Sequence Analysis '17: lecture 4. Molecular evolution Global, semi-global and local Affine gap penalty
|
|
|
- Anna Booth
- 6 years ago
- Views:
Transcription
1 Sequence Analysis '17: lecture 4 Molecular evolution Global, semi-global and local Affine gap penalty
2 How sequences evolve point mutations (single base changes) deletion (loss of residues within the sequence) insertion (gain of residue within the sequence) truncation (loss of either end) extension (gain of residues at either end) Mechanisms of insertion or extension: duplication or whole gene or domain polymerase "stutter" transposable element more??
3 How evolution is scored point mutations... substitution matrix deletion... gap penalty insertion... gap penalty truncation... end gap penalty extension... end gap penalty
4 Yes, an alignment algorithm is really A Model for Sequence Evolution! The way we do alignment should model how sequences evolve. How much evolutionary time is represented by... point mutations... relatively frequent, usually bad deletion... infrequent, always bad, location dependent insertion... infrequent, always bad, location dependent truncation... frequent, not so bad extension... frequent, not so bad
5 Extension/truncation, domains : end gaps Example: here is an alignment of mouse nitric oxide synthase (thick black line). It has multiple domains which are homologous to several shorter proteins. If we penalize end gaps, what happens to the score of the true alignment? Did "end gaps" evolve the same way as internal gaps? (no!) Unless the two proteins are known to be single domains, it makes more sense NOT to penalize end gaps.
6 A structure-based multiple sequence alignment : fluorescent proteins. Note the... ragged N-terminal edge, small number of indels, large number of point substitutions distributed unevenly over the sequence. Note conserved regions, vs. hot spots. 6
7 To make a ragged edge: don't penalize end gaps First: To ignore starting gap penalties, set gap rows to zero (keep the traceback arrows). G T T C A G C T T T C A C T
8 How to NOT penalize end gaps Second: To ignore ending gap penalty, start the traceback with the MAX score at the end of either sequence. (i.e. use last row or column) G T T C A G C T T T C A C T
9 Semi-global: no end gaps If we penalize end gaps in neither sequence, we are asking for the best alignment that contains at least two of the 4 termini. Good for identifying terminal domains in two multi-domain proteins. G T T C A G C T T T C A C T
10 Semi-global: Penalize end gaps on one side If we penalize end gaps in sequence 2 but not in sequence 1, we are asking for an alignment that contains all of sequence 2 within sequence 1. G T T C A G C T T T C A C T
11 Semi-global: Penalize end gaps on one side If we penalize end gaps in sequence 1 but not in sequence 2, we are asking for an alignment that contains all of sequence 1 within sequence 2. G T T C A G C T T T C A C T
12 Local Alignment A local alignment can start anywhere and end anywhere in the alignment matrix. start P G T S F E P A T S F M A(i,j) = MAX end AT..TSFEP...PGTSF..M end A(i-1,j-1) + S(i,j) A(i,j-1) + gap A(i-1,j) + gap + S(i,j) start is the maximum score anywhere in the matrix.
13 Local Alignment Asks for largest domain, sub-domain, or set of contiguous domains that are in common between two sequences. Worst score is always zero () for no alignment It's possible to have multiple local alignments between two proteins More appropriate than global or semiglobal when there are no assumptions about the sequence relationship. Used for database searches. Required to obtain e-values (stay tuned)
14 Global, semi-global, and local alignment The choice of alignment method makes a statement about how the sequences are related. Was one sequence inserted into the other? Global alignment (end gaps) requires that all 4 termini are counted. In general, the two sequences are about the same length. Semi-global (no end gaps in 1 or both seqs) requires that one of the two sequences be completely contained in the other or that 2 or the 4 the termini be included. Local alignment finds subsequences in both. Does not require that the termini be included in the alignment.
15 The optimal alignment may be no alignment If the maximum score in the alignment matrix is <., then the optimal local alignment has score = and looks like this: ATSFM~~~~~~~ ~~~~~PGTSFEP
16 A structure-based multiple sequence alignment : fluorescent proteins. Note the... lengths of gaps Longer gaps are not so much less frequent than shorter ones. But very long gaps are rare. 16
17 Affine gap penalty-- theory Each gap represents an evolutionary event (duplication, polymerase stutter, deletion/ligation, etc.) If the alignment has "evolutionary distance" meaning, then the gap penalty score should be proportional to the number of gaps. Q: Are long gaps proportionally less likely? A: Less likely, yes, but proportionally, no.
18 Which alignment is intuitively better? AGGCTACT~T~TCA GGCTACTATATCA AGGCTACTTT~~CA GGCTACTATATCA
19 Linear versus Affine gap penalty. Making alignments agree with biological intuition. linear gap penalty Gap penalty for the whole sequence is just the total number of gap characters times a constant. affine gap penalty initiation length of gap length of gap extension Gap penalty for the whole sequence is the function. N*(gap initiation penalty) + E*(gap extension penalty) where N is the number of gap initiation characters, E is the number of gap extension characters
20 Example: affine gap gap initiation = -5 gap extension = AGGCTACT~T~TCA GGCTACTATATCA AGGCTACTTT~~CA GGCTACTATATCA -6
21 Affine Gap DP, two ways. Either this You can have 5 types of arrows, instead of just three. (1) Match (2) Open a gap in first sequence. (3) Open a gap in second sequence. (4) Extend a gap in first sequence. (5) Extend a gap in second sequence. or this Variable length arrows. no fixed number. Arrows carry length info.
22 Affine gap DP algorithm using variable length arrows A K L K L D Q F G P A D P Q F G S i,j = max { S i-1,j-1 + s(i,j), n S i-1-n,j-1 +s(i,j) - ginit - (n-1) gext, S i-1,j-1-n +s(i,j) - ginit - (n-1) gext }...where s(i,j) is the substiution score, n is the length of the gap, ginit is the gap initiation penalty, and gext is the gap extension penalty. Notes: All arrows end in match. Gap-to-gap not possible. Local or semi-global only. End-gaps not scored. Arrows still translate to an alignment. Still optimal.
23 Traceback for linear gap penalty Each arrow advances either one sequence or both, by 1. Each column has one arrow. A D P Q F G A ~ ~ ~ ~ D P Q F G ~ A K L K L D ~ Q F G P A K L K L D Q F G P
24 Traceback for affine gap DP with variable length arrows Each arrow advances one sequence by 1, the other sequence by n. Output of one arrow is n columns. n = arrowlength. Number of arrows is number of columns. arrowlength A ~ ~ ~ ~ D P Q F G ~ A K L K L D ~ Q F G P A K L K L D Q F G P A D P Q F G End gaps are ignored in local alignment
25 Does deletion-to-insertion make sense??? Special rules may apply for going from I to D and D to I. AGGCTACT~TATCA GGCTACTA~ATCA If you think this alignment does not make sense, then D to I and I to D can simply be disallowed in the DP algorithm. Most programs do this. [Exception: For a global alignment, D-to-I or I-to-D arrows are allowed at the ends of alignments because there is no other way to complete the matrix.]
