Gibbs Sampling. Stephen F. Altschul. National Center for Biotechnology Information National Library of Medicine National Institutes of Health
|
|
|
- Philip Miller
- 5 years ago
- Views:
Transcription
1 Gibbs Sampling Stephen F. Altschul National Center for Biotechnology Information National Library of Medicine National Institutes of Health
2 Optimization in High-Dimensional Space Smooth and simple landscapes Relatively easy to find optimum. Algorithms: Newton s method; gradient descent. Random landscapes Finding optimal solution intractable. Algorithms: Brute force enumeration. Rough but correlated landscapes Difficult to find provably optimum solution. Fairly effective heuristic methods available. Algorithms: Simulated annealing; Gibbs sampling. Success depends on details of landscape. Difficulties: Local optima. Images courtesy of the internet
3 Local Multiple Alignment Simple Version of Problem Input: sequence; pattern width. Problem: Find highest-scoring, ungapped local multiple alignment, involving one segment of length from each sequence. Search space: One may score the local alignment in various ways. Here, we will use BILD scores. Gibbs sampling: Given an alignment, one may easily derive a profile or scoring matrix. Given a profile, one may easily calculate its likelihoods, implied by various segments within a sequence. Gibbs sampling alternates between generating profiles from given alignments, and sampling alignment positions based on given profile, until convergence. Lawrence, C.E., et al. (1993) Science 262:
4 1. Initialization Choose random length- segments from within the input sequences: MTQPSKTTKLTKDEVDRLISDYQTKQDEQAQETLVRVYTNLVDMLAKKYSKGKSFHEDLRQVGMIGLLGAIKRYD PVVGKSFEAFAIPTIIGEIKRFLRDKTWSVHVPRRIKELGPRIKMAVDQLTTETQRSPKVEEIAEFLDVSEEEVL ETMEMGKSYQALSVDHSIEADSDGSTVTILDIVGSQEDGYERVNQQLMLQSVLHVLSDREKQIIDLTYIQNKSQK ETGDILGISQMHVSRLQRKAVKKLREALIEDPSMELM MPPLFVMNNEILMHLRALKKTKKDVSLHDPIGQDKEGNEISLIDVLKSENEDVIDTIQLNMELEKVKQYIDILDD REKEVIVGRFGLDLKKEKTQREIAKELGISRSYVSRIEKRALMKMFHEFYRAEKEKRKKAKGK MELRDLDLNLLVVFNQLLVDRRVSITAENLGLTQPAVSNALKRLRTSLQDPLFVRTHQGMEPTPYAAHLAEPVTS AMHALRNALQHHESFDPLTSERTFTLAMTDIGEIYFMPRLMDVLAHQAPNCVISTVRDSSMSLMQALQNGTVDLA VGLLPNLQTGFFQRRLLQNHYVCLCRKDHPVTREPLTLERFCSYGHVRVIAAGTGHGEVDTYMTRVGIRRDIRLE VPHFAAVGHILQRTDLLATVPIRLADCCVEPFGLSALPHPVVLPEIAINMFWHAKYHKDLANIWLRQLMFDLFTD MNAYTVSRLALDAGVSVHIVRDYLLRGLLRPVACTTGGYGLFDDAALQRLCFVRAAFEAGIGLGALARLCRALDA ANCDETAAQLAVLRQFVERRREALANLEVQLAAMPTAPAQHAESLP
5 2. Remove one segment from alignment Select a sequence at random from among the input sequences, and remove its segment from the multiple alignment: VRVYTNLVDMLAKKY ALMKMFHEFYRAEKE TDIGEIYFMPRLMDV LAVLRQFVERRREAL PSPLYPWMRSQFGKC DDTAIRTVLNQALSR QWERGDSEPTGKNLF YHHIKKEKSPKGKSS RIESALLNKIAMLGT
6 3. Construct a profile from the remaining alignment Multiple alignment: VRVYTNLVDMLAKKY ALMKMFHEFYRAEKE TDIGEIYFMPRLMDV LAVLRQFVERRREAL Log-odd scores: A: C: D: E:
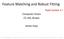
7 4. Calculate relative likelihoods at all positions, and sample Sequence : 0.25 w = Normalized likelihoods: ( 2 ) Probability Position Sample a random position from sequence, weighted by normalized likelihoods. Add the segment at this position to the multiple local alignment. If this new alignment is better than any so far seen, remember it. If there has been no improvement in the last iterations, stop. Otherwise, return to step 2, and remove a new segment.
8 Why Does the Algorithm Work? When no common pattern is represented in the multiple alignment, the positions in sequence to be sampled have roughly equal likelihoods, so the algorithm performs a random walk through the solution space. Once a single segment is chosen that is similar to segments found in most or all sequences, these other segments are slightly favored, and a second related segment may well be sampled. As more related segments are found, the process accelerates, converging on a locally optimal solution. If there are no other good local optima, this solution has a good chance or being the global optimum.
9 Behavior of the Objective Function w = 15 Input 30 HTH sequences Pattern width 15 Pattern score Search space ~10 Time < 1.0 seconds Iteration
10 The Evolving Multiple Alignment 1 iteration 10 iterations 480 iterations 1680 iterations
11 Phase Shifts The Gibbs sampling algorithm may easily converge on a local optimum that is a phase-shifted version of the global optimum. Why? Optimal solution: Solution found: One remedy is to add a separate phase-shift sampling step. No segments are removed, but likelihoods are calculated for the current alignment and several phase-shifted alternatives. These alignments are then sampled among. This can be understood as changing the topology, of definition of distance, on the underlying alignment space. SQKETGDILGISQMHVSRLQRKAVKKL TQREIAKELGISRSYVSRIEKRALMKM RVSITAENLGLTQPAVSNALKRLRTSL CFVRAAFEAGIGLGALARLCRALDAAN RRIEIAHALCLTERQIKIWFQNRRMKW NQIRAADLLGLNRNTLRKKIRDLDIQV TQRSLAKALKISHVSVSQWERGDSEPT EKEEVAKKCGITPLQVRVWFINKRMRS GTEKTAEAVGVDKSQISRWKRDWIPKF
12 How does one choose pattern width? Pattern Width Choosing too small discards available information for locating a pattern, while choosing too large adds unnecessary noise. The Gibbs sampling algorithm, however, should be fairly robust to deviations that are not too far from the optimal. What is a reasonable criterion for optimal pattern width? It can be difficult to compare multiple alignment scores directly for different choices of, especially when all column scores are positive. One criterion for selecting is the Minimum Description Length Principle. For ungapped local multiple alignments, this is equivalent to optimizing the BILD score along a single high-dimensional diagonal, which can be achieved using a variation of the Smith-Waterman algorithm. Employing the criterion of optimal BILD score, may be modified dynamically, within a Gibbs sampling program. Grunwald, P.D. (2007) The Minimum Description Length Principle. MIT Press, Cambridge, MA. Altschul, S.F., et al. (2010) "The construction and use of log-odds substitution scores for multiple sequence alignment." PLoS Comput. Biol. 6:e
13 Close Sequences If two input sequences are too similar to one another, they can cause each other to stick during the sampling stage. In other words, even when they are misaligned, the current position in one sequence will cause the equivalent position in the other sequence to be selected, and vice versa. Possible remedies One may remove extra copies of sequences that are too similar to one another from the input set, and add them back in at a later stage. Paradoxically, this suggests that the most distantly related sequences should be aligned first. Alternatively, one may employ a strategy analogous to the realignment stage in MUSCLE. The relative alignment of a set of closely related sequences can be fixed. Then segments from these sequences can be removed in tandem from the multiple alignment, and new segments (in their previously-fixed relative alignment) sampled in one pass.
