Biological Image Analysis. Dr. Rob Clements -
|
|
|
- Christian McCormick
- 5 years ago
- Views:
Transcription
1 Biological Image Analysis Dr. Rob Clements - rclement@kent.edu
2 Course Overview Design methods to automatically analyze microscope images Group project based Presentations/discussions Final products: Presentation Working plugin or macro for imagej Report..lab openings...
3 Imaging Projects Plugins Creation MICROSCOPY 1) Tiled data reconstruction and correction 2) Astrocyte Cell classification 3) Microvessel analysis
4 Fluorescence Microscopy -specimen can be labeled with a fluorophore (green fluorescent protein-gfp) directed at an object of Interest -specimen is illuminated with light of a specific wavelength which is absorbed by the fluorophores -fluorphore emits longer wavelength (different color) -green fluorophore absorbs a blue excitation photon and an electron changes from the ground state to an excited state. -electron then decays back from the excited state to the ground state and releases a green photon
5 Fluorescence Microscopy -Possible to use concurrent fluorophores as long as excitation/emission wavelength different -commercially available bound to organelle probes and antibodies -permits probing multiple structures concurrently
6 Fluorescence Microscopy Wavelength Filtering * Excitation filter - the excitation path just prior to the dichroic mirror to select the excitation wavelength * Emission filter - to select the emission wavelength from the sample and remove excitation light, it is placed beneath the dichroic mirror.
7 Confocal Microscopy -laser line to illuminate and scan samples, with series of mirrors and filters -uses point illumination and pinhole to eliminate out-of-focus light
8 Confocal Microscopy Permits 3D data collection by creating image stacks and 3D reconstruction -far better resolution by imaging single planes and recombining Problems Acquisition time, brightness, Axial Blurring Point Spread Function
9 Imaging Projects Plugins Creation MICROSCOPY 1) Tiled data reconstruction and correction 2) Astrocyte Cell classification 3) Microvessel analysis
10 Imaging Projects 1) Tiled data reconstruction and correction -accurately stitch multiple images together -correct brightness across images -existing plugin usage? -olympus data format input -OIB
11 Imaging Projects 1) Tiled data reconstruction and correction -accurately stitch multiple images together -multichannel OIB and MATL input rgbtiff/select output -stage drift-->predictable?
12 Imaging Projects 2) Tiled data reconstruction and correction TISSUE RECONSTRUCTION 4.35 mm x 2.6mm x 1.4 mm voxel resolution; 1.2 microns 3508x2072x700 pixels optical sections
13
14 Imaging Projects 2) Tiled data reconstruction and correction -correct brightness across images/seems -use adjacent tiles, (calibration image?) -different channel issues?
15 Existing Tools -grid collection stitching plugin (no brightness correction, need to read MATL file)
16 Imaging Projects Plugins Creation MICROSCOPY 1) Tiled data reconstruction and correction 2) Multiple Astrocyte Cell classification 3) Microvessel analysis
17 Types of Neurons
18 Types of Neurons
19 Other Cell Types in the Brain Glail Cells -microglia, astrocytes, oligodendrocytes Endothelial Mast Cells
20 Imaging Projects 2) Astrocyte analysis -automatically segment Individual Cells in 3D -classify cells and provide volume and surface area -extract and count individual glial cells from larger data -use and extend existing plugins -beginning point from existing course
21 Imaging Projects 2) Astrocyte analysis -automatically segment Individual Cells in 3D -intelligent thresholding brightness differences? -segmentation use of seed points -extendable to larger data sets with multiple astrocytes -using seed points -overlapping/touching cell structures?
22 Imaging Projects 2) Astrocyte analysis -classify cells and provide volume and surface area -existing tools for quantification -describe number/length of cell extensions -define cell body -describe number of bifurcations -group similar cells for classification -use existing plugins (---skeletonize3d/analyze skeleton?) plugins?
23 Imaging Projects 2) Astrocyte analysis -extract, classify and count individual glial cells in larger image -find seed points to start segmentation -use other channel information (nuclear stain) -overlapping glial cells? -where to terminate segmentation?
24 Imaging Projects 2) Astrocyte analysis -automatically segment Individual Glial Cells in 3D -classify astrocytes and provide volume and surface area -extract and count individual glial cells -measure degree of overlap with vessels -use and extend existing plugins
25 Existing Tools -works on single cells -implementation? -seed point identification? -Work on multiple data?
26 Imaging Projects Plugins Creation MICROSCOPY 1) Tiled data reconstruction and correction 2) Astrocyte Cell classification 3) Microvessel analysis
27 Imaging Projects 3) Microvessel analysis -segment microvessels in 3D -classify vessel segments -surface area/volume -segment width, density -identify branching patterns
28 Imaging Projects 3) Microvessel analysis Segment microvessels in 3D -intelligent thresholding -resistent to partial staining techniques -transected segments?
29 Imaging Projects 3) Microvessel analysis Classify vessel segments in 3D -number of segments -surface area/volume -existing tools -segment width -vessel density -identify branching patterns -existing plugins
30 Imaging Projects 3) Microvessel analysis Test on Multiple datasets Provide relevant metric output
31 Functional Units in the Brain Blood Brain Barrier
32 Imaging Projects 3) Microvessel analysis -measure degree of astrocyte overlap with vessels in secondary channel
33 Existing Tools -vessels of multiple sizes? -degree of overlap with additional channel? (mask creation)
34
35 Existing Tools DATABASES Cell Centered Database (CCDB), SenseLab Project L-Neuron Neuromorph brainmaps.org Neuroscience Information Framework (NIF) Connectome Project. Free and Commercial Software cellprofiler Xvoxtrace Neurolucida Imaris Amira SSECRETT and NeuroTrace ImageJ/FIJI NeuronJ, Simple Neurite Tracer Find Connected Regions Neuron Morphology, TrakEM2 Simple Interactive Object Extraction (SIOX) Advanced Weka Segmentation Trainable Segmentation plugin.
36 Existing Tools DATABASES Cell Centered Database (CCDB), SenseLab Project L-Neuron Neuromorph brainmaps.org Neuroscience Information Framework (NIF) Connectome Project. ImageJ/FIJI NeuronJ, Simple Neurite Tracer Find Connected Regions Neuron Morphology, TrakEM2 Simple Interactive Object Extraction (SIOX) Advanced Weka Segmentation Trainable Segmentation plugin.
37 IMAGEJ Plugins and Macros Examples Forum
38 Introduction to the Main Menu Of these, we ll concentrate on: Image Process Analyze Plugins Help
39 Image Menu
40 Process Menu
41 Analyze Menu
42 Plugins Menu
43 FIJI Fiji is an image processing package. It can be described as a distribution of ImageJ (and soon ImageJ2) together with Java, Java 3D and a lot of plugins organized into a coherent menu structure. Fiji compares to ImageJ as Ubuntu compares to Linux.