26 If it doesn t make biological sense, your program shouldn t do it! 26
27 Q: How do I know what the gap penalties should be? A: Optimize the gap penalties against gold standard alignments. 27
28 The gold standard for alignments: BAliBase A database of curated multiple sequence alignments derived from structure-based alignments. 28
29 Example of a "gold standard alignment A structure-based alignment. Based on D structures. Two similar structures may be superimposed 3D. The parts that overlay well are the matches (purple and green), and the parts that do not overlay well are the insertions (yellow) and deletions (red). Aligned positions have similar chemical 3D environment. Unaligned positions deviate in 3D, therefore their environment is different.
30 Jargon word: indel Since deletions are just the flip-side of insertions, often we want a word that lumps both insertions and deletions together -- indel. Say it when you don't care whether it's an insertion or a deletion. Its another word for gap. 3
31 Example of a "gold standard alignment A structure-based alignment is a sequence alignment that comes from a protein structure superposition. 2DRC:A 1/2 MISLIAALAVDRVIGMENAM-PFNLPADLAWFKRNTL DKPVIMGRHTWESIG- 1DRF:_ 3/4 SLNCIVAVSQNMGIGKNGDLPWPPLRNEFRYFQRMTTTSSVEGKQNLVIMGKKTWFSIPE 2DRC:A 52/53 --RPLPGRKNIILSSQP--GTDDRVTWVKSVDEAIAACG------DVPEIMVIGGGRVYE 1DRF:_ 63/64 KNRPLKGRINLVLSRELKEPPQGAHFLSRSLDDALKLTEQPELANKVDMVWIVGGSSVYK 2DRC:A 12/13 1DRF:_ 123/124 2DRC:A 154/155 1DRF:_ 18/181 QFLPK--AQKLYLTHIDAEVEGDTHFPDYEPDDWESVF------SEFHDADAQNSHSYCF EAMNHPGHLKLFVTRIMQDFESDTFFPEIDLEKYKLLPEYPGVLSDVQEE---KGIKYKF EILERR EVYEKN What do you see? Lots of mismatches (%id=29), A few indels (8), Indels are usually > 1 residue.
32 Gap penalties should be numbers that consistently reproduce the gold standard alignments. 32
33 How we do alignment research 1. Hypothesize a scoring function that makes better alignments. 2. Implement it. 3. Train it. 4. Publish it iff it works. 33
34 How do we "train it"? ANSWER: machine learning. Ingredients for machine learning: 1) A training set 2) A test set 3) An objective function 4) A parameter space 5) A learning algorithm 34
35 Machine learning for alignment Create a database of sequence alignments (BAliBase) Divide database into Training set and Test set. Alignments must be non-redundant. No duplicates. Alignments must be representative. Enough of them that adding more won t change the results. Test set must have no overlap with Training set. Define an objective function, ε ε is defined as a function of alignments, given parameters. ε reaches an optimum as alignments approach the truth for members of the Training set. Explore parameter space (try all reasonable values) may use exhaustive search, gradient descent, Monte Carlo, bracketing, etc. Cross-validate when all training is done. Report the accuracy on a Test set not on the Training set. 35
36 Possible objective functions for machine learning an alignment function "number of gaps" metric (minimized) ε = Σ Ngap(test) - Ngap(gold std) "gold standard matches" metric (maximized) ε = ΣN(gold std aligned positions found in test alignments) 36
37 Bioinformatics research story: Bioinformatics 22(4):413-42, 26 Improved pairwise alignments of proteins in the Twilight Zone using local structure predictions Yao-ming Huang and * Christopher Bystroff Center for Bioinformatics, Dept of Biology, Rensselaer Polytechnic Institute, Troy, New York 1218 USA Objective function ε used in Huang & Bystroff: ε = total number of correct matches, based on balibase. sequence 2 Fig. 6-a sequence 1 BLOSUM4 SDM HMMSUM-D 3 HMMSUM-D 3+NS HMMSUM-D HMMSUM-D NS BAliBASE 1R69 1 SISSRVKSKRIQLGLNQAELAQKVGTTQQSIE-Q-LENGKTKRPRFLPELASALGVSVDWLLNGT 1NEQ GLKKRKLSLSALSRQFGYAPTTLANA- 1NEQ QFGYAPTTLANALERHWPKGE-QIIANALETKPEVI NEQ 13 DVIAGLKKRKLSL----SALSRQFGYAPTTLANA-LERHWPKGEQII---ANALETKPEVIWPSR 1NEQ 13 DVIAGLKKRKLSLSALSRQFGYAPTTLANALE-R HWPKGEQIIANALETKPEVIWPSR 1NEQ VIAGLKKRKLSLSALSRQFGYAPTTLA-N-ALERHWPKGEQIIANALETKPEVIWPSR-- 1NEQ 11 --RADVIAGLKKRKLSLSALSRQFGYAPTTLA-N-ALERHWPKGEQI -IANALETKPEVIWPSR 1NEQ 9 WHRADVIAGLKKRKLSLSALSRQFGYAPTTLA-N-ALER--HWPKGEQIIANALETKPEVIWPSR Different substitution matrices (left) gave different alignments when sequence similarity is in the Twilight zone (very difficult alignments). 37
38 Ex 4: optimizing gap penalty In a browser, goto to NCBI. Search Protein database for 1DRF. Save. Then 2DRC. Save. Select both. Save as Alignment. Select alignment.align using Kalign. Gop=5, Gep=8, Free end gaps (terminal gap=). No bonus. Get... G = # of indels - 8. (8 indels in gold std, see Slide 3) ID = percent identity. (mouse-over-r-clk/statistics/distance, identity, percent) Optimize ε = ID - 5 G (number of gaps should count for more than %id) What is the optimal Gop, Gep? Write the Gop, Gep, G and ID on a piece of paper with your name on it and turn it in next time. 38
39 Kalign parameter window in UGENE Causes fewer indels Causes shorter indels Forces/ignores alignmen of termini Causes more matches* *Increasing the bonus is essentially the same as decreasing the combined gap penalties. 39
40 Ex 4: exhaustive approach gap opening penalty gap extension penalty , 35% +5, 4% , 5% -8, 5% In each box record: G =# of gaps - 8, ID = % Identity
41 Ex 4: bracketing Fix Gep. Guess Gop. Align. Calculate ε. Increase Gop by 1. Calculate ε. Increase Gop by another 1. Calculate ε. Plot it. If it is going up, go the other way. If it is going down, keep going. If it goes down then up, you have bracketed the solution. ε Gop 41
42 Some things to ponder How does scoring approximate the evolutionary distance? When should you use semi-global alignment? What is a evolutionary hot spot? How is dynamic programming different for local alignment? Is the affine gap penalty more biologically relevant than a linear gap penalty? Why? Why are structure-based alignments considered the gold standard of sequence alignment? What does it mean for a deletion to follow immediately after an insertion, evolutionarilly? Structurally? 42
Bioinformatics 1: lecture 4. Followup of lecture 3? Molecular evolution Global, semi-global and local Affine gap penalty
 Bioinformatics 1: lecture 4 Followup of lecture 3? Molecular evolution Global, semi-global and local Affine gap penalty How sequences evolve point mutations (single base changes) deletion (loss of residues
Bioinformatics 1: lecture 4 Followup of lecture 3? Molecular evolution Global, semi-global and local Affine gap penalty How sequences evolve point mutations (single base changes) deletion (loss of residues
Pairwise Sequence Alignment: Dynamic Programming Algorithms. COMP Spring 2015 Luay Nakhleh, Rice University
 Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 - Spring 2015 Luay Nakhleh, Rice University DP Algorithms for Pairwise Alignment The number of all possible pairwise alignments (if
Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 - Spring 2015 Luay Nakhleh, Rice University DP Algorithms for Pairwise Alignment The number of all possible pairwise alignments (if
Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 Luay Nakhleh, Rice University
 1 Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 Luay Nakhleh, Rice University DP Algorithms for Pairwise Alignment 2 The number of all possible pairwise alignments (if gaps are allowed)