14 Several Generalizations of the Problem Some sequences may be missing the pattern. Some sequences may have multiple copies of the pattern. The sequences may contain multiple distinct patterns, either consistently ordered or in arbitrary order. The best alignment between the consensus pattern and its occurrences within the sequences may contain gaps.
An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST
 An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST Alexander Chan 5075504 Biochemistry 218 Final Project An Analysis of Pairwise
An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST Alexander Chan 5075504 Biochemistry 218 Final Project An Analysis of Pairwise
Data Mining Chapter 8: Search and Optimization Methods Fall 2011 Ming Li Department of Computer Science and Technology Nanjing University
 Data Mining Chapter 8: Search and Optimization Methods Fall 2011 Ming Li Department of Computer Science and Technology Nanjing University Search & Optimization Search and Optimization method deals with
Data Mining Chapter 8: Search and Optimization Methods Fall 2011 Ming Li Department of Computer Science and Technology Nanjing University Search & Optimization Search and Optimization method deals with
Biology 644: Bioinformatics
 Find the best alignment between 2 sequences with lengths n and m, respectively Best alignment is very dependent upon the substitution matrix and gap penalties The Global Alignment Problem tries to find
Find the best alignment between 2 sequences with lengths n and m, respectively Best alignment is very dependent upon the substitution matrix and gap penalties The Global Alignment Problem tries to find
x n+1 = x n f(x n) f (x n ), (1)
 1 Optimization The field of optimization is large and vastly important, with a deep history in computer science (among other places). Generally, an optimization problem is defined by having a score function
1 Optimization The field of optimization is large and vastly important, with a deep history in computer science (among other places). Generally, an optimization problem is defined by having a score function
Central Issues in Biological Sequence Comparison
 Central Issues in Biological Sequence Comparison Definitions: What is one trying to find or optimize? Algorithms: Can one find the proposed object optimally or in reasonable time optimize? Statistics:
Central Issues in Biological Sequence Comparison Definitions: What is one trying to find or optimize? Algorithms: Can one find the proposed object optimally or in reasonable time optimize? Statistics:
Today s Lecture. Multiple sequence alignment. Improved scoring of pairwise alignments. Affine gap penalties Profiles
 Today s Lecture Multiple sequence alignment Improved scoring of pairwise alignments Affine gap penalties Profiles 1 The Edit Graph for a Pair of Sequences G A C G T T G A A T G A C C C A C A T G A C G
Today s Lecture Multiple sequence alignment Improved scoring of pairwise alignments Affine gap penalties Profiles 1 The Edit Graph for a Pair of Sequences G A C G T T G A A T G A C C C A C A T G A C G
Linear Equation Systems Iterative Methods
 Linear Equation Systems Iterative Methods Content Iterative Methods Jacobi Iterative Method Gauss Seidel Iterative Method Iterative Methods Iterative methods are those that produce a sequence of successive
Linear Equation Systems Iterative Methods Content Iterative Methods Jacobi Iterative Method Gauss Seidel Iterative Method Iterative Methods Iterative methods are those that produce a sequence of successive
COS 551: Introduction to Computational Molecular Biology Lecture: Oct 17, 2000 Lecturer: Mona Singh Scribe: Jacob Brenner 1. Database Searching
 COS 551: Introduction to Computational Molecular Biology Lecture: Oct 17, 2000 Lecturer: Mona Singh Scribe: Jacob Brenner 1 Database Searching In database search, we typically have a large sequence database
COS 551: Introduction to Computational Molecular Biology Lecture: Oct 17, 2000 Lecturer: Mona Singh Scribe: Jacob Brenner 1 Database Searching In database search, we typically have a large sequence database
Supervised vs. Unsupervised Learning
 Clustering Supervised vs. Unsupervised Learning So far we have assumed that the training samples used to design the classifier were labeled by their class membership (supervised learning) We assume now
Clustering Supervised vs. Unsupervised Learning So far we have assumed that the training samples used to design the classifier were labeled by their class membership (supervised learning) We assume now
HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT
 HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT - Swarbhanu Chatterjee. Hidden Markov models are a sophisticated and flexible statistical tool for the study of protein models. Using HMMs to analyze proteins
HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT - Swarbhanu Chatterjee. Hidden Markov models are a sophisticated and flexible statistical tool for the study of protein models. Using HMMs to analyze proteins
Chapter 6. Multiple sequence alignment (week 10)
 Course organization Introduction ( Week 1,2) Part I: Algorithms for Sequence Analysis (Week 1-11) Chapter 1-3, Models and theories» Probability theory and Statistics (Week 3)» Algorithm complexity analysis
Course organization Introduction ( Week 1,2) Part I: Algorithms for Sequence Analysis (Week 1-11) Chapter 1-3, Models and theories» Probability theory and Statistics (Week 3)» Algorithm complexity analysis
Chapter 8 Multiple sequence alignment. Chaochun Wei Spring 2018
 1896 1920 1987 2006 Chapter 8 Multiple sequence alignment Chaochun Wei Spring 2018 Contents 1. Reading materials 2. Multiple sequence alignment basic algorithms and tools how to improve multiple alignment
1896 1920 1987 2006 Chapter 8 Multiple sequence alignment Chaochun Wei Spring 2018 Contents 1. Reading materials 2. Multiple sequence alignment basic algorithms and tools how to improve multiple alignment
3.4 Multiple sequence alignment
 3.4 Multiple sequence alignment Why produce a multiple sequence alignment? Using more than two sequences results in a more convincing alignment by revealing conserved regions in ALL of the sequences Aligned
3.4 Multiple sequence alignment Why produce a multiple sequence alignment? Using more than two sequences results in a more convincing alignment by revealing conserved regions in ALL of the sequences Aligned
FastA & the chaining problem
 FastA & the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 1 Sources for this lecture: Lectures by Volker Heun, Daniel Huson and Knut Reinert,
FastA & the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 1 Sources for this lecture: Lectures by Volker Heun, Daniel Huson and Knut Reinert,
FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:
 FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:56 4001 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem
FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:56 4001 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem
Non-Stationary Covariance Models for Discontinuous Functions as Applied to Aircraft Design Problems
 Non-Stationary Covariance Models for Discontinuous Functions as Applied to Aircraft Design Problems Trent Lukaczyk December 1, 2012 1 Introduction 1.1 Application The NASA N+2 Supersonic Aircraft Project
Non-Stationary Covariance Models for Discontinuous Functions as Applied to Aircraft Design Problems Trent Lukaczyk December 1, 2012 1 Introduction 1.1 Application The NASA N+2 Supersonic Aircraft Project
Profiles and Multiple Alignments. COMP 571 Luay Nakhleh, Rice University
 Profiles and Multiple Alignments COMP 571 Luay Nakhleh, Rice University Outline Profiles and sequence logos Profile hidden Markov models Aligning profiles Multiple sequence alignment by gradual sequence
Profiles and Multiple Alignments COMP 571 Luay Nakhleh, Rice University Outline Profiles and sequence logos Profile hidden Markov models Aligning profiles Multiple sequence alignment by gradual sequence
Lectures by Volker Heun, Daniel Huson and Knut Reinert, in particular last years lectures
 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 4.1 Sources for this lecture Lectures by Volker Heun, Daniel Huson and Knut
4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 4.1 Sources for this lecture Lectures by Volker Heun, Daniel Huson and Knut
Using the DATAMINE Program
 6 Using the DATAMINE Program 304 Using the DATAMINE Program This chapter serves as a user s manual for the DATAMINE program, which demonstrates the algorithms presented in this book. Each menu selection
6 Using the DATAMINE Program 304 Using the DATAMINE Program This chapter serves as a user s manual for the DATAMINE program, which demonstrates the algorithms presented in this book. Each menu selection
CISC 889 Bioinformatics (Spring 2003) Multiple Sequence Alignment
 CISC 889 Bioinformatics (Spring 2003) Multiple Sequence Alignment Courtesy of jalview 1 Motivations Collective statistic Protein families Identification and representation of conserved sequence features
CISC 889 Bioinformatics (Spring 2003) Multiple Sequence Alignment Courtesy of jalview 1 Motivations Collective statistic Protein families Identification and representation of conserved sequence features
Perceptron: This is convolution!
 Perceptron: This is convolution! v v v Shared weights v Filter = local perceptron. Also called kernel. By pooling responses at different locations, we gain robustness to the exact spatial location of image
Perceptron: This is convolution! v v v Shared weights v Filter = local perceptron. Also called kernel. By pooling responses at different locations, we gain robustness to the exact spatial location of image
Clustering. Supervised vs. Unsupervised Learning
 Clustering Supervised vs. Unsupervised Learning So far we have assumed that the training samples used to design the classifier were labeled by their class membership (supervised learning) We assume now
Clustering Supervised vs. Unsupervised Learning So far we have assumed that the training samples used to design the classifier were labeled by their class membership (supervised learning) We assume now
PROTEIN MULTIPLE ALIGNMENT MOTIVATION: BACKGROUND: Marina Sirota
 Marina Sirota MOTIVATION: PROTEIN MULTIPLE ALIGNMENT To study evolution on the genetic level across a wide range of organisms, biologists need accurate tools for multiple sequence alignment of protein
Marina Sirota MOTIVATION: PROTEIN MULTIPLE ALIGNMENT To study evolution on the genetic level across a wide range of organisms, biologists need accurate tools for multiple sequence alignment of protein
Lecture 4. Convexity Robust cost functions Optimizing non-convex functions. 3B1B Optimization Michaelmas 2017 A. Zisserman
 Lecture 4 3B1B Optimization Michaelmas 2017 A. Zisserman Convexity Robust cost functions Optimizing non-convex functions grid search branch and bound simulated annealing evolutionary optimization The Optimization
Lecture 4 3B1B Optimization Michaelmas 2017 A. Zisserman Convexity Robust cost functions Optimizing non-convex functions grid search branch and bound simulated annealing evolutionary optimization The Optimization
Bioinformatics explained: Smith-Waterman
 Bioinformatics Explained Bioinformatics explained: Smith-Waterman May 1, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com
Bioinformatics Explained Bioinformatics explained: Smith-Waterman May 1, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com
Today. Golden section, discussion of error Newton s method. Newton s method, steepest descent, conjugate gradient
 Optimization Last time Root finding: definition, motivation Algorithms: Bisection, false position, secant, Newton-Raphson Convergence & tradeoffs Example applications of Newton s method Root finding in
Optimization Last time Root finding: definition, motivation Algorithms: Bisection, false position, secant, Newton-Raphson Convergence & tradeoffs Example applications of Newton s method Root finding in
Representation is Everything: Towards Efficient and Adaptable Similarity Measures for Biological Data
 Representation is Everything: Towards Efficient and Adaptable Similarity Measures for Biological Data Charu C. Aggarwal IBM T. J. Watson Research Center charu@us.ibm.com Abstract Distance function computation
Representation is Everything: Towards Efficient and Adaptable Similarity Measures for Biological Data Charu C. Aggarwal IBM T. J. Watson Research Center charu@us.ibm.com Abstract Distance function computation
BLAST, Profile, and PSI-BLAST
 BLAST, Profile, and PSI-BLAST Jianlin Cheng, PhD School of Electrical Engineering and Computer Science University of Central Florida 26 Free for academic use Copyright @ Jianlin Cheng & original sources
BLAST, Profile, and PSI-BLAST Jianlin Cheng, PhD School of Electrical Engineering and Computer Science University of Central Florida 26 Free for academic use Copyright @ Jianlin Cheng & original sources
Efficient Iterative Semi-supervised Classification on Manifold
 . Efficient Iterative Semi-supervised Classification on Manifold... M. Farajtabar, H. R. Rabiee, A. Shaban, A. Soltani-Farani Sharif University of Technology, Tehran, Iran. Presented by Pooria Joulani
. Efficient Iterative Semi-supervised Classification on Manifold... M. Farajtabar, H. R. Rabiee, A. Shaban, A. Soltani-Farani Sharif University of Technology, Tehran, Iran. Presented by Pooria Joulani
Theoretical Concepts of Machine Learning
 Theoretical Concepts of Machine Learning Part 2 Institute of Bioinformatics Johannes Kepler University, Linz, Austria Outline 1 Introduction 2 Generalization Error 3 Maximum Likelihood 4 Noise Models 5
Theoretical Concepts of Machine Learning Part 2 Institute of Bioinformatics Johannes Kepler University, Linz, Austria Outline 1 Introduction 2 Generalization Error 3 Maximum Likelihood 4 Noise Models 5
Computer Vision. Exercise 3 Panorama Stitching 09/12/2013. Compute Vision : Exercise 3 Panorama Stitching