44 dir1 = getdirectory("choose a Directory "); list = getfilelist(dir1); ddir1 =dir1+"\\tiffs\\"; IMAGEJ Writing Macros File.makeDirectory(ddir1); (use the macro-recorder under for (i=0; i<list.length; i++) { plugins macro--record to get started) file = list[i]; file2 = dir1 + list[i]; open(file2); //name1 = list[i] + " - C=0"; //name2 = list[i] + " - C=1"; //name3 = list[i] + " - C=2"; //selectwindow(name1); run("brightness/contrast..."); setminandmax(0, 4095); //selectwindow(name2); run("brightness/contrast..."); setminandmax(0, 4095); //selectwindow(name3); setminandmax(0, 4095); run("brightness/contrast..."); setminandmax(0, 4095); //run("concatenate...", "stack1=name1 stack2=name2 title=stack"); //run("concatenate...", "stack1=name3 stack2=stack title=stacks"); name1 = ddir1 + i; saveas("tiff", name1); close(); }
45 Dialog.create("TILER Z-Project") numcols = 3; numrows = 3; Dialog.addNumber("ROWS", numcols); Dialog.addNumber("COLUMNS", numrows); Dialog.show(); IMAGEJ Writing Macros numslices = nslices; numrows2 = numrows*numcols; last=nslices*numrows2; run("brightness/contrast..."); setminandmax(0, 4095); call("ij.imageplus.setdefault16bitrange", 0); run("make Montage...", "columns=numslices rows=numrows2 scale=1 first=1 last=last increment=1 border=0 font=12"); run("montage to Stack...", "images_per_row=numslices images_per_column=1 border=0"); run("z Project...", "start=1 stop=numslices projection=[max Intensity]"); run("montage to Stack...", "images_per_row=1 images_per_column=numrows2 border=0"); run("make Montage...", "columns=numcols rows=numrows scale=1 first=1 last=numrows2 increment=1 border=0 font=12");
46 //TRIPLE STAIN RGB ANALYZER dir1 = getdirectory("choose a Directory "); list = getfilelist(dir1); ddir1 =dir1+"\\sum\\"; File.makeDirectory(ddir1); ddir2 =dir1+"\\max\\"; File.makeDirectory(ddir2); for (i=0; i<list.length; i++) { file = list[i]; file2 = dir1 + list[i]; open(file2); run("brightness/contrast..."); setminandmax(0, 4095); call("ij.imageplus.setdefault16bitrange", 0); run("stack Splitter", "number=2"); r = "(stk_0002_)" + file; g = "(stk_0001_)" + file; b = "(none)" + file; selectwindow(file); run("close"); run("merge Channels...", "red=r green=g blue=*none* gray=*none*"); run("z Project...", " projection=[sum Slices]"); name1 = ddir1 + "SUM_RGB" + file; saveas("tiff", name1); selectwindow("rgb"); run("z Project...", " projection=[max Intensity]"); name1 = ddir2 + "MAX_RGB" + file ; saveas("tiff", name1); close(); close(); close(); } dir2 = dir1+"\\max\\"; list = getfilelist(dir2); for (i=0; i<list.length; i++) { file = dir2 + list[i]; match = indexof(file, "RGB"); file = list[i]; if (match > 0) { open(dir2 + list[i]); run("split Channels"); selectwindow(file+ " (green)"); setthreshold(94, 255); run("convert to Mask"); selectwindow(file+ " (green)"); wait(2000); run("invert"); wait(1000); selectwindow(file+ " (green)"); wait(1000); imagecalculator("subtract create", file + " (red)",file + " (green)"); run("measure"); IMAGEJ Writing Macros selectwindow("results"); name = dir2 + "Results.xls"; run("text...", "save=name"); run("clear Results"); //print(f, list[2] + "\t"); //selectimage(-3); //saveas("raw Data", "./2.raw"); //selectimage(-3); //saveas("raw Data", "./3.raw"); //selectimage(-4); //saveas("raw Data", "./3.raw"); dir2 = dir1+"\\sum\\"; list = getfilelist(dir2); for (i=0; i<list.length; i++) { file = dir2 + list[i]; match = indexof(file, "RGB"); file = list[i]; if (match > 0) { open(dir2 + list[i]); run("split Channels"); selectwindow(file+ " (green)"); setthreshold(94, 255); run("convert to Mask"); selectwindow(file+ " (green)"); Wait(2000); run("invert"); wait(1000); selectwindow(file+ " (green)"); wait(1000); imagecalculator("subtract create", file + " (red)",file + " (green)"); run("measure"); run("close All"); } } selectwindow("results"); name = dir2 + "Results.xls"; run("text...", "save=name"); run("clear Results"); }
47 import ij.*; import ij.process.*; import ij.gui.*; import ij.plugin.*; public class Demontager_ implements PlugIn { /** This plugin converts a montaged image to an image stack based on the number of rows and columns input by the user. This is the opposite of what the Image/Stacks/Make Montage command does. */ public void run(string arg) { ImagePlus imp = IJ.getImage(); GenericDialog gd = new GenericDialog("Demontager"); gd.addnumericfield("columns:",1,0); gd.addnumericfield("rows:",1,0); gd.showdialog(); if (gd.wascanceled()) return; int columns = (int)gd.getnextnumber(); int rows = (int)gd.getnextnumber(); ImageProcessor ip = imp.getprocessor(); ImageStack stack = demontage(ip, columns, rows); new ImagePlus("Demontage", stack).show(); } IMAGEJ Writing Plugins public ImageStack demontage(imageprocessor ip, int columns, int rows) { int width = ip.getwidth(); int height = ip.getheight(); int cropwidth = width/columns; int cropheight = height/rows; ImageStack stack = new ImageStack(cropwidth, cropheight); for (int j=1; j<=rows; j++) { for (int i = 1; i<=columns; i++) { int x = i * cropwidth - cropwidth; int y = j * cropheight - cropheight; ip.setroi(x, y, cropwidth, cropheight); stack.addslice(null, ip.crop()); } } return stack; } }
48 Automating/ Calling ImageJ from other Programs Call it from the command line Plugins located in same directory: java -cp ij.jar:. analyze blobs.tif > results.txt (Unix) java -cp ij.jar;. analyze blobs.tif > results.txt (Windows) Or Define the Directory: java -Dplugins.dir=/usr/local/ImageJ -cp /usr/local/ij.jar:. analyze blobs.tif > results.txt (Unix) java -Dplugins.dir=C:\ImageJ -cp C:\ImageJ\ij.jar;. analyze blobs.tif > results.txt (Windows) Run it in Batch mode no GUI -batch path [arg] Runs a macro or script in batch (no GUI) mode, passing it an optional argument. ImageJ exits when the macro finishes.
49 Confounding Factors in Data Brightness inhomogeneities (laterally and axially (PSF) confocal) Sample auto fluorescence Poorly/incompletely stained Transected structures due to physical sectioning Alignment and recombination (tiled data or badly aligned image stacks) Dense overlapping structures Sample variability Size Processing time Error? confidence estimates
50 ALL PROJECTS Implemented/useable as ImageJ plugin or macro Call from command line? Extendable to other data Test/works on all provided data automated minimal user input to trigger Simple Graphical Interface Sensible feedback of results Non-destructive image overlay Table of results/graphs Error warnings Don't reinvent the Wheel!!!
