shortest paths in a grid Counting the number of shortest paths in a grid with blocked intersections Finding paths in a weighted grid Sequence alignment 9/18/2018 EECS 4425, Fall
6 Setting up DP in practice The optimal solution should be computable as a (recursive) function of the solution to sub-problems Solve sub-problems systematically and store solutions (to avoid duplication of work). 9/18/2018 EECS 4425, Fall
7 Number of paths in a grid Combinatorial approach DP approach: how can we decompose the problem into sub-problems? 9/18/2018 EECS 4425, Fall
8 Number of paths in a grid with blocked intersections Combinatorial approach? DP approach: how can we decompose the problem into sub-problems? 9/18/2018 EECS 4425, Fall
9 Manhattan Tourist Problem (MTP) Imagine seeking a path (from source to sink) to travel (only eastward and southward) with the most number of attractions (*) in the Manhattan grid Source * * * * * * * * * * * * Sink 9/18/2018 EECS 4425, Fall
10 Manhattan Tourist Problem (MTP) Imagine seeking a path (from source to sink) to travel (only eastward and southward) with the most number of attractions (*) in the Manhattan grid Source * * * * * * * * * * * * Sink 9/18/2018 EECS 4425, Fall
11 Manhattan Tourist Problem: Formulation Goal: Find the best path in a weighted grid. Input: A weighted grid G with two distinct vertices, one labeled source and the other labeled sink Output: A best path in G from source to sink 9/18/2018 EECS 4425, Fall
12 MTP: An Example source j coordinate i coordinate sink 9/18/ EECS 4425, Fall
13 MTP: Greedy Algorithm Is Not Optimal source promising start, but leads to bad choices! sink 9/18/2018 EECS 4425, Fall
14 MTP: Recurrence Computing the score for a point (i,j) by the recurrence relation: s i, j = max s i-1, j + weight of the edge between (i-1, j) and (i, j) s i, j-1 + weight of the edge between (i, j-1) and (i, j) The running time is n x m for a n by m grid (n = # of rows, m = # of columns) 9/18/2018 EECS 4425, Fall
15 MTP: Simple Recursive Program MT(n,m) if n=0 or m=0 What s wrong with this return Line(n,m) approach? x MT(n-1,m)+ length of the edge from (n- 1,m) to (n,m) y MT(n,m-1)+ length of the edge from (n,m-1) to (n,m) return max{x,y} 9/18/2018 EECS 4425, Fall
16 Manhattan Is Not A Perfect Grid A 2 A 3 A 1 B What about diagonals? The score at point B is given by: s B = max of s A1 + weight of the edge (A 1, B) s A2 + weight of the edge (A 2, B) s A3 + weight of the edge (A 3, B) 9/18/2018 EECS 4425, Fall
17 Manhattan Is Not A Perfect Grid (cont d) Computing the score for point x is given by the recurrence relation: s x = max of s y + weight of vertex (y, x) where y є Predecessors(x) Predecessors (x) set of vertices that have edges leading to x The running time for a graph G(V, E) (V is the set of all vertices and E is the set of all edges) is O(E) since each edge is evaluated once 9/18/2018 EECS 4425, Fall
18 Traveling on the Grid The only hitch is that one must decide on the order in which visit the vertices By the time the vertex x is analyzed, the values s y for all its predecessors y should be computed otherwise we are in trouble. We need to traverse the vertices in some order Try to find such order for a directed cycle??? 9/18/2018 EECS 4425, Fall
19 DAG: Directed Acyclic Graph Since Manhattan is not a perfect regular grid, we represent it as a DAG DAG for Dressing in the morning problem 9/18/2018 EECS 4425, Fall
20 Topological Ordering A numbering of vertices of the graph is called topological ordering of the DAG if every edge of the DAG connects a vertex with a smaller label to a vertex with a larger label In other words, if vertices are positioned on a line in an increasing order of labels then all edges go from left to right. 9/18/2018 EECS 4425, Fall
21 Topological ordering 2 different topological orderings of the DAG 9/18/2018 EECS 4425, Fall
22 Longest Path in DAG Problem Goal: Find a longest path between two vertices in a weighted DAG Input: A weighted DAG G with source and sink vertices Output: A longest path in G from source to sink 9/18/2018 EECS 4425, Fall
23 Longest Path in DAG: Dynamic Programming Suppose vertex v has indegree 3 and predecessors {u 1, u 2, u 3 } Longest path to v from source is: su 1 + weight of edge from u 1 to v s v = max of su 2 + weight of edge from u 2 to v su 3 + weight of edge from u 3 to v In General: s v = max u (s u + weight of edge from u to v) 9/18/2018 EECS 4425, Fall
24 Traversing the Manhattan Grid 3 different strategies: a) Column by column b) Row by row c) Along diagonals c) a) b) 9/18/2018 EECS 4425, Fall
25 Alignment: 2 row representation Given 2 DNA sequences v and w: v : w : AT CT GAT T GCAT A m = 7 n = 6 Alignment : 2 * k matrix ( k > m, n ) letters of v letters of w A T -- G T T A T -- A T C G T -- A -- C 4 matches 2 insertions 2 deletions 9/18/2018 EECS 4425, Fall
26 Aligning DNA Sequences V = ATCTGATG W = TGCATAC match n = 8 m = 7 mismatch matches mismatches insertions deletions V W A T C T G A T G T G C A T A C indels deletion insertion 9/18/2018 EECS 4425, Fall
27 Aligning DNA Sequences - 2 Brute force is infeasible. Number of alignments of X[1..n],Y[1..m], n<m is ( m+n) n For m=n, this is about 2 2n /πn 9/18/2018 EECS 4425, Fall
28 Longest Common Subsequence (LCS) Alignment without Mismatches Given two sequences v = v 1 v 2 v m and w = w 1 w 2 w n The LCS of v and w is a sequence of positions in v: 1 < i 1 < i 2 < < i t < m and a sequence of positions in w: 1 < j 1 < j 2 < < j t < n such that i t -th letter of v equals to j t -letter of w and t is maximal 9/18/2018 EECS 4425, Fall
29 LCS: Example i coords: 0 elements of v elements of w j coords: A T -- C -- T G A T C -- T G C A T -- A -- C (0,0) (1,0) (2,1) (2,2) (3,3) (3,4) (4,5) (5,5) (6,6) (7,6) (8,7) positions in v: 2 < 3 < 4 < 6 < 8 Matches shown in red positions in w: 1 < 3 < 5 < 6 < 7 Every common subsequence is a path in 2-D grid 9/18/2018 EECS 4425, Fall
30 LCS Problem as Manhattan Tourist Problem i T 0 1 j A T C T G A T C G C A T 5 A C 6 7 9/18/2018 EECS 4425, Fall
31 i T 0 1 j Edit Graph for LCS Problem A T C T G A T C G C A T 5 A C 6 7 9/18/2018 EECS 4425, Fall
32 Edit Graph for LCS Problem i T G C A T A C j A T C T G A T C Every path is a common subsequence. Every diagonal edge adds an extra element to common subsequence LCS Problem: Find a path with maximum number of diagonal edges 9/18/2018 EECS 4425, Fall
33 Computing LCS Let v i = prefix of v of length i: v 1 v i and w j = prefix of w of length j: w 1 w j The length of LCS(v i,w j ) is computed by: s i, j = max s i-1, j s i, j-1 s i-1, j if v i = w j 9/18/2018 EECS 4425, Fall
34 Computing LCS (cont d) i-1,j -1 i-1,j s i,j = MAX 1 0 s i-1,j s i,j i,j -1 i,j s i-1,j , if v i = w j 9/18/2018 EECS 4425, Fall
35 Every Path in the Grid Corresponds to an Alignment W A T C G V A T G T V = A T - G T W= A T C G /18/2018 EECS 4425, Fall
36 Aligning Sequences without Insertions and Deletions: Hamming Distance Given two DNA sequences v and w : v : w : AT AT AT AT T AT AT AT A The Hamming distance: d H (v, w) = 8 is large but the sequences are very similar 9/18/2018 EECS 4425, Fall