 Computer Vision Exercise 3 Panorama Stitching 09/12/2013 Compute Vision : Exercise 3 Panorama Stitching The task Compute Vision : Exercise 3 Panorama Stitching 09/12/2013 2 Pipeline Compute Vision : Exercise
Computer Vision Exercise 3 Panorama Stitching 09/12/2013 Compute Vision : Exercise 3 Panorama Stitching The task Compute Vision : Exercise 3 Panorama Stitching 09/12/2013 2 Pipeline Compute Vision : Exercise
Modern Methods of Data Analysis - WS 07/08
 Modern Methods of Data Analysis Lecture XV (04.02.08) Contents: Function Minimization (see E. Lohrmann & V. Blobel) Optimization Problem Set of n independent variables Sometimes in addition some constraints
Modern Methods of Data Analysis Lecture XV (04.02.08) Contents: Function Minimization (see E. Lohrmann & V. Blobel) Optimization Problem Set of n independent variables Sometimes in addition some constraints
Lecture Overview. Sequence search & alignment. Searching sequence databases. Sequence Alignment & Search. Goals: Motivations:
 Lecture Overview Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating
Lecture Overview Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating
3D Statistical Shape Model Building using Consistent Parameterization
 3D Statistical Shape Model Building using Consistent Parameterization Matthias Kirschner, Stefan Wesarg Graphisch Interaktive Systeme, TU Darmstadt matthias.kirschner@gris.tu-darmstadt.de Abstract. We
3D Statistical Shape Model Building using Consistent Parameterization Matthias Kirschner, Stefan Wesarg Graphisch Interaktive Systeme, TU Darmstadt matthias.kirschner@gris.tu-darmstadt.de Abstract. We
Sequence Alignment & Search
 Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating the first version
Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating the first version
A NEW GENERATION OF HOMOLOGY SEARCH TOOLS BASED ON PROBABILISTIC INFERENCE
 205 A NEW GENERATION OF HOMOLOGY SEARCH TOOLS BASED ON PROBABILISTIC INFERENCE SEAN R. EDDY 1 eddys@janelia.hhmi.org 1 Janelia Farm Research Campus, Howard Hughes Medical Institute, 19700 Helix Drive,
205 A NEW GENERATION OF HOMOLOGY SEARCH TOOLS BASED ON PROBABILISTIC INFERENCE SEAN R. EDDY 1 eddys@janelia.hhmi.org 1 Janelia Farm Research Campus, Howard Hughes Medical Institute, 19700 Helix Drive,
CS 581. Tandy Warnow
 CS 581 Tandy Warnow This week Maximum parsimony: solving it on small datasets Maximum Likelihood optimization problem Felsenstein s pruning algorithm Bayesian MCMC methods Research opportunities Maximum
CS 581 Tandy Warnow This week Maximum parsimony: solving it on small datasets Maximum Likelihood optimization problem Felsenstein s pruning algorithm Bayesian MCMC methods Research opportunities Maximum
Unsupervised Learning : Clustering
 Unsupervised Learning : Clustering Things to be Addressed Traditional Learning Models. Cluster Analysis K-means Clustering Algorithm Drawbacks of traditional clustering algorithms. Clustering as a complex
Unsupervised Learning : Clustering Things to be Addressed Traditional Learning Models. Cluster Analysis K-means Clustering Algorithm Drawbacks of traditional clustering algorithms. Clustering as a complex
Sequence alignment theory and applications Session 3: BLAST algorithm
 Sequence alignment theory and applications Session 3: BLAST algorithm Introduction to Bioinformatics online course : IBT Sonal Henson Learning Objectives Understand the principles of the BLAST algorithm
Sequence alignment theory and applications Session 3: BLAST algorithm Introduction to Bioinformatics online course : IBT Sonal Henson Learning Objectives Understand the principles of the BLAST algorithm
Non-Linearity of Scorecard Log-Odds
 Non-Linearity of Scorecard Log-Odds Ross McDonald, Keith Smith, Matthew Sturgess, Edward Huang Retail Decision Science, Lloyds Banking Group Edinburgh Credit Scoring Conference 6 th August 9 Lloyds Banking
Non-Linearity of Scorecard Log-Odds Ross McDonald, Keith Smith, Matthew Sturgess, Edward Huang Retail Decision Science, Lloyds Banking Group Edinburgh Credit Scoring Conference 6 th August 9 Lloyds Banking
Supplementary Material: The Emergence of. Organizing Structure in Conceptual Representation
 Supplementary Material: The Emergence of Organizing Structure in Conceptual Representation Brenden M. Lake, 1,2 Neil D. Lawrence, 3 Joshua B. Tenenbaum, 4,5 1 Center for Data Science, New York University
Supplementary Material: The Emergence of Organizing Structure in Conceptual Representation Brenden M. Lake, 1,2 Neil D. Lawrence, 3 Joshua B. Tenenbaum, 4,5 1 Center for Data Science, New York University
INDEPENDENT COMPONENT ANALYSIS WITH QUANTIZING DENSITY ESTIMATORS. Peter Meinicke, Helge Ritter. Neuroinformatics Group University Bielefeld Germany
 INDEPENDENT COMPONENT ANALYSIS WITH QUANTIZING DENSITY ESTIMATORS Peter Meinicke, Helge Ritter Neuroinformatics Group University Bielefeld Germany ABSTRACT We propose an approach to source adaptivity in
INDEPENDENT COMPONENT ANALYSIS WITH QUANTIZING DENSITY ESTIMATORS Peter Meinicke, Helge Ritter Neuroinformatics Group University Bielefeld Germany ABSTRACT We propose an approach to source adaptivity in
Feature Matching and Robust Fitting
 Feature Matching and Robust Fitting Computer Vision CS 143, Brown Read Szeliski 4.1 James Hays Acknowledgment: Many slides from Derek Hoiem and Grauman&Leibe 2008 AAAI Tutorial Project 2 questions? This
Feature Matching and Robust Fitting Computer Vision CS 143, Brown Read Szeliski 4.1 James Hays Acknowledgment: Many slides from Derek Hoiem and Grauman&Leibe 2008 AAAI Tutorial Project 2 questions? This
The Effect of Inverse Document Frequency Weights on Indexed Sequence Retrieval. Kevin C. O'Kane. Department of Computer Science
 The Effect of Inverse Document Frequency Weights on Indexed Sequence Retrieval Kevin C. O'Kane Department of Computer Science The University of Northern Iowa Cedar Falls, Iowa okane@cs.uni.edu http://www.cs.uni.edu/~okane
The Effect of Inverse Document Frequency Weights on Indexed Sequence Retrieval Kevin C. O'Kane Department of Computer Science The University of Northern Iowa Cedar Falls, Iowa okane@cs.uni.edu http://www.cs.uni.edu/~okane
24 Grundlagen der Bioinformatik, SS 10, D. Huson, April 26, This lecture is based on the following papers, which are all recommended reading:
 24 Grundlagen der Bioinformatik, SS 10, D. Huson, April 26, 2010 3 BLAST and FASTA This lecture is based on the following papers, which are all recommended reading: D.J. Lipman and W.R. Pearson, Rapid
24 Grundlagen der Bioinformatik, SS 10, D. Huson, April 26, 2010 3 BLAST and FASTA This lecture is based on the following papers, which are all recommended reading: D.J. Lipman and W.R. Pearson, Rapid
Distributed Protein Sequence Alignment
 Distributed Protein Sequence Alignment ABSTRACT J. Michael Meehan meehan@wwu.edu James Hearne hearne@wwu.edu Given the explosive growth of biological sequence databases and the computational complexity