51 The Data! -multichannnel data, not all channels necessary -some operations require different channels -use 3D data (not 2D projections) 1) Tiled data reconstruction and correction 2-tiled data/cortex/test _ tiled data/spinalcord/test _ ) Astrocyte Cell classification 3-astrocyte classification/single -only use astrocyte data channel 3-astrocyte classification/multiple -astrocyte data for segmenting, nuclei for seed points 3) Microvessel analysis 4-microvessel analysis/big 4-microvessel analysis/small
52 Imaging Projects Data examples -show how to split channels -brightness adjust or reset 1) Tiled data reconstruction and correction -show projection image and MATL file 2) Astrocyte Cell classification -show projection image and sample data 3) Microvessel analysis -show projection image and channel data
A Cluster-based Neuroinformatics System for Efficient Segmentation, Management and Analysis of Very Large Microscopy Data
 A Cluster-based Neuroinformatics System for Efficient Segmentation, Management and Analysis of Very Large Microscopy Data http://drosophila.biology.kent.edu/users/rclement/resource/jan17.pdf rclement@kent.edu
A Cluster-based Neuroinformatics System for Efficient Segmentation, Management and Analysis of Very Large Microscopy Data http://drosophila.biology.kent.edu/users/rclement/resource/jan17.pdf rclement@kent.edu
Supplementary Figure 1 Guide stars of progressively longer wavelengths can be used for direct wavefront sensing at increasingly large depth in the
 Supplementary Figure 1 Guide stars of progressively longer wavelengths can be used for direct wavefront sensing at increasingly large depth in the cortex of the living mouse. Typical SH images of guide
Supplementary Figure 1 Guide stars of progressively longer wavelengths can be used for direct wavefront sensing at increasingly large depth in the cortex of the living mouse. Typical SH images of guide
WCIF COLOCALIS ATION PLUGINS
 WCIF COLOCALIS ATION PLUGINS Colocalisation Test Background When a coefficient is calculated for two images, it is often unclear quite what this means, in particular for intermediate values. This raises
WCIF COLOCALIS ATION PLUGINS Colocalisation Test Background When a coefficient is calculated for two images, it is often unclear quite what this means, in particular for intermediate values. This raises
Last class. This class. Single molecule imaging Deconvolution. FLIM Confocal
 FLIM, Confocal Last class Single molecule imaging Deconvolution This class FLIM Confocal FLIM Fluorescence Lifetime IMaging FLIM Fluorescence lifetime imaging Easier and more accurate quantitation Much
FLIM, Confocal Last class Single molecule imaging Deconvolution This class FLIM Confocal FLIM Fluorescence Lifetime IMaging FLIM Fluorescence lifetime imaging Easier and more accurate quantitation Much
Bosch Institute Advanced Microscopy Facility Workshops Summer/Autumn 2016
 Bosch Institute Advanced Microscopy Facility Workshops Summer/Autumn 2016 Presented by Dr Louise Cole and Dr Cathy Payne, Advanced Microscopy Facility, Bosch Institute, School of Medical Sciences, The
Bosch Institute Advanced Microscopy Facility Workshops Summer/Autumn 2016 Presented by Dr Louise Cole and Dr Cathy Payne, Advanced Microscopy Facility, Bosch Institute, School of Medical Sciences, The
Fast, reliable and automatic 3D alignment of confocal image stacks: Practical
 Fast, reliable and automatic 3D alignment of confocal image stacks: Practical Dept. of Infection Biology, Biozentrum, University of Basel mario@xuvtools.org 30.04.2010 Overview Motivation Existing Commercial
Fast, reliable and automatic 3D alignment of confocal image stacks: Practical Dept. of Infection Biology, Biozentrum, University of Basel mario@xuvtools.org 30.04.2010 Overview Motivation Existing Commercial
9 Tumor Blood Vessels: 3-D Tubular Network Analysis
 Chap. 9 2015/8/20 14:02 page 1 1 9 Tumor Blood Vessels: 3-D Tubular Network Analysis Chap. 9 2015/8/20 14:02 page 2 2 Chap. 9 2015/8/20 14:02 page 3 3 Contents 9 Tumor Blood Vessels: 3-D Tubular Network
Chap. 9 2015/8/20 14:02 page 1 1 9 Tumor Blood Vessels: 3-D Tubular Network Analysis Chap. 9 2015/8/20 14:02 page 2 2 Chap. 9 2015/8/20 14:02 page 3 3 Contents 9 Tumor Blood Vessels: 3-D Tubular Network
Wide Guy: Inverted Widefield Microscope
 Wide Guy: Inverted Widefield Microscope Kyle Marchuk Adam Fries Jordan Briscoe August 2017 Contents 1 Introduction 2 2 Initial Setup 3 2.1 Hardware Startup........................................... 3
Wide Guy: Inverted Widefield Microscope Kyle Marchuk Adam Fries Jordan Briscoe August 2017 Contents 1 Introduction 2 2 Initial Setup 3 2.1 Hardware Startup........................................... 3
Introductory Guide to Light Microscopy - Biomedical Confocal Microscopy
 Introductory Guide to Light Microscopy - Biomedical Confocal Microscopy 7 May 2007 Michael Hooker Microscopy Facility Michael Chua microscopy@unc.edu 843-3268 6007 Thurston Bowles Wendy Salmon wendy_salmon@med.unc.edu
Introductory Guide to Light Microscopy - Biomedical Confocal Microscopy 7 May 2007 Michael Hooker Microscopy Facility Michael Chua microscopy@unc.edu 843-3268 6007 Thurston Bowles Wendy Salmon wendy_salmon@med.unc.edu
LSM 5 MP, LSM 510 and LSM 510 META Laser Scanning Microscopes
 LSM 5 MP, LSM 510 and LSM 510 META Laser Scanning Microscopes Brief Operating Manual Release 4.2 January 2007 Contents Page Starting the System...3 Setting the microscope...6 Configuring the beam path
LSM 5 MP, LSM 510 and LSM 510 META Laser Scanning Microscopes Brief Operating Manual Release 4.2 January 2007 Contents Page Starting the System...3 Setting the microscope...6 Configuring the beam path
Neuron Crawler: An Automatic Tracing Algorithm for Very Large Neuron Images
 Neuron Crawler: An Automatic Tracing Algorithm for Very Large Neuron Images Zhi Zhou, Staci A. Sorensen, and Hanchuan Peng* Allen Institute for Brain Science, Seattle, WA 98103. * Corresponding author.
Neuron Crawler: An Automatic Tracing Algorithm for Very Large Neuron Images Zhi Zhou, Staci A. Sorensen, and Hanchuan Peng* Allen Institute for Brain Science, Seattle, WA 98103. * Corresponding author.
2, the coefficient of variation R 2, and properties of the photon counts traces
 Supplementary Figure 1 Quality control of FCS traces. (a) Typical trace that passes the quality control (QC) according to the parameters shown in f. The QC is based on thresholds applied to fitting parameters
Supplementary Figure 1 Quality control of FCS traces. (a) Typical trace that passes the quality control (QC) according to the parameters shown in f. The QC is based on thresholds applied to fitting parameters
Neural Circuit Tracer
 2014 Neural Circuit Tracer Software for automated tracing of neurites from light microscopy stacks of images User Guide Version 4.0 Contents 1. Introduction... 2 2. System requirements and installation...
2014 Neural Circuit Tracer Software for automated tracing of neurites from light microscopy stacks of images User Guide Version 4.0 Contents 1. Introduction... 2 2. System requirements and installation...
1. ABOUT INSTALLATION COMPATIBILITY SURESIM WORKFLOWS a. Workflow b. Workflow SURESIM TUTORIAL...
 SuReSim manual 1. ABOUT... 2 2. INSTALLATION... 2 3. COMPATIBILITY... 2 4. SURESIM WORKFLOWS... 2 a. Workflow 1... 3 b. Workflow 2... 4 5. SURESIM TUTORIAL... 5 a. Import Data... 5 b. Parameter Selection...
SuReSim manual 1. ABOUT... 2 2. INSTALLATION... 2 3. COMPATIBILITY... 2 4. SURESIM WORKFLOWS... 2 a. Workflow 1... 3 b. Workflow 2... 4 5. SURESIM TUTORIAL... 5 a. Import Data... 5 b. Parameter Selection...
please download and install.
 Scripting using ImageJ Macro - a quick 1 hour tutorial - We use Fiji. http://fiji.sc please download and install. Kota Miura @ CMCI EMBL 2-1 Why do we write macro? Aim: Students acquire ImageJ macro programming
Scripting using ImageJ Macro - a quick 1 hour tutorial - We use Fiji. http://fiji.sc please download and install. Kota Miura @ CMCI EMBL 2-1 Why do we write macro? Aim: Students acquire ImageJ macro programming
Nonlinear optics and two photon microscopy. Table of contents. Sam Whiteley and Seth Parker PHYS 173/BGGN 266 July 13, 2014
 Nonlinear optics and two photon microscopy Sam Whiteley and Seth Parker PHYS 173/BGGN 266 July 13, 2014 Table of contents 1. Introduction 2. Optical setup 3. Initial images and troubleshooting 4. Determining
Nonlinear optics and two photon microscopy Sam Whiteley and Seth Parker PHYS 173/BGGN 266 July 13, 2014 Table of contents 1. Introduction 2. Optical setup 3. Initial images and troubleshooting 4. Determining
Micro-Magellan Install and User Guide
 Micro-Magellan Install and User Guide Download and install Micro-Manager, which includes Micro-Magellan: Build at time of publication: Windows 64 bit: http://valelab4.ucsf.edu/~mm/nightlybuilds/1.4/windows/mmsetup_64bit_1.4.23_20160807.
Micro-Magellan Install and User Guide Download and install Micro-Manager, which includes Micro-Magellan: Build at time of publication: Windows 64 bit: http://valelab4.ucsf.edu/~mm/nightlybuilds/1.4/windows/mmsetup_64bit_1.4.23_20160807.