1 Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 Luay Nakhleh, Rice University DP Algorithms for Pairwise Alignment 2 The number of all possible pairwise alignments (if gaps are allowed)
Sequence alignment algorithms
 Sequence alignment algorithms Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 23 rd 27 After this lecture, you can decide when to use local and global sequence alignments
Sequence alignment algorithms Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 23 rd 27 After this lecture, you can decide when to use local and global sequence alignments
Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD
 Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD Assumptions: Biological sequences evolved by evolution. Micro scale changes: For short sequences (e.g. one
Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD Assumptions: Biological sequences evolved by evolution. Micro scale changes: For short sequences (e.g. one
Lecture Overview. Sequence search & alignment. Searching sequence databases. Sequence Alignment & Search. Goals: Motivations:
 Lecture Overview Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating
Lecture Overview Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014
 Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
Sequence Alignment & Search
 Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating the first version
Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating the first version
Computational Genomics and Molecular Biology, Fall
 Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
Algorithmic Approaches for Biological Data, Lecture #20
 Algorithmic Approaches for Biological Data, Lecture #20 Katherine St. John City University of New York American Museum of Natural History 20 April 2016 Outline Aligning with Gaps and Substitution Matrices
Algorithmic Approaches for Biological Data, Lecture #20 Katherine St. John City University of New York American Museum of Natural History 20 April 2016 Outline Aligning with Gaps and Substitution Matrices
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 04: Variations of sequence alignments http://www.pitt.edu/~mcs2/teaching/biocomp/tutorials/global.html Slides adapted from Dr. Shaojie Zhang (University
EECS730: Introduction to Bioinformatics Lecture 04: Variations of sequence alignments http://www.pitt.edu/~mcs2/teaching/biocomp/tutorials/global.html Slides adapted from Dr. Shaojie Zhang (University
Principles of Bioinformatics. BIO540/STA569/CSI660 Fall 2010
 Principles of Bioinformatics BIO540/STA569/CSI660 Fall 2010 Lecture 11 Multiple Sequence Alignment I Administrivia Administrivia The midterm examination will be Monday, October 18 th, in class. Closed
Principles of Bioinformatics BIO540/STA569/CSI660 Fall 2010 Lecture 11 Multiple Sequence Alignment I Administrivia Administrivia The midterm examination will be Monday, October 18 th, in class. Closed
Computational Molecular Biology
 Computational Molecular Biology Erwin M. Bakker Lecture 3, mainly from material by R. Shamir [2] and H.J. Hoogeboom [4]. 1 Pairwise Sequence Alignment Biological Motivation Algorithmic Aspect Recursive
Computational Molecular Biology Erwin M. Bakker Lecture 3, mainly from material by R. Shamir [2] and H.J. Hoogeboom [4]. 1 Pairwise Sequence Alignment Biological Motivation Algorithmic Aspect Recursive
Global Alignment Scoring Matrices Local Alignment Alignment with Affine Gap Penalties
 Global Alignment Scoring Matrices Local Alignment Alignment with Affine Gap Penalties From LCS to Alignment: Change the Scoring The Longest Common Subsequence (LCS) problem the simplest form of sequence
Global Alignment Scoring Matrices Local Alignment Alignment with Affine Gap Penalties From LCS to Alignment: Change the Scoring The Longest Common Subsequence (LCS) problem the simplest form of sequence
Lecture 3: February Local Alignment: The Smith-Waterman Algorithm
 CSCI1820: Sequence Alignment Spring 2017 Lecture 3: February 7 Lecturer: Sorin Istrail Scribe: Pranavan Chanthrakumar Note: LaTeX template courtesy of UC Berkeley EECS dept. Notes are also adapted from
CSCI1820: Sequence Alignment Spring 2017 Lecture 3: February 7 Lecturer: Sorin Istrail Scribe: Pranavan Chanthrakumar Note: LaTeX template courtesy of UC Berkeley EECS dept. Notes are also adapted from
Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA.
 Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA. Fasta is used to compare a protein or DNA sequence to all of the
Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA. Fasta is used to compare a protein or DNA sequence to all of the
Computational Molecular Biology
 Computational Molecular Biology Erwin M. Bakker Lecture 2 Materials used from R. Shamir [2] and H.J. Hoogeboom [4]. 1 Molecular Biology Sequences DNA A, T, C, G RNA A, U, C, G Protein A, R, D, N, C E,
Computational Molecular Biology Erwin M. Bakker Lecture 2 Materials used from R. Shamir [2] and H.J. Hoogeboom [4]. 1 Molecular Biology Sequences DNA A, T, C, G RNA A, U, C, G Protein A, R, D, N, C E,
PROTEIN MULTIPLE ALIGNMENT MOTIVATION: BACKGROUND: Marina Sirota
 Marina Sirota MOTIVATION: PROTEIN MULTIPLE ALIGNMENT To study evolution on the genetic level across a wide range of organisms, biologists need accurate tools for multiple sequence alignment of protein
Marina Sirota MOTIVATION: PROTEIN MULTIPLE ALIGNMENT To study evolution on the genetic level across a wide range of organisms, biologists need accurate tools for multiple sequence alignment of protein
BLAST MCDB 187. Friday, February 8, 13
 BLAST MCDB 187 BLAST Basic Local Alignment Sequence Tool Uses shortcut to compute alignments of a sequence against a database very quickly Typically takes about a minute to align a sequence against a database
BLAST MCDB 187 BLAST Basic Local Alignment Sequence Tool Uses shortcut to compute alignments of a sequence against a database very quickly Typically takes about a minute to align a sequence against a database
Brief review from last class
 Sequence Alignment Brief review from last class DNA is has direction, we will use only one (5 -> 3 ) and generate the opposite strand as needed. DNA is a 3D object (see lecture 1) but we will model it
Sequence Alignment Brief review from last class DNA is has direction, we will use only one (5 -> 3 ) and generate the opposite strand as needed. DNA is a 3D object (see lecture 1) but we will model it
Today s Lecture. Multiple sequence alignment. Improved scoring of pairwise alignments. Affine gap penalties Profiles
 Today s Lecture Multiple sequence alignment Improved scoring of pairwise alignments Affine gap penalties Profiles 1 The Edit Graph for a Pair of Sequences G A C G T T G A A T G A C C C A C A T G A C G
Today s Lecture Multiple sequence alignment Improved scoring of pairwise alignments Affine gap penalties Profiles 1 The Edit Graph for a Pair of Sequences G A C G T T G A A T G A C C C A C A T G A C G
Bioinformatics for Biologists
 Bioinformatics for Biologists Sequence Analysis: Part I. Pairwise alignment and database searching Fran Lewitter, Ph.D. Director Bioinformatics & Research Computing Whitehead Institute Topics to Cover
Bioinformatics for Biologists Sequence Analysis: Part I. Pairwise alignment and database searching Fran Lewitter, Ph.D. Director Bioinformatics & Research Computing Whitehead Institute Topics to Cover
FASTA. Besides that, FASTA package provides SSEARCH, an implementation of the optimal Smith- Waterman algorithm.