Distributed Protein Sequence Alignment ABSTRACT J. Michael Meehan meehan@wwu.edu James Hearne hearne@wwu.edu Given the explosive growth of biological sequence databases and the computational complexity
CS201: Computer Vision Introduction to Tracking
 CS201: Computer Vision Introduction to Tracking John Magee 18 November 2014 Slides courtesy of: Diane H. Theriault Question of the Day How can we represent and use motion in images? 1 What is Motion? Change
CS201: Computer Vision Introduction to Tracking John Magee 18 November 2014 Slides courtesy of: Diane H. Theriault Question of the Day How can we represent and use motion in images? 1 What is Motion? Change
Chapter 2 Basic Structure of High-Dimensional Spaces
 Chapter 2 Basic Structure of High-Dimensional Spaces Data is naturally represented geometrically by associating each record with a point in the space spanned by the attributes. This idea, although simple,
Chapter 2 Basic Structure of High-Dimensional Spaces Data is naturally represented geometrically by associating each record with a point in the space spanned by the attributes. This idea, although simple,
Simplex of Nelder & Mead Algorithm
 Simplex of N & M Simplex of Nelder & Mead Algorithm AKA the Amoeba algorithm In the class of direct search methods Unconstrained (although constraints can be added as part of error function) nonlinear
Simplex of N & M Simplex of Nelder & Mead Algorithm AKA the Amoeba algorithm In the class of direct search methods Unconstrained (although constraints can be added as part of error function) nonlinear
CHAPTER 2 CONVENTIONAL AND NON-CONVENTIONAL TECHNIQUES TO SOLVE ORPD PROBLEM
 20 CHAPTER 2 CONVENTIONAL AND NON-CONVENTIONAL TECHNIQUES TO SOLVE ORPD PROBLEM 2.1 CLASSIFICATION OF CONVENTIONAL TECHNIQUES Classical optimization methods can be classified into two distinct groups:
20 CHAPTER 2 CONVENTIONAL AND NON-CONVENTIONAL TECHNIQUES TO SOLVE ORPD PROBLEM 2.1 CLASSIFICATION OF CONVENTIONAL TECHNIQUES Classical optimization methods can be classified into two distinct groups:
Lecture 19: November 5
 0-725/36-725: Convex Optimization Fall 205 Lecturer: Ryan Tibshirani Lecture 9: November 5 Scribes: Hyun Ah Song Note: LaTeX template courtesy of UC Berkeley EECS dept. Disclaimer: These notes have not
0-725/36-725: Convex Optimization Fall 205 Lecturer: Ryan Tibshirani Lecture 9: November 5 Scribes: Hyun Ah Song Note: LaTeX template courtesy of UC Berkeley EECS dept. Disclaimer: These notes have not
Stephen Scott.
 1 / 33 sscott@cse.unl.edu 2 / 33 Start with a set of sequences In each column, residues are homolgous Residues occupy similar positions in 3D structure Residues diverge from a common ancestral residue
1 / 33 sscott@cse.unl.edu 2 / 33 Start with a set of sequences In each column, residues are homolgous Residues occupy similar positions in 3D structure Residues diverge from a common ancestral residue
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014
 Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
Lecture 6 - Multivariate numerical optimization
 Lecture 6 - Multivariate numerical optimization Björn Andersson (w/ Jianxin Wei) Department of Statistics, Uppsala University February 13, 2014 1 / 36 Table of Contents 1 Plotting functions of two variables
Lecture 6 - Multivariate numerical optimization Björn Andersson (w/ Jianxin Wei) Department of Statistics, Uppsala University February 13, 2014 1 / 36 Table of Contents 1 Plotting functions of two variables
CPSC 340: Machine Learning and Data Mining. Principal Component Analysis Fall 2016
 CPSC 340: Machine Learning and Data Mining Principal Component Analysis Fall 2016 A2/Midterm: Admin Grades/solutions will be posted after class. Assignment 4: Posted, due November 14. Extra office hours:
CPSC 340: Machine Learning and Data Mining Principal Component Analysis Fall 2016 A2/Midterm: Admin Grades/solutions will be posted after class. Assignment 4: Posted, due November 14. Extra office hours:
From Smith-Waterman to BLAST
 From Smith-Waterman to BLAST Jeremy Buhler July 23, 2015 Smith-Waterman is the fundamental tool that we use to decide how similar two sequences are. Isn t that all that BLAST does? In principle, it is
From Smith-Waterman to BLAST Jeremy Buhler July 23, 2015 Smith-Waterman is the fundamental tool that we use to decide how similar two sequences are. Isn t that all that BLAST does? In principle, it is
JET 2 User Manual 1 INSTALLATION 2 EXECUTION AND FUNCTIONALITIES. 1.1 Download. 1.2 System requirements. 1.3 How to install JET 2
 JET 2 User Manual 1 INSTALLATION 1.1 Download The JET 2 package is available at www.lcqb.upmc.fr/jet2. 1.2 System requirements JET 2 runs on Linux or Mac OS X. The program requires some external tools
JET 2 User Manual 1 INSTALLATION 1.1 Download The JET 2 package is available at www.lcqb.upmc.fr/jet2. 1.2 System requirements JET 2 runs on Linux or Mac OS X. The program requires some external tools
Phylogenetics. Introduction to Bioinformatics Dortmund, Lectures: Sven Rahmann. Exercises: Udo Feldkamp, Michael Wurst
 Phylogenetics Introduction to Bioinformatics Dortmund, 16.-20.07.2007 Lectures: Sven Rahmann Exercises: Udo Feldkamp, Michael Wurst 1 Phylogenetics phylum = tree phylogenetics: reconstruction of evolutionary
Phylogenetics Introduction to Bioinformatics Dortmund, 16.-20.07.2007 Lectures: Sven Rahmann Exercises: Udo Feldkamp, Michael Wurst 1 Phylogenetics phylum = tree phylogenetics: reconstruction of evolutionary
Behavioral Data Mining. Lecture 18 Clustering
 Behavioral Data Mining Lecture 18 Clustering Outline Why? Cluster quality K-means Spectral clustering Generative Models Rationale Given a set {X i } for i = 1,,n, a clustering is a partition of the X i
Behavioral Data Mining Lecture 18 Clustering Outline Why? Cluster quality K-means Spectral clustering Generative Models Rationale Given a set {X i } for i = 1,,n, a clustering is a partition of the X i
Research Article International Journals of Advanced Research in Computer Science and Software Engineering ISSN: X (Volume-7, Issue-6)
 International Journals of Advanced Research in Computer Science and Software Engineering ISSN: 77-18X (Volume-7, Issue-6) Research Article June 017 DDGARM: Dotlet Driven Global Alignment with Reduced Matrix
International Journals of Advanced Research in Computer Science and Software Engineering ISSN: 77-18X (Volume-7, Issue-6) Research Article June 017 DDGARM: Dotlet Driven Global Alignment with Reduced Matrix
Bioinformatics for Biologists
 Bioinformatics for Biologists Sequence Analysis: Part I. Pairwise alignment and database searching Fran Lewitter, Ph.D. Director Bioinformatics & Research Computing Whitehead Institute Topics to Cover
Bioinformatics for Biologists Sequence Analysis: Part I. Pairwise alignment and database searching Fran Lewitter, Ph.D. Director Bioinformatics & Research Computing Whitehead Institute Topics to Cover
Computer Vision for HCI. Topics of This Lecture
 Computer Vision for HCI Interest Points Topics of This Lecture Local Invariant Features Motivation Requirements, Invariances Keypoint Localization Features from Accelerated Segment Test (FAST) Harris Shi-Tomasi