What should I know about the 3D Restoration module? Improvision Technical Note No. 142 Last updated: 28 June, 2000
 What should I know about the 3D Restoration module? Improvision Technical Note No. 142 Last updated: 28 June, 2000 Topic This technical note discusses some aspects of iterative deconvolution and how to
What should I know about the 3D Restoration module? Improvision Technical Note No. 142 Last updated: 28 June, 2000 Topic This technical note discusses some aspects of iterative deconvolution and how to
3-D. Here red spheres show the location of gold nanoparticles inside/around a cell nucleus.
 3-D The CytoViva 3-D System allows the user can locate objects of interest in a 3-D space. It does this by acquiring multiple Z planes and performing our custom software routines to locate and observe
3-D The CytoViva 3-D System allows the user can locate objects of interest in a 3-D space. It does this by acquiring multiple Z planes and performing our custom software routines to locate and observe
Optical Sectioning. Bo Huang. Pharmaceutical Chemistry
 Optical Sectioning Bo Huang Pharmaceutical Chemistry Approaches to 3D imaging Physical cutting Technical difficulty Highest resolution Highest sensitivity Optical sectioning Simple sample prep. No physical
Optical Sectioning Bo Huang Pharmaceutical Chemistry Approaches to 3D imaging Physical cutting Technical difficulty Highest resolution Highest sensitivity Optical sectioning Simple sample prep. No physical
Independent Resolution Test of
 Independent Resolution Test of as conducted and published by Dr. Adam Puche, PhD University of Maryland June 2005 as presented by (formerly Thales Optem Inc.) www.qioptiqimaging.com Independent Resolution
Independent Resolution Test of as conducted and published by Dr. Adam Puche, PhD University of Maryland June 2005 as presented by (formerly Thales Optem Inc.) www.qioptiqimaging.com Independent Resolution
Setting up Micro-Magellan for Device control
 Setting up Micro-Magellan for Device control From the plug-in menu, select Micro-Magellan and the unpopulated window opens Select Configure device control The window below will open, select devices to
Setting up Micro-Magellan for Device control From the plug-in menu, select Micro-Magellan and the unpopulated window opens Select Configure device control The window below will open, select devices to
Confocal vs. Deconvolution
 Confocal vs. Deconvolution Cesare Covino ALEMBIC Advanced Light and Electron Bio-Imaging Center Istituto Scientifico San Raffaele (Milano) www.hsr.it/research/alembic Fluorescence high contrast high sensibility
Confocal vs. Deconvolution Cesare Covino ALEMBIC Advanced Light and Electron Bio-Imaging Center Istituto Scientifico San Raffaele (Milano) www.hsr.it/research/alembic Fluorescence high contrast high sensibility
Textbook de.nbi Bioimage Analysis Workshop
 BIOIMAGE ANALYSIS WORKSHOP WITHIN THE FRAMEWORK OF HD-HUB Textbook de.nbi Bioimage Analysis Workshop An introduction into KNIME image analysis for life scientists Jürgen Reymann ViroQuant-CellNetworks
BIOIMAGE ANALYSIS WORKSHOP WITHIN THE FRAMEWORK OF HD-HUB Textbook de.nbi Bioimage Analysis Workshop An introduction into KNIME image analysis for life scientists Jürgen Reymann ViroQuant-CellNetworks
Zeiss Universal SIM. A structured illumination system based on a Zeiss Universal microscope stand. by Nidhogg
 Zeiss Universal SIM A structured illumination system based on a Zeiss Universal microscope stand. by Nidhogg Structured illumination Structured illumination applied to fluorescence microscopy is a super
Zeiss Universal SIM A structured illumination system based on a Zeiss Universal microscope stand. by Nidhogg Structured illumination Structured illumination applied to fluorescence microscopy is a super
Bosch Institute Advanced Microscopy Facility Workshops Summer/Autumn 2017
 Bosch Institute Advanced Microscopy Facility Workshops Summer/Autumn 2017 Presented by Dr Louise Cole, Advanced Microscopy Facility, Bosch Institute, School of Medical Sciences, The University of Sydney.
Bosch Institute Advanced Microscopy Facility Workshops Summer/Autumn 2017 Presented by Dr Louise Cole, Advanced Microscopy Facility, Bosch Institute, School of Medical Sciences, The University of Sydney.
Using Virtual Slides in Medical Education with the Virtual Slice System. Jack Glaser, President MicroBrightField, Inc.
 Using Virtual Slides in Medical Education with the Virtual Slice System Jack Glaser, President MicroBrightField, Inc. Advantages of Virtual Slides Overview 0.08x Single section 0.63x Entire 2 x3 inch
Using Virtual Slides in Medical Education with the Virtual Slice System Jack Glaser, President MicroBrightField, Inc. Advantages of Virtual Slides Overview 0.08x Single section 0.63x Entire 2 x3 inch
Presentation and analysis of multidimensional data sets
 Presentation and analysis of multidimensional data sets Overview 1. 3D data visualisation Multidimensional images Data pre-processing Visualisation methods Multidimensional images wavelength time 3D image
Presentation and analysis of multidimensional data sets Overview 1. 3D data visualisation Multidimensional images Data pre-processing Visualisation methods Multidimensional images wavelength time 3D image
TissueFAXS SL Confocal high throughput configuration (actual appearance of the product may differ)
 TISSUEFAXS CONFOCAL TissueFAXS SL Confocal high throughput configuration (actual appearance of the product may differ) TissueFAXS Confocal provides a unique combination of digital slide scanning and laser
TISSUEFAXS CONFOCAL TissueFAXS SL Confocal high throughput configuration (actual appearance of the product may differ) TissueFAXS Confocal provides a unique combination of digital slide scanning and laser
Algorithm User Guide:
 Algorithm User Guide: Membrane Quantification Use the Aperio algorithms to adjust (tune) the parameters until the quantitative results are sufficiently accurate for the purpose for which you intend to
Algorithm User Guide: Membrane Quantification Use the Aperio algorithms to adjust (tune) the parameters until the quantitative results are sufficiently accurate for the purpose for which you intend to
Imaris 3D and 4D interactive analysis and visualization solutions for the life sciences
 Imaris 3D and 4D interactive analysis and visualization solutions for the life sciences Imaris Imaris State of the Art Image Visualization and Analysis Over the last 25 years Imaris has continuously improved
Imaris 3D and 4D interactive analysis and visualization solutions for the life sciences Imaris Imaris State of the Art Image Visualization and Analysis Over the last 25 years Imaris has continuously improved
Bulletin 80C01C01-01E
 Bulletin 80C01C01-01E *1 *2 The CSU-X1 is the advanced model of our CSU-series, which are widely recognized as the most powerful tools for live cell imaging. Faster The world's fastest scanning speed (up
Bulletin 80C01C01-01E *1 *2 The CSU-X1 is the advanced model of our CSU-series, which are widely recognized as the most powerful tools for live cell imaging. Faster The world's fastest scanning speed (up
Leica Microsystems Intelligent Structured Illumination Microscopy
 Widefield Mouse kidney section. Maximum projection of a stack containing 65 planes. Green: glomeruli and convoluted tubules (wheat germ agglutinin Alexa Fluor 488) Blue: Nuclei (DAPI). Structured Illumination
Widefield Mouse kidney section. Maximum projection of a stack containing 65 planes. Green: glomeruli and convoluted tubules (wheat germ agglutinin Alexa Fluor 488) Blue: Nuclei (DAPI). Structured Illumination
Colin Paul Updated 14 November Preparation of publication-quality videos using ImageJ
 Preparation of publication-quality videos using ImageJ Statements made in scientific papers are often much easier to understand if they are supplemented by representative videos. In the best case, these
Preparation of publication-quality videos using ImageJ Statements made in scientific papers are often much easier to understand if they are supplemented by representative videos. In the best case, these
OBCOL. (Organelle Based CO-Localisation) Users Guide
 OBCOL (Organelle Based CO-Localisation) Users Guide INTRODUCTION OBCOL is an ImageJ plugin designed to autonomously detect objects within an image (or image stack) and analyse them separately as individual
OBCOL (Organelle Based CO-Localisation) Users Guide INTRODUCTION OBCOL is an ImageJ plugin designed to autonomously detect objects within an image (or image stack) and analyse them separately as individual
