Automated CT marker segmentation for image registration in radionuclide therapy
|
|
|
- Charlotte Sharp
- 5 years ago
- Views:
Transcription
1 INSTITUTE OF PHYSICSPUBLISHING Phys. Med. Biol. 46 (2001) N269 N279 PHYSICS INMEDICINE AND BIOLOGY PII: S (01)26583-X NOTE Automated CT marker segmentation for image registration in radionuclide therapy Periklis Papavasileiou, Glenn D Flux, Maggie A Flower and Matthew J Guy Joint Department of Physics, Institute of Cancer Research and Royal Marsden NHS Trust, Downs Road Sutton, SM2 5PT, Surrey, UK periklis@icr.ac.uk Received 6 July 2001, in final form 12 September 2001 Published 14 November 2001 Online at stacks.iop.org/pmb/46/n269 Abstract In this paper a novel, automated CT marker segmentation technique for image registration is described. The technique, which is based on analysing each CT slice contour individually, treats the cross sections of the external markers as protrusions of the slice contour. Knowledge-based criteria, using the shape and dimensions of the markers, are defined to enable marker identification and segmentation. Following segmentation, the three-dimensional (3D) markers centroids are localized using an intensity-weighted algorithm. Finally, image registration is performed using a least-squares fit algorithm. The technique was applied to both simulated and patient studies. The patients were undergoing 131 I-mIBG radionuclide therapy with each study comprising several 99m Tc single photon emission computed tomography (SPECT) scans and one CT marker scan. The mean residual 3D registration errors (±1 SD) computed for the simulated and patient studies were 1.8 ± 0.3 mm and 4.3 ± 0.5 mm respectively. 1. Introduction Different imaging modalities provide different but complementary information. Single photon emission computed tomography (SPECT) images provide functional information whereas computed tomography (CT) images are superior in revealing the anatomy. Both functional and anatomical information are combined to aid diagnosis and improve the outcome of interventional and treatment-planning procedures. Image registration is required to enable mapping between the co-ordinates of one imaging space to the other. In biologically targeted radionuclide therapy (BTRT), administration of a 131 I-labelled tumour-specific agent is followed by a series of sequential SPECT scans to characterize the dose distribution to both tumour and normal organs. Registration of the sequential 131 I SPECT scans is essential to allow 3D dose calculations to be generated. CT scans are acquired, /01/ $ IOP Publishing Ltd Printed in the UK N269
2 N270 P Papavasileiou et al prior to the administration of the radioactivity, to allow delineation of volumes of interest (VOIs) around tumour sites and normal organs for which 3D dosimetry is to be performed. Furthermore, 3D dosimetry requires accurate SPECT quantification and registered CT scans are potentially useful for attenuation correction of the SPECT scans. Several image registration methods have been developed and used in medical imaging including voxel-based and point-based methods (Maes et al 1997, Eberl et al 1996, Yu et al 1995, Woods et al 1993, Hill et al 1991). Voxel-based methods rely on computing the transformation parameters by optimizing an appropriate similarity criterion, whereas pointbased approaches require the determination of two sets of corresponding points in order to compute the transformation matrix by a least-squares fit. Those points can be either intrinsic, e.g. anatomical/functional landmark points or extrinsic, e.g. centroids of external markers. Point-based registration based on external markers is widely used in radionuclide therapy mainly because the lack of anatomical information in the SPECT images renders voxelbased registration difficult to implement. External markers are attached to the patient s skin and the process involves localization of the 3D markers centroids on both scans to be registered. Localization of the markers centroids is usually done manually. A semi-automated method was described by Flux (1995) which consisted of providing an initial estimate for the centroid and then employed an intensity-weighted algorithm to compute the centroid. An automated marker centroid localization process has recently been proposed for SPECT marker centroid localization (Papavasileiou et al 2001). Wang et al (1996) described an automated method based on automated segmentation of markers fixed on the patient s skull (CT and MRI head scans) during interventional neurosurgical procedures. In this paper, we describe a new automated segmentation method for CT external markers attached to the patient s skin. The method analyses each CT slice contour individually and treats the markers cross sections as protrusions of the slice contours. Knowledge-based criteria are used to identify the markers and localization of the markers centroids is carried out using an intensity-weighted algorithm. The technique is validated on both simulated and patient studies. The algorithms were written in C and IDL (Research Systems Inc., Colorado, USA) and are included in the Royal Marsden Dosimetry Package (RMDP) (Guy 2000). 2. Methods 2.1. Markers The external markers used for CT scanning consist of Perspex discs, 15 mm in diameter and 12 mm deep, containing a spherical cavity of 10 mm diameter, which is stopped with a small nylon screw and is filled with BaCl 2. Two contrast agents are available in our department, BaCl 2 and BaSO 4. Even though BaCl 2 is poisonous, it does not precipitate like BaSO 4 does. Therefore, BaCl 2 was chosen to be the more appropriate as a CT marker contrast agent. The diameter of marker cavity matched the slice thickness of the CT scans. The size of the SPECT markers is smaller than that of those used on CT, though the geometry and the shape are the same. The reason for that discrepancy is that the scatter and the poor resolution of the SPECT scans result in the marker being shown on several slices even with smaller marker size. Skin markings are made on the patient skin to allow accurate and reproducible placement of the external markers for the 99m Tc SPECT acquisition. The markers are taped on the patient s skin before each scan. The number and the distribution of the markers have to ensure that the tumour as well as normal organs, for which dose distributions have to be generated,
3 Automated CT marker segmentation for image registration in radionuclide therapy N271 START for each slice Generation of the 3D markers connected components Computation of the 3D marker centroids. END N Grey-level thresholding Contour extraction if contour sampling incomplete Y Define a contour segment of 10 mm length. Apply the Collinearity criterion to test if the contour-segment voxels lie on a line. N if Collinearity Y Marker identification criterion. - Define a vector perpendicular to the sampled contour segment. - Define 4 artificial segments along the perpendicular vector parallel to the linear contour segment. The intersections of the parallel segments with the contour are counted. If #intersections is above a threshold of 6, a marker is identified. Figure 1. Flowchart of the automated CT marker segmentation process. are included within the patient volume defined by the markers. The size of the patient is also a factor affecting the number of markers; 5 10 markers are used for marker registration at our centre CT marker segmentation The automated marker segmentation algorithm is an iterative algorithm analysing each CT slice individually. The marker segmentation and centroid localization process consists of the following steps, which are illustrated in the flowchart in figure 1: 1. Grey-level thresholding. The patient image is separated from the background using greylevel thresholding. The threshold is computed automatically for each individual slice using a method described by Bae et al (1993). The main assumption of this approach is that the grey levels within the patient region differ significantly from those in the background. The mean intensity of the voxels, along a horizontal line, from the edge to the centre of the image is computed cumulatively. Since the intensity of the patientregion voxels is significantly higher than that of the background, the mean intensity is relatively constant until the patient-region voxels are included. The mean intensity of the voxels from the centre to the edge is also computed and it is relatively constant until the background voxels are included. At the boundary between the patient region and the background the maximum difference between the two mean intensities occurs. At that point, the mean intensities of the patient and background regions are computed and the threshold is given by the average of those two intensities. All voxels above the threshold
4 N272 P Papavasileiou et al are assigned an intensity of 1, whereas all voxel below threshold are assigned an intensity of Contour extraction. Following grey-level thresholding, the contour of each slice is extracted based on an edge-tracking algorithm first described by Henrich (1979). The location of the starting patient-region edge voxel is found by sampling a horizontal line, on the thresholded image, from the right-hand image edge to the centre and identifying the first non-zero voxel. That voxel is the starting point for contour extraction. Edge tracking is carried out anti-clockwise. The eight-voxel neighbourhood of the starting voxel is examined anti-clockwise for the first non-zero voxel; this voxel is marked as the next contour voxel and the searching process is repeated in its eight-voxel neighbourhood. The algorithm follows the patient-region edge until it reaches the starting voxel (i.e. a closed contour is obtained). 3. Contour sampling. The markers are treated as protrusions of the slice contour with a flat surface parallel to skin surface. The shape of the markers cross sections is close to rectangular but the exact height, width and shape depends on the orientation of the marker relative to the CT x-ray beam. The height of the markers cross sections ranges between 12.0 and 13.0 mm, whereas their width ranges from 9.0 to 12.0 mm. From each contour voxel, a contour segment of 10.0 mm length is defined; the number of voxels comprising the sampled contour segment depends on the image digital resolution (voxel size) and on the curvature of the sampled segment. 4. Collinearity criterion. A collinearity criterion is applied (Pavlidis, 1982) to test if the sampled contour-segment voxels lie on a straight line. A line is drawn between the end-points of the sampled segment and is given by the equation, x(y j y k ) + y(x k x j ) + y k x j y j x k = 0 (1) where (x j,y j ) and (x k,y k ) are co-ordinates of the two end-points of the segment. If a contour voxel with co-ordinates (u, v) does not lie on the line given by equation 1, then its distance from the line is equal to the magnitude d/l where d and L are given by: d = u(y j y k ) + v(x k x j ) + y k x j y j x k (2) L = (y j y k ) 2 + (x k x j ) 2. (3) The arc length, L a, of the sampled segment is also computed: k L a = (xi x i 1 ) 2 + (y i y i 1 ) 2. (4) i=j+1 Collinearity can be established by evaluating either the d/l, that is, setting a maximum threshold on the distance of any point from the fitted, or the L/L a, that is, comparing the length of the arc, along the point sequence, with the length of the line. We chose the L/L a criterion because it takes into consideration all the points in the segment. If the ratio L a /L is less than 1.1, collinearity is confirmed (Pavlidis, 1982) and the process continues to step 5. Otherwise the process returns to step 3 to sample another contour segment of 10.0 mm length. An L a /L value of 1.1 means a deviation of 10% from a perfect linear segment (L a /L value of 1.0). 5. Marker identification criterion. A vector perpendicular to the linear contour segment is defined. Artificial segments parallel to the contour segment are then generated along the perpendicular vector as shown in figure 2(a). Those segments are generated only on one side of the sampled contour segment; the choice of side is pre-defined since the direction of contour generation is known. The number and the length of the artificial segments depend
5 Automated CT marker segmentation for image registration in radionuclide therapy N273 contrast-filled spherical cavity A B C D body contour intersection normal to segment AB (a) A B E S End point of the 2D marker polygon Starting point of the 2D marker polygon body contour (b) Figure 2. (a) Graphic illustration of the CT marker identification criterion. AB and CD are contour segments, which satisfy both the length and the collinearity criteria. Only AB belongs to a marker because it satisfies the intersection criterion. (b) Generation of the 2D marker polygon defined by the sampled segment ABand the intersections, S and E, of the artificial segment furthest away from AB. on the height and width of the marker respectively; four parallel, artificial segments, 3.0 mm apart, with length of 14.0 mm are defined for the linear contour segment. The intersections of these artificial segments with the slice contour are counted and if their number exceeds a threshold of six (three pairs of intersections), a marker is identified. The voxels at the intersection furthest away from the linear contour segment mark the endpoints of the two-dimensional (2D) marker representation. A 2D polygon is defined by the sampled contour segment and the above two endpoints as shown in figure 2(b). The voxels enclosed in that polygon are saved. The process returns to step 3 to continue the contour sampling from the contour voxel corresponding to the endpoint of the 2D polygon. When contour sampling is completed the process continues to step Generation of 3D marker connected component. Following analysis of all CT slices, the segmented markers voxels are copied to a temporary binary file where all marker voxels are assigned an intensity of 1. 3D six-neighbourhood connectivity is employed to allow generation of the marker 3D-connected components; the four immediate 2D neighbours within the slice and the two immediate ones in the neighbouring slices are considered to establish connectivity.
6 N274 P Papavasileiou et al a) b) c) d) Figure 3. (a) Slice through a CT marker. (b) Slice contour. (c) The linear marker segment and the perpendicular vector are superimposed on the slice contour. The two highlighted contour voxels correspond to the start and end of the 2D segmented marker polygon. (d) The segmented marker. 7. Marker centroid computation. The marker centroid is determined using the intensities in the original image of the voxels comprising the marker 3D-connected component. Initially, the arithmetic centroid of the segmented marker is computed which serves as an initial estimate of the geometric centroid. Then, an intensity-weighted algorithm is applied to calculate the marker centroid more accurately. An example of the automated marker segmentation process is shown in figure 3, where the main steps of the algorithm are illustrated Marker registration One-to-one correspondence between the two CT and SPECT marker centroid sets (marker matching) was established based on an automated method (Papavasileiou et al 2001). The least-squares fit algorithm developed by Arun et al (1987) was then employed to compute the six transformation parameters (X, Y, Z translation, XY, XZ and YZ rotation) required to
7 Automated CT marker segmentation for image registration in radionuclide therapy N275 register the two marker sets. Scaling of the image data was carried out automatically using the known voxel sizes of the scans to be registered Simulated study In order to validate the CT marker segmentation process, a simulated study was generated. Nine copies of a CT marker scan, which was acquired as part of a patient therapy study and in which six markers were used, were generated. These were subjected to a range of transformation parameters ( mm and 0 10 for translation and rotation respectively). The 3D matrix dimensions of the CT scan were ( mm 3 ). Each of the ten scans of the simulated study was registered to the remaining nine resulting in 45, i.e. (n 1 (n 2 1))/2, registration pairs, where n 1 = n 2 = 10. The robustness of the collinearity criterion was also assessed with the simulated study. The segmentation process was repeated for L a /L value of 1.05 and 1.08, respectively, and the number of markers segmented for each of the above values was computed Patient studies Two different patient studies (A and B) were used to test the automated CT marker segmentation algorithm described in this paper. Both studies analysed were abdominal studies. Both patients had received therapeutic doses of iodine-131 metaiodobenzylguanidine ( 131 I-mIBG) for treatment of neuroblastoma (Hoefnagel et al 1987). Each patient study consisted of one CT and five 99m Tc-SPECT marker scans. The CT scans were acquired, prior to the administration of the therapeutic activity, on a Siemens DR Somatom CT scanner. The 99m Tc-SPECT scans (matrix , voxel dimensions mm 3 ) were acquired on a General Electric STARCAM camera (General Electric Medical Systems, Milwaukee, WI) with a highenergy, high-resolution collimator; the high-energy collimator is required because an 131 I scan is acquired after the 99m Tc marker scan. The SPECT scans were then interpolated to matrices ( mm 3 ). The matrix dimensions of each CT slice were ; the voxel dimensions were mm 3 and mm 3 for studies A and B respectively. The CT image planes were contiguous. SPECT marker segmentation and centroid computation were carried out using an automated approach (Papavasileiou et al 2001). Each 99m Tc-SPECT marker scan was registered to the CT scan. Seven and six markers were used for studies A and B respectively. 3. Results 3.1. Simulated study The automated segmentation process resulted in all six markers being segmented on each registration pair. In order to test the accuracy of the registration process, the 3D registration residual error was computed by measuring the root-mean-square (rms) distance between coordinates of pairs of corresponding marker centroids following image registration. Subvoxel registration accuracy was achieved, with a mean 3D residual error (±1 SD)forthe 45 registration pairs of 1.8 ± 0.3 mm. The value of the collinearity criterion was shown to affect the number of identified markers (figure 4) with a value less than 1.1 resulting in failure of the algorithm to identify all the markers.
8 N276 P Papavasileiou et al Number of segmented markers Scan index Figure 4. Number of segmented markers in the simulated study for three different values of the collinearity criterion, L a /L, 1.05 ( ), 1.08 ( ) and 1.10 ( ) respectively. Scan index refers to the corresponding scan analysed in the simulated study Patient studies All 13 markers, 7 for study A and 6 for study B, were segmented by the automated segmentation algorithm. The difference in CT voxel sizes on the two studies, mm 3 and mm 3 for studies A and B respectively, did not affect the sensitivity of the segmentation process. As with the simulated study, the accuracy of the segmentation process was assessed by computing the 3D residual error of the registration process. A value of 4.3 ± 0.5 mm (mean ±SD) for the 3D registration error was calculated. The computation of the 3D residual error in patient studies made the necessary assumption that the markers are in the same position irrespective of the modality used and/or the time of the scan. Even though we were confident with the process of repositioning the markers, for either SPECT or CT acquisition, contributions to the registration error due to the difference in the scanner beds could not be avoided. However, as shown by the mean residual error value, those contributions were very small. 4. Discussion The automated CT marker segmentation technique described in this paper is based on the assumption that the shape of the 2D marker cross sections is not distorted by the choice of the threshold. The automated thresholding approach employed (Bae et al 1993) neither produced erroneous threshold values nor distorted the shape of the marker cross sections on the abdominal studies analysed; the studies analysed in this paper were abdominal studies. The above thresholding technique will produce erroneous threshold values in situations where the mean background value is significantly higher than zero and/or when the voxel intensity values along the sampled line is not significantly higher than those of the background (e.g. lungs, intestine). In the latter case, manual selection of the threshold might be necessary. The importance of thresholding in automated marker segmentation was reiterated by Wang et al (1996). They employed thresholding to segment the markers and the patient region (i.e. patient s skull) from the background. Their segmentation approach relied on the shape and dimensions of the contrast-filled marker cavity. As a result, the choice of threshold employed was more critical because the threshold had to be sufficiently higher than the background and at the same time sufficiently low in order to exploit the intensity information within the marker. The above approach required that the thresholding resulted in the head volume being segmented, thus leaving only the markers. Should a threshold associated with the BaCl 2 be applied to the abdominal scans, it would result in several anatomical structures,
9 Automated CT marker segmentation for image registration in radionuclide therapy N277 i.e. bones, present in the thresholded image, which would have to be analysed as potential markers. Consequently, the segmentation process would be slower. The algorithm described in this paper relies on knowledge of the shape and dimensions of the marker cross sections, thus being less vulnerable to the threshold value. The main requirement for the thresholding process is that the marker cross sections are not distorted so that the collinearity criterion is applicable. The value for the above criterion is important, as is shown in figure 4; a value less than 1.1 was found to be insufficient for the identification of every marker in the simulated study. The above failure was expected since the contour is not continuous but discrete and since thresholding might result in a particular segment deviating from perfect linearity. The choice of image digital resolution (i.e. voxel size) in conjunction with the size of the field-of-view (FOV) might affect the smallest detectable contour feature. For a specific FOV, the smaller the dimension of the contour feature to be identified, the highest the digital resolution required (Kita and Shirai 1991). The digital resolution (i.e. voxel size) has to be such that the number of voxels comprising the sampled contour segment is sufficient for the collinearity criterion to be applied and for an accurate definition of the vector perpendicular to the segment. The matrix size of ( ) employed in this paper, was found to provide sufficient digital resolution for the dimensions of the markers even though the FOV dimensions employed in the two studies were different (FOV of 200 and 400 mm for studies A and B respectively). The algorithm s sensitivity depends on the number and the frequency of the artificial segments parallel to the linear marker segment. Four artificial segments, with segment separation of 3.0 mm, were defined for each potential contour marker segment. The number and frequency of the artificial segments depend on the marker s height and voxel size; segment separation must be at least equal to the voxel (x,y) dimension. The algorithm is not dependent on the marker cross section being rectangular. The shape of the cross sections depends mainly on the orientation of the marker relative to the CT x-ray beam but also on the threshold used. This irregularity in shape would render marker segmentation using morphological operators (e.g. opening, closing) difficult. Segmentation based on morphological operators requires the definition of a structural element (e.g. rectangle) which is then convolved with the 2D binary image to identify potential matches,that is potential markers. Definition of more than one structural element would be required to segment external markers placed on the skin. The requirement for segmentation is that the identification criteria, i.e. linear segment and intersections between the artificial segments and the contour, are satisfied. The orientation of the marker with respect to the CT x-ray beam, i.e. the out-ofplane rotation, will have an effect on the shape of the cross section (figure 5). Moreover, it might result in a marker cross section not being identified as shown in figure 5(b). An exampleof the above situation is shown in figure 6; the cross sections of markers m A and m C are correctly identified wheras the cross sections of m B is not. In order to overcome the above problem, prior to the computation of the marker centroid the volume of the 3D component is enlarged by adding two slices, one below and one above. Therefore, all the information available for the marker is taken into account, making the segmentation process robust to out-of-plane rotations and not dependent on the slice thickness, i.e. the number slices. The appearance of the markers in more than one slice also allows computation of the marker centroid with sub-slice accuracy. The registration technique described in this paper has been shown to be accurate. In order to use the registered CT/SPECT scans in the dosimetric process, e.g. delineation of the anatomical volumes on CT and transfer of those on the SPECT to allow 3D dose calculations, the position of the skin markers to the internal organs should be constant between scans.
10 N278 P Papavasileiou et al a) b) Figure 5. Simple schematic diagrams of the 3D marker (top) and the potential shape of the marker cross sections (bottom). In (a) the marker top-surface is perpendicular to the CT slice. In (b), the out-of-plane rotation results in the marker cross sections shape not being rectangular. Also, one of the marker cross sections is not identified because the marker identification criteria are not satisfied. m C m B m A a) b) Figure 6. (a) Slice through a CT marker. (b) Slice contour. Marker, m A, is correctly identified with its cross section being rectangular. The cross section of marker m B is not identified whereas the cross section of m C is correctly identified despite the fact that it is not rectangular. Provided that this assumption is valid, internal organ movement due to different scanner beds between CT and SPECT scans will be reflected in the residual marker registration error. However, internal organ movement with respect to the skin surface, with the skin marker geometry remaining constant, is not reflected. 5. Conclusion An automated technique to segment external CT markers has been developed for image registration. Aprioriinformation on the shape and dimensions of the external markers is employed to localize and segment the markers. The method is very sensitive and robust. The technique has been applied to radionuclide therapy studies resulting in a 3D residual
11 Automated CT marker segmentation for image registration in radionuclide therapy N279 registration error of 4.3 mm. The technique is applicable to any diagnostic and/or therapeutic study (e.g. registration of single CT scans, each being acquired prior to the delivery of external beam therapy during fractionated radiotherapy), which involves external CT markers. Acknowledgments We thank Professor Robert J Ott for his useful comments on the manuscript. This work was funded by the Association of International Cancer Research, the Medical Research Council and the Royal Marsden NHS Trust Charitable. References Arun K S, Huang T S and Blostein S D 1987 Least-squares fitting of two 3-D point sets IEEE Trans. Pattern Anal. Machine Intell Bae KT, Giger ML, Chen C-T and Kahn CE Jr 1993 Automatic segmentation of liver structure in CT images Med. Phys Eberl S, Kanno I, Fulton R R, Ryan A, Hutton B F and Fulham M J 1996 Automated interstudy image registration technique for SPECT and PET J. Nucl. Med Flux G D 1995 Multimodality image registration and its application to the dosimetry of intralesional radionuclide therapy PhD Thesis University of London, London, UK Guy M J 2000 The application of quantitative single photon emission tomography to targeted radionuclide therapy PhD Thesis University of London, London, UK Henrich G, Mai N and Backmund H 1979 Preprocessing in computed tomography picture analysis: a bone-deleting algorithm J. Comput. Assist. Tomogr Hill D L G, Hawkes D J, Crossman J E, Gleeson M J, Cox T C S, Bracey E E C M L, Strong A J and Graves P 1991 Registration of MR and CT images for skull base surgery using point-like anatomical features Br. J. Radiol Hoefnagel C A, Voûte P A, de Kraker J and Marcuse H R 1987 Radionuclide diagnosis and therapy of neural crest tumours using iodine-131 metaiodobenzylguanidine J. Nucl. Med Kita Y and Shirai Y 1991 Extraction of accurate stomach contours from x-ray images of barium-filled stomachs and its application to detect potential abnormalities CVGP:Graphical Models Imaging Process Maes F, Collignon A, Vandermeulen D, Marchal and Suetens P 1997 Multimodality image registration by maximisation of mutual information IEEE Trans. Med. Imaging Papavasileiou P, Flux G D, Flower M A and Guy M J 2001 An automated technique for SPECT marker-based image registration in radionuclide therapy Phys. Med. Biol Pavlidis T 1982 Algorithms for Graphics and Image Processing (Berlin: Springer) Wang M Y, Maurer Jr C R, Fitzpatrick J M and Maciunas R J 1996 An automatic technique for finding and localizing externally attached markers in CT and MR volume images of the head IEEE Trans. Biomed. Eng Woods R P, Mazziota J C and Cherry S R 1993 MRI-PET registration with automated algorithm J. Comput. Assist. Tomogr Yu J N, Fahey F H, Gage H D, Eades C G, Harkness B A, Pelizzari C A and Keys Jr J W 1995 Intermodality, retrospective image registration in the thorax J. Nucl. Med
Abstract. 1. Introduction
 A New Automated Method for Three- Dimensional Registration of Medical Images* P. Kotsas, M. Strintzis, D.W. Piraino Department of Electrical and Computer Engineering, Aristotelian University, 54006 Thessaloniki,
A New Automated Method for Three- Dimensional Registration of Medical Images* P. Kotsas, M. Strintzis, D.W. Piraino Department of Electrical and Computer Engineering, Aristotelian University, 54006 Thessaloniki,
3/27/2012 WHY SPECT / CT? SPECT / CT Basic Principles. Advantages of SPECT. Advantages of CT. Dr John C. Dickson, Principal Physicist UCLH
 3/27/212 Advantages of SPECT SPECT / CT Basic Principles Dr John C. Dickson, Principal Physicist UCLH Institute of Nuclear Medicine, University College London Hospitals and University College London john.dickson@uclh.nhs.uk
3/27/212 Advantages of SPECT SPECT / CT Basic Principles Dr John C. Dickson, Principal Physicist UCLH Institute of Nuclear Medicine, University College London Hospitals and University College London john.dickson@uclh.nhs.uk
Reproducibility of interactive registration of 3D CT and MR pediatric treatment planning head images
 JOURNAL OF APPLIED CLINICAL MEDICAL PHYSICS, VOLUME 2, NUMBER 3, SUMMER 2001 Reproducibility of interactive registration of 3D CT and MR pediatric treatment planning head images Jaap Vaarkamp* Department
JOURNAL OF APPLIED CLINICAL MEDICAL PHYSICS, VOLUME 2, NUMBER 3, SUMMER 2001 Reproducibility of interactive registration of 3D CT and MR pediatric treatment planning head images Jaap Vaarkamp* Department
Automated segmentation methods for liver analysis in oncology applications
 University of Szeged Department of Image Processing and Computer Graphics Automated segmentation methods for liver analysis in oncology applications Ph. D. Thesis László Ruskó Thesis Advisor Dr. Antal
University of Szeged Department of Image Processing and Computer Graphics Automated segmentation methods for liver analysis in oncology applications Ph. D. Thesis László Ruskó Thesis Advisor Dr. Antal
OnDemand3D Fusion Technology
 CYBERMED INC., ONDEMAND3D TECHNOLOGY INC. OnDemand3D Fusion Technology White Paper December 2009 USA Republic of Korea www.ondemand3d.com Introduction OnDemand3D TM Fusion is registration technology to
CYBERMED INC., ONDEMAND3D TECHNOLOGY INC. OnDemand3D Fusion Technology White Paper December 2009 USA Republic of Korea www.ondemand3d.com Introduction OnDemand3D TM Fusion is registration technology to
A Radiometry Tolerant Method for Direct 3D/2D Registration of Computed Tomography Data to X-ray Images
 A Radiometry Tolerant Method for Direct 3D/2D Registration of Computed Tomography Data to X-ray Images Transfer Function Independent Registration Boris Peter Selby 1, Georgios Sakas 2, Stefan Walter 1,
A Radiometry Tolerant Method for Direct 3D/2D Registration of Computed Tomography Data to X-ray Images Transfer Function Independent Registration Boris Peter Selby 1, Georgios Sakas 2, Stefan Walter 1,
Quantitative imaging for clinical dosimetry
 Quantitative imaging for clinical dosimetry Irène Buvat Laboratoire d Imagerie Fonctionnelle U678 INSERM - UPMC CHU Pitié-Salpêtrière, Paris buvat@imed.jussieu.fr http://www.guillemet.org/irene Methodology
Quantitative imaging for clinical dosimetry Irène Buvat Laboratoire d Imagerie Fonctionnelle U678 INSERM - UPMC CHU Pitié-Salpêtrière, Paris buvat@imed.jussieu.fr http://www.guillemet.org/irene Methodology
Technical aspects of SPECT and SPECT-CT. John Buscombe
 Technical aspects of SPECT and SPECT-CT John Buscombe What does the clinician need to know? For SPECT What factors affect SPECT How those factors should be sought Looking for artefacts For SPECT-CT Issues
Technical aspects of SPECT and SPECT-CT John Buscombe What does the clinician need to know? For SPECT What factors affect SPECT How those factors should be sought Looking for artefacts For SPECT-CT Issues
Joint CI-JAI advanced accelerator lecture series Imaging and detectors for medical physics Lecture 1: Medical imaging
 Joint CI-JAI advanced accelerator lecture series Imaging and detectors for medical physics Lecture 1: Medical imaging Dr Barbara Camanzi barbara.camanzi@stfc.ac.uk Course layout Day AM 09.30 11.00 PM 15.30
Joint CI-JAI advanced accelerator lecture series Imaging and detectors for medical physics Lecture 1: Medical imaging Dr Barbara Camanzi barbara.camanzi@stfc.ac.uk Course layout Day AM 09.30 11.00 PM 15.30
Computational Medical Imaging Analysis Chapter 4: Image Visualization
 Computational Medical Imaging Analysis Chapter 4: Image Visualization Jun Zhang Laboratory for Computational Medical Imaging & Data Analysis Department of Computer Science University of Kentucky Lexington,
Computational Medical Imaging Analysis Chapter 4: Image Visualization Jun Zhang Laboratory for Computational Medical Imaging & Data Analysis Department of Computer Science University of Kentucky Lexington,
2D Rigid Registration of MR Scans using the 1d Binary Projections
 2D Rigid Registration of MR Scans using the 1d Binary Projections Panos D. Kotsas Abstract This paper presents the application of a signal intensity independent registration criterion for 2D rigid body
2D Rigid Registration of MR Scans using the 1d Binary Projections Panos D. Kotsas Abstract This paper presents the application of a signal intensity independent registration criterion for 2D rigid body
Annales UMCS Informatica AI 1 (2003) UMCS. Registration of CT and MRI brain images. Karol Kuczyński, Paweł Mikołajczak
 Annales Informatica AI 1 (2003) 149-156 Registration of CT and MRI brain images Karol Kuczyński, Paweł Mikołajczak Annales Informatica Lublin-Polonia Sectio AI http://www.annales.umcs.lublin.pl/ Laboratory
Annales Informatica AI 1 (2003) 149-156 Registration of CT and MRI brain images Karol Kuczyński, Paweł Mikołajczak Annales Informatica Lublin-Polonia Sectio AI http://www.annales.umcs.lublin.pl/ Laboratory
Determination of Three-Dimensional Voxel Sensitivity for Two- and Three-Headed Coincidence Imaging
 IEEE TRANSACTIONS ON NUCLEAR SCIENCE, VOL. 50, NO. 3, JUNE 2003 405 Determination of Three-Dimensional Voxel Sensitivity for Two- and Three-Headed Coincidence Imaging Edward J. Soares, Kevin W. Germino,
IEEE TRANSACTIONS ON NUCLEAR SCIENCE, VOL. 50, NO. 3, JUNE 2003 405 Determination of Three-Dimensional Voxel Sensitivity for Two- and Three-Headed Coincidence Imaging Edward J. Soares, Kevin W. Germino,
Lucy Phantom MR Grid Evaluation
 Lucy Phantom MR Grid Evaluation Anil Sethi, PhD Loyola University Medical Center, Maywood, IL 60153 November 2015 I. Introduction: The MR distortion grid, used as an insert with Lucy 3D QA phantom, is
Lucy Phantom MR Grid Evaluation Anil Sethi, PhD Loyola University Medical Center, Maywood, IL 60153 November 2015 I. Introduction: The MR distortion grid, used as an insert with Lucy 3D QA phantom, is
Medical Image Registration by Maximization of Mutual Information
 Medical Image Registration by Maximization of Mutual Information EE 591 Introduction to Information Theory Instructor Dr. Donald Adjeroh Submitted by Senthil.P.Ramamurthy Damodaraswamy, Umamaheswari Introduction
Medical Image Registration by Maximization of Mutual Information EE 591 Introduction to Information Theory Instructor Dr. Donald Adjeroh Submitted by Senthil.P.Ramamurthy Damodaraswamy, Umamaheswari Introduction
Digital Image Processing
 Digital Image Processing SPECIAL TOPICS CT IMAGES Hamid R. Rabiee Fall 2015 What is an image? 2 Are images only about visual concepts? We ve already seen that there are other kinds of image. In this lecture
Digital Image Processing SPECIAL TOPICS CT IMAGES Hamid R. Rabiee Fall 2015 What is an image? 2 Are images only about visual concepts? We ve already seen that there are other kinds of image. In this lecture
Constructing System Matrices for SPECT Simulations and Reconstructions
 Constructing System Matrices for SPECT Simulations and Reconstructions Nirantha Balagopal April 28th, 2017 M.S. Report The University of Arizona College of Optical Sciences 1 Acknowledgement I would like
Constructing System Matrices for SPECT Simulations and Reconstructions Nirantha Balagopal April 28th, 2017 M.S. Report The University of Arizona College of Optical Sciences 1 Acknowledgement I would like
Semi-Automatic Segmentation of the Patellar Cartilage in MRI
 Semi-Automatic Segmentation of the Patellar Cartilage in MRI Lorenz König 1, Martin Groher 1, Andreas Keil 1, Christian Glaser 2, Maximilian Reiser 2, Nassir Navab 1 1 Chair for Computer Aided Medical
Semi-Automatic Segmentation of the Patellar Cartilage in MRI Lorenz König 1, Martin Groher 1, Andreas Keil 1, Christian Glaser 2, Maximilian Reiser 2, Nassir Navab 1 1 Chair for Computer Aided Medical
3D Voxel-Based Volumetric Image Registration with Volume-View Guidance
 3D Voxel-Based Volumetric Image Registration with Volume-View Guidance Guang Li*, Huchen Xie, Holly Ning, Deborah Citrin, Jacek Copala, Barbara Arora, Norman Coleman, Kevin Camphausen, and Robert Miller
3D Voxel-Based Volumetric Image Registration with Volume-View Guidance Guang Li*, Huchen Xie, Holly Ning, Deborah Citrin, Jacek Copala, Barbara Arora, Norman Coleman, Kevin Camphausen, and Robert Miller
Medical Image Analysis
 Computer assisted Image Analysis VT04 29 april 2004 Medical Image Analysis Lecture 10 (part 1) Xavier Tizon Medical Image Processing Medical imaging modalities XRay,, CT Ultrasound MRI PET, SPECT Generic
Computer assisted Image Analysis VT04 29 april 2004 Medical Image Analysis Lecture 10 (part 1) Xavier Tizon Medical Image Processing Medical imaging modalities XRay,, CT Ultrasound MRI PET, SPECT Generic
Deformable Registration Using Scale Space Keypoints
 Deformable Registration Using Scale Space Keypoints Mehdi Moradi a, Purang Abolmaesoumi a,b and Parvin Mousavi a a School of Computing, Queen s University, Kingston, Ontario, Canada K7L 3N6; b Department
Deformable Registration Using Scale Space Keypoints Mehdi Moradi a, Purang Abolmaesoumi a,b and Parvin Mousavi a a School of Computing, Queen s University, Kingston, Ontario, Canada K7L 3N6; b Department
Motion artifact detection in four-dimensional computed tomography images
 Motion artifact detection in four-dimensional computed tomography images G Bouilhol 1,, M Ayadi, R Pinho, S Rit 1, and D Sarrut 1, 1 University of Lyon, CREATIS; CNRS UMR 5; Inserm U144; INSA-Lyon; University
Motion artifact detection in four-dimensional computed tomography images G Bouilhol 1,, M Ayadi, R Pinho, S Rit 1, and D Sarrut 1, 1 University of Lyon, CREATIS; CNRS UMR 5; Inserm U144; INSA-Lyon; University
Workshop on Quantitative SPECT and PET Brain Studies January, 2013 PUCRS, Porto Alegre, Brasil Corrections in SPECT and PET
 Workshop on Quantitative SPECT and PET Brain Studies 14-16 January, 2013 PUCRS, Porto Alegre, Brasil Corrections in SPECT and PET Físico João Alfredo Borges, Me. Corrections in SPECT and PET SPECT and
Workshop on Quantitative SPECT and PET Brain Studies 14-16 January, 2013 PUCRS, Porto Alegre, Brasil Corrections in SPECT and PET Físico João Alfredo Borges, Me. Corrections in SPECT and PET SPECT and
Optimization of CT Simulation Imaging. Ingrid Reiser Dept. of Radiology The University of Chicago
 Optimization of CT Simulation Imaging Ingrid Reiser Dept. of Radiology The University of Chicago Optimization of CT imaging Goal: Achieve image quality that allows to perform the task at hand (diagnostic
Optimization of CT Simulation Imaging Ingrid Reiser Dept. of Radiology The University of Chicago Optimization of CT imaging Goal: Achieve image quality that allows to perform the task at hand (diagnostic
Using Probability Maps for Multi organ Automatic Segmentation
 Using Probability Maps for Multi organ Automatic Segmentation Ranveer Joyseeree 1,2, Óscar Jiménez del Toro1, and Henning Müller 1,3 1 University of Applied Sciences Western Switzerland (HES SO), Sierre,
Using Probability Maps for Multi organ Automatic Segmentation Ranveer Joyseeree 1,2, Óscar Jiménez del Toro1, and Henning Müller 1,3 1 University of Applied Sciences Western Switzerland (HES SO), Sierre,
Ch. 4 Physical Principles of CT
 Ch. 4 Physical Principles of CT CLRS 408: Intro to CT Department of Radiation Sciences Review: Why CT? Solution for radiography/tomography limitations Superimposition of structures Distinguishing between
Ch. 4 Physical Principles of CT CLRS 408: Intro to CT Department of Radiation Sciences Review: Why CT? Solution for radiography/tomography limitations Superimposition of structures Distinguishing between
Medical Image Registration
 Medical Image Registration Submitted by NAREN BALRAJ SINGH SB ID# 105299299 Introduction Medical images are increasingly being used within healthcare for diagnosis, planning treatment, guiding treatment
Medical Image Registration Submitted by NAREN BALRAJ SINGH SB ID# 105299299 Introduction Medical images are increasingly being used within healthcare for diagnosis, planning treatment, guiding treatment
2 Michael E. Leventon and Sarah F. F. Gibson a b c d Fig. 1. (a, b) Two MR scans of a person's knee. Both images have high resolution in-plane, but ha
 Model Generation from Multiple Volumes using Constrained Elastic SurfaceNets Michael E. Leventon and Sarah F. F. Gibson 1 MIT Artificial Intelligence Laboratory, Cambridge, MA 02139, USA leventon@ai.mit.edu
Model Generation from Multiple Volumes using Constrained Elastic SurfaceNets Michael E. Leventon and Sarah F. F. Gibson 1 MIT Artificial Intelligence Laboratory, Cambridge, MA 02139, USA leventon@ai.mit.edu
Copyright 2009 Society of Photo Optical Instrumentation Engineers. This paper was published in Proceedings of SPIE, vol. 7260, Medical Imaging 2009:
 Copyright 2009 Society of Photo Optical Instrumentation Engineers. This paper was published in Proceedings of SPIE, vol. 7260, Medical Imaging 2009: Computer Aided Diagnosis and is made available as an
Copyright 2009 Society of Photo Optical Instrumentation Engineers. This paper was published in Proceedings of SPIE, vol. 7260, Medical Imaging 2009: Computer Aided Diagnosis and is made available as an
Image Registration. Prof. Dr. Lucas Ferrari de Oliveira UFPR Informatics Department
 Image Registration Prof. Dr. Lucas Ferrari de Oliveira UFPR Informatics Department Introduction Visualize objects inside the human body Advances in CS methods to diagnosis, treatment planning and medical
Image Registration Prof. Dr. Lucas Ferrari de Oliveira UFPR Informatics Department Introduction Visualize objects inside the human body Advances in CS methods to diagnosis, treatment planning and medical
Three Dimensional Dosimetry Analyses In Radionuclide Therapy Using IDL And MCNP-based Software Tools
 Three Dimensional Dosimetry Analyses In Radionuclide Therapy Using IDL And MCNP-based Software Tools M. G. Stabin 1, H. Yoriyaz 2, R. Li 1, A. B. Brill 1 1 Vanderbilt University, Nashville, TN, USA 2 Instituto
Three Dimensional Dosimetry Analyses In Radionuclide Therapy Using IDL And MCNP-based Software Tools M. G. Stabin 1, H. Yoriyaz 2, R. Li 1, A. B. Brill 1 1 Vanderbilt University, Nashville, TN, USA 2 Instituto
8/3/2017. Contour Assessment for Quality Assurance and Data Mining. Objective. Outline. Tom Purdie, PhD, MCCPM
 Contour Assessment for Quality Assurance and Data Mining Tom Purdie, PhD, MCCPM Objective Understand the state-of-the-art in contour assessment for quality assurance including data mining-based techniques
Contour Assessment for Quality Assurance and Data Mining Tom Purdie, PhD, MCCPM Objective Understand the state-of-the-art in contour assessment for quality assurance including data mining-based techniques
Deviceless respiratory motion correction in PET imaging exploring the potential of novel data driven strategies
 g Deviceless respiratory motion correction in PET imaging exploring the potential of novel data driven strategies Presented by Adam Kesner, Ph.D., DABR Assistant Professor, Division of Radiological Sciences,
g Deviceless respiratory motion correction in PET imaging exploring the potential of novel data driven strategies Presented by Adam Kesner, Ph.D., DABR Assistant Professor, Division of Radiological Sciences,
Automatic 3D Registration of Lung Surfaces in Computed Tomography Scans
 Automatic 3D Registration of Lung Surfaces in Computed Tomography Scans Margrit Betke, PhD 1, Harrison Hong, BA 1, and Jane P. Ko, MD 2 1 Computer Science Department Boston University, Boston, MA 02215,
Automatic 3D Registration of Lung Surfaces in Computed Tomography Scans Margrit Betke, PhD 1, Harrison Hong, BA 1, and Jane P. Ko, MD 2 1 Computer Science Department Boston University, Boston, MA 02215,
RADIOMICS: potential role in the clinics and challenges
 27 giugno 2018 Dipartimento di Fisica Università degli Studi di Milano RADIOMICS: potential role in the clinics and challenges Dr. Francesca Botta Medical Physicist Istituto Europeo di Oncologia (Milano)
27 giugno 2018 Dipartimento di Fisica Università degli Studi di Milano RADIOMICS: potential role in the clinics and challenges Dr. Francesca Botta Medical Physicist Istituto Europeo di Oncologia (Milano)
Diagnostic imaging techniques. Krasznai Zoltán. University of Debrecen Medical and Health Science Centre Department of Biophysics and Cell Biology
 Diagnostic imaging techniques Krasznai Zoltán University of Debrecen Medical and Health Science Centre Department of Biophysics and Cell Biology 1. Computer tomography (CT) 2. Gamma camera 3. Single Photon
Diagnostic imaging techniques Krasznai Zoltán University of Debrecen Medical and Health Science Centre Department of Biophysics and Cell Biology 1. Computer tomography (CT) 2. Gamma camera 3. Single Photon
ANALYSIS OF PULMONARY FIBROSIS IN MRI, USING AN ELASTIC REGISTRATION TECHNIQUE IN A MODEL OF FIBROSIS: Scleroderma
 ANALYSIS OF PULMONARY FIBROSIS IN MRI, USING AN ELASTIC REGISTRATION TECHNIQUE IN A MODEL OF FIBROSIS: Scleroderma ORAL DEFENSE 8 th of September 2017 Charlotte MARTIN Supervisor: Pr. MP REVEL M2 Bio Medical
ANALYSIS OF PULMONARY FIBROSIS IN MRI, USING AN ELASTIC REGISTRATION TECHNIQUE IN A MODEL OF FIBROSIS: Scleroderma ORAL DEFENSE 8 th of September 2017 Charlotte MARTIN Supervisor: Pr. MP REVEL M2 Bio Medical
A Study of Medical Image Analysis System
 Indian Journal of Science and Technology, Vol 8(25), DOI: 10.17485/ijst/2015/v8i25/80492, October 2015 ISSN (Print) : 0974-6846 ISSN (Online) : 0974-5645 A Study of Medical Image Analysis System Kim Tae-Eun
Indian Journal of Science and Technology, Vol 8(25), DOI: 10.17485/ijst/2015/v8i25/80492, October 2015 ISSN (Print) : 0974-6846 ISSN (Online) : 0974-5645 A Study of Medical Image Analysis System Kim Tae-Eun
Comparison of Vessel Segmentations using STAPLE
 Comparison of Vessel Segmentations using STAPLE Julien Jomier, Vincent LeDigarcher, and Stephen R. Aylward Computer-Aided Diagnosis and Display Lab The University of North Carolina at Chapel Hill, Department
Comparison of Vessel Segmentations using STAPLE Julien Jomier, Vincent LeDigarcher, and Stephen R. Aylward Computer-Aided Diagnosis and Display Lab The University of North Carolina at Chapel Hill, Department
Reduction of Metal Artifacts in Computed Tomographies for the Planning and Simulation of Radiation Therapy
 Reduction of Metal Artifacts in Computed Tomographies for the Planning and Simulation of Radiation Therapy T. Rohlfing a, D. Zerfowski b, J. Beier a, P. Wust a, N. Hosten a, R. Felix a a Department of
Reduction of Metal Artifacts in Computed Tomographies for the Planning and Simulation of Radiation Therapy T. Rohlfing a, D. Zerfowski b, J. Beier a, P. Wust a, N. Hosten a, R. Felix a a Department of
Skull Segmentation of MR images based on texture features for attenuation correction in PET/MR
 Skull Segmentation of MR images based on texture features for attenuation correction in PET/MR CHAIBI HASSEN, NOURINE RACHID ITIO Laboratory, Oran University Algeriachaibih@yahoo.fr, nourine@yahoo.com
Skull Segmentation of MR images based on texture features for attenuation correction in PET/MR CHAIBI HASSEN, NOURINE RACHID ITIO Laboratory, Oran University Algeriachaibih@yahoo.fr, nourine@yahoo.com
Automatic Subthalamic Nucleus Targeting for Deep Brain Stimulation. A Validation Study
 Automatic Subthalamic Nucleus Targeting for Deep Brain Stimulation. A Validation Study F. Javier Sánchez Castro a, Claudio Pollo a,b, Jean-Guy Villemure b, Jean-Philippe Thiran a a École Polytechnique
Automatic Subthalamic Nucleus Targeting for Deep Brain Stimulation. A Validation Study F. Javier Sánchez Castro a, Claudio Pollo a,b, Jean-Guy Villemure b, Jean-Philippe Thiran a a École Polytechnique
Geometry Practice. 1. Angles located next to one another sharing a common side are called angles.
 Geometry Practice Name 1. Angles located next to one another sharing a common side are called angles. 2. Planes that meet to form right angles are called planes. 3. Lines that cross are called lines. 4.
Geometry Practice Name 1. Angles located next to one another sharing a common side are called angles. 2. Planes that meet to form right angles are called planes. 3. Lines that cross are called lines. 4.
An Automated Image-based Method for Multi-Leaf Collimator Positioning Verification in Intensity Modulated Radiation Therapy
 An Automated Image-based Method for Multi-Leaf Collimator Positioning Verification in Intensity Modulated Radiation Therapy Chenyang Xu 1, Siemens Corporate Research, Inc., Princeton, NJ, USA Xiaolei Huang,
An Automated Image-based Method for Multi-Leaf Collimator Positioning Verification in Intensity Modulated Radiation Therapy Chenyang Xu 1, Siemens Corporate Research, Inc., Princeton, NJ, USA Xiaolei Huang,
Utilizing Salient Region Features for 3D Multi-Modality Medical Image Registration
 Utilizing Salient Region Features for 3D Multi-Modality Medical Image Registration Dieter Hahn 1, Gabriele Wolz 2, Yiyong Sun 3, Frank Sauer 3, Joachim Hornegger 1, Torsten Kuwert 2 and Chenyang Xu 3 1
Utilizing Salient Region Features for 3D Multi-Modality Medical Image Registration Dieter Hahn 1, Gabriele Wolz 2, Yiyong Sun 3, Frank Sauer 3, Joachim Hornegger 1, Torsten Kuwert 2 and Chenyang Xu 3 1
Iterative and analytical reconstruction algorithms for varying-focal-length cone-beam
 Home Search Collections Journals About Contact us My IOPscience Iterative and analytical reconstruction algorithms for varying-focal-length cone-beam projections This content has been downloaded from IOPscience.
Home Search Collections Journals About Contact us My IOPscience Iterative and analytical reconstruction algorithms for varying-focal-length cone-beam projections This content has been downloaded from IOPscience.
Comparison of Vessel Segmentations Using STAPLE
 Comparison of Vessel Segmentations Using STAPLE Julien Jomier, Vincent LeDigarcher, and Stephen R. Aylward Computer-Aided Diagnosis and Display Lab, The University of North Carolina at Chapel Hill, Department
Comparison of Vessel Segmentations Using STAPLE Julien Jomier, Vincent LeDigarcher, and Stephen R. Aylward Computer-Aided Diagnosis and Display Lab, The University of North Carolina at Chapel Hill, Department
REMOVAL OF THE EFFECT OF COMPTON SCATTERING IN 3-D WHOLE BODY POSITRON EMISSION TOMOGRAPHY BY MONTE CARLO
 REMOVAL OF THE EFFECT OF COMPTON SCATTERING IN 3-D WHOLE BODY POSITRON EMISSION TOMOGRAPHY BY MONTE CARLO Abstract C.S. Levin, Y-C Tai, E.J. Hoffman, M. Dahlbom, T.H. Farquhar UCLA School of Medicine Division
REMOVAL OF THE EFFECT OF COMPTON SCATTERING IN 3-D WHOLE BODY POSITRON EMISSION TOMOGRAPHY BY MONTE CARLO Abstract C.S. Levin, Y-C Tai, E.J. Hoffman, M. Dahlbom, T.H. Farquhar UCLA School of Medicine Division
2D-3D Registration using Gradient-based MI for Image Guided Surgery Systems
 2D-3D Registration using Gradient-based MI for Image Guided Surgery Systems Yeny Yim 1*, Xuanyi Chen 1, Mike Wakid 1, Steve Bielamowicz 2, James Hahn 1 1 Department of Computer Science, The George Washington
2D-3D Registration using Gradient-based MI for Image Guided Surgery Systems Yeny Yim 1*, Xuanyi Chen 1, Mike Wakid 1, Steve Bielamowicz 2, James Hahn 1 1 Department of Computer Science, The George Washington
Mutual information based CT registration of the lung at exhale and inhale breathing states using thin-plate splines
 Mutual information based CT registration of the lung at exhale and inhale breathing states using thin-plate splines Martha M. Coselmon, a) James M. Balter, Daniel L. McShan, and Marc L. Kessler Department
Mutual information based CT registration of the lung at exhale and inhale breathing states using thin-plate splines Martha M. Coselmon, a) James M. Balter, Daniel L. McShan, and Marc L. Kessler Department
Advanced Image Reconstruction Methods for Photoacoustic Tomography
 Advanced Image Reconstruction Methods for Photoacoustic Tomography Mark A. Anastasio, Kun Wang, and Robert Schoonover Department of Biomedical Engineering Washington University in St. Louis 1 Outline Photoacoustic/thermoacoustic
Advanced Image Reconstruction Methods for Photoacoustic Tomography Mark A. Anastasio, Kun Wang, and Robert Schoonover Department of Biomedical Engineering Washington University in St. Louis 1 Outline Photoacoustic/thermoacoustic
Overview of Proposed TG-132 Recommendations
 Overview of Proposed TG-132 Recommendations Kristy K Brock, Ph.D., DABR Associate Professor Department of Radiation Oncology, University of Michigan Chair, AAPM TG 132: Image Registration and Fusion Conflict
Overview of Proposed TG-132 Recommendations Kristy K Brock, Ph.D., DABR Associate Professor Department of Radiation Oncology, University of Michigan Chair, AAPM TG 132: Image Registration and Fusion Conflict
Image-based Monte Carlo calculations for dosimetry
 Image-based Monte Carlo calculations for dosimetry Irène Buvat Imagerie et Modélisation en Neurobiologie et Cancérologie UMR 8165 CNRS Universités Paris 7 et Paris 11 Orsay, France buvat@imnc.in2p3.fr
Image-based Monte Carlo calculations for dosimetry Irène Buvat Imagerie et Modélisation en Neurobiologie et Cancérologie UMR 8165 CNRS Universités Paris 7 et Paris 11 Orsay, France buvat@imnc.in2p3.fr
Computed tomography of simple objects. Related topics. Principle. Equipment TEP Beam hardening, artefacts, and algorithms
 Related topics Beam hardening, artefacts, and algorithms Principle The CT principle is demonstrated with the aid of simple objects. In the case of very simple targets, only a few images need to be taken
Related topics Beam hardening, artefacts, and algorithms Principle The CT principle is demonstrated with the aid of simple objects. In the case of very simple targets, only a few images need to be taken
Automatic Vascular Tree Formation Using the Mahalanobis Distance
 Automatic Vascular Tree Formation Using the Mahalanobis Distance Julien Jomier, Vincent LeDigarcher, and Stephen R. Aylward Computer-Aided Diagnosis and Display Lab, Department of Radiology The University
Automatic Vascular Tree Formation Using the Mahalanobis Distance Julien Jomier, Vincent LeDigarcher, and Stephen R. Aylward Computer-Aided Diagnosis and Display Lab, Department of Radiology The University
Computed tomography (Item No.: P )
 Computed tomography (Item No.: P2550100) Curricular Relevance Area of Expertise: Biology Education Level: University Topic: Modern Imaging Methods Subtopic: X-ray Imaging Experiment: Computed tomography
Computed tomography (Item No.: P2550100) Curricular Relevance Area of Expertise: Biology Education Level: University Topic: Modern Imaging Methods Subtopic: X-ray Imaging Experiment: Computed tomography
Determination of rotations in three dimensions using two-dimensional portal image registration
 Determination of rotations in three dimensions using two-dimensional portal image registration Anthony E. Lujan, a) James M. Balter, and Randall K. Ten Haken Department of Nuclear Engineering and Radiological
Determination of rotations in three dimensions using two-dimensional portal image registration Anthony E. Lujan, a) James M. Balter, and Randall K. Ten Haken Department of Nuclear Engineering and Radiological
MEDICAL EQUIPMENT: COMPUTED TOMOGRAPHY. Prof. Yasser Mostafa Kadah
 MEDICAL EQUIPMENT: COMPUTED TOMOGRAPHY Prof. Yasser Mostafa Kadah www.k-space.org Recommended Textbook X-Ray Computed Tomography in Biomedical Engineering, by Robert Cierniak, Springer, 211 Computed Tomography
MEDICAL EQUIPMENT: COMPUTED TOMOGRAPHY Prof. Yasser Mostafa Kadah www.k-space.org Recommended Textbook X-Ray Computed Tomography in Biomedical Engineering, by Robert Cierniak, Springer, 211 Computed Tomography
Fiber Selection from Diffusion Tensor Data based on Boolean Operators
 Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhof 1, G. Greiner 2, M. Buchfelder 3, C. Nimsky 4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhof 1, G. Greiner 2, M. Buchfelder 3, C. Nimsky 4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Spiral CT. Protocol Optimization & Quality Assurance. Ge Wang, Ph.D. Department of Radiology University of Iowa Iowa City, Iowa 52242, USA
 Spiral CT Protocol Optimization & Quality Assurance Ge Wang, Ph.D. Department of Radiology University of Iowa Iowa City, Iowa 52242, USA Spiral CT Protocol Optimization & Quality Assurance Protocol optimization
Spiral CT Protocol Optimization & Quality Assurance Ge Wang, Ph.D. Department of Radiology University of Iowa Iowa City, Iowa 52242, USA Spiral CT Protocol Optimization & Quality Assurance Protocol optimization
Interactive Boundary Detection for Automatic Definition of 2D Opacity Transfer Function
 Interactive Boundary Detection for Automatic Definition of 2D Opacity Transfer Function Martin Rauberger, Heinrich Martin Overhoff Medical Engineering Laboratory, University of Applied Sciences Gelsenkirchen,
Interactive Boundary Detection for Automatic Definition of 2D Opacity Transfer Function Martin Rauberger, Heinrich Martin Overhoff Medical Engineering Laboratory, University of Applied Sciences Gelsenkirchen,
Key words: automated registration, voxel similarity, multiresolution optimization, magnetic resonance, positron emission tomography
 Automated three-dimensional registration of magnetic resonance and positron emission tomography brain images by multiresolution optimization of voxel similarity measures Colin Studholme, a) Derek L. G.
Automated three-dimensional registration of magnetic resonance and positron emission tomography brain images by multiresolution optimization of voxel similarity measures Colin Studholme, a) Derek L. G.
Medical Imaging BMEN Spring 2016
 Name Medical Imaging BMEN 420-501 Spring 2016 Homework #4 and Nuclear Medicine Notes All questions are from the introductory Powerpoint (based on Chapter 7) and text Medical Imaging Signals and Systems,
Name Medical Imaging BMEN 420-501 Spring 2016 Homework #4 and Nuclear Medicine Notes All questions are from the introductory Powerpoint (based on Chapter 7) and text Medical Imaging Signals and Systems,
UvA-DARE (Digital Academic Repository) Motion compensation for 4D PET/CT Kruis, M.F. Link to publication
 UvA-DARE (Digital Academic Repository) Motion compensation for 4D PET/CT Kruis, M.F. Link to publication Citation for published version (APA): Kruis, M. F. (2014). Motion compensation for 4D PET/CT General
UvA-DARE (Digital Academic Repository) Motion compensation for 4D PET/CT Kruis, M.F. Link to publication Citation for published version (APA): Kruis, M. F. (2014). Motion compensation for 4D PET/CT General
Validation of GEANT4 for Accurate Modeling of 111 In SPECT Acquisition
 Validation of GEANT4 for Accurate Modeling of 111 In SPECT Acquisition Bernd Schweizer, Andreas Goedicke Philips Technology Research Laboratories, Aachen, Germany bernd.schweizer@philips.com Abstract.
Validation of GEANT4 for Accurate Modeling of 111 In SPECT Acquisition Bernd Schweizer, Andreas Goedicke Philips Technology Research Laboratories, Aachen, Germany bernd.schweizer@philips.com Abstract.
Design and performance characteristics of a Cone Beam CT system for Leksell Gamma Knife Icon
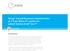 Design and performance characteristics of a Cone Beam CT system for Leksell Gamma Knife Icon WHITE PAPER Introduction Introducing an image guidance system based on Cone Beam CT (CBCT) and a mask immobilization
Design and performance characteristics of a Cone Beam CT system for Leksell Gamma Knife Icon WHITE PAPER Introduction Introducing an image guidance system based on Cone Beam CT (CBCT) and a mask immobilization
3D Volume Mesh Generation of Human Organs Using Surface Geometries Created from the Visible Human Data Set
 3D Volume Mesh Generation of Human Organs Using Surface Geometries Created from the Visible Human Data Set John M. Sullivan, Jr., Ziji Wu, and Anand Kulkarni Worcester Polytechnic Institute Worcester,
3D Volume Mesh Generation of Human Organs Using Surface Geometries Created from the Visible Human Data Set John M. Sullivan, Jr., Ziji Wu, and Anand Kulkarni Worcester Polytechnic Institute Worcester,
Dynamic digital phantoms
 Dynamic digital phantoms In radiation research the term phantom is used to describe an inanimate object or system used to tune the performance of radiation imaging or radiotherapeutic devices. A wide range
Dynamic digital phantoms In radiation research the term phantom is used to describe an inanimate object or system used to tune the performance of radiation imaging or radiotherapeutic devices. A wide range
SPECT QA and QC. Bruce McBride St. Vincent s Hospital Sydney.
 SPECT QA and QC Bruce McBride St. Vincent s Hospital Sydney. SPECT QA and QC What is needed? Why? How often? Who says? QA and QC in Nuclear Medicine QA - collective term for all the efforts made to produce
SPECT QA and QC Bruce McBride St. Vincent s Hospital Sydney. SPECT QA and QC What is needed? Why? How often? Who says? QA and QC in Nuclear Medicine QA - collective term for all the efforts made to produce
VALIDATION OF DIR. Raj Varadhan, PhD, DABMP Minneapolis Radiation Oncology
 VALIDATION OF DIR Raj Varadhan, PhD, DABMP Minneapolis Radiation Oncology Overview Basics: Registration Framework, Theory Discuss Validation techniques Using Synthetic CT data & Phantoms What metrics to
VALIDATION OF DIR Raj Varadhan, PhD, DABMP Minneapolis Radiation Oncology Overview Basics: Registration Framework, Theory Discuss Validation techniques Using Synthetic CT data & Phantoms What metrics to
PROSTATE CANCER DETECTION USING LABEL IMAGE CONSTRAINED MULTIATLAS SELECTION
 PROSTATE CANCER DETECTION USING LABEL IMAGE CONSTRAINED MULTIATLAS SELECTION Ms. Vaibhavi Nandkumar Jagtap 1, Mr. Santosh D. Kale 2 1 PG Scholar, 2 Assistant Professor, Department of Electronics and Telecommunication,
PROSTATE CANCER DETECTION USING LABEL IMAGE CONSTRAINED MULTIATLAS SELECTION Ms. Vaibhavi Nandkumar Jagtap 1, Mr. Santosh D. Kale 2 1 PG Scholar, 2 Assistant Professor, Department of Electronics and Telecommunication,
coding of various parts showing different features, the possibility of rotation or of hiding covering parts of the object's surface to gain an insight
 Three-Dimensional Object Reconstruction from Layered Spatial Data Michael Dangl and Robert Sablatnig Vienna University of Technology, Institute of Computer Aided Automation, Pattern Recognition and Image
Three-Dimensional Object Reconstruction from Layered Spatial Data Michael Dangl and Robert Sablatnig Vienna University of Technology, Institute of Computer Aided Automation, Pattern Recognition and Image
Automatic Cerebral Aneurysm Detection in Multimodal Angiographic Images
 Automatic Cerebral Aneurysm Detection in Multimodal Angiographic Images Clemens M. Hentschke, Oliver Beuing, Rosa Nickl and Klaus D. Tönnies Abstract We propose a system to automatically detect cerebral
Automatic Cerebral Aneurysm Detection in Multimodal Angiographic Images Clemens M. Hentschke, Oliver Beuing, Rosa Nickl and Klaus D. Tönnies Abstract We propose a system to automatically detect cerebral
PET-CT in Radiation Treatment Planning
 PET-CT in Radiation Treatment Planning TA van de Water, Radiotherapeutic Institute Friesland, Leeuwarden JA van Dalen, Isala, Zwolle ACKNOWLEDGEMENTS A. van der Schaaf (University of Groningen, University
PET-CT in Radiation Treatment Planning TA van de Water, Radiotherapeutic Institute Friesland, Leeuwarden JA van Dalen, Isala, Zwolle ACKNOWLEDGEMENTS A. van der Schaaf (University of Groningen, University
REAL-TIME ADAPTIVITY IN HEAD-AND-NECK AND LUNG CANCER RADIOTHERAPY IN A GPU ENVIRONMENT
 REAL-TIME ADAPTIVITY IN HEAD-AND-NECK AND LUNG CANCER RADIOTHERAPY IN A GPU ENVIRONMENT Anand P Santhanam Assistant Professor, Department of Radiation Oncology OUTLINE Adaptive radiotherapy for head and
REAL-TIME ADAPTIVITY IN HEAD-AND-NECK AND LUNG CANCER RADIOTHERAPY IN A GPU ENVIRONMENT Anand P Santhanam Assistant Professor, Department of Radiation Oncology OUTLINE Adaptive radiotherapy for head and
Shadow casting. What is the problem? Cone Beam Computed Tomography THE OBJECTIVES OF DIAGNOSTIC IMAGING IDEAL DIAGNOSTIC IMAGING STUDY LIMITATIONS
 Cone Beam Computed Tomography THE OBJECTIVES OF DIAGNOSTIC IMAGING Reveal pathology Reveal the anatomic truth Steven R. Singer, DDS srs2@columbia.edu IDEAL DIAGNOSTIC IMAGING STUDY Provides desired diagnostic
Cone Beam Computed Tomography THE OBJECTIVES OF DIAGNOSTIC IMAGING Reveal pathology Reveal the anatomic truth Steven R. Singer, DDS srs2@columbia.edu IDEAL DIAGNOSTIC IMAGING STUDY Provides desired diagnostic
Performance Evaluation of radionuclide imaging systems
 Performance Evaluation of radionuclide imaging systems Nicolas A. Karakatsanis STIR Users meeting IEEE Nuclear Science Symposium and Medical Imaging Conference 2009 Orlando, FL, USA Geant4 Application
Performance Evaluation of radionuclide imaging systems Nicolas A. Karakatsanis STIR Users meeting IEEE Nuclear Science Symposium and Medical Imaging Conference 2009 Orlando, FL, USA Geant4 Application
PET/CT multimodality imaging for radiotherapy planning in lung cancer The medical physicist point of view Isabelle Gardin Rouen CHB and Quant.I.
 PET/CT multimodality imaging for radiotherapy planning in lung cancer The medical physicist point of view Isabelle Gardin Rouen CHB and Quant.I.F (EA4108 FR CNRS 3638) Outline Specific acquisition conditions
PET/CT multimodality imaging for radiotherapy planning in lung cancer The medical physicist point of view Isabelle Gardin Rouen CHB and Quant.I.F (EA4108 FR CNRS 3638) Outline Specific acquisition conditions
Is deformable image registration a solved problem?
 Is deformable image registration a solved problem? Marcel van Herk On behalf of the imaging group of the RT department of NKI/AVL Amsterdam, the Netherlands DIR 1 Image registration Find translation.deformation
Is deformable image registration a solved problem? Marcel van Herk On behalf of the imaging group of the RT department of NKI/AVL Amsterdam, the Netherlands DIR 1 Image registration Find translation.deformation
Good Morning! Thank you for joining us
 Good Morning! Thank you for joining us Deformable Registration, Contour Propagation and Dose Mapping: 101 and 201 Marc Kessler, PhD, FAAPM The University of Michigan Conflict of Interest I receive direct
Good Morning! Thank you for joining us Deformable Registration, Contour Propagation and Dose Mapping: 101 and 201 Marc Kessler, PhD, FAAPM The University of Michigan Conflict of Interest I receive direct
Moore Catholic High School Math Department
 Moore Catholic High School Math Department Geometry Vocabulary The following is a list of terms and properties which are necessary for success in a Geometry class. You will be tested on these terms during
Moore Catholic High School Math Department Geometry Vocabulary The following is a list of terms and properties which are necessary for success in a Geometry class. You will be tested on these terms during
A Registration-Based Atlas Propagation Framework for Automatic Whole Heart Segmentation
 A Registration-Based Atlas Propagation Framework for Automatic Whole Heart Segmentation Xiahai Zhuang (PhD) Centre for Medical Image Computing University College London Fields-MITACS Conference on Mathematics
A Registration-Based Atlas Propagation Framework for Automatic Whole Heart Segmentation Xiahai Zhuang (PhD) Centre for Medical Image Computing University College London Fields-MITACS Conference on Mathematics
Multimodality image integration for radiotherapy treatment, an easy approach
 Multimodality image integration for radiotherapy treatment, an easy approach ASantos a, J Pascau b,mdesco b,jasantos b,fcalvo b,cbenito b, P Garcia-Barreno b a Universidad Politécnica de Madrid, E-28040
Multimodality image integration for radiotherapy treatment, an easy approach ASantos a, J Pascau b,mdesco b,jasantos b,fcalvo b,cbenito b, P Garcia-Barreno b a Universidad Politécnica de Madrid, E-28040
Virtual Phantoms for IGRT QA
 TM Virtual Phantoms for IGRT QA Why ImSimQA? ImSimQA was developed to overcome the limitations of physical phantoms for testing modern medical imaging and radiation therapy software systems, when there
TM Virtual Phantoms for IGRT QA Why ImSimQA? ImSimQA was developed to overcome the limitations of physical phantoms for testing modern medical imaging and radiation therapy software systems, when there
ADVANCING CANCER TREATMENT
 3 ADVANCING CANCER TREATMENT SUPPORTING CLINICS WORLDWIDE RaySearch is advancing cancer treatment through pioneering software. We believe software has un limited potential, and that it is now the driving
3 ADVANCING CANCER TREATMENT SUPPORTING CLINICS WORLDWIDE RaySearch is advancing cancer treatment through pioneering software. We believe software has un limited potential, and that it is now the driving
Automated Image Analysis Software for Quality Assurance of a Radiotherapy CT Simulator
 Automated Image Analysis Software for Quality Assurance of a Radiotherapy CT Simulator Andrew J Reilly Imaging Physicist Oncology Physics Edinburgh Cancer Centre Western General Hospital EDINBURGH EH4
Automated Image Analysis Software for Quality Assurance of a Radiotherapy CT Simulator Andrew J Reilly Imaging Physicist Oncology Physics Edinburgh Cancer Centre Western General Hospital EDINBURGH EH4
Neurosurgery for functional disorders guided by multimodality imaging
 Neurosurgery for functional disorders guided by multimodality imaging ASantos a, J Pascau b,mdesco b,cbenito b, F García-Salazar b, R Carrillo b, P Garcia-Barreno b a Universidad Politécnica de Madrid,
Neurosurgery for functional disorders guided by multimodality imaging ASantos a, J Pascau b,mdesco b,cbenito b, F García-Salazar b, R Carrillo b, P Garcia-Barreno b a Universidad Politécnica de Madrid,
Image Acquisition Systems
 Image Acquisition Systems Goals and Terminology Conventional Radiography Axial Tomography Computer Axial Tomography (CAT) Magnetic Resonance Imaging (MRI) PET, SPECT Ultrasound Microscopy Imaging ITCS
Image Acquisition Systems Goals and Terminology Conventional Radiography Axial Tomography Computer Axial Tomography (CAT) Magnetic Resonance Imaging (MRI) PET, SPECT Ultrasound Microscopy Imaging ITCS
COMPARISON OF DOSE CALCULATION ALGORITHMS FOR LEKSELL GAMMA KNIFE PERFEXION USING MONTE CARLO VOXEL PHANTOMS
 COMPARISON OF DOSE CALCULATION ALGORITHMS FOR LEKSELL GAMMA KNIFE PERFEXION USING MONTE CARLO VOXEL PHANTOMS Jan Pipek 1, Josef Novotný Jr. 1,2,3, Josef Novotný 1, Petra Kozubíková 1 1 Faculty of Nuclear
COMPARISON OF DOSE CALCULATION ALGORITHMS FOR LEKSELL GAMMA KNIFE PERFEXION USING MONTE CARLO VOXEL PHANTOMS Jan Pipek 1, Josef Novotný Jr. 1,2,3, Josef Novotný 1, Petra Kozubíková 1 1 Faculty of Nuclear
MEDICAL IMAGE ANALYSIS
 SECOND EDITION MEDICAL IMAGE ANALYSIS ATAM P. DHAWAN g, A B IEEE Engineering in Medicine and Biology Society, Sponsor IEEE Press Series in Biomedical Engineering Metin Akay, Series Editor +IEEE IEEE PRESS
SECOND EDITION MEDICAL IMAGE ANALYSIS ATAM P. DHAWAN g, A B IEEE Engineering in Medicine and Biology Society, Sponsor IEEE Press Series in Biomedical Engineering Metin Akay, Series Editor +IEEE IEEE PRESS
ML reconstruction for CT
 ML reconstruction for CT derivation of MLTR rigid motion correction resolution modeling polychromatic ML model dual energy ML model Bruno De Man, Katrien Van Slambrouck, Maarten Depypere, Frederik Maes,
ML reconstruction for CT derivation of MLTR rigid motion correction resolution modeling polychromatic ML model dual energy ML model Bruno De Man, Katrien Van Slambrouck, Maarten Depypere, Frederik Maes,
Multi-modal Image Registration Using the Generalized Survival Exponential Entropy
 Multi-modal Image Registration Using the Generalized Survival Exponential Entropy Shu Liao and Albert C.S. Chung Lo Kwee-Seong Medical Image Analysis Laboratory, Department of Computer Science and Engineering,
Multi-modal Image Registration Using the Generalized Survival Exponential Entropy Shu Liao and Albert C.S. Chung Lo Kwee-Seong Medical Image Analysis Laboratory, Department of Computer Science and Engineering,
Respiratory Motion Compensation for Simultaneous PET/MR Based on Strongly Undersampled Radial MR Data
 Respiratory Motion Compensation for Simultaneous PET/MR Based on Strongly Undersampled Radial MR Data Christopher M Rank 1, Thorsten Heußer 1, Andreas Wetscherek 1, and Marc Kachelrieß 1 1 German Cancer
Respiratory Motion Compensation for Simultaneous PET/MR Based on Strongly Undersampled Radial MR Data Christopher M Rank 1, Thorsten Heußer 1, Andreas Wetscherek 1, and Marc Kachelrieß 1 1 German Cancer
COMPARATIVE STUDIES OF DIFFERENT SYSTEM MODELS FOR ITERATIVE CT IMAGE RECONSTRUCTION
 COMPARATIVE STUDIES OF DIFFERENT SYSTEM MODELS FOR ITERATIVE CT IMAGE RECONSTRUCTION BY CHUANG MIAO A Thesis Submitted to the Graduate Faculty of WAKE FOREST UNIVERSITY GRADUATE SCHOOL OF ARTS AND SCIENCES
COMPARATIVE STUDIES OF DIFFERENT SYSTEM MODELS FOR ITERATIVE CT IMAGE RECONSTRUCTION BY CHUANG MIAO A Thesis Submitted to the Graduate Faculty of WAKE FOREST UNIVERSITY GRADUATE SCHOOL OF ARTS AND SCIENCES
A software tool for the quantitative evaluation of 3D dose calculation algorithms
 A software tool for the quantitative evaluation of 3D dose calculation algorithms William B. Harms, Sr., Daniel A. Low, John W. Wong, a) and James A. Purdy Washington University School of Medicine, Mallinckrodt
A software tool for the quantitative evaluation of 3D dose calculation algorithms William B. Harms, Sr., Daniel A. Low, John W. Wong, a) and James A. Purdy Washington University School of Medicine, Mallinckrodt
Whole Body MRI Intensity Standardization
 Whole Body MRI Intensity Standardization Florian Jäger 1, László Nyúl 1, Bernd Frericks 2, Frank Wacker 2 and Joachim Hornegger 1 1 Institute of Pattern Recognition, University of Erlangen, {jaeger,nyul,hornegger}@informatik.uni-erlangen.de
Whole Body MRI Intensity Standardization Florian Jäger 1, László Nyúl 1, Bernd Frericks 2, Frank Wacker 2 and Joachim Hornegger 1 1 Institute of Pattern Recognition, University of Erlangen, {jaeger,nyul,hornegger}@informatik.uni-erlangen.de
Adaptive Local Multi-Atlas Segmentation: Application to Heart Segmentation in Chest CT Scans
 Adaptive Local Multi-Atlas Segmentation: Application to Heart Segmentation in Chest CT Scans Eva M. van Rikxoort, Ivana Isgum, Marius Staring, Stefan Klein and Bram van Ginneken Image Sciences Institute,
Adaptive Local Multi-Atlas Segmentation: Application to Heart Segmentation in Chest CT Scans Eva M. van Rikxoort, Ivana Isgum, Marius Staring, Stefan Klein and Bram van Ginneken Image Sciences Institute,
INTRODUCTION TO MEDICAL IMAGING- 3D LOCALIZATION LAB MANUAL 1. Modifications for P551 Fall 2013 Medical Physics Laboratory
 INTRODUCTION TO MEDICAL IMAGING- 3D LOCALIZATION LAB MANUAL 1 Modifications for P551 Fall 2013 Medical Physics Laboratory Introduction Following the introductory lab 0, this lab exercise the student through
INTRODUCTION TO MEDICAL IMAGING- 3D LOCALIZATION LAB MANUAL 1 Modifications for P551 Fall 2013 Medical Physics Laboratory Introduction Following the introductory lab 0, this lab exercise the student through
arxiv: v1 [cs.cv] 6 Jun 2017
![arxiv: v1 [cs.cv] 6 Jun 2017 arxiv: v1 [cs.cv] 6 Jun 2017](/thumbs/96/128307097.jpg) Volume Calculation of CT lung Lesions based on Halton Low-discrepancy Sequences Liansheng Wang a, Shusheng Li a, and Shuo Li b a Department of Computer Science, Xiamen University, Xiamen, China b Dept.
Volume Calculation of CT lung Lesions based on Halton Low-discrepancy Sequences Liansheng Wang a, Shusheng Li a, and Shuo Li b a Department of Computer Science, Xiamen University, Xiamen, China b Dept.
Estimating 3D Respiratory Motion from Orbiting Views
 Estimating 3D Respiratory Motion from Orbiting Views Rongping Zeng, Jeffrey A. Fessler, James M. Balter The University of Michigan Oct. 2005 Funding provided by NIH Grant P01 CA59827 Motivation Free-breathing
Estimating 3D Respiratory Motion from Orbiting Views Rongping Zeng, Jeffrey A. Fessler, James M. Balter The University of Michigan Oct. 2005 Funding provided by NIH Grant P01 CA59827 Motivation Free-breathing