Computer Vision for HCI Interest Points Topics of This Lecture Local Invariant Features Motivation Requirements, Invariances Keypoint Localization Features from Accelerated Segment Test (FAST) Harris Shi-Tomasi
Clustering. CE-717: Machine Learning Sharif University of Technology Spring Soleymani
 Clustering CE-717: Machine Learning Sharif University of Technology Spring 2016 Soleymani Outline Clustering Definition Clustering main approaches Partitional (flat) Hierarchical Clustering validation
Clustering CE-717: Machine Learning Sharif University of Technology Spring 2016 Soleymani Outline Clustering Definition Clustering main approaches Partitional (flat) Hierarchical Clustering validation
Multiple sequence alignment. November 20, 2018
 Multiple sequence alignment November 20, 2018 Why do multiple alignment? Gain insight into evolutionary history Can assess time of divergence by looking at the number of mutations needed to change one
Multiple sequence alignment November 20, 2018 Why do multiple alignment? Gain insight into evolutionary history Can assess time of divergence by looking at the number of mutations needed to change one
Chapter 7: Competitive learning, clustering, and self-organizing maps
 Chapter 7: Competitive learning, clustering, and self-organizing maps António R. C. Paiva EEL 6814 Spring 2008 Outline Competitive learning Clustering Self-Organizing Maps What is competition in neural
Chapter 7: Competitive learning, clustering, and self-organizing maps António R. C. Paiva EEL 6814 Spring 2008 Outline Competitive learning Clustering Self-Organizing Maps What is competition in neural
Neural Network Learning. Today s Lecture. Continuation of Neural Networks. Artificial Neural Networks. Lecture 24: Learning 3. Victor R.
 Lecture 24: Learning 3 Victor R. Lesser CMPSCI 683 Fall 2010 Today s Lecture Continuation of Neural Networks Artificial Neural Networks Compose of nodes/units connected by links Each link has a numeric
Lecture 24: Learning 3 Victor R. Lesser CMPSCI 683 Fall 2010 Today s Lecture Continuation of Neural Networks Artificial Neural Networks Compose of nodes/units connected by links Each link has a numeric
Mouse, Human, Chimpanzee
 More Alignments 1 Mouse, Human, Chimpanzee Mouse to Human Chimpanzee to Human 2 Mouse v.s. Human Chromosome X of Mouse to Human 3 Local Alignment Given: two sequences S and T Find: substrings of S and
More Alignments 1 Mouse, Human, Chimpanzee Mouse to Human Chimpanzee to Human 2 Mouse v.s. Human Chromosome X of Mouse to Human 3 Local Alignment Given: two sequences S and T Find: substrings of S and
GLOBEX Bioinformatics (Summer 2015) Multiple Sequence Alignment
 GLOBEX Bioinformatics (Summer 2015) Multiple Sequence Alignment Scoring Dynamic Programming algorithms Heuristic algorithms CLUSTAL W Courtesy of jalview Motivations Collective (or aggregate) statistic
GLOBEX Bioinformatics (Summer 2015) Multiple Sequence Alignment Scoring Dynamic Programming algorithms Heuristic algorithms CLUSTAL W Courtesy of jalview Motivations Collective (or aggregate) statistic
Kapitel 5: Local Search
 Inhalt: Kapitel 5: Local Search Gradient Descent (Hill Climbing) Metropolis Algorithm and Simulated Annealing Local Search in Hopfield Neural Networks Local Search for Max-Cut Single-flip neighborhood
Inhalt: Kapitel 5: Local Search Gradient Descent (Hill Climbing) Metropolis Algorithm and Simulated Annealing Local Search in Hopfield Neural Networks Local Search for Max-Cut Single-flip neighborhood
Gene regulation. DNA is merely the blueprint Shared spatially (among all tissues) and temporally But cells manage to differentiate
 Gene regulation DNA is merely the blueprint Shared spatially (among all tissues) and temporally But cells manage to differentiate Especially but not only during developmental stage And cells respond to
Gene regulation DNA is merely the blueprint Shared spatially (among all tissues) and temporally But cells manage to differentiate Especially but not only during developmental stage And cells respond to
NEURAL NETWORK VISUALIZATION
 Neural Network Visualization 465 NEURAL NETWORK VISUALIZATION Jakub Wejchert Gerald Tesauro IB M Research T.J. Watson Research Center Yorktown Heights NY 10598 ABSTRACT We have developed graphics to visualize
Neural Network Visualization 465 NEURAL NETWORK VISUALIZATION Jakub Wejchert Gerald Tesauro IB M Research T.J. Watson Research Center Yorktown Heights NY 10598 ABSTRACT We have developed graphics to visualize
10703 Deep Reinforcement Learning and Control
 10703 Deep Reinforcement Learning and Control Russ Salakhutdinov Machine Learning Department rsalakhu@cs.cmu.edu Policy Gradient I Used Materials Disclaimer: Much of the material and slides for this lecture
10703 Deep Reinforcement Learning and Control Russ Salakhutdinov Machine Learning Department rsalakhu@cs.cmu.edu Policy Gradient I Used Materials Disclaimer: Much of the material and slides for this lecture
Comparison of Sequence Similarity Measures for Distant Evolutionary Relationships
 Comparison of Sequence Similarity Measures for Distant Evolutionary Relationships Abhishek Majumdar, Peter Z. Revesz Department of Computer Science and Engineering, University of Nebraska-Lincoln, Lincoln,
Comparison of Sequence Similarity Measures for Distant Evolutionary Relationships Abhishek Majumdar, Peter Z. Revesz Department of Computer Science and Engineering, University of Nebraska-Lincoln, Lincoln,
Level-set MCMC Curve Sampling and Geometric Conditional Simulation
 Level-set MCMC Curve Sampling and Geometric Conditional Simulation Ayres Fan John W. Fisher III Alan S. Willsky February 16, 2007 Outline 1. Overview 2. Curve evolution 3. Markov chain Monte Carlo 4. Curve
Level-set MCMC Curve Sampling and Geometric Conditional Simulation Ayres Fan John W. Fisher III Alan S. Willsky February 16, 2007 Outline 1. Overview 2. Curve evolution 3. Markov chain Monte Carlo 4. Curve
Image segmentation using an annealed Hopfield neural network
 Iowa State University From the SelectedWorks of Sarah A. Rajala December 16, 1992 Image segmentation using an annealed Hopfield neural network Yungsik Kim, North Carolina State University Sarah A. Rajala,
Iowa State University From the SelectedWorks of Sarah A. Rajala December 16, 1992 Image segmentation using an annealed Hopfield neural network Yungsik Kim, North Carolina State University Sarah A. Rajala,
Efficient Robust Shape Optimization for Crashworthiness
 10 th World Congress on Structural and Multidisciplinary Optimization May 19-24, 2013, Orlando, Florida, USA Efficient Robust Shape Optimization for Crashworthiness Milan Rayamajhi 1, Stephan Hunkeler
10 th World Congress on Structural and Multidisciplinary Optimization May 19-24, 2013, Orlando, Florida, USA Efficient Robust Shape Optimization for Crashworthiness Milan Rayamajhi 1, Stephan Hunkeler
Robust Regression. Robust Data Mining Techniques By Boonyakorn Jantaranuson