Imaris 3D and 4D interactive analysis and visualization solutions for the life sciences
 Imaris 3D and 4D interactive analysis and visualization solutions for the life sciences Imaris A Brief History Imaris was created 25 years ago to focus purely on the visualization of 3D fluorescence images
Imaris 3D and 4D interactive analysis and visualization solutions for the life sciences Imaris A Brief History Imaris was created 25 years ago to focus purely on the visualization of 3D fluorescence images
Course 4 Advanced Image Analysis & Processing. April 2015
 Course 4 Advanced Image Analysis & Processing April 2015 1 From Survey Results Heterogenous audience Day 1 : ImageJ -Macro : language, batch, use of ActionBar. Day2 : ImageJ - Plugins : install, writing,
Course 4 Advanced Image Analysis & Processing April 2015 1 From Survey Results Heterogenous audience Day 1 : ImageJ -Macro : language, batch, use of ActionBar. Day2 : ImageJ - Plugins : install, writing,
Multi Time Series Rev
 Multi Time Series Rev. 4.0.12 The Multi Time Series program is designed to provide flexible programming of automated time dependent experiments. The basic programming unit is a single Time Series Block
Multi Time Series Rev. 4.0.12 The Multi Time Series program is designed to provide flexible programming of automated time dependent experiments. The basic programming unit is a single Time Series Block
1. Getting Started: Brief Step-By-Step Guide (PDF File)
 Page 1 of 5 1. Getting Started: Brief Step-By-Step Guide (PDF File) Leica SP2 confocal microscope is controlled via software LCS (Leica Confocal Software). The latest version is V2.5. Bellowing is a brief
Page 1 of 5 1. Getting Started: Brief Step-By-Step Guide (PDF File) Leica SP2 confocal microscope is controlled via software LCS (Leica Confocal Software). The latest version is V2.5. Bellowing is a brief
Lecture 19 - Applied Image Analysis
 Lecture 19 - Applied Image Analysis Graeme Ball Three Questions 1. Why should you be interested in image processing & analysis? 2. What are the key concepts & techniques? 3. How can you apply them most
Lecture 19 - Applied Image Analysis Graeme Ball Three Questions 1. Why should you be interested in image processing & analysis? 2. What are the key concepts & techniques? 3. How can you apply them most
NDD FLIM Systems for Leica SP2 MP and SP5 MP Multiphoton Microscopes
 NDD FLIM Systems for Leica SP2 MP and SP5 MP Multiphoton Microscopes bh FLIM systems for the confocal and the multiphoton versions of the Leica SP2 and SP5 microscopes are available since 2002 [4]. These
NDD FLIM Systems for Leica SP2 MP and SP5 MP Multiphoton Microscopes bh FLIM systems for the confocal and the multiphoton versions of the Leica SP2 and SP5 microscopes are available since 2002 [4]. These
Cell Image Analyzer - A visual scripting interface for ImageJ and its usage at the microscopy facility Montpellier RIO Imaging
 Cell Image Analyzer - A visual scripting interface for ImageJ and its usage at the microscopy facility Montpellier RIO Imaging Volker Baecker a and Pierre Travo a a Montpellier RIO Imaging, CNRS, 1919,
Cell Image Analyzer - A visual scripting interface for ImageJ and its usage at the microscopy facility Montpellier RIO Imaging Volker Baecker a and Pierre Travo a a Montpellier RIO Imaging, CNRS, 1919,
IMARIS 3D and 4D interactive analysis and visualization solutions for the life sciences
 IMARIS 3D and 4D interactive analysis and visualization solutions for the life sciences IMARIS A Brief History For over 20 years Bitplane has offered enabling scientific software tools for the life science
IMARIS 3D and 4D interactive analysis and visualization solutions for the life sciences IMARIS A Brief History For over 20 years Bitplane has offered enabling scientific software tools for the life science
PHASEQUANT (User Manual)
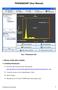 PHASEQUANT (User Manual) ROI Pane Toolbar Threshold Bar Canvas Threshold Display Training Pane Buttons Fig 1. PhaseQuant GUI 1. INSTALLATION AND LOADING 1.1 Installing PhaseQuant Download the zip file
PHASEQUANT (User Manual) ROI Pane Toolbar Threshold Bar Canvas Threshold Display Training Pane Buttons Fig 1. PhaseQuant GUI 1. INSTALLATION AND LOADING 1.1 Installing PhaseQuant Download the zip file
Samples Carolina sample slides (pollen, algae, ). Clean off oil with lens paper then OpticPad around lens (metal not glass) when done.
 Bi/BE 227 Winter 2018 Assignment #1 Widefield and confocal laser scanning microscopy Schedule: Jan 3: Lecture Jan 3-12: Students get trained on how to use scopes, start on assignment Jan 3-17: Carrying
Bi/BE 227 Winter 2018 Assignment #1 Widefield and confocal laser scanning microscopy Schedule: Jan 3: Lecture Jan 3-12: Students get trained on how to use scopes, start on assignment Jan 3-17: Carrying
COLOCALISATION. Alexia Loynton-Ferrand. Imaging Core Facility Biozentrum Basel
 COLOCALISATION Alexia Loynton-Ferrand Imaging Core Facility Biozentrum Basel OUTLINE Introduction How to best prepare your samples for colocalisation How to acquire the images for colocalisation How to
COLOCALISATION Alexia Loynton-Ferrand Imaging Core Facility Biozentrum Basel OUTLINE Introduction How to best prepare your samples for colocalisation How to acquire the images for colocalisation How to
Algorithm User Guide:
 Algorithm User Guide: Microvessel Analysis Use the Aperio algorithms to adjust (tune) the parameters until the quantitative results are sufficiently accurate for the purpose for which you intend to use
Algorithm User Guide: Microvessel Analysis Use the Aperio algorithms to adjust (tune) the parameters until the quantitative results are sufficiently accurate for the purpose for which you intend to use
Colocalization in Confocal Microscopy. Quantitative Colocalization Analysis
 Quantitative Colocalization Analysis 1 Quantitative Colocalization Analysis Colocalization of fluorescently labeled molecules (immunostainings, fluorescent proteins, etc.) is often used as the first indicator
Quantitative Colocalization Analysis 1 Quantitative Colocalization Analysis Colocalization of fluorescently labeled molecules (immunostainings, fluorescent proteins, etc.) is often used as the first indicator
Colocalisation/Correlation
 Colocalisation/Correlation The past: I see yellow - therefore there is colocalisation but published images look over exposed. No colocalisation definition + No stats = No Science. From Now On: 3D. Quantification.
Colocalisation/Correlation The past: I see yellow - therefore there is colocalisation but published images look over exposed. No colocalisation definition + No stats = No Science. From Now On: 3D. Quantification.
DeltaVision OMX SR super-resolution microscope. Super-resolution doesn t need to be complicated
 DeltaVision OMX SR super-resolution microscope Super-resolution doesn t need to be complicated Live-cell super-resolution microscopy DeltaVision OMX SR is a compact super-resolution microscope system optimized
DeltaVision OMX SR super-resolution microscope Super-resolution doesn t need to be complicated Live-cell super-resolution microscopy DeltaVision OMX SR is a compact super-resolution microscope system optimized
Building Your Own 2-Photon Microscope: Challenges, Advantages and Limitations
 Building Your Own 2-Photon : Challenges, Advantages and Limitations Roberto Weigert, Ph.D. Intracellular Membrane Trafficking Unit Oral and Pharyngeal Cancer Branch NIDCR-NIH Building Your Own 2-Photon
Building Your Own 2-Photon : Challenges, Advantages and Limitations Roberto Weigert, Ph.D. Intracellular Membrane Trafficking Unit Oral and Pharyngeal Cancer Branch NIDCR-NIH Building Your Own 2-Photon
Chemical Characterization of Diverse Pharmaceutical Samples by Confocal Raman Microscopy
 Whitepaper Chemical Characterization of Diverse Pharmaceutical Samples by Confocal Raman Microscopy WITec GmbH, Lise-Meitner-Str. 6, 89081 Ulm, Germany, www.witec.de Introduction The development and production
Whitepaper Chemical Characterization of Diverse Pharmaceutical Samples by Confocal Raman Microscopy WITec GmbH, Lise-Meitner-Str. 6, 89081 Ulm, Germany, www.witec.de Introduction The development and production
ONBI Practical 7: Comparison of techniques
 ONBI Practical 7: Comparison of techniques RM Parton 2014 Aims of practical 7: One of the most common issues confronting people new to microscopy is the confusing array of different techniques available.
ONBI Practical 7: Comparison of techniques RM Parton 2014 Aims of practical 7: One of the most common issues confronting people new to microscopy is the confusing array of different techniques available.
HCImage. Image Acquisition and Analysis Software. Light... Camera... Acquisition...
 HCImage Image Acquisition and Analysis Software Light... Camera... Acquisition... HCImage Analysis HCImage Analysis provides comprehensive control of Hamamatsu cameras, microscopes, stages and other peripheral
HCImage Image Acquisition and Analysis Software Light... Camera... Acquisition... HCImage Analysis HCImage Analysis provides comprehensive control of Hamamatsu cameras, microscopes, stages and other peripheral
EPFL SV PTBIOP BIOP COURSE 2015 OPTICAL SLICING METHODS
 BIOP COURSE 2015 OPTICAL SLICING METHODS OPTICAL SLICING METHODS Scanning Methods Wide field Methods Point Scanning Deconvolution Line Scanning Multiple Beam Scanning Single Photon Multiple Photon Total
BIOP COURSE 2015 OPTICAL SLICING METHODS OPTICAL SLICING METHODS Scanning Methods Wide field Methods Point Scanning Deconvolution Line Scanning Multiple Beam Scanning Single Photon Multiple Photon Total
Quantitative Image Analysis and 3-D Digital Reconstruction of Aortic Valve Leaflet
 Quantitative Image Analysis and 3-D Digital Reconstruction of Aortic Valve Leaflet Chi Zheng 1 Mentors: John A. Stella 2 and Michael S. Sacks, Ph. D. 2 1 Bioengineering and Bioinformatics Summer Institute,
Quantitative Image Analysis and 3-D Digital Reconstruction of Aortic Valve Leaflet Chi Zheng 1 Mentors: John A. Stella 2 and Michael S. Sacks, Ph. D. 2 1 Bioengineering and Bioinformatics Summer Institute,
Zeiss LSM 510 Meta inverted confocal manual
 Power Up Zeiss LSM 510 Meta inverted confocal manual *adapted from the Duke University Light Microscopy Core Facility manual 1. Turn on the mercury lamp (if required) and sign in the log book, recording
Power Up Zeiss LSM 510 Meta inverted confocal manual *adapted from the Duke University Light Microscopy Core Facility manual 1. Turn on the mercury lamp (if required) and sign in the log book, recording
Olympus IX-70 Imaging Protocol
 Olympus IX-70 Imaging Protocol 1) System Startup F Please note our sign-up policy. You must inform the facility at least 24 hours beforehand if you can t come; otherwise, you will receive a charge for
Olympus IX-70 Imaging Protocol 1) System Startup F Please note our sign-up policy. You must inform the facility at least 24 hours beforehand if you can t come; otherwise, you will receive a charge for
Confocal 6 Guide. Switching On Components of the system should be switched on in the following order: 1. Lasers Argon laser (488nm and 514nm)
 Confocal 6 Guide The system is kept at 37⁰C. If you require the system at a lower temperature book at least one hour before the beginning of your session to allow the system to cool down. It is your responsibility
Confocal 6 Guide The system is kept at 37⁰C. If you require the system at a lower temperature book at least one hour before the beginning of your session to allow the system to cool down. It is your responsibility
Colocalization tutorial
 Colocalization tutorial I. Introduction Fluorescence microscopy is helpful for determining when two proteins colocalize in the cell. At the most basic level, images can be inspected by eye. Quantitative
Colocalization tutorial I. Introduction Fluorescence microscopy is helpful for determining when two proteins colocalize in the cell. At the most basic level, images can be inspected by eye. Quantitative
SynPAnal User Manual. Eric Danielson and Sang H. Lee Medical College of Wisconsin
 SynPAnal User Manual Eric Danielson and Sang H. Lee Medical College of Wisconsin Installation Instructions Make sure the most recent version of JAVA is installed and is functioning. Unzip the SynPAnal_Installation.zip
SynPAnal User Manual Eric Danielson and Sang H. Lee Medical College of Wisconsin Installation Instructions Make sure the most recent version of JAVA is installed and is functioning. Unzip the SynPAnal_Installation.zip
Virtual Frap User Guide
 Virtual Frap User Guide http://wiki.vcell.uchc.edu/twiki/bin/view/vcell/vfrap Center for Cell Analysis and Modeling University of Connecticut Health Center 2010-1 - 1 Introduction Flourescence Photobleaching
Virtual Frap User Guide http://wiki.vcell.uchc.edu/twiki/bin/view/vcell/vfrap Center for Cell Analysis and Modeling University of Connecticut Health Center 2010-1 - 1 Introduction Flourescence Photobleaching
COLOCALISATION. Alexia Ferrand. Imaging Core Facility Biozentrum Basel
 COLOCALISATION Alexia Ferrand Imaging Core Facility Biozentrum Basel OUTLINE Introduction How to best prepare your samples for colocalisation How to acquire the images for colocalisation How to analyse
COLOCALISATION Alexia Ferrand Imaging Core Facility Biozentrum Basel OUTLINE Introduction How to best prepare your samples for colocalisation How to acquire the images for colocalisation How to analyse
Image processing and quantitative image analysis
 CMB 551 Module 1A Image processing and quantitative image analysis Sam Johnson Benjamin Carlson http://microscopy.duke.edu/ benjamin.carlson@duke.edu Quantification principles Images are collections of
CMB 551 Module 1A Image processing and quantitative image analysis Sam Johnson Benjamin Carlson http://microscopy.duke.edu/ benjamin.carlson@duke.edu Quantification principles Images are collections of
cief Data Analysis Chapter Overview Chapter 12:
 page 285 Chapter 12: cief Data Analysis Chapter Overview Analysis Screen Overview Opening Run Files How Run Data is Displayed Viewing Run Data Data Notifications and Warnings Checking Your Results Group
page 285 Chapter 12: cief Data Analysis Chapter Overview Analysis Screen Overview Opening Run Files How Run Data is Displayed Viewing Run Data Data Notifications and Warnings Checking Your Results Group
BRAT Manual V1.1. May 6th, Christian Göschl & Wolfgang Busch. for correspondence:
 BRAT Manual V1.1 May 6th, 2014 Christian Göschl & Wolfgang Busch Email for correspondence: wolfgang.busch@gmi.oeaw.ac.at Gregor Mendel Institute Dr. Bohr-Gasse 3 1030 Vienna Austria 1 2 Brat Segmentation
BRAT Manual V1.1 May 6th, 2014 Christian Göschl & Wolfgang Busch Email for correspondence: wolfgang.busch@gmi.oeaw.ac.at Gregor Mendel Institute Dr. Bohr-Gasse 3 1030 Vienna Austria 1 2 Brat Segmentation
The Paper Project Guide to Confocal Imaging at Home & in the Classroom
 The Paper Project Guide to Confocal Imaging at Home & in the Classroom http://lifesciences.asu.edu/paperproject The Paper Project Guide to Confocal Imaging at Home & in the Classroom CJ Kazilek & Dennis
The Paper Project Guide to Confocal Imaging at Home & in the Classroom http://lifesciences.asu.edu/paperproject The Paper Project Guide to Confocal Imaging at Home & in the Classroom CJ Kazilek & Dennis
ND Processing Tools in NIS-Elements
 ND Processing Tools in NIS-Elements Overview This technical note describes basic uses of the ND processing tools available in NIS-Elements. These tools are specifically designed for arithmetic functions
ND Processing Tools in NIS-Elements Overview This technical note describes basic uses of the ND processing tools available in NIS-Elements. These tools are specifically designed for arithmetic functions
ImageJ2 Directions & Goals
 ImageJ2 Directions & Goals Strengths of ImageJ Cross-platform and public domain Macro language makes batch processing easy Wealth of existing image processing plugins Large science-oriented community of
ImageJ2 Directions & Goals Strengths of ImageJ Cross-platform and public domain Macro language makes batch processing easy Wealth of existing image processing plugins Large science-oriented community of
3. Open ImageJ, select Plugin Macro Run and then select NeuronRead, which will start immediately.
 NeuronRead Tutorial Setup The NeuronRead 1 macro runs on ImageJ software and is supported by three main plugins: MorphoLibJ, Skeletonize3D and Analyze Skeleton. For the smoothest and fastest application
NeuronRead Tutorial Setup The NeuronRead 1 macro runs on ImageJ software and is supported by three main plugins: MorphoLibJ, Skeletonize3D and Analyze Skeleton. For the smoothest and fastest application
Basic Users Guide for the LSM 510 Confocal Microscope
 Basic Users Guide for the LSM 510 Confocal Microscope Ian Jones and Adam Westmacott Department of Biology & Biochemistry, University of Bath, Bath, UK Updated 01 October 2003 This guide describes the basic
Basic Users Guide for the LSM 510 Confocal Microscope Ian Jones and Adam Westmacott Department of Biology & Biochemistry, University of Bath, Bath, UK Updated 01 October 2003 This guide describes the basic
IN Cell Analyzer 1000 Analysis Modules Multi Target Analysis Training
 IN Cell Analyzer 1000 Analysis Modules Multi Target Analysis Training Multi Target Analysis Neurite Outgrowth Membrane Translocation Analysis Object Intensity Dual Area Object Analysis Granularity Morphology
IN Cell Analyzer 1000 Analysis Modules Multi Target Analysis Training Multi Target Analysis Neurite Outgrowth Membrane Translocation Analysis Object Intensity Dual Area Object Analysis Granularity Morphology
Visualization on BioHPC
 Visualization on BioHPC [web] [email] portal.biohpc.swmed.edu biohpc-help@utsouthwestern.edu 1 Updated for 2015-09-16 Outline What is Visualization - Scientific Visualization - Work flow for Visualization
Visualization on BioHPC [web] [email] portal.biohpc.swmed.edu biohpc-help@utsouthwestern.edu 1 Updated for 2015-09-16 Outline What is Visualization - Scientific Visualization - Work flow for Visualization
CFIM MICROSCOPY COURSE TIMETABLE PRINCIPLES OF MICROSCOPY MONDAY 6 TH OF JANUARY 2014 FRIDAY 10 TH OF JANUARY 2014
 MICROSCOPY COURSE TIMETABLE PRINCIPLES OF MICROSCOPY MONDAY 6 TH OF JANUARY 2014 FRIDAY 10 TH OF JANUARY 2014 CONFOCAL AND FLUORESCENCE MICROSCOPY MONDAY 20 TH OF JANUARY 2014 FRIDAY 24 TH OF JANUARY 2014
MICROSCOPY COURSE TIMETABLE PRINCIPLES OF MICROSCOPY MONDAY 6 TH OF JANUARY 2014 FRIDAY 10 TH OF JANUARY 2014 CONFOCAL AND FLUORESCENCE MICROSCOPY MONDAY 20 TH OF JANUARY 2014 FRIDAY 24 TH OF JANUARY 2014
Olympus DeltaVision Microscope Start-Up and Shut-Down Instructions.
 DeltaVision Instructions: 1. Start-up (Olympus DV) 2. Basic operation (Olympus DV) 2.1 set-up 2.2 designing and running an experiment 3. Transferring files via FTP 4. Deconvolving files 5. File conversion
DeltaVision Instructions: 1. Start-up (Olympus DV) 2. Basic operation (Olympus DV) 2.1 set-up 2.2 designing and running an experiment 3. Transferring files via FTP 4. Deconvolving files 5. File conversion
Automatically Improving 3D Neuron Segmentations for Expansion Microscopy Connectomics. by Albert Gerovitch
 Automatically Improving 3D Neuron Segmentations for Expansion Microscopy Connectomics by Albert Gerovitch 1 Abstract Understanding the geometry of neurons and their connections is key to comprehending
Automatically Improving 3D Neuron Segmentations for Expansion Microscopy Connectomics by Albert Gerovitch 1 Abstract Understanding the geometry of neurons and their connections is key to comprehending
LEXT 3D Measuring LASER Microscope
 LEXT 3D Measuring LASER Microscope Warning: This instrument may only be operated by those who have been trained by AAF staff and have read and signed the AAF laboratory policies. A) STARTUP 1. Computer
LEXT 3D Measuring LASER Microscope Warning: This instrument may only be operated by those who have been trained by AAF staff and have read and signed the AAF laboratory policies. A) STARTUP 1. Computer
High%Throughput,Microscopy,Course:,P11, Kota,Miura,&,Andrea,Boni, EMBL,Heidelberg, ,20,
 High%ThroughputMicroscopyCourse:P11 KotaMiura&AndreaBoni EMBLHeidelberg 2012.10.1920 Task TASK:AutomaPcmeasurementofintensity changesoverpmeatthevicinityofnuclear membrane. Trytheprogramworking. OpentheImage(2chstack)
High%ThroughputMicroscopyCourse:P11 KotaMiura&AndreaBoni EMBLHeidelberg 2012.10.1920 Task TASK:AutomaPcmeasurementofintensity changesoverpmeatthevicinityofnuclear membrane. Trytheprogramworking. OpentheImage(2chstack)
Technology and equipment at the Bioimaging Center
 Technology and equipment at the Bioimaging Center Light microscopy / Widefields Name Type Applications and equipment LEICA AF6000LX Nikon BioStation Timelaps imaging, CO2 and temperature controlled (14
Technology and equipment at the Bioimaging Center Light microscopy / Widefields Name Type Applications and equipment LEICA AF6000LX Nikon BioStation Timelaps imaging, CO2 and temperature controlled (14
Introduction. Loading Images
 Introduction CellProfiler is a free Open Source software for automated image analysis. Versions for Mac, Windows and Linux are available and can be downloaded at: http://www.cellprofiler.org/. CellProfiler
Introduction CellProfiler is a free Open Source software for automated image analysis. Versions for Mac, Windows and Linux are available and can be downloaded at: http://www.cellprofiler.org/. CellProfiler
Characteristic Quantities of Microvascular Structures in CLSM Volume Datasets K. Winter¹, L. H.-W. Metz, J.-P. Kuska², B. Frerich³
 Characteristic Quantities of Microvascular Structures in CLSM Volume Datasets K. Winter¹, L. H.-W. Metz, J.-P. Kuska², B. Frerich³ ¹Translational Centre for Regenerative Medicine (TRM-Leipzig), University
Characteristic Quantities of Microvascular Structures in CLSM Volume Datasets K. Winter¹, L. H.-W. Metz, J.-P. Kuska², B. Frerich³ ¹Translational Centre for Regenerative Medicine (TRM-Leipzig), University
SPCM Software Runs Online-FLIM at 10 Images per Second
 SPCM Software Runs Online-FLIM at 10 Images per Second Abstract: Version 9.72 SPCM software of the bh TCSPC/FLIM systems displays fluorescence lifetime images at a rate of 10 images per second. The calculation
SPCM Software Runs Online-FLIM at 10 Images per Second Abstract: Version 9.72 SPCM software of the bh TCSPC/FLIM systems displays fluorescence lifetime images at a rate of 10 images per second. The calculation
Supplementary Figures
 Supplementary Figures re co rd ed ch annels darkf ield MPM2 PI cell cycle phases G1/S/G2 prophase metaphase anaphase telophase bri ghtfield 55 pixel Supplementary Figure 1 Images of Jurkat cells captured
Supplementary Figures re co rd ed ch annels darkf ield MPM2 PI cell cycle phases G1/S/G2 prophase metaphase anaphase telophase bri ghtfield 55 pixel Supplementary Figure 1 Images of Jurkat cells captured
MAT 17C - DISCUSSION #4, Counting Proteins
 MAT 17C - DISCUSSION #4, Counting Proteins Visualization of molecules inside a living cell is one of the most important tools of molecular biology in the 21 st century. In fact, it was the subject of the
MAT 17C - DISCUSSION #4, Counting Proteins Visualization of molecules inside a living cell is one of the most important tools of molecular biology in the 21 st century. In fact, it was the subject of the
MetaMorph Standard Operation Protocol Basic Application
 MetaMorph Standard Operation Protocol Basic Application Contents Basic Navigation and Image Handling... 2 Opening Images... 2 Separating Multichannel Images... 2 Cropping an Image... 3 Changing an 8 bit
MetaMorph Standard Operation Protocol Basic Application Contents Basic Navigation and Image Handling... 2 Opening Images... 2 Separating Multichannel Images... 2 Cropping an Image... 3 Changing an 8 bit
LSM510 Confocal Microscope Standard Operation Protocol Basic Operation
 LSM510 Confocal Microscope Standard Operation Protocol Basic Operation Please make sure that the COMPRESSED AIR has been TURNED ON prior to the use of the equipment. Kindly inform the administrator if
LSM510 Confocal Microscope Standard Operation Protocol Basic Operation Please make sure that the COMPRESSED AIR has been TURNED ON prior to the use of the equipment. Kindly inform the administrator if
Find the Correspondences
 Advanced Topics in BioImage Analysis - 3. Find the Correspondences from Registration to Motion Estimation CellNetworks Math-Clinic Qi Gao 03.02.2017 Math-Clinic core facility Provide services on bioinformatics
Advanced Topics in BioImage Analysis - 3. Find the Correspondences from Registration to Motion Estimation CellNetworks Math-Clinic Qi Gao 03.02.2017 Math-Clinic core facility Provide services on bioinformatics
Fluorescence Tomography Source Reconstruction and Analysis
 TECHNICAL NOTE Pre-clinical in vivo imaging Fluorescence Tomography Source Reconstruction and Analysis Note: This Technical Note is part of a series for Fluorescence Imaging Tomography (FLIT). The user
TECHNICAL NOTE Pre-clinical in vivo imaging Fluorescence Tomography Source Reconstruction and Analysis Note: This Technical Note is part of a series for Fluorescence Imaging Tomography (FLIT). The user
Simplified model of ray propagation and outcoupling in TFs.
 Supplementary Figure 1 Simplified model of ray propagation and outcoupling in TFs. After total reflection at the core boundary with an angle α, a ray entering the taper (blue line) hits the taper sidewalls
Supplementary Figure 1 Simplified model of ray propagation and outcoupling in TFs. After total reflection at the core boundary with an angle α, a ray entering the taper (blue line) hits the taper sidewalls
Effect of OTF attenuation on single-slice SR-SIM reconstruction
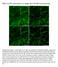 Effect of OTF attenuation on single-slice SR-SIM reconstruction Supplementary Figure 1: Actin filaments in U2OS cells, labelled with Phalloidin-Atto488, measured on a DeltaVision OMX, excited at 488 nm
Effect of OTF attenuation on single-slice SR-SIM reconstruction Supplementary Figure 1: Actin filaments in U2OS cells, labelled with Phalloidin-Atto488, measured on a DeltaVision OMX, excited at 488 nm
Introduction to Advanced Instruments
 Introduction to Advanced Instruments Multiphoton and Confocal Microscope System 黃兆祺 國立交通大學生物科技學院分子醫學與生物工程研究所 Phone: 03-5712121 ext 56968 (office) 56969 (Lab) E-mail: hwangeric@mail.nctu.edu.tw 1 Fluorescence
Introduction to Advanced Instruments Multiphoton and Confocal Microscope System 黃兆祺 國立交通大學生物科技學院分子醫學與生物工程研究所 Phone: 03-5712121 ext 56968 (office) 56969 (Lab) E-mail: hwangeric@mail.nctu.edu.tw 1 Fluorescence
PH880 Topics in Physics
 PH880 Topics in Physics Modern Optical Imaging (Fall 2010) Overview of week 8 Monday Nonlinear Microscopy Wednesday No class (Mid term week) Quantum Optics Electro- magnetic Optics Wave Optics Ray Optics
PH880 Topics in Physics Modern Optical Imaging (Fall 2010) Overview of week 8 Monday Nonlinear Microscopy Wednesday No class (Mid term week) Quantum Optics Electro- magnetic Optics Wave Optics Ray Optics
A Model Based Neuron Detection Approach Using Sparse Location Priors
 A Model Based Neuron Detection Approach Using Sparse Location Priors Electronic Imaging, Burlingame, CA 30 th January 2017 Soumendu Majee 1 Dong Hye Ye 1 Gregery T. Buzzard 2 Charles A. Bouman 1 1 Department
A Model Based Neuron Detection Approach Using Sparse Location Priors Electronic Imaging, Burlingame, CA 30 th January 2017 Soumendu Majee 1 Dong Hye Ye 1 Gregery T. Buzzard 2 Charles A. Bouman 1 1 Department
SIVIC GUI Overview. SIVIC GUI Layout Overview
 SIVIC GUI Overview SIVIC GUI Layout Overview At the top of the SIVIC GUI is a row of buttons called the Toolbar. It is a quick interface for loading datasets, controlling how the mouse manipulates the
SIVIC GUI Overview SIVIC GUI Layout Overview At the top of the SIVIC GUI is a row of buttons called the Toolbar. It is a quick interface for loading datasets, controlling how the mouse manipulates the
amira 5 Visualize Analyze Present
 amira 5 Visualize Analyze Present Visioneering Your Ideas Engineering Your Vision Your research is inspired by knowledge, intuition, and a powerful vision. Yet you are facing many challenges when moving
amira 5 Visualize Analyze Present Visioneering Your Ideas Engineering Your Vision Your research is inspired by knowledge, intuition, and a powerful vision. Yet you are facing many challenges when moving
Essentials of Biological Image Analysis
 Essentials of Biological Image Analysis Volker Baecker INSERM BioCampus Montpellier (UMS 3426) MRI TIGR 27/04/2012 Overview 1. Digital image 2. Basic Image Analysis 1. Point Operations 2. Local Filtering
Essentials of Biological Image Analysis Volker Baecker INSERM BioCampus Montpellier (UMS 3426) MRI TIGR 27/04/2012 Overview 1. Digital image 2. Basic Image Analysis 1. Point Operations 2. Local Filtering
TrakEM2 workshop. Albert Cardona
 TrakEM2 workshop Albert Cardona 1 Introduction 2 Motivation Our current work in neuroscience requires detailed anatomical descriptions of dendrites and axons (the wires), and synapses (the contact points).
TrakEM2 workshop Albert Cardona 1 Introduction 2 Motivation Our current work in neuroscience requires detailed anatomical descriptions of dendrites and axons (the wires), and synapses (the contact points).
Image restoration by deconvolution
 Image restoration by deconvolution chong.zhang@bioquant.uni-heidelberg.de 17/12/2014 (part) Slides courtesy: Sébastien Tosi (IRB Barcelona) A few concepts related to the topic Convolution Deconvolution
Image restoration by deconvolution chong.zhang@bioquant.uni-heidelberg.de 17/12/2014 (part) Slides courtesy: Sébastien Tosi (IRB Barcelona) A few concepts related to the topic Convolution Deconvolution
Image Analysis & Cell Phenotyping
 Image Analysis & Cell Phenotyping Biological Image Processing Programs Feature ImageJ CellProfiler MetaMorph Definiens Matlab Neurolucida Price Free Free $$ $$$$ $ $$ Flexibility ++++ + ++ Optimized for
Image Analysis & Cell Phenotyping Biological Image Processing Programs Feature ImageJ CellProfiler MetaMorph Definiens Matlab Neurolucida Price Free Free $$ $$$$ $ $$ Flexibility ++++ + ++ Optimized for
Fiji Is Just ImageJ an open source platform for biological image analysis
 Fiji Is Just ImageJ an open source platform for biological image analysis Pavel Tomancak Max Planck Institute of Molecular Cell Biology and Genetics in Dresden (MPI-CBG) OME meeting, Paris June 15th Why
Fiji Is Just ImageJ an open source platform for biological image analysis Pavel Tomancak Max Planck Institute of Molecular Cell Biology and Genetics in Dresden (MPI-CBG) OME meeting, Paris June 15th Why