 FASTA INTRODUCTION Definition (by David J. Lipman and William R. Pearson in 1985) - Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence
FASTA INTRODUCTION Definition (by David J. Lipman and William R. Pearson in 1985) - Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence
Sequence Alignment. part 2
 Sequence Alignment part 2 Dynamic programming with more realistic scoring scheme Using the same initial sequences, we ll look at a dynamic programming example with a scoring scheme that selects for matches
Sequence Alignment part 2 Dynamic programming with more realistic scoring scheme Using the same initial sequences, we ll look at a dynamic programming example with a scoring scheme that selects for matches
EECS 4425: Introductory Computational Bioinformatics Fall Suprakash Datta
 EECS 4425: Introductory Computational Bioinformatics Fall 2018 Suprakash Datta datta [at] cse.yorku.ca Office: CSEB 3043 Phone: 416-736-2100 ext 77875 Course page: http://www.cse.yorku.ca/course/4425 Many
EECS 4425: Introductory Computational Bioinformatics Fall 2018 Suprakash Datta datta [at] cse.yorku.ca Office: CSEB 3043 Phone: 416-736-2100 ext 77875 Course page: http://www.cse.yorku.ca/course/4425 Many
Today s Lecture. Edit graph & alignment algorithms. Local vs global Computational complexity of pairwise alignment Multiple sequence alignment
 Today s Lecture Edit graph & alignment algorithms Smith-Waterman algorithm Needleman-Wunsch algorithm Local vs global Computational complexity of pairwise alignment Multiple sequence alignment 1 Sequence
Today s Lecture Edit graph & alignment algorithms Smith-Waterman algorithm Needleman-Wunsch algorithm Local vs global Computational complexity of pairwise alignment Multiple sequence alignment 1 Sequence
Stephen Scott.
 1 / 33 sscott@cse.unl.edu 2 / 33 Start with a set of sequences In each column, residues are homolgous Residues occupy similar positions in 3D structure Residues diverge from a common ancestral residue
1 / 33 sscott@cse.unl.edu 2 / 33 Start with a set of sequences In each column, residues are homolgous Residues occupy similar positions in 3D structure Residues diverge from a common ancestral residue
Sequence analysis Pairwise sequence alignment
 UMF11 Introduction to bioinformatics, 25 Sequence analysis Pairwise sequence alignment 1. Sequence alignment Lecturer: Marina lexandersson 12 September, 25 here are two types of sequence alignments, global
UMF11 Introduction to bioinformatics, 25 Sequence analysis Pairwise sequence alignment 1. Sequence alignment Lecturer: Marina lexandersson 12 September, 25 here are two types of sequence alignments, global
From Smith-Waterman to BLAST
 From Smith-Waterman to BLAST Jeremy Buhler July 23, 2015 Smith-Waterman is the fundamental tool that we use to decide how similar two sequences are. Isn t that all that BLAST does? In principle, it is
From Smith-Waterman to BLAST Jeremy Buhler July 23, 2015 Smith-Waterman is the fundamental tool that we use to decide how similar two sequences are. Isn t that all that BLAST does? In principle, it is
Biology 644: Bioinformatics
 Find the best alignment between 2 sequences with lengths n and m, respectively Best alignment is very dependent upon the substitution matrix and gap penalties The Global Alignment Problem tries to find
Find the best alignment between 2 sequences with lengths n and m, respectively Best alignment is very dependent upon the substitution matrix and gap penalties The Global Alignment Problem tries to find
Sequence Comparison: Dynamic Programming. Genome 373 Genomic Informatics Elhanan Borenstein
 Sequence omparison: Dynamic Programming Genome 373 Genomic Informatics Elhanan Borenstein quick review: hallenges Find the best global alignment of two sequences Find the best global alignment of multiple
Sequence omparison: Dynamic Programming Genome 373 Genomic Informatics Elhanan Borenstein quick review: hallenges Find the best global alignment of two sequences Find the best global alignment of multiple
Reconstructing long sequences from overlapping sequence fragment. Searching databases for related sequences and subsequences
 SEQUENCE ALIGNMENT ALGORITHMS 1 Why compare sequences? Reconstructing long sequences from overlapping sequence fragment Searching databases for related sequences and subsequences Storing, retrieving and
SEQUENCE ALIGNMENT ALGORITHMS 1 Why compare sequences? Reconstructing long sequences from overlapping sequence fragment Searching databases for related sequences and subsequences Storing, retrieving and
BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio CS 466 Saurabh Sinha
 BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio. 1990. CS 466 Saurabh Sinha Motivation Sequence homology to a known protein suggest function of newly sequenced protein Bioinformatics
BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio. 1990. CS 466 Saurabh Sinha Motivation Sequence homology to a known protein suggest function of newly sequenced protein Bioinformatics
CISC 889 Bioinformatics (Spring 2003) Multiple Sequence Alignment
 CISC 889 Bioinformatics (Spring 2003) Multiple Sequence Alignment Courtesy of jalview 1 Motivations Collective statistic Protein families Identification and representation of conserved sequence features
CISC 889 Bioinformatics (Spring 2003) Multiple Sequence Alignment Courtesy of jalview 1 Motivations Collective statistic Protein families Identification and representation of conserved sequence features
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 06: Multiple Sequence Alignment https://upload.wikimedia.org/wikipedia/commons/thumb/7/79/rplp0_90_clustalw_aln.gif/575px-rplp0_90_clustalw_aln.gif Slides
EECS730: Introduction to Bioinformatics Lecture 06: Multiple Sequence Alignment https://upload.wikimedia.org/wikipedia/commons/thumb/7/79/rplp0_90_clustalw_aln.gif/575px-rplp0_90_clustalw_aln.gif Slides
TCCAGGTG-GAT TGCAAGTGCG-T. Local Sequence Alignment & Heuristic Local Aligners. Review: Probabilistic Interpretation. Chance or true homology?
 Local Sequence Alignment & Heuristic Local Aligners Lectures 18 Nov 28, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20 Johnson Hall
Local Sequence Alignment & Heuristic Local Aligners Lectures 18 Nov 28, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20 Johnson Hall
Programming assignment for the course Sequence Analysis (2006)
 Programming assignment for the course Sequence Analysis (2006) Original text by John W. Romein, adapted by Bart van Houte (bart@cs.vu.nl) Introduction Please note: This assignment is only obligatory for
Programming assignment for the course Sequence Analysis (2006) Original text by John W. Romein, adapted by Bart van Houte (bart@cs.vu.nl) Introduction Please note: This assignment is only obligatory for
Outline. Sequence Alignment. Types of Sequence Alignment. Genomics & Computational Biology. Section 2. How Computers Store Information
 enomics & omputational Biology Section Lan Zhang Sep. th, Outline How omputers Store Information Sequence lignment Dot Matrix nalysis Dynamic programming lobal: NeedlemanWunsch lgorithm Local: SmithWaterman
enomics & omputational Biology Section Lan Zhang Sep. th, Outline How omputers Store Information Sequence lignment Dot Matrix nalysis Dynamic programming lobal: NeedlemanWunsch lgorithm Local: SmithWaterman
Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it.
 Sequence Alignments Overview Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it. Sequence alignment means arranging
Sequence Alignments Overview Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it. Sequence alignment means arranging
Dynamic Programming: Sequence alignment. CS 466 Saurabh Sinha
 Dynamic Programming: Sequence alignment CS 466 Saurabh Sinha DNA Sequence Comparison: First Success Story Finding sequence similarities with genes of known function is a common approach to infer a newly
Dynamic Programming: Sequence alignment CS 466 Saurabh Sinha DNA Sequence Comparison: First Success Story Finding sequence similarities with genes of known function is a common approach to infer a newly
GLOBEX Bioinformatics (Summer 2015) Multiple Sequence Alignment
 GLOBEX Bioinformatics (Summer 2015) Multiple Sequence Alignment Scoring Dynamic Programming algorithms Heuristic algorithms CLUSTAL W Courtesy of jalview Motivations Collective (or aggregate) statistic
GLOBEX Bioinformatics (Summer 2015) Multiple Sequence Alignment Scoring Dynamic Programming algorithms Heuristic algorithms CLUSTAL W Courtesy of jalview Motivations Collective (or aggregate) statistic
An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST
 An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST Alexander Chan 5075504 Biochemistry 218 Final Project An Analysis of Pairwise
An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST Alexander Chan 5075504 Biochemistry 218 Final Project An Analysis of Pairwise
Basic Local Alignment Search Tool (BLAST)
 BLAST 26.04.2018 Basic Local Alignment Search Tool (BLAST) BLAST (Altshul-1990) is an heuristic Pairwise Alignment composed by six-steps that search for local similarities. The most used access point to
BLAST 26.04.2018 Basic Local Alignment Search Tool (BLAST) BLAST (Altshul-1990) is an heuristic Pairwise Alignment composed by six-steps that search for local similarities. The most used access point to
BGGN 213 Foundations of Bioinformatics Barry Grant
 BGGN 213 Foundations of Bioinformatics Barry Grant http://thegrantlab.org/bggn213 Recap From Last Time: 25 Responses: https://tinyurl.com/bggn213-02-f17 Why ALIGNMENT FOUNDATIONS Why compare biological
BGGN 213 Foundations of Bioinformatics Barry Grant http://thegrantlab.org/bggn213 Recap From Last Time: 25 Responses: https://tinyurl.com/bggn213-02-f17 Why ALIGNMENT FOUNDATIONS Why compare biological
Profiles and Multiple Alignments. COMP 571 Luay Nakhleh, Rice University
 Profiles and Multiple Alignments COMP 571 Luay Nakhleh, Rice University Outline Profiles and sequence logos Profile hidden Markov models Aligning profiles Multiple sequence alignment by gradual sequence
Profiles and Multiple Alignments COMP 571 Luay Nakhleh, Rice University Outline Profiles and sequence logos Profile hidden Markov models Aligning profiles Multiple sequence alignment by gradual sequence
FastA & the chaining problem
 FastA & the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 1 Sources for this lecture: Lectures by Volker Heun, Daniel Huson and Knut Reinert,
FastA & the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 1 Sources for this lecture: Lectures by Volker Heun, Daniel Huson and Knut Reinert,
BLAST, Profile, and PSI-BLAST
 BLAST, Profile, and PSI-BLAST Jianlin Cheng, PhD School of Electrical Engineering and Computer Science University of Central Florida 26 Free for academic use Copyright @ Jianlin Cheng & original sources
BLAST, Profile, and PSI-BLAST Jianlin Cheng, PhD School of Electrical Engineering and Computer Science University of Central Florida 26 Free for academic use Copyright @ Jianlin Cheng & original sources
FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:
 FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:56 4001 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem
FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:56 4001 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem
Bioinformatics explained: Smith-Waterman
 Bioinformatics Explained Bioinformatics explained: Smith-Waterman May 1, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com
Bioinformatics Explained Bioinformatics explained: Smith-Waterman May 1, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com
DNA Alignment With Affine Gap Penalties
 DNA Alignment With Affine Gap Penalties Laurel Schuster Why Use Affine Gap Penalties? When aligning two DNA sequences, one goal may be to infer the mutations that made them different. Though it s impossible
DNA Alignment With Affine Gap Penalties Laurel Schuster Why Use Affine Gap Penalties? When aligning two DNA sequences, one goal may be to infer the mutations that made them different. Though it s impossible
Sequence Alignment Heuristics
 Sequence Alignment Heuristics Some slides from: Iosif Vaisman, GMU mason.gmu.edu/~mmasso/binf630alignment.ppt Serafim Batzoglu, Stanford http://ai.stanford.edu/~serafim/ Geoffrey J. Barton, Oxford Protein
Sequence Alignment Heuristics Some slides from: Iosif Vaisman, GMU mason.gmu.edu/~mmasso/binf630alignment.ppt Serafim Batzoglu, Stanford http://ai.stanford.edu/~serafim/ Geoffrey J. Barton, Oxford Protein
Multiple Sequence Alignment (MSA)
 I519 Introduction to Bioinformatics, Fall 2013 Multiple Sequence Alignment (MSA) Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Outline Multiple sequence alignment (MSA) Generalize
I519 Introduction to Bioinformatics, Fall 2013 Multiple Sequence Alignment (MSA) Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Outline Multiple sequence alignment (MSA) Generalize
Lectures by Volker Heun, Daniel Huson and Knut Reinert, in particular last years lectures
 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 4.1 Sources for this lecture Lectures by Volker Heun, Daniel Huson and Knut
4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 4.1 Sources for this lecture Lectures by Volker Heun, Daniel Huson and Knut
Multiple sequence alignment. November 20, 2018
 Multiple sequence alignment November 20, 2018 Why do multiple alignment? Gain insight into evolutionary history Can assess time of divergence by looking at the number of mutations needed to change one
Multiple sequence alignment November 20, 2018 Why do multiple alignment? Gain insight into evolutionary history Can assess time of divergence by looking at the number of mutations needed to change one
Scoring and heuristic methods for sequence alignment CG 17
 Scoring and heuristic methods for sequence alignment CG 17 Amino Acid Substitution Matrices Used to score alignments. Reflect evolution of sequences. Unitary Matrix: M ij = 1 i=j { 0 o/w Genetic Code Matrix:
Scoring and heuristic methods for sequence alignment CG 17 Amino Acid Substitution Matrices Used to score alignments. Reflect evolution of sequences. Unitary Matrix: M ij = 1 i=j { 0 o/w Genetic Code Matrix:
Pairwise Sequence Alignment. Zhongming Zhao, PhD
 Pairwise Sequence Alignment Zhongming Zhao, PhD Email: zhongming.zhao@vanderbilt.edu http://bioinfo.mc.vanderbilt.edu/ Sequence Similarity match mismatch A T T A C G C G T A C C A T A T T A T G C G A T
Pairwise Sequence Alignment Zhongming Zhao, PhD Email: zhongming.zhao@vanderbilt.edu http://bioinfo.mc.vanderbilt.edu/ Sequence Similarity match mismatch A T T A C G C G T A C C A T A T T A T G C G A T
C E N T R. Introduction to bioinformatics 2007 E B I O I N F O R M A T I C S V U F O R I N T. Lecture 13 G R A T I V. Iterative homology searching,
 C E N T R E F O R I N T E G R A T I V E B I O I N F O R M A T I C S V U Introduction to bioinformatics 2007 Lecture 13 Iterative homology searching, PSI (Position Specific Iterated) BLAST basic idea use
C E N T R E F O R I N T E G R A T I V E B I O I N F O R M A T I C S V U Introduction to bioinformatics 2007 Lecture 13 Iterative homology searching, PSI (Position Specific Iterated) BLAST basic idea use
Heuristic methods for pairwise alignment:
 Bi03c_1 Unit 03c: Heuristic methods for pairwise alignment: k-tuple-methods k-tuple-methods for alignment of pairs of sequences Bi03c_2 dynamic programming is too slow for large databases Use heuristic
Bi03c_1 Unit 03c: Heuristic methods for pairwise alignment: k-tuple-methods k-tuple-methods for alignment of pairs of sequences Bi03c_2 dynamic programming is too slow for large databases Use heuristic
Multiple Sequence Alignment. Mark Whitsitt - NCSA
 Multiple Sequence Alignment Mark Whitsitt - NCSA What is a Multiple Sequence Alignment (MA)? GMHGTVYANYAVDSSDLLLAFGVRFDDRVTGKLEAFASRAKIVHIDIDSAEIGKNKQPHV GMHGTVYANYAVEHSDLLLAFGVRFDDRVTGKLEAFASRAKIVHIDIDSAEIGKNKTPHV
Multiple Sequence Alignment Mark Whitsitt - NCSA What is a Multiple Sequence Alignment (MA)? GMHGTVYANYAVDSSDLLLAFGVRFDDRVTGKLEAFASRAKIVHIDIDSAEIGKNKQPHV GMHGTVYANYAVEHSDLLLAFGVRFDDRVTGKLEAFASRAKIVHIDIDSAEIGKNKTPHV
In this section we describe how to extend the match refinement to the multiple case and then use T-Coffee to heuristically compute a multiple trace.
 5 Multiple Match Refinement and T-Coffee In this section we describe how to extend the match refinement to the multiple case and then use T-Coffee to heuristically compute a multiple trace. This exposition
5 Multiple Match Refinement and T-Coffee In this section we describe how to extend the match refinement to the multiple case and then use T-Coffee to heuristically compute a multiple trace. This exposition
As of August 15, 2008, GenBank contained bases from reported sequences. The search procedure should be
 48 Bioinformatics I, WS 09-10, S. Henz (script by D. Huson) November 26, 2009 4 BLAST and BLAT Outline of the chapter: 1. Heuristics for the pairwise local alignment of two sequences 2. BLAST: search and
48 Bioinformatics I, WS 09-10, S. Henz (script by D. Huson) November 26, 2009 4 BLAST and BLAT Outline of the chapter: 1. Heuristics for the pairwise local alignment of two sequences 2. BLAST: search and
1. R. Durbin, S. Eddy, A. Krogh und G. Mitchison: Biological sequence analysis, Cambridge, 1998
 7 Multiple Sequence Alignment The exposition was prepared by Clemens GrÃP pl, based on earlier versions by Daniel Huson, Knut Reinert, and Gunnar Klau. It is based on the following sources, which are all
7 Multiple Sequence Alignment The exposition was prepared by Clemens GrÃP pl, based on earlier versions by Daniel Huson, Knut Reinert, and Gunnar Klau. It is based on the following sources, which are all
CS313 Exercise 4 Cover Page Fall 2017
 CS313 Exercise 4 Cover Page Fall 2017 Due by the start of class on Thursday, October 12, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
CS313 Exercise 4 Cover Page Fall 2017 Due by the start of class on Thursday, October 12, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
Bioinformatics III Structural Bioinformatics and Genome Analysis
 Bioinformatics III Structural Bioinformatics and Genome Analysis Chapter 3 Structural Comparison and Alignment 3.1 Introduction 1. Basic algorithms review Dynamic programming Distance matrix 2. SARF2,
Bioinformatics III Structural Bioinformatics and Genome Analysis Chapter 3 Structural Comparison and Alignment 3.1 Introduction 1. Basic algorithms review Dynamic programming Distance matrix 2. SARF2,
CPSC 340: Machine Learning and Data Mining. More Regularization Fall 2017
 CPSC 340: Machine Learning and Data Mining More Regularization Fall 2017 Assignment 3: Admin Out soon, due Friday of next week. Midterm: You can view your exam during instructor office hours or after class
CPSC 340: Machine Learning and Data Mining More Regularization Fall 2017 Assignment 3: Admin Out soon, due Friday of next week. Midterm: You can view your exam during instructor office hours or after class
Dynamic Programming & Smith-Waterman algorithm
 m m Seminar: Classical Papers in Bioinformatics May 3rd, 2010 m m 1 2 3 m m Introduction m Definition is a method of solving problems by breaking them down into simpler steps problem need to contain overlapping
m m Seminar: Classical Papers in Bioinformatics May 3rd, 2010 m m 1 2 3 m m Introduction m Definition is a method of solving problems by breaking them down into simpler steps problem need to contain overlapping
Sequence Alignment (chapter 6) p The biological problem p Global alignment p Local alignment p Multiple alignment
 Sequence lignment (chapter 6) p The biological problem p lobal alignment p Local alignment p Multiple alignment Local alignment: rationale p Otherwise dissimilar proteins may have local regions of similarity
Sequence lignment (chapter 6) p The biological problem p lobal alignment p Local alignment p Multiple alignment Local alignment: rationale p Otherwise dissimilar proteins may have local regions of similarity
Lecture 9: Core String Edits and Alignments
 Biosequence Algorithms, Spring 2005 Lecture 9: Core String Edits and Alignments Pekka Kilpeläinen University of Kuopio Department of Computer Science BSA Lecture 9: String Edits and Alignments p.1/30 III:
Biosequence Algorithms, Spring 2005 Lecture 9: Core String Edits and Alignments Pekka Kilpeläinen University of Kuopio Department of Computer Science BSA Lecture 9: String Edits and Alignments p.1/30 III:
Special course in Computer Science: Advanced Text Algorithms
 Special course in Computer Science: Advanced Text Algorithms Lecture 6: Alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Special course in Computer Science: Advanced Text Algorithms Lecture 6: Alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Similarity Searches on Sequence Databases
 Similarity Searches on Sequence Databases Lorenza Bordoli Swiss Institute of Bioinformatics EMBnet Course, Zürich, October 2004 Swiss Institute of Bioinformatics Swiss EMBnet node Outline Importance of
Similarity Searches on Sequence Databases Lorenza Bordoli Swiss Institute of Bioinformatics EMBnet Course, Zürich, October 2004 Swiss Institute of Bioinformatics Swiss EMBnet node Outline Importance of
Chapter 8 Multiple sequence alignment. Chaochun Wei Spring 2018
 1896 1920 1987 2006 Chapter 8 Multiple sequence alignment Chaochun Wei Spring 2018 Contents 1. Reading materials 2. Multiple sequence alignment basic algorithms and tools how to improve multiple alignment
1896 1920 1987 2006 Chapter 8 Multiple sequence alignment Chaochun Wei Spring 2018 Contents 1. Reading materials 2. Multiple sequence alignment basic algorithms and tools how to improve multiple alignment
Lecture 10. Sequence alignments
 Lecture 10 Sequence alignments Alignment algorithms: Overview Given a scoring system, we need to have an algorithm for finding an optimal alignment for a pair of sequences. We want to maximize the score
Lecture 10 Sequence alignments Alignment algorithms: Overview Given a scoring system, we need to have an algorithm for finding an optimal alignment for a pair of sequences. We want to maximize the score
Pairwise Sequence alignment Basic Algorithms
 Pairwise Sequence alignment Basic Algorithms Agenda - Previous Lesson: Minhala - + Biological Story on Biomolecular Sequences - + General Overview of Problems in Computational Biology - Reminder: Dynamic
Pairwise Sequence alignment Basic Algorithms Agenda - Previous Lesson: Minhala - + Biological Story on Biomolecular Sequences - + General Overview of Problems in Computational Biology - Reminder: Dynamic
Sequence alignment theory and applications Session 3: BLAST algorithm
 Sequence alignment theory and applications Session 3: BLAST algorithm Introduction to Bioinformatics online course : IBT Sonal Henson Learning Objectives Understand the principles of the BLAST algorithm
Sequence alignment theory and applications Session 3: BLAST algorithm Introduction to Bioinformatics online course : IBT Sonal Henson Learning Objectives Understand the principles of the BLAST algorithm
1. R. Durbin, S. Eddy, A. Krogh und G. Mitchison: Biological sequence analysis, Cambridge, 1998
 7 Multiple Sequence Alignment The exposition was prepared by Clemens Gröpl, based on earlier versions by Daniel Huson, Knut Reinert, and Gunnar Klau. It is based on the following sources, which are all
7 Multiple Sequence Alignment The exposition was prepared by Clemens Gröpl, based on earlier versions by Daniel Huson, Knut Reinert, and Gunnar Klau. It is based on the following sources, which are all
Lecture 10: Local Alignments
 Lecture 10: Local Alignments Study Chapter 6.8-6.10 1 Outline Edit Distances Longest Common Subsequence Global Sequence Alignment Scoring Matrices Local Sequence Alignment Alignment with Affine Gap Penalties
Lecture 10: Local Alignments Study Chapter 6.8-6.10 1 Outline Edit Distances Longest Common Subsequence Global Sequence Alignment Scoring Matrices Local Sequence Alignment Alignment with Affine Gap Penalties
Chapter 6. Multiple sequence alignment (week 10)
 Course organization Introduction ( Week 1,2) Part I: Algorithms for Sequence Analysis (Week 1-11) Chapter 1-3, Models and theories» Probability theory and Statistics (Week 3)» Algorithm complexity analysis
Course organization Introduction ( Week 1,2) Part I: Algorithms for Sequence Analysis (Week 1-11) Chapter 1-3, Models and theories» Probability theory and Statistics (Week 3)» Algorithm complexity analysis
Lectures 12 and 13 Dynamic programming: weighted interval scheduling
 Lectures 12 and 13 Dynamic programming: weighted interval scheduling COMP 523: Advanced Algorithmic Techniques Lecturer: Dariusz Kowalski Lectures 12-13: Dynamic Programming 1 Overview Last week: Graph
Lectures 12 and 13 Dynamic programming: weighted interval scheduling COMP 523: Advanced Algorithmic Techniques Lecturer: Dariusz Kowalski Lectures 12-13: Dynamic Programming 1 Overview Last week: Graph
24 Grundlagen der Bioinformatik, SS 10, D. Huson, April 26, This lecture is based on the following papers, which are all recommended reading:
 24 Grundlagen der Bioinformatik, SS 10, D. Huson, April 26, 2010 3 BLAST and FASTA This lecture is based on the following papers, which are all recommended reading: D.J. Lipman and W.R. Pearson, Rapid
24 Grundlagen der Bioinformatik, SS 10, D. Huson, April 26, 2010 3 BLAST and FASTA This lecture is based on the following papers, which are all recommended reading: D.J. Lipman and W.R. Pearson, Rapid
Research Article Aligning Sequences by Minimum Description Length
 Hindawi Publishing Corporation EURASIP Journal on Bioinformatics and Systems Biology Volume 2007, Article ID 72936, 14 pages doi:10.1155/2007/72936 Research Article Aligning Sequences by Minimum Description
Hindawi Publishing Corporation EURASIP Journal on Bioinformatics and Systems Biology Volume 2007, Article ID 72936, 14 pages doi:10.1155/2007/72936 Research Article Aligning Sequences by Minimum Description
Sequence comparison: Local alignment
 Sequence comparison: Local alignment Genome 559: Introuction to Statistical an Computational Genomics Prof. James H. Thomas http://faculty.washington.eu/jht/gs559_217/ Review global alignment en traceback
Sequence comparison: Local alignment Genome 559: Introuction to Statistical an Computational Genomics Prof. James H. Thomas http://faculty.washington.eu/jht/gs559_217/ Review global alignment en traceback
Basics on bioinforma-cs Lecture 4. Concita Cantarella
 Basics on bioinforma-cs Lecture 4 Concita Cantarella concita.cantarella@entecra.it; concita.cantarella@gmail.com Why compare sequences Sequence comparison is a way of arranging the sequences of DNA, RNA
Basics on bioinforma-cs Lecture 4 Concita Cantarella concita.cantarella@entecra.it; concita.cantarella@gmail.com Why compare sequences Sequence comparison is a way of arranging the sequences of DNA, RNA
Computational Biology Lecture 4: Overlap detection, Local Alignment, Space Efficient Needleman-Wunsch Saad Mneimneh
 Computational Biology Lecture 4: Overlap detection, Local Alignment, Space Efficient Needleman-Wunsch Saad Mneimneh Overlap detection: Semi-Global Alignment An overlap of two sequences is considered an
Computational Biology Lecture 4: Overlap detection, Local Alignment, Space Efficient Needleman-Wunsch Saad Mneimneh Overlap detection: Semi-Global Alignment An overlap of two sequences is considered an
Local Alignment & Gap Penalties CMSC 423
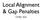 Local Alignment & ap Penalties CMSC 423 lobal, Semi-global, Local Alignments Last time, we saw a dynamic programming algorithm for global alignment: both strings s and t must be completely matched: s t
Local Alignment & ap Penalties CMSC 423 lobal, Semi-global, Local Alignments Last time, we saw a dynamic programming algorithm for global alignment: both strings s and t must be completely matched: s t
Accelerating the Prediction of Protein Interactions
 Accelerating the Prediction of Protein Interactions Alex Rodionov, Jonathan Rose, Elisabeth R.M. Tillier, Alexandr Bezginov October 21 21 Motivation The human genome is sequenced, but we don't know what
Accelerating the Prediction of Protein Interactions Alex Rodionov, Jonathan Rose, Elisabeth R.M. Tillier, Alexandr Bezginov October 21 21 Motivation The human genome is sequenced, but we don't know what
Biological Sequence Matching Using Fuzzy Logic
 International Journal of Scientific & Engineering Research Volume 2, Issue 7, July-2011 1 Biological Sequence Matching Using Fuzzy Logic Nivit Gill, Shailendra Singh Abstract: Sequence alignment is the
International Journal of Scientific & Engineering Research Volume 2, Issue 7, July-2011 1 Biological Sequence Matching Using Fuzzy Logic Nivit Gill, Shailendra Singh Abstract: Sequence alignment is the
Bioinformatics. Sequence alignment BLAST Significance. Next time Protein Structure
 Bioinformatics Sequence alignment BLAST Significance Next time Protein Structure 1 Experimental origins of sequence data The Sanger dideoxynucleotide method F Each color is one lane of an electrophoresis
Bioinformatics Sequence alignment BLAST Significance Next time Protein Structure 1 Experimental origins of sequence data The Sanger dideoxynucleotide method F Each color is one lane of an electrophoresis
COS 551: Introduction to Computational Molecular Biology Lecture: Oct 17, 2000 Lecturer: Mona Singh Scribe: Jacob Brenner 1. Database Searching
 COS 551: Introduction to Computational Molecular Biology Lecture: Oct 17, 2000 Lecturer: Mona Singh Scribe: Jacob Brenner 1 Database Searching In database search, we typically have a large sequence database
COS 551: Introduction to Computational Molecular Biology Lecture: Oct 17, 2000 Lecturer: Mona Singh Scribe: Jacob Brenner 1 Database Searching In database search, we typically have a large sequence database
Notes on Dynamic-Programming Sequence Alignment
 Notes on Dynamic-Programming Sequence Alignment Introduction. Following its introduction by Needleman and Wunsch (1970), dynamic programming has become the method of choice for rigorous alignment of DNA
Notes on Dynamic-Programming Sequence Alignment Introduction. Following its introduction by Needleman and Wunsch (1970), dynamic programming has become the method of choice for rigorous alignment of DNA
CS273: Algorithms for Structure Handout # 4 and Motion in Biology Stanford University Thursday, 8 April 2004
 CS273: Algorithms for Structure Handout # 4 and Motion in Biology Stanford University Thursday, 8 April 2004 Lecture #4: 8 April 2004 Topics: Sequence Similarity Scribe: Sonil Mukherjee 1 Introduction
CS273: Algorithms for Structure Handout # 4 and Motion in Biology Stanford University Thursday, 8 April 2004 Lecture #4: 8 April 2004 Topics: Sequence Similarity Scribe: Sonil Mukherjee 1 Introduction
Small Libraries of Protein Fragments through Clustering
 Small Libraries of Protein Fragments through Clustering Varun Ganapathi Department of Computer Science Stanford University June 8, 2005 Abstract When trying to extract information from the information
Small Libraries of Protein Fragments through Clustering Varun Ganapathi Department of Computer Science Stanford University June 8, 2005 Abstract When trying to extract information from the information
BLAST. Basic Local Alignment Search Tool. Used to quickly compare a protein or DNA sequence to a database.
 BLAST Basic Local Alignment Search Tool Used to quickly compare a protein or DNA sequence to a database. There is no such thing as a free lunch BLAST is fast and highly sensitive compared to competitors.
BLAST Basic Local Alignment Search Tool Used to quickly compare a protein or DNA sequence to a database. There is no such thing as a free lunch BLAST is fast and highly sensitive compared to competitors.
Machine Learning. Computational biology: Sequence alignment and profile HMMs
 10-601 Machine Learning Computational biology: Sequence alignment and profile HMMs Central dogma DNA CCTGAGCCAACTATTGATGAA transcription mrna CCUGAGCCAACUAUUGAUGAA translation Protein PEPTIDE 2 Growth
10-601 Machine Learning Computational biology: Sequence alignment and profile HMMs Central dogma DNA CCTGAGCCAACTATTGATGAA transcription mrna CCUGAGCCAACUAUUGAUGAA translation Protein PEPTIDE 2 Growth
Computational Biology Lecture 6: Affine gap penalty function, multiple sequence alignment Saad Mneimneh
 Computational Biology Lecture 6: Affine gap penalty function, multiple sequence alignment Saad Mneimneh We saw earlier how we can use a concave gap penalty function γ, i.e. one that satisfies γ(x+1) γ(x)
Computational Biology Lecture 6: Affine gap penalty function, multiple sequence alignment Saad Mneimneh We saw earlier how we can use a concave gap penalty function γ, i.e. one that satisfies γ(x+1) γ(x)
Comparison of Sequence Similarity Measures for Distant Evolutionary Relationships
 Comparison of Sequence Similarity Measures for Distant Evolutionary Relationships Abhishek Majumdar, Peter Z. Revesz Department of Computer Science and Engineering, University of Nebraska-Lincoln, Lincoln,
Comparison of Sequence Similarity Measures for Distant Evolutionary Relationships Abhishek Majumdar, Peter Z. Revesz Department of Computer Science and Engineering, University of Nebraska-Lincoln, Lincoln,
Data Mining Technologies for Bioinformatics Sequences
 Data Mining Technologies for Bioinformatics Sequences Deepak Garg Computer Science and Engineering Department Thapar Institute of Engineering & Tecnology, Patiala Abstract Main tool used for sequence alignment
Data Mining Technologies for Bioinformatics Sequences Deepak Garg Computer Science and Engineering Department Thapar Institute of Engineering & Tecnology, Patiala Abstract Main tool used for sequence alignment
MULTIPLE SEQUENCE ALIGNMENT SOLUTIONS AND APPLICATIONS
 MULTIPLE SEQUENCE ALIGNMENT SOLUTIONS AND APPLICATIONS By XU ZHANG A DISSERTATION PRESENTED TO THE GRADUATE SCHOOL OF THE UNIVERSITY OF FLORIDA IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE
MULTIPLE SEQUENCE ALIGNMENT SOLUTIONS AND APPLICATIONS By XU ZHANG A DISSERTATION PRESENTED TO THE GRADUATE SCHOOL OF THE UNIVERSITY OF FLORIDA IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE
Pairwise alignment II
 Pairwise alignment II Agenda - Previous Lesson: Minhala + Introduction - Review Dynamic Programming - Pariwise Alignment Biological Motivation Today: - Quick Review: Sequence Alignment (Global, Local,
Pairwise alignment II Agenda - Previous Lesson: Minhala + Introduction - Review Dynamic Programming - Pariwise Alignment Biological Motivation Today: - Quick Review: Sequence Alignment (Global, Local,
DynamicProgramming. September 17, 2018
 DynamicProgramming September 17, 2018 1 Lecture 11: Dynamic Programming CBIO (CSCI) 4835/6835: Introduction to Computational Biology 1.1 Overview and Objectives We ve so far discussed sequence alignment
DynamicProgramming September 17, 2018 1 Lecture 11: Dynamic Programming CBIO (CSCI) 4835/6835: Introduction to Computational Biology 1.1 Overview and Objectives We ve so far discussed sequence alignment
Alignment of Pairs of Sequences
 Bi03a_1 Unit 03a: Alignment of Pairs of Sequences Partners for alignment Bi03a_2 Protein 1 Protein 2 =amino-acid sequences (20 letter alphabeth + gap) LGPSSKQTGKGS-SRIWDN LN-ITKSAGKGAIMRLGDA -------TGKG--------
Bi03a_1 Unit 03a: Alignment of Pairs of Sequences Partners for alignment Bi03a_2 Protein 1 Protein 2 =amino-acid sequences (20 letter alphabeth + gap) LGPSSKQTGKGS-SRIWDN LN-ITKSAGKGAIMRLGDA -------TGKG--------