 Robust Regression Robust Data Mining Techniques By Boonyakorn Jantaranuson Outline Introduction OLS and important terminology Least Median of Squares (LMedS) M-estimator Penalized least squares What is
Robust Regression Robust Data Mining Techniques By Boonyakorn Jantaranuson Outline Introduction OLS and important terminology Least Median of Squares (LMedS) M-estimator Penalized least squares What is
Jyoti Lakhani 1, Ajay Khunteta 2, Dharmesh Harwani *3 1 Poornima University, Jaipur & Maharaja Ganga Singh University, Bikaner, Rajasthan, India
 International Journal of Scientific Research in Computer Science, Engineering and Information Technology 2017 IJSRCSEIT Volume 2 Issue 6 ISSN : 2456-3307 Improvisation of Global Pairwise Sequence Alignment
International Journal of Scientific Research in Computer Science, Engineering and Information Technology 2017 IJSRCSEIT Volume 2 Issue 6 ISSN : 2456-3307 Improvisation of Global Pairwise Sequence Alignment
Heuristic methods for pairwise alignment:
 Bi03c_1 Unit 03c: Heuristic methods for pairwise alignment: k-tuple-methods k-tuple-methods for alignment of pairs of sequences Bi03c_2 dynamic programming is too slow for large databases Use heuristic
Bi03c_1 Unit 03c: Heuristic methods for pairwise alignment: k-tuple-methods k-tuple-methods for alignment of pairs of sequences Bi03c_2 dynamic programming is too slow for large databases Use heuristic
Sequence analysis Pairwise sequence alignment
 UMF11 Introduction to bioinformatics, 25 Sequence analysis Pairwise sequence alignment 1. Sequence alignment Lecturer: Marina lexandersson 12 September, 25 here are two types of sequence alignments, global
UMF11 Introduction to bioinformatics, 25 Sequence analysis Pairwise sequence alignment 1. Sequence alignment Lecturer: Marina lexandersson 12 September, 25 here are two types of sequence alignments, global
mywbut.com Informed Search Strategies-II
 Informed Search Strategies-II 1 3.3 Iterative-Deepening A* 3.3.1 IDA* Algorithm Iterative deepening A* or IDA* is similar to iterative-deepening depth-first, but with the following modifications: The depth
Informed Search Strategies-II 1 3.3 Iterative-Deepening A* 3.3.1 IDA* Algorithm Iterative deepening A* or IDA* is similar to iterative-deepening depth-first, but with the following modifications: The depth
Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it.
 Sequence Alignments Overview Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it. Sequence alignment means arranging
Sequence Alignments Overview Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it. Sequence alignment means arranging
BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio CS 466 Saurabh Sinha
 BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio. 1990. CS 466 Saurabh Sinha Motivation Sequence homology to a known protein suggest function of newly sequenced protein Bioinformatics
BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio. 1990. CS 466 Saurabh Sinha Motivation Sequence homology to a known protein suggest function of newly sequenced protein Bioinformatics
A Brief Look at Optimization
 A Brief Look at Optimization CSC 412/2506 Tutorial David Madras January 18, 2018 Slides adapted from last year s version Overview Introduction Classes of optimization problems Linear programming Steepest
A Brief Look at Optimization CSC 412/2506 Tutorial David Madras January 18, 2018 Slides adapted from last year s version Overview Introduction Classes of optimization problems Linear programming Steepest
MRF-based Algorithms for Segmentation of SAR Images
 This paper originally appeared in the Proceedings of the 998 International Conference on Image Processing, v. 3, pp. 770-774, IEEE, Chicago, (998) MRF-based Algorithms for Segmentation of SAR Images Robert
This paper originally appeared in the Proceedings of the 998 International Conference on Image Processing, v. 3, pp. 770-774, IEEE, Chicago, (998) MRF-based Algorithms for Segmentation of SAR Images Robert
University of Florida CISE department Gator Engineering. Clustering Part 2
 Clustering Part 2 Dr. Sanjay Ranka Professor Computer and Information Science and Engineering University of Florida, Gainesville Partitional Clustering Original Points A Partitional Clustering Hierarchical
Clustering Part 2 Dr. Sanjay Ranka Professor Computer and Information Science and Engineering University of Florida, Gainesville Partitional Clustering Original Points A Partitional Clustering Hierarchical
Lecture 18: March 23
 0-725/36-725: Convex Optimization Spring 205 Lecturer: Ryan Tibshirani Lecture 8: March 23 Scribes: James Duyck Note: LaTeX template courtesy of UC Berkeley EECS dept. Disclaimer: These notes have not
0-725/36-725: Convex Optimization Spring 205 Lecturer: Ryan Tibshirani Lecture 8: March 23 Scribes: James Duyck Note: LaTeX template courtesy of UC Berkeley EECS dept. Disclaimer: These notes have not
Tuning. Philipp Koehn presented by Gaurav Kumar. 28 September 2017
 Tuning Philipp Koehn presented by Gaurav Kumar 28 September 2017 The Story so Far: Generative Models 1 The definition of translation probability follows a mathematical derivation argmax e p(e f) = argmax
Tuning Philipp Koehn presented by Gaurav Kumar 28 September 2017 The Story so Far: Generative Models 1 The definition of translation probability follows a mathematical derivation argmax e p(e f) = argmax
LAGAN and Multi-LAGAN: Efficient Tools for Large-Scale Multiple Alignment of Genomic DNA
 LAGAN and Multi-LAGAN: Efficient Tools for Large-Scale Multiple Alignment of Genomic DNA Michael Brudno, Chuong B. Do, Gregory M. Cooper, et al. Presented by Xuebei Yang About Alignments Pairwise Alignments
LAGAN and Multi-LAGAN: Efficient Tools for Large-Scale Multiple Alignment of Genomic DNA Michael Brudno, Chuong B. Do, Gregory M. Cooper, et al. Presented by Xuebei Yang About Alignments Pairwise Alignments
A Self-Organizing Binary System*
 212 1959 PROCEEDINGS OF THE EASTERN JOINT COMPUTER CONFERENCE A Self-Organizing Binary System* RICHARD L. MATTSONt INTRODUCTION ANY STIMULUS to a system such as described in this paper can be coded into
212 1959 PROCEEDINGS OF THE EASTERN JOINT COMPUTER CONFERENCE A Self-Organizing Binary System* RICHARD L. MATTSONt INTRODUCTION ANY STIMULUS to a system such as described in this paper can be coded into
On-line Estimation of Power System Security Limits
 Bulk Power System Dynamics and Control V, August 26-31, 2001, Onomichi, Japan On-line Estimation of Power System Security Limits K. Tomsovic A. Sittithumwat J. Tong School of Electrical Engineering and
Bulk Power System Dynamics and Control V, August 26-31, 2001, Onomichi, Japan On-line Estimation of Power System Security Limits K. Tomsovic A. Sittithumwat J. Tong School of Electrical Engineering and
CS570: Introduction to Data Mining
 CS570: Introduction to Data Mining Classification Advanced Reading: Chapter 8 & 9 Han, Chapters 4 & 5 Tan Anca Doloc-Mihu, Ph.D. Slides courtesy of Li Xiong, Ph.D., 2011 Han, Kamber & Pei. Data Mining.