37 Aligning Sequences with Insertions and Deletions By shifting one sequence over one position: v : AT AT AT AT -- w : -- T AT AT AT A The edit distance: d H (v, w) = 2. Hamming distance neglects insertions and deletions in DNA 9/18/2018 EECS 4425, Fall
38 Edit Distance Levenshtein (1966) introduced edit distance between two strings as the minimum number of elementary operations (insertions, deletions, and substitutions) to transform one string into the other d(v,w) = MIN number of elementary operations to transform v w 9/18/2018 EECS 4425, Fall
39 Edit Distance vs Hamming Distance Hamming distance always compares i -th letter of v with i -th letter of w V = ATATATAT W = TATATATA Hamming distance: d(v, w)=8 Computing Hamming distance is a trivial task. 9/18/2018 EECS 4425, Fall
40 Edit Distance vs Hamming Distance Hamming distance always compares i -th letter of v with i -th letter of w V = ATATATAT W = TATATATA Hamming distance: d(v, w)=8 Just one shift Make it all line up Computing Hamming distance is a trivial task Edit distance may compare i -th letter of v with j -th letter of w V = - ATATATAT W = TATATATA Edit distance: d(v, w)=2 Computing edit distance is a non-trivial task 9/18/2018 EECS 4425, Fall
41 Edit Distance vs Hamming Distance Hamming distance always compares i -th letter of v with i -th letter of w V = ATATATAT W = TATATATA Hamming distance: d(v, w)=8 Edit distance may compare i -th letter of v with j -th letter of w V = - ATATATAT W = TATATATA Edit distance: d(v, w)=2 (one insertion and one deletion) How to find what j goes with what i??? 9/18/2018 EECS 4425, Fall
42 Edit Distance: Example TGCATAT ATCCGAT in 5 steps TGCATAT (delete last T) TGCATA (delete last A) TGCAT (insert A at front) ATGCAT (substitute C for 3 rd G) ATCCAT (insert G before last A) ATCCGAT (Done) 9/18/2018 EECS 4425, Fall
43 Edit Distance: Example TGCATAT ATCCGAT in 5 steps TGCATAT (delete last T) TGCATA (delete last A) TGCAT (insert A at front) ATGCAT (substitute C for 3 rd G) ATCCAT (insert G before last A) ATCCGAT (Done) What is the edit distance? 5? 9/18/2018 EECS 4425, Fall
44 Edit Distance: Example (cont d) TGCATAT ATCCGAT in 4 steps TGCATAT (insert A at front) ATGCATAT (delete 6 th T) ATGCATA (substitute G for 5 th A) ATGCGTA (substitute C for 3 rd G) ATCCGAT (Done) 9/18/2018 EECS 4425, Fall
45 Edit Distance: Example (cont d) TGCATAT ATCCGAT in 4 steps TGCATAT (insert A at front) ATGCATAT (delete 6 th T) ATGCATA (substitute G for 5 th A) ATGCGTA (substitute C for 3 rd G) ATCCGAT (Done) Can it be done in 3 steps??? 9/18/2018 EECS 4425, Fall
46 The Alignment Grid Every alignment path is from source to sink 9/18/2018 EECS 4425, Fall
47 Alignment as a Path in the Edit Graph A T _ G T T A T _ A T C G T _ A _ C Corresponding path - (0,0), (1,1), (2,2), (2,3), (3,4), (4,5), (5,5), (6,6), (7,6), (7,7) 9/18/2018 EECS 4425, Fall
48 Alignments in Edit Graph (cont d) and represent indels in v and w with score 0. represent matches with score 1. The score of the alignment path is 5. 9/18/2018 EECS 4425, Fall
49 Alignment as a Path in the Edit Graph Every path in the edit graph corresponds to an alignment: 9/18/2018 EECS 4425, Fall
50 Alignment as a Path in the Edit Graph Old Alignment v= AT_GTTAT_ w= ATCGT_A_C New Alignment v= AT_GTTAT_ w= ATCG_TA_C /18/2018 EECS 4425, Fall
51 Alignment as a Path in the Edit Graph v= AT_GTTAT_ w= ATCGT_A_C (0,0), (1,1), (2,2), (2,3), (3,4), (4,5), (5,5), (6,6), (7,6), (7,7) 9/18/2018 EECS 4425, Fall
52 Alignment: Dynamic Programming s i,j = s i-1, j-1 +1 if v i = w j max s i-1, j s i, j-1 9/18/2018 EECS 4425, Fall
53 Dynamic Programming Example Initialize 1 st row and 1 st column to be all zeroes. Or, to be more precise, initialize 0 th row and 0 th column to be all zeroes. 9/18/2018 EECS 4425, Fall
54 Dynamic Programming Example S i,j = S i-1, j-1 max S i-1, j S i, j-1 value from NW +1, if v i = w j value from North (top) value from West (left) 9/18/2018 EECS 4425, Fall
55 Alignment: tracing paths Arrows show where the score originated from. if from the top if from the left if v i = w j 9/18/2018 EECS 4425, Fall
56 Path tracing example Find a match in row and column 2. i=2, j=2,5 is a match (T). j=2, i=4,5,7 is a match (T). Since v i = w j, s i,j = s i-1,j-1 +1 s 2,2 = [s 1,1 = 1] + 1 s 2,5 = [s 1,4 = 1] + 1 s 4,2 = [s 3,1 = 1] + 1 s 5,2 = [s 4,1 = 1] + 1 s 7,2 = [s 6,1 = 1] + 1 9/18/2018 EECS 4425, Fall
57 Path tracing example Continuing with the dynamic programming algorithm gives this result. 9/18/2018 EECS 4425, Fall
58 Alignment: Dynamic Programming s i,j = s i-1, j-1 +1 if v i = w j max s i-1, j s i, j-1 9/18/2018 EECS 4425, Fall
59 Alignment: Dynamic Programming s i,j = s i-1, j-1 +1 if v i = w j max s i-1, j +0 s i, j-1 +0 This recurrence corresponds to the Manhattan Tourist problem (three incoming edges into a vertex) with all horizontal and vertical edges weighted by zero. 9/18/2018 EECS 4425, Fall
60 1. LCS(v,w) 2. for i 1 to n 3. s i, for j 1 to m 5. s 0,j 0 6. for i 1 to n 7. for j 1 to m LCS Algorithm 8. s i-1,j 9. s i,j max s i,j s i-1,j-1 + 1, if v i = w j 11. if s i,j = s i-1,j b i,j if s i,j = s i,j-1 if s i,j = s i-1,j return (s n,m, b) 9/18/2018 EECS 4425, Fall
61 Now What? LCS(v,w) created the alignment grid Now we need a way to read the best alignment of v and w Follow the arrows backwards from sink 9/18/2018 EECS 4425, Fall
62 Printing LCS: Backtracking 1. PrintLCS(b,v,i,j) 2. if i = 0 or j = 0 3. return 4. if b i,j = 5. PrintLCS(b,v,i-1,j-1) 6. print v i 7. else 8. if b i,j = 9. PrintLCS(b,v,i-1,j) 10. else 11. PrintLCS(b,v,i,j-1) 9/18/2018 EECS 4425, Fall
63 LCS Runtime It takes O(nm) time to fill in the nxm dynamic programming matrix. Why O(nm)? The pseudocode consists of a nested for loop inside of another for loop to set up a nxm matrix. 9/18/2018 EECS 4425, Fall
64 Why does DP work? Avoids re-computing the same subproblems Limits the amount of work done in each step 9/18/2018 EECS 4425, Fall
65 When is DP applicable? Optimal substructure: Optimal solution to problem (instance) contains optimal solutions to sub-problems Overlapping sub-problems: Limited number of distinct sub-problems, repeated many many times 9/18/2018 EECS 4425, Fall
66 Next: Sequence Alignment Global Alignment Scoring Matrices Local Alignment Alignment with Affine Gap Penalties 9/18/2018 EECS 4425, Fall
67 From LCS to Alignment The Longest Common Subsequence (LCS) problem the simplest form of sequence alignment allows only insertions and deletions (no mismatches). In the LCS Problem, we scored 1 for matches and 0 for indels Consider penalizing indels and mismatches with negative scores Simplest scoring schema: +1 : match premium -μ : mismatch penalty -σ : indel penalty 9/18/2018 EECS 4425, Fall
68 Simple Scoring When mismatches are penalized by μ, indels are penalized by σ, and matches are rewarded with +1, the resulting score is: #matches μ(#mismatches) σ (#indels) 9/18/2018 EECS 4425, Fall
69 The Global Alignment Problem Find the best alignment between two strings under a given scoring schema Input : Strings v and w and a scoring schema Output : Alignment of maximum score = -б = 1 if match = -µ if mismatch s i-1,j-1 +1 if v i = w j s i,j = max s i-1,j-1 -µ if v i w j s i-1,j - σ s i,j-1 - σ µ : mismatch penalty σ : indel penalty 9/18/2018 EECS 4425, Fall
70 Scoring Matrices To generalize scoring, consider a (4+1) x(4+1) scoring matrix δ. In the case of an amino acid sequence alignment, the scoring matrix would be a (20+1)x(20+1) size. The addition of 1 is to include the score for comparison of a gap character -. This will simplify the algorithm as follows: s i-1,j-1 + δ (v i, w j ) s i,j = max s i-1,j + δ (v i, -) s i,j-1 + δ (-, w j ) 9/18/2018 EECS 4425, Fall
71 Measuring Similarity Measuring the extent of similarity between two sequences Based on percent sequence identity Based on conservation 9/18/2018 EECS 4425, Fall
72 Percent Sequence Identity The extent to which two nucleotide or amino acid sequences are invariant A C C T G A G A G A C G T G G C A G mismatch 70% identical indel 9/18/2018 EECS 4425, Fall
73 Making a Scoring Matrix Scoring matrices are created based on biological evidence. Alignments can be thought of as two sequences that differ due to mutations. Some of these mutations have little effect on the protein s function, therefore some penalties, δ(v i, w j ), will be less harsh than others. 9/18/2018 EECS 4425, Fall
74 Scoring Matrix: Example A R N K A R N K Notice that although R and K are different amino acids, they have a positive score. Why? They are both positively charged amino acids will not greatly change function of protein. 9/18/2018 EECS 4425, Fall
75 Conservation Amino acid changes that tend to preserve the physico-chemical properties of the original residue Polar to polar aspartate glutamate Nonpolar to nonpolar alanine valine Similarly behaving residues leucine to isoleucine 9/18/2018 EECS 4425, Fall
76 Scoring matrices Amino acid substitution matrices PAM BLOSUM DNA substitution matrices DNA is less conserved than protein sequences Less effective to compare coding regions at nucleotide level 9/18/2018 EECS 4425, Fall
Dynamic Programming: Sequence alignment. CS 466 Saurabh Sinha
 Dynamic Programming: Sequence alignment CS 466 Saurabh Sinha DNA Sequence Comparison: First Success Story Finding sequence similarities with genes of known function is a common approach to infer a newly
Dynamic Programming: Sequence alignment CS 466 Saurabh Sinha DNA Sequence Comparison: First Success Story Finding sequence similarities with genes of known function is a common approach to infer a newly
Finding sequence similarities with genes of known function is a common approach to infer a newly sequenced gene s function
 Outline DNA Sequence Comparison: First Success Stories Change Problem Manhattan Tourist Problem Longest Paths in Graphs Sequence Alignment Edit Distance Longest Common Subsequence Problem Dot Matrices
Outline DNA Sequence Comparison: First Success Stories Change Problem Manhattan Tourist Problem Longest Paths in Graphs Sequence Alignment Edit Distance Longest Common Subsequence Problem Dot Matrices
Dynamic Programming Part I: Examples. Bioinfo I (Institut Pasteur de Montevideo) Dynamic Programming -class4- July 25th, / 77
 Dynamic Programming Part I: Examples Bioinfo I (Institut Pasteur de Montevideo) Dynamic Programming -class4- July 25th, 2011 1 / 77 Dynamic Programming Recall: the Change Problem Other problems: Manhattan
Dynamic Programming Part I: Examples Bioinfo I (Institut Pasteur de Montevideo) Dynamic Programming -class4- July 25th, 2011 1 / 77 Dynamic Programming Recall: the Change Problem Other problems: Manhattan
Global Alignment Scoring Matrices Local Alignment Alignment with Affine Gap Penalties
 Global Alignment Scoring Matrices Local Alignment Alignment with Affine Gap Penalties From LCS to Alignment: Change the Scoring The Longest Common Subsequence (LCS) problem the simplest form of sequence
Global Alignment Scoring Matrices Local Alignment Alignment with Affine Gap Penalties From LCS to Alignment: Change the Scoring The Longest Common Subsequence (LCS) problem the simplest form of sequence
1. Basic idea: Use smaller instances of the problem to find the solution of larger instances
 Chapter 8. Dynamic Programming CmSc Intro to Algorithms. Basic idea: Use smaller instances of the problem to find the solution of larger instances Example : Fibonacci numbers F = F = F n = F n- + F n-
Chapter 8. Dynamic Programming CmSc Intro to Algorithms. Basic idea: Use smaller instances of the problem to find the solution of larger instances Example : Fibonacci numbers F = F = F n = F n- + F n-
Pairwise Sequence alignment Basic Algorithms
 Pairwise Sequence alignment Basic Algorithms Agenda - Previous Lesson: Minhala - + Biological Story on Biomolecular Sequences - + General Overview of Problems in Computational Biology - Reminder: Dynamic
Pairwise Sequence alignment Basic Algorithms Agenda - Previous Lesson: Minhala - + Biological Story on Biomolecular Sequences - + General Overview of Problems in Computational Biology - Reminder: Dynamic
Dynamic Programming Comp 122, Fall 2004
 Dynamic Programming Comp 122, Fall 2004 Review: the previous lecture Principles of dynamic programming: optimization problems, optimal substructure property, overlapping subproblems, trade space for time,
Dynamic Programming Comp 122, Fall 2004 Review: the previous lecture Principles of dynamic programming: optimization problems, optimal substructure property, overlapping subproblems, trade space for time,
9/29/09 Comp /Comp Fall
 9/29/9 Comp 9-9/Comp 79-9 Fall 29 1 So far we ve tried: A greedy algorithm that does not work for all inputs (it is incorrect) An exhaustive search algorithm that is correct, but can take a long time A
9/29/9 Comp 9-9/Comp 79-9 Fall 29 1 So far we ve tried: A greedy algorithm that does not work for all inputs (it is incorrect) An exhaustive search algorithm that is correct, but can take a long time A
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 04: Variations of sequence alignments http://www.pitt.edu/~mcs2/teaching/biocomp/tutorials/global.html Slides adapted from Dr. Shaojie Zhang (University
EECS730: Introduction to Bioinformatics Lecture 04: Variations of sequence alignments http://www.pitt.edu/~mcs2/teaching/biocomp/tutorials/global.html Slides adapted from Dr. Shaojie Zhang (University
Dynamic Programming: Edit Distance
 Dynamic Programming: Edit Distance Outline. DNA Sequence Comparison and CF. Change Problem. Manhattan Tourist Problem. Longest Paths in Graphs. Sequence Alignment 6. Edit Distance Section : DNA Sequence
Dynamic Programming: Edit Distance Outline. DNA Sequence Comparison and CF. Change Problem. Manhattan Tourist Problem. Longest Paths in Graphs. Sequence Alignment 6. Edit Distance Section : DNA Sequence
3/5/2018 Lecture15. Comparing Sequences. By Miguel Andrade at English Wikipedia.
 3/5/2018 Lecture15 Comparing Sequences By Miguel Andrade at English Wikipedia 1 http://localhost:8888/notebooks/comp555s18/lecture15.ipynb# 1/1 Sequence Similarity A common problem in Biology Insulin Protein
3/5/2018 Lecture15 Comparing Sequences By Miguel Andrade at English Wikipedia 1 http://localhost:8888/notebooks/comp555s18/lecture15.ipynb# 1/1 Sequence Similarity A common problem in Biology Insulin Protein
DynamicProgramming. September 17, 2018
 DynamicProgramming September 17, 2018 1 Lecture 11: Dynamic Programming CBIO (CSCI) 4835/6835: Introduction to Computational Biology 1.1 Overview and Objectives We ve so far discussed sequence alignment
DynamicProgramming September 17, 2018 1 Lecture 11: Dynamic Programming CBIO (CSCI) 4835/6835: Introduction to Computational Biology 1.1 Overview and Objectives We ve so far discussed sequence alignment
Sequence Alignment: Mo1va1on and Algorithms. Lecture 2: August 23, 2012
 Sequence Alignment: Mo1va1on and Algorithms Lecture 2: August 23, 2012 Mo1va1on and Introduc1on Importance of Sequence Alignment For DNA, RNA and amino acid sequences, high sequence similarity usually
Sequence Alignment: Mo1va1on and Algorithms Lecture 2: August 23, 2012 Mo1va1on and Introduc1on Importance of Sequence Alignment For DNA, RNA and amino acid sequences, high sequence similarity usually
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 06: Multiple Sequence Alignment https://upload.wikimedia.org/wikipedia/commons/thumb/7/79/rplp0_90_clustalw_aln.gif/575px-rplp0_90_clustalw_aln.gif Slides
EECS730: Introduction to Bioinformatics Lecture 06: Multiple Sequence Alignment https://upload.wikimedia.org/wikipedia/commons/thumb/7/79/rplp0_90_clustalw_aln.gif/575px-rplp0_90_clustalw_aln.gif Slides
6.1 The Power of DNA Sequence Comparison
 6 Dynamic Programming Algorithms We introduced dynamic programming in chapter 2 with the Rocks problem. While the Rocks problem does not appear to be related to bioinformatics, the algorithm that we described
6 Dynamic Programming Algorithms We introduced dynamic programming in chapter 2 with the Rocks problem. While the Rocks problem does not appear to be related to bioinformatics, the algorithm that we described
Computational Genomics and Molecular Biology, Fall
 Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
DNA Alignment With Affine Gap Penalties
 DNA Alignment With Affine Gap Penalties Laurel Schuster Why Use Affine Gap Penalties? When aligning two DNA sequences, one goal may be to infer the mutations that made them different. Though it s impossible
DNA Alignment With Affine Gap Penalties Laurel Schuster Why Use Affine Gap Penalties? When aligning two DNA sequences, one goal may be to infer the mutations that made them different. Though it s impossible
Sequence Alignment: Mo1va1on and Algorithms
 Sequence Alignment: Mo1va1on and Algorithms Mo1va1on and Introduc1on Importance of Sequence Alignment For DNA, RNA and amino acid sequences, high sequence similarity usually implies significant func1onal
Sequence Alignment: Mo1va1on and Algorithms Mo1va1on and Introduc1on Importance of Sequence Alignment For DNA, RNA and amino acid sequences, high sequence similarity usually implies significant func1onal
Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD
 Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD Assumptions: Biological sequences evolved by evolution. Micro scale changes: For short sequences (e.g. one
Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD Assumptions: Biological sequences evolved by evolution. Micro scale changes: For short sequences (e.g. one
Computational Molecular Biology
 Computational Molecular Biology Erwin M. Bakker Lecture 2 Materials used from R. Shamir [2] and H.J. Hoogeboom [4]. 1 Molecular Biology Sequences DNA A, T, C, G RNA A, U, C, G Protein A, R, D, N, C E,
Computational Molecular Biology Erwin M. Bakker Lecture 2 Materials used from R. Shamir [2] and H.J. Hoogeboom [4]. 1 Molecular Biology Sequences DNA A, T, C, G RNA A, U, C, G Protein A, R, D, N, C E,
Lecture 10: Local Alignments
 Lecture 10: Local Alignments Study Chapter 6.8-6.10 1 Outline Edit Distances Longest Common Subsequence Global Sequence Alignment Scoring Matrices Local Sequence Alignment Alignment with Affine Gap Penalties
Lecture 10: Local Alignments Study Chapter 6.8-6.10 1 Outline Edit Distances Longest Common Subsequence Global Sequence Alignment Scoring Matrices Local Sequence Alignment Alignment with Affine Gap Penalties
Today s Lecture. Multiple sequence alignment. Improved scoring of pairwise alignments. Affine gap penalties Profiles
 Today s Lecture Multiple sequence alignment Improved scoring of pairwise alignments Affine gap penalties Profiles 1 The Edit Graph for a Pair of Sequences G A C G T T G A A T G A C C C A C A T G A C G
Today s Lecture Multiple sequence alignment Improved scoring of pairwise alignments Affine gap penalties Profiles 1 The Edit Graph for a Pair of Sequences G A C G T T G A A T G A C C C A C A T G A C G
Divide and Conquer. Bioinformatics: Issues and Algorithms. CSE Fall 2007 Lecture 12
 Divide and Conquer Bioinformatics: Issues and Algorithms CSE 308-408 Fall 007 Lecture 1 Lopresti Fall 007 Lecture 1-1 - Outline MergeSort Finding mid-point in alignment matrix in linear space Linear space
Divide and Conquer Bioinformatics: Issues and Algorithms CSE 308-408 Fall 007 Lecture 1 Lopresti Fall 007 Lecture 1-1 - Outline MergeSort Finding mid-point in alignment matrix in linear space Linear space
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014
 Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
Biology 644: Bioinformatics
 Find the best alignment between 2 sequences with lengths n and m, respectively Best alignment is very dependent upon the substitution matrix and gap penalties The Global Alignment Problem tries to find
Find the best alignment between 2 sequences with lengths n and m, respectively Best alignment is very dependent upon the substitution matrix and gap penalties The Global Alignment Problem tries to find
Lectures 12 and 13 Dynamic programming: weighted interval scheduling
 Lectures 12 and 13 Dynamic programming: weighted interval scheduling COMP 523: Advanced Algorithmic Techniques Lecturer: Dariusz Kowalski Lectures 12-13: Dynamic Programming 1 Overview Last week: Graph
Lectures 12 and 13 Dynamic programming: weighted interval scheduling COMP 523: Advanced Algorithmic Techniques Lecturer: Dariusz Kowalski Lectures 12-13: Dynamic Programming 1 Overview Last week: Graph
Lecture 3: February Local Alignment: The Smith-Waterman Algorithm
 CSCI1820: Sequence Alignment Spring 2017 Lecture 3: February 7 Lecturer: Sorin Istrail Scribe: Pranavan Chanthrakumar Note: LaTeX template courtesy of UC Berkeley EECS dept. Notes are also adapted from
CSCI1820: Sequence Alignment Spring 2017 Lecture 3: February 7 Lecturer: Sorin Istrail Scribe: Pranavan Chanthrakumar Note: LaTeX template courtesy of UC Berkeley EECS dept. Notes are also adapted from
BLAST - Basic Local Alignment Search Tool
 Lecture for ic Bioinformatics (DD2450) April 11, 2013 Searching 1. Input: Query Sequence 2. Database of sequences 3. Subject Sequence(s) 4. Output: High Segment Pairs (HSPs) Sequence Similarity Measures:
Lecture for ic Bioinformatics (DD2450) April 11, 2013 Searching 1. Input: Query Sequence 2. Database of sequences 3. Subject Sequence(s) 4. Output: High Segment Pairs (HSPs) Sequence Similarity Measures:
IN101: Algorithmic techniques Vladimir-Alexandru Paun ENSTA ParisTech
 IN101: Algorithmic techniques Vladimir-Alexandru Paun ENSTA ParisTech License CC BY-NC-SA 2.0 http://creativecommons.org/licenses/by-nc-sa/2.0/fr/ Outline Previously on IN101 Python s anatomy Functions,
IN101: Algorithmic techniques Vladimir-Alexandru Paun ENSTA ParisTech License CC BY-NC-SA 2.0 http://creativecommons.org/licenses/by-nc-sa/2.0/fr/ Outline Previously on IN101 Python s anatomy Functions,
Divide & Conquer Algorithms
 Divide & Conquer Algorithms Outline 1. MergeSort 2. Finding the middle vertex 3. Linear space sequence alignment 4. Block alignment 5. Four-Russians speedup 6. LCS in sub-quadratic time Section 1: MergeSort
Divide & Conquer Algorithms Outline 1. MergeSort 2. Finding the middle vertex 3. Linear space sequence alignment 4. Block alignment 5. Four-Russians speedup 6. LCS in sub-quadratic time Section 1: MergeSort
Divide & Conquer Algorithms
 Divide & Conquer Algorithms Outline MergeSort Finding the middle point in the alignment matrix in linear space Linear space sequence alignment Block Alignment Four-Russians speedup Constructing LCS in
Divide & Conquer Algorithms Outline MergeSort Finding the middle point in the alignment matrix in linear space Linear space sequence alignment Block Alignment Four-Russians speedup Constructing LCS in
Principles of Bioinformatics. BIO540/STA569/CSI660 Fall 2010
 Principles of Bioinformatics BIO540/STA569/CSI660 Fall 2010 Lecture 11 Multiple Sequence Alignment I Administrivia Administrivia The midterm examination will be Monday, October 18 th, in class. Closed
Principles of Bioinformatics BIO540/STA569/CSI660 Fall 2010 Lecture 11 Multiple Sequence Alignment I Administrivia Administrivia The midterm examination will be Monday, October 18 th, in class. Closed
.. Fall 2011 CSC 570: Bioinformatics Alexander Dekhtyar..
 .. Fall 2011 CSC 570: Bioinformatics Alexander Dekhtyar.. PAM and BLOSUM Matrices Prepared by: Jason Banich and Chris Hoover Background As DNA sequences change and evolve, certain amino acids are more
.. Fall 2011 CSC 570: Bioinformatics Alexander Dekhtyar.. PAM and BLOSUM Matrices Prepared by: Jason Banich and Chris Hoover Background As DNA sequences change and evolve, certain amino acids are more
Pairwise Sequence Alignment: Dynamic Programming Algorithms. COMP Spring 2015 Luay Nakhleh, Rice University
 Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 - Spring 2015 Luay Nakhleh, Rice University DP Algorithms for Pairwise Alignment The number of all possible pairwise alignments (if
Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 - Spring 2015 Luay Nakhleh, Rice University DP Algorithms for Pairwise Alignment The number of all possible pairwise alignments (if
Memoization/Dynamic Programming. The String reconstruction problem. CS124 Lecture 11 Spring 2018
 CS124 Lecture 11 Spring 2018 Memoization/Dynamic Programming Today s lecture discusses memoization, which is a method for speeding up algorithms based on recursion, by using additional memory to remember
CS124 Lecture 11 Spring 2018 Memoization/Dynamic Programming Today s lecture discusses memoization, which is a method for speeding up algorithms based on recursion, by using additional memory to remember
Lecture 12: Divide and Conquer Algorithms
 Lecture 12: Divide and Conquer Algorithms Study Chapter 7.1 7.4 1 Divide and Conquer Algorithms Divide problem into sub-problems Conquer by solving sub-problems recursively. If the sub-problems are small
Lecture 12: Divide and Conquer Algorithms Study Chapter 7.1 7.4 1 Divide and Conquer Algorithms Divide problem into sub-problems Conquer by solving sub-problems recursively. If the sub-problems are small
Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it.
 Sequence Alignments Overview Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it. Sequence alignment means arranging
Sequence Alignments Overview Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it. Sequence alignment means arranging
As of August 15, 2008, GenBank contained bases from reported sequences. The search procedure should be
 48 Bioinformatics I, WS 09-10, S. Henz (script by D. Huson) November 26, 2009 4 BLAST and BLAT Outline of the chapter: 1. Heuristics for the pairwise local alignment of two sequences 2. BLAST: search and
48 Bioinformatics I, WS 09-10, S. Henz (script by D. Huson) November 26, 2009 4 BLAST and BLAT Outline of the chapter: 1. Heuristics for the pairwise local alignment of two sequences 2. BLAST: search and
Special course in Computer Science: Advanced Text Algorithms
 Special course in Computer Science: Advanced Text Algorithms Lecture 6: Alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Special course in Computer Science: Advanced Text Algorithms Lecture 6: Alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Dynamic Programming & Smith-Waterman algorithm
 m m Seminar: Classical Papers in Bioinformatics May 3rd, 2010 m m 1 2 3 m m Introduction m Definition is a method of solving problems by breaking them down into simpler steps problem need to contain overlapping
m m Seminar: Classical Papers in Bioinformatics May 3rd, 2010 m m 1 2 3 m m Introduction m Definition is a method of solving problems by breaking them down into simpler steps problem need to contain overlapping
Lecture Overview. Sequence search & alignment. Searching sequence databases. Sequence Alignment & Search. Goals: Motivations:
 Lecture Overview Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating
Lecture Overview Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating
Lectures by Volker Heun, Daniel Huson and Knut Reinert, in particular last years lectures
 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 4.1 Sources for this lecture Lectures by Volker Heun, Daniel Huson and Knut
4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 4.1 Sources for this lecture Lectures by Volker Heun, Daniel Huson and Knut
Presentation for use with the textbook, Algorithm Design and Applications, by M. T. Goodrich and R. Tamassia, Wiley, Dynamic Programming
 Presentation for use with the textbook, Algorithm Design and Applications, by M. T. Goodrich and R. Tamassia, Wiley, 25 Dynamic Programming Terrible Fibonacci Computation Fibonacci sequence: f = f(n) 2
Presentation for use with the textbook, Algorithm Design and Applications, by M. T. Goodrich and R. Tamassia, Wiley, 25 Dynamic Programming Terrible Fibonacci Computation Fibonacci sequence: f = f(n) 2
Lecture 10. Sequence alignments
 Lecture 10 Sequence alignments Alignment algorithms: Overview Given a scoring system, we need to have an algorithm for finding an optimal alignment for a pair of sequences. We want to maximize the score
Lecture 10 Sequence alignments Alignment algorithms: Overview Given a scoring system, we need to have an algorithm for finding an optimal alignment for a pair of sequences. We want to maximize the score
Sequence Alignment & Search
 Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating the first version
Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating the first version
CMPS 2200 Fall Dynamic Programming. Carola Wenk. Slides courtesy of Charles Leiserson with changes and additions by Carola Wenk
 CMPS 00 Fall 04 Dynamic Programming Carola Wenk Slides courtesy of Charles Leiserson with changes and additions by Carola Wenk 9/30/4 CMPS 00 Intro. to Algorithms Dynamic programming Algorithm design technique
CMPS 00 Fall 04 Dynamic Programming Carola Wenk Slides courtesy of Charles Leiserson with changes and additions by Carola Wenk 9/30/4 CMPS 00 Intro. to Algorithms Dynamic programming Algorithm design technique
BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio CS 466 Saurabh Sinha
 BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio. 1990. CS 466 Saurabh Sinha Motivation Sequence homology to a known protein suggest function of newly sequenced protein Bioinformatics
BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio. 1990. CS 466 Saurabh Sinha Motivation Sequence homology to a known protein suggest function of newly sequenced protein Bioinformatics
Dynamic Programming II
 June 9, 214 DP: Longest common subsequence biologists often need to find out how similar are 2 DNA sequences DNA sequences are strings of bases: A, C, T and G how to define similarity? DP: Longest common
June 9, 214 DP: Longest common subsequence biologists often need to find out how similar are 2 DNA sequences DNA sequences are strings of bases: A, C, T and G how to define similarity? DP: Longest common
Notes on Dynamic-Programming Sequence Alignment
 Notes on Dynamic-Programming Sequence Alignment Introduction. Following its introduction by Needleman and Wunsch (1970), dynamic programming has become the method of choice for rigorous alignment of DNA
Notes on Dynamic-Programming Sequence Alignment Introduction. Following its introduction by Needleman and Wunsch (1970), dynamic programming has become the method of choice for rigorous alignment of DNA
FastA & the chaining problem
 FastA & the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 1 Sources for this lecture: Lectures by Volker Heun, Daniel Huson and Knut Reinert,
FastA & the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 1 Sources for this lecture: Lectures by Volker Heun, Daniel Huson and Knut Reinert,
5.1 The String reconstruction problem
 CS125 Lecture 5 Fall 2014 5.1 The String reconstruction problem The greedy approach doesn t always work, as we have seen. It lacks flexibility; if at some point, it makes a wrong choice, it becomes stuck.
CS125 Lecture 5 Fall 2014 5.1 The String reconstruction problem The greedy approach doesn t always work, as we have seen. It lacks flexibility; if at some point, it makes a wrong choice, it becomes stuck.
CSE 417 Dynamic Programming (pt 5) Multiple Inputs
 CSE 417 Dynamic Programming (pt 5) Multiple Inputs Reminders > HW5 due Wednesday Dynamic Programming Review > Apply the steps... optimal substructure: (small) set of solutions, constructed from solutions
CSE 417 Dynamic Programming (pt 5) Multiple Inputs Reminders > HW5 due Wednesday Dynamic Programming Review > Apply the steps... optimal substructure: (small) set of solutions, constructed from solutions
FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:
 FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:56 4001 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem
FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:56 4001 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem
CS 170 DISCUSSION 8 DYNAMIC PROGRAMMING. Raymond Chan raychan3.github.io/cs170/fa17.html UC Berkeley Fall 17
 CS 170 DISCUSSION 8 DYNAMIC PROGRAMMING Raymond Chan raychan3.github.io/cs170/fa17.html UC Berkeley Fall 17 DYNAMIC PROGRAMMING Recursive problems uses the subproblem(s) solve the current one. Dynamic
CS 170 DISCUSSION 8 DYNAMIC PROGRAMMING Raymond Chan raychan3.github.io/cs170/fa17.html UC Berkeley Fall 17 DYNAMIC PROGRAMMING Recursive problems uses the subproblem(s) solve the current one. Dynamic
Dynamic Programming Course: A structure based flexible search method for motifs in RNA. By: Veksler, I., Ziv-Ukelson, M., Barash, D.
 Dynamic Programming Course: A structure based flexible search method for motifs in RNA By: Veksler, I., Ziv-Ukelson, M., Barash, D., Kedem, K Outline Background Motivation RNA s structure representations
Dynamic Programming Course: A structure based flexible search method for motifs in RNA By: Veksler, I., Ziv-Ukelson, M., Barash, D., Kedem, K Outline Background Motivation RNA s structure representations
Lecture 3: Graphs and flows
 Chapter 3 Lecture 3: Graphs and flows Graphs: a useful combinatorial structure. Definitions: graph, directed and undirected graph, edge as ordered pair, path, cycle, connected graph, strongly connected
Chapter 3 Lecture 3: Graphs and flows Graphs: a useful combinatorial structure. Definitions: graph, directed and undirected graph, edge as ordered pair, path, cycle, connected graph, strongly connected
Shortest Path Algorithm
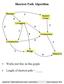 Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
Dynamic Programming. Lecture Overview Introduction
 Lecture 12 Dynamic Programming 12.1 Overview Dynamic Programming is a powerful technique that allows one to solve many different types of problems in time O(n 2 ) or O(n 3 ) for which a naive approach
Lecture 12 Dynamic Programming 12.1 Overview Dynamic Programming is a powerful technique that allows one to solve many different types of problems in time O(n 2 ) or O(n 3 ) for which a naive approach
PROTEIN MULTIPLE ALIGNMENT MOTIVATION: BACKGROUND: Marina Sirota
 Marina Sirota MOTIVATION: PROTEIN MULTIPLE ALIGNMENT To study evolution on the genetic level across a wide range of organisms, biologists need accurate tools for multiple sequence alignment of protein
Marina Sirota MOTIVATION: PROTEIN MULTIPLE ALIGNMENT To study evolution on the genetic level across a wide range of organisms, biologists need accurate tools for multiple sequence alignment of protein
Computational Molecular Biology
 Computational Molecular Biology Erwin M. Bakker Lecture 3, mainly from material by R. Shamir [2] and H.J. Hoogeboom [4]. 1 Pairwise Sequence Alignment Biological Motivation Algorithmic Aspect Recursive
Computational Molecular Biology Erwin M. Bakker Lecture 3, mainly from material by R. Shamir [2] and H.J. Hoogeboom [4]. 1 Pairwise Sequence Alignment Biological Motivation Algorithmic Aspect Recursive
TCCAGGTG-GAT TGCAAGTGCG-T. Local Sequence Alignment & Heuristic Local Aligners. Review: Probabilistic Interpretation. Chance or true homology?
 Local Sequence Alignment & Heuristic Local Aligners Lectures 18 Nov 28, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20 Johnson Hall
Local Sequence Alignment & Heuristic Local Aligners Lectures 18 Nov 28, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20 Johnson Hall
Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 Luay Nakhleh, Rice University
 1 Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 Luay Nakhleh, Rice University DP Algorithms for Pairwise Alignment 2 The number of all possible pairwise alignments (if gaps are allowed)
1 Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 Luay Nakhleh, Rice University DP Algorithms for Pairwise Alignment 2 The number of all possible pairwise alignments (if gaps are allowed)
Longest Common Subsequence, Knapsack, Independent Set Scribe: Wilbur Yang (2016), Mary Wootters (2017) Date: November 6, 2017
 CS161 Lecture 13 Longest Common Subsequence, Knapsack, Independent Set Scribe: Wilbur Yang (2016), Mary Wootters (2017) Date: November 6, 2017 1 Overview Last lecture, we talked about dynamic programming
CS161 Lecture 13 Longest Common Subsequence, Knapsack, Independent Set Scribe: Wilbur Yang (2016), Mary Wootters (2017) Date: November 6, 2017 1 Overview Last lecture, we talked about dynamic programming
Sequence Alignment (chapter 6) p The biological problem p Global alignment p Local alignment p Multiple alignment
 Sequence lignment (chapter 6) p The biological problem p lobal alignment p Local alignment p Multiple alignment Local alignment: rationale p Otherwise dissimilar proteins may have local regions of similarity
Sequence lignment (chapter 6) p The biological problem p lobal alignment p Local alignment p Multiple alignment Local alignment: rationale p Otherwise dissimilar proteins may have local regions of similarity
Algorithms for Data Science
 Algorithms for Data Science CSOR W4246 Eleni Drinea Computer Science Department Columbia University Thursday, October 1, 2015 Outline 1 Recap 2 Shortest paths in graphs with non-negative edge weights (Dijkstra
Algorithms for Data Science CSOR W4246 Eleni Drinea Computer Science Department Columbia University Thursday, October 1, 2015 Outline 1 Recap 2 Shortest paths in graphs with non-negative edge weights (Dijkstra
Chapter 6. Dynamic Programming
 Chapter 6 Dynamic Programming CS 573: Algorithms, Fall 203 September 2, 203 6. Maximum Weighted Independent Set in Trees 6..0. Maximum Weight Independent Set Problem Input Graph G = (V, E) and weights
Chapter 6 Dynamic Programming CS 573: Algorithms, Fall 203 September 2, 203 6. Maximum Weighted Independent Set in Trees 6..0. Maximum Weight Independent Set Problem Input Graph G = (V, E) and weights
Lecture 9: Core String Edits and Alignments
 Biosequence Algorithms, Spring 2005 Lecture 9: Core String Edits and Alignments Pekka Kilpeläinen University of Kuopio Department of Computer Science BSA Lecture 9: String Edits and Alignments p.1/30 III:
Biosequence Algorithms, Spring 2005 Lecture 9: Core String Edits and Alignments Pekka Kilpeläinen University of Kuopio Department of Computer Science BSA Lecture 9: String Edits and Alignments p.1/30 III:
Lecture 2. Dynammic Programming Notes.txt Page 1
 Lecture 2. Dynammic Programming Notes.txt Page 1 Dynamic Programming Michael Schatz (mschatz@cshl.edu) =============================================================================== Motivation: Many optimization
Lecture 2. Dynammic Programming Notes.txt Page 1 Dynamic Programming Michael Schatz (mschatz@cshl.edu) =============================================================================== Motivation: Many optimization
An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST
 An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST Alexander Chan 5075504 Biochemistry 218 Final Project An Analysis of Pairwise
An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST Alexander Chan 5075504 Biochemistry 218 Final Project An Analysis of Pairwise
Alignment of Pairs of Sequences
 Bi03a_1 Unit 03a: Alignment of Pairs of Sequences Partners for alignment Bi03a_2 Protein 1 Protein 2 =amino-acid sequences (20 letter alphabeth + gap) LGPSSKQTGKGS-SRIWDN LN-ITKSAGKGAIMRLGDA -------TGKG--------
Bi03a_1 Unit 03a: Alignment of Pairs of Sequences Partners for alignment Bi03a_2 Protein 1 Protein 2 =amino-acid sequences (20 letter alphabeth + gap) LGPSSKQTGKGS-SRIWDN LN-ITKSAGKGAIMRLGDA -------TGKG--------
Divide and Conquer Algorithms. Problem Set #3 is graded Problem Set #4 due on Thursday
 Divide and Conquer Algorithms Problem Set #3 is graded Problem Set #4 due on Thursday 1 The Essence of Divide and Conquer Divide problem into sub-problems Conquer by solving sub-problems recursively. If
Divide and Conquer Algorithms Problem Set #3 is graded Problem Set #4 due on Thursday 1 The Essence of Divide and Conquer Divide problem into sub-problems Conquer by solving sub-problems recursively. If
15-451/651: Design & Analysis of Algorithms January 26, 2015 Dynamic Programming I last changed: January 28, 2015
 15-451/651: Design & Analysis of Algorithms January 26, 2015 Dynamic Programming I last changed: January 28, 2015 Dynamic Programming is a powerful technique that allows one to solve many different types
15-451/651: Design & Analysis of Algorithms January 26, 2015 Dynamic Programming I last changed: January 28, 2015 Dynamic Programming is a powerful technique that allows one to solve many different types
Reconstructing long sequences from overlapping sequence fragment. Searching databases for related sequences and subsequences
 SEQUENCE ALIGNMENT ALGORITHMS 1 Why compare sequences? Reconstructing long sequences from overlapping sequence fragment Searching databases for related sequences and subsequences Storing, retrieving and
SEQUENCE ALIGNMENT ALGORITHMS 1 Why compare sequences? Reconstructing long sequences from overlapping sequence fragment Searching databases for related sequences and subsequences Storing, retrieving and
Bioinformatics for Biologists
 Bioinformatics for Biologists Sequence Analysis: Part I. Pairwise alignment and database searching Fran Lewitter, Ph.D. Director Bioinformatics & Research Computing Whitehead Institute Topics to Cover
Bioinformatics for Biologists Sequence Analysis: Part I. Pairwise alignment and database searching Fran Lewitter, Ph.D. Director Bioinformatics & Research Computing Whitehead Institute Topics to Cover
Alignment of Long Sequences
 Alignment of Long Sequences BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2009 Mark Craven craven@biostat.wisc.edu Pairwise Whole Genome Alignment: Task Definition Given a pair of genomes (or other large-scale
Alignment of Long Sequences BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2009 Mark Craven craven@biostat.wisc.edu Pairwise Whole Genome Alignment: Task Definition Given a pair of genomes (or other large-scale
Sequence alignment algorithms
 Sequence alignment algorithms Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 23 rd 27 After this lecture, you can decide when to use local and global sequence alignments
Sequence alignment algorithms Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 23 rd 27 After this lecture, you can decide when to use local and global sequence alignments
Sequence Alignment 1
 Sequence Alignment 1 Nucleotide and Base Pairs Purine: A and G Pyrimidine: T and C 2 DNA 3 For this course DNA is double-helical, with two complementary strands. Complementary bases: Adenine (A) - Thymine
Sequence Alignment 1 Nucleotide and Base Pairs Purine: A and G Pyrimidine: T and C 2 DNA 3 For this course DNA is double-helical, with two complementary strands. Complementary bases: Adenine (A) - Thymine
Analysis of Algorithms, I
 Analysis of Algorithms, I CSOR W4231.002 Eleni Drinea Computer Science Department Columbia University Thursday, March 8, 2016 Outline 1 Recap Single-source shortest paths in graphs with real edge weights:
Analysis of Algorithms, I CSOR W4231.002 Eleni Drinea Computer Science Department Columbia University Thursday, March 8, 2016 Outline 1 Recap Single-source shortest paths in graphs with real edge weights:
Algorithmic Approaches for Biological Data, Lecture #20
 Algorithmic Approaches for Biological Data, Lecture #20 Katherine St. John City University of New York American Museum of Natural History 20 April 2016 Outline Aligning with Gaps and Substitution Matrices
Algorithmic Approaches for Biological Data, Lecture #20 Katherine St. John City University of New York American Museum of Natural History 20 April 2016 Outline Aligning with Gaps and Substitution Matrices
Sequence analysis Pairwise sequence alignment
 UMF11 Introduction to bioinformatics, 25 Sequence analysis Pairwise sequence alignment 1. Sequence alignment Lecturer: Marina lexandersson 12 September, 25 here are two types of sequence alignments, global
UMF11 Introduction to bioinformatics, 25 Sequence analysis Pairwise sequence alignment 1. Sequence alignment Lecturer: Marina lexandersson 12 September, 25 here are two types of sequence alignments, global
Dynamic Programming (Part #2)
 Dynamic Programming (Part #) Introduction to Algorithms MIT Press (Chapter 5) Matrix-Chain Multiplication Problem: given a sequence A, A,, A n, compute the product: A A A n Matrix compatibility: C = A
Dynamic Programming (Part #) Introduction to Algorithms MIT Press (Chapter 5) Matrix-Chain Multiplication Problem: given a sequence A, A,, A n, compute the product: A A A n Matrix compatibility: C = A
Solutions to Problem 1 of Homework 3 (10 (+6) Points)
 Solutions to Problem 1 of Homework 3 (10 (+6) Points) Sometimes, computing extra information can lead to more efficient divide-and-conquer algorithms. As an example, we will improve on the solution to
Solutions to Problem 1 of Homework 3 (10 (+6) Points) Sometimes, computing extra information can lead to more efficient divide-and-conquer algorithms. As an example, we will improve on the solution to
Dynamic Programming 1
 Dynamic Programming 1 Jie Wang University of Massachusetts Lowell Department of Computer Science 1 I thank Prof. Zachary Kissel of Merrimack College for sharing his lecture notes with me; some of the examples
Dynamic Programming 1 Jie Wang University of Massachusetts Lowell Department of Computer Science 1 I thank Prof. Zachary Kissel of Merrimack College for sharing his lecture notes with me; some of the examples
Multiple Sequence Alignment (MSA)
 I519 Introduction to Bioinformatics, Fall 2013 Multiple Sequence Alignment (MSA) Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Outline Multiple sequence alignment (MSA) Generalize
I519 Introduction to Bioinformatics, Fall 2013 Multiple Sequence Alignment (MSA) Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Outline Multiple sequence alignment (MSA) Generalize
Subsequence Definition. CS 461, Lecture 8. Today s Outline. Example. Assume given sequence X = x 1, x 2,..., x m. Jared Saia University of New Mexico
 Subsequence Definition CS 461, Lecture 8 Jared Saia University of New Mexico Assume given sequence X = x 1, x 2,..., x m Let Z = z 1, z 2,..., z l Then Z is a subsequence of X if there exists a strictly
Subsequence Definition CS 461, Lecture 8 Jared Saia University of New Mexico Assume given sequence X = x 1, x 2,..., x m Let Z = z 1, z 2,..., z l Then Z is a subsequence of X if there exists a strictly
CSE 373 Analysis of Algorithms, Fall Homework #3 Solutions Due Monday, October 18, 2003
 Piyush Kumar CSE 373 Analysis of Algorithms, Fall 2003 Homework #3 Solutions Due Monday, October 18, 2003 Problem 1 Find an optimal parenthesization of a matrix chain product whose sequence of dimensions
Piyush Kumar CSE 373 Analysis of Algorithms, Fall 2003 Homework #3 Solutions Due Monday, October 18, 2003 Problem 1 Find an optimal parenthesization of a matrix chain product whose sequence of dimensions
Basics on bioinforma-cs Lecture 4. Concita Cantarella
 Basics on bioinforma-cs Lecture 4 Concita Cantarella concita.cantarella@entecra.it; concita.cantarella@gmail.com Why compare sequences Sequence comparison is a way of arranging the sequences of DNA, RNA
Basics on bioinforma-cs Lecture 4 Concita Cantarella concita.cantarella@entecra.it; concita.cantarella@gmail.com Why compare sequences Sequence comparison is a way of arranging the sequences of DNA, RNA
Algorithms IV. Dynamic Programming. Guoqiang Li. School of Software, Shanghai Jiao Tong University
 Algorithms IV Dynamic Programming Guoqiang Li School of Software, Shanghai Jiao Tong University Dynamic Programming Shortest Paths in Dags, Revisited Shortest Paths in Dags, Revisited The special distinguishing
Algorithms IV Dynamic Programming Guoqiang Li School of Software, Shanghai Jiao Tong University Dynamic Programming Shortest Paths in Dags, Revisited Shortest Paths in Dags, Revisited The special distinguishing
Dynamic Programming in 3-D Progressive Alignment Profile Progressive Alignment (ClustalW) Scoring Multiple Alignments Entropy Sum of Pairs Alignment
 Dynamic Programming in 3-D Progressive Alignment Profile Progressive Alignment (ClustalW) Scoring Multiple Alignments Entropy Sum of Pairs Alignment Partial Order Alignment (POA) A-Bruijin (ABA) Approach
Dynamic Programming in 3-D Progressive Alignment Profile Progressive Alignment (ClustalW) Scoring Multiple Alignments Entropy Sum of Pairs Alignment Partial Order Alignment (POA) A-Bruijin (ABA) Approach
Mouse, Human, Chimpanzee
 More Alignments 1 Mouse, Human, Chimpanzee Mouse to Human Chimpanzee to Human 2 Mouse v.s. Human Chromosome X of Mouse to Human 3 Local Alignment Given: two sequences S and T Find: substrings of S and
More Alignments 1 Mouse, Human, Chimpanzee Mouse to Human Chimpanzee to Human 2 Mouse v.s. Human Chromosome X of Mouse to Human 3 Local Alignment Given: two sequences S and T Find: substrings of S and
Algorithms for Computational Biology. String Distance Measures
 Algorithms for Computational Biology Zsuzsanna Lipták Masters in Molecular and Medical Biotechnology a.a. 2015/16, fall term String Distance Measures Similarity vs. distance Two ways of measuring the same
Algorithms for Computational Biology Zsuzsanna Lipták Masters in Molecular and Medical Biotechnology a.a. 2015/16, fall term String Distance Measures Similarity vs. distance Two ways of measuring the same
Algorithms. Lecture Notes 5
 Algorithms. Lecture Notes 5 Dynamic Programming for Sequence Comparison The linear structure of the Sequence Comparison problem immediately suggests a dynamic programming approach. Naturally, our sub-instances
Algorithms. Lecture Notes 5 Dynamic Programming for Sequence Comparison The linear structure of the Sequence Comparison problem immediately suggests a dynamic programming approach. Naturally, our sub-instances
Computer Science & Engineering 423/823 Design and Analysis of Algorithms
 Computer Science & Engineering 423/823 Design and Analysis of Algorithms Lecture 07 Single-Source Shortest Paths (Chapter 24) Stephen Scott and Vinodchandran N. Variyam sscott@cse.unl.edu 1/36 Introduction
Computer Science & Engineering 423/823 Design and Analysis of Algorithms Lecture 07 Single-Source Shortest Paths (Chapter 24) Stephen Scott and Vinodchandran N. Variyam sscott@cse.unl.edu 1/36 Introduction
Introduction to Algorithms
 Introduction to Algorithms 6.046J/18.401J LECTURE 12 Dynamic programming Longest common subsequence Optimal substructure Overlapping subproblems Prof. Charles E. Leiserson Dynamic programming Design technique,
Introduction to Algorithms 6.046J/18.401J LECTURE 12 Dynamic programming Longest common subsequence Optimal substructure Overlapping subproblems Prof. Charles E. Leiserson Dynamic programming Design technique,
FASTA. Besides that, FASTA package provides SSEARCH, an implementation of the optimal Smith- Waterman algorithm.
 FASTA INTRODUCTION Definition (by David J. Lipman and William R. Pearson in 1985) - Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence
FASTA INTRODUCTION Definition (by David J. Lipman and William R. Pearson in 1985) - Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence
Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA.
 Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA. Fasta is used to compare a protein or DNA sequence to all of the
Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA. Fasta is used to compare a protein or DNA sequence to all of the
Sequence Alignment. part 2
 Sequence Alignment part 2 Dynamic programming with more realistic scoring scheme Using the same initial sequences, we ll look at a dynamic programming example with a scoring scheme that selects for matches
Sequence Alignment part 2 Dynamic programming with more realistic scoring scheme Using the same initial sequences, we ll look at a dynamic programming example with a scoring scheme that selects for matches
CMPSCI 311: Introduction to Algorithms Practice Final Exam
 CMPSCI 311: Introduction to Algorithms Practice Final Exam Name: ID: Instructions: Answer the questions directly on the exam pages. Show all your work for each question. Providing more detail including
CMPSCI 311: Introduction to Algorithms Practice Final Exam Name: ID: Instructions: Answer the questions directly on the exam pages. Show all your work for each question. Providing more detail including
Algorithm Design and Analysis
 Algorithm Design and Analysis LECTURE 16 Dynamic Programming Least Common Subsequence Saving space Adam Smith Least Common Subsequence A.k.a. sequence alignment edit distance Longest Common Subsequence
Algorithm Design and Analysis LECTURE 16 Dynamic Programming Least Common Subsequence Saving space Adam Smith Least Common Subsequence A.k.a. sequence alignment edit distance Longest Common Subsequence
Evolutionary tree reconstruction (Chapter 10)
 Evolutionary tree reconstruction (Chapter 10) Early Evolutionary Studies Anatomical features were the dominant criteria used to derive evolutionary relationships between species since Darwin till early
Evolutionary tree reconstruction (Chapter 10) Early Evolutionary Studies Anatomical features were the dominant criteria used to derive evolutionary relationships between species since Darwin till early