CS570: Introduction to Data Mining Classification Advanced Reading: Chapter 8 & 9 Han, Chapters 4 & 5 Tan Anca Doloc-Mihu, Ph.D. Slides courtesy of Li Xiong, Ph.D., 2011 Han, Kamber & Pei. Data Mining.
Sequence Alignment. Ulf Leser
 Sequence Alignment Ulf Leser his Lecture Approximate String Matching Edit distance and alignment Computing global alignments Local alignment Ulf Leser: Bioinformatics, Summer Semester 2016 2 ene Function
Sequence Alignment Ulf Leser his Lecture Approximate String Matching Edit distance and alignment Computing global alignments Local alignment Ulf Leser: Bioinformatics, Summer Semester 2016 2 ene Function
Multiple Sequence Alignment. Mark Whitsitt - NCSA
 Multiple Sequence Alignment Mark Whitsitt - NCSA What is a Multiple Sequence Alignment (MA)? GMHGTVYANYAVDSSDLLLAFGVRFDDRVTGKLEAFASRAKIVHIDIDSAEIGKNKQPHV GMHGTVYANYAVEHSDLLLAFGVRFDDRVTGKLEAFASRAKIVHIDIDSAEIGKNKTPHV
Multiple Sequence Alignment Mark Whitsitt - NCSA What is a Multiple Sequence Alignment (MA)? GMHGTVYANYAVDSSDLLLAFGVRFDDRVTGKLEAFASRAKIVHIDIDSAEIGKNKQPHV GMHGTVYANYAVEHSDLLLAFGVRFDDRVTGKLEAFASRAKIVHIDIDSAEIGKNKTPHV
CISC 636 Computational Biology & Bioinformatics (Fall 2016)
 CISC 636 Computational Biology & Bioinformatics (Fall 2016) Sequence pairwise alignment Score statistics: E-value and p-value Heuristic algorithms: BLAST and FASTA Database search: gene finding and annotations
CISC 636 Computational Biology & Bioinformatics (Fall 2016) Sequence pairwise alignment Score statistics: E-value and p-value Heuristic algorithms: BLAST and FASTA Database search: gene finding and annotations
Guided Image Super-Resolution: A New Technique for Photogeometric Super-Resolution in Hybrid 3-D Range Imaging
 Guided Image Super-Resolution: A New Technique for Photogeometric Super-Resolution in Hybrid 3-D Range Imaging Florin C. Ghesu 1, Thomas Köhler 1,2, Sven Haase 1, Joachim Hornegger 1,2 04.09.2014 1 Pattern
Guided Image Super-Resolution: A New Technique for Photogeometric Super-Resolution in Hybrid 3-D Range Imaging Florin C. Ghesu 1, Thomas Köhler 1,2, Sven Haase 1, Joachim Hornegger 1,2 04.09.2014 1 Pattern
Random Search Report An objective look at random search performance for 4 problem sets
 Random Search Report An objective look at random search performance for 4 problem sets Dudon Wai Georgia Institute of Technology CS 7641: Machine Learning Atlanta, GA dwai3@gatech.edu Abstract: This report
Random Search Report An objective look at random search performance for 4 problem sets Dudon Wai Georgia Institute of Technology CS 7641: Machine Learning Atlanta, GA dwai3@gatech.edu Abstract: This report
Supplementary Material for The Generalized PatchMatch Correspondence Algorithm
 Supplementary Material for The Generalized PatchMatch Correspondence Algorithm Connelly Barnes 1, Eli Shechtman 2, Dan B Goldman 2, Adam Finkelstein 1 1 Princeton University, 2 Adobe Systems 1 Overview
Supplementary Material for The Generalized PatchMatch Correspondence Algorithm Connelly Barnes 1, Eli Shechtman 2, Dan B Goldman 2, Adam Finkelstein 1 1 Princeton University, 2 Adobe Systems 1 Overview
Sparse Matrices Reordering using Evolutionary Algorithms: A Seeded Approach
 1 Sparse Matrices Reordering using Evolutionary Algorithms: A Seeded Approach David Greiner, Gustavo Montero, Gabriel Winter Institute of Intelligent Systems and Numerical Applications in Engineering (IUSIANI)
1 Sparse Matrices Reordering using Evolutionary Algorithms: A Seeded Approach David Greiner, Gustavo Montero, Gabriel Winter Institute of Intelligent Systems and Numerical Applications in Engineering (IUSIANI)
Convex Optimization CMU-10725
 Convex Optimization CMU-10725 Conjugate Direction Methods Barnabás Póczos & Ryan Tibshirani Conjugate Direction Methods 2 Books to Read David G. Luenberger, Yinyu Ye: Linear and Nonlinear Programming Nesterov:
Convex Optimization CMU-10725 Conjugate Direction Methods Barnabás Póczos & Ryan Tibshirani Conjugate Direction Methods 2 Books to Read David G. Luenberger, Yinyu Ye: Linear and Nonlinear Programming Nesterov:
